Abstract
Eph-related receptor tyrosine kinases (RTKs) have been implicated in intercellular communication during embryonic development. To elucidate their signal transduction pathways, we applied the yeast two-hybrid system. We could demonstrate that the carboxyl termini of the Eph-related RTKs EphA7, EphB2, EphB3, EphB5, and EphB6 interact with the PDZ domain of the ras-binding protein AF6. A mutational analysis revealed that six C-terminal residues of the receptors are involved in binding to the PDZ domain of AF6 in a sequence-specific fashion. Moreover, this PDZ domain also interacts with C-terminal sequences derived from other transmembrane receptors such as neurexins and the Notch ligand Jagged. In contrast to the association of EphB3 to the PDZ domain of AF6, the interaction with full-length AF6 clearly depends on the kinase activity of EphB3, suggesting a regulated mechanism for the PDZ-domain-mediated interaction. These data gave rise to the idea that the binding of AF6 to EphB3 occurs in a cooperative fashion because of synergistic effects involving different epitopes of both proteins. Moreover, in NIH 3T3 and NG108 cells endogenous AF6 is phosphorylated specifically by EphB3 and EphB2 in a ligand-dependent fashion. Our observations add the PDZ domain to the group of conserved protein modules such as Src-homology-2 (SH2) and phosphotyrosine-binding (PTB) domains that regulate signal transduction through their ability to mediate the interaction with RTKs.
The effects of many growth factors and cytokines are mediated by high-affinity binding to receptor tyrosine kinases (RTKs) resulting in autophosphorylation of the cytoplasmic domain (1). Phosphotyrosine residues serve as binding sites for downstream signaling proteins that establish a complex network of interactions within the cell. Modular structures involved in binding of activated RTKs identified so far include Src-homology-2 (SH2) and phosphotyrosine-binding (PTB) domains, which both bind specifically to phosphorylated tyrosine residues in a sequence-specific fashion (2).
Recently, we isolated two Eph-related RTKs, named EphB3 and EphB2 (refs. 3 and 4; B.B., U.H., T.K., and K.S., unpublished data), which both bind specifically to the transmembrane subgroup of Eph-receptor ligands (5, 6). The genetic analysis of EphB2 and EphB3 revealed a physiological requirement of both receptors for pathfinding of specific commissural axons in the central nervous system (7, 8). To date, the information about signaling molecules mediating Eph-receptor-specific responses is still limited to a few family members. Interactions have been described for the activated Eph-family members EphA2, EphA4, and EphB1 and the SH2-domain-containing proteins p85 subunit of phosphatidylinositol 3-kinase, the adapter protein SLAP, Grb2, Grb10, and Fyn (8–11). Lately, we found that activated EphB3 interacts with Crk, Fyn, and rasGAP in a SH2-dependent manner (12): Autophosphorylation of Tyr-614 in the juxtamembrane region of the receptor generates a multi-docking-site for these interactions. Moreover, we reported that EphB2 binds rasGAP and phosphorylates p62doc, which in turn associates with rasGAP and Nck (13).
In an attempt to identify yet-unknown proteins capable of interacting with Eph-related RTKs, we performed two-hybrid screenings. We found that the C-terminal tails of various Eph-related receptors (EphA7, EphB2, EphB3, EphB5 and EphB6) interact with the PDZ domain of the ras-binding protein AF6. The interaction is mediated in a sequence-specific fashion by the PDZ domain of AF6 and six C-terminal amino acids of the receptors. The binding of the PDZ domain does not depend on the catalytic activity of the receptors. In contrast, the kinase activity of EphB3 is required for high-affinity binding of full-length AF6, suggesting a novel regulated mechanism for PDZ-domain-mediated interactions. Furthermore, we could demonstrate that in NIH 3T3 and NG108 cells endogenously expressed AF6 is phosphorylated by EphB3 and EphB2 in an ephrine-B1-dependent fashion. Our data give rise to the idea that PDZ domains are modular structures involved in signal transduction of RTKs.
MATERIALS AND METHODS
Cloning Procedures.
pRc/CMV-EphB3 and pRc/CMV-EphB3K665R have been described (5). Site-directed mutagenesis was performed with QuikChange (Stratagene). A cDNA fragment of AF6 (amino acids 36–1608) was inserted into pEF-BOS-HA (14) to generate HA-AF6, which contains an N-terminal tag of influenza virus hemagglutinin. cDNAs encoding the intracellular domains of either EphB2 (amino acids 552–970) or EphB3 (amino acids 582–998) were inserted into pBTM116URA (15, 16) for the generation of bait constructs. A partial cDNA clone of EphB3 encoding the intracellular domain except its termination codon was inserted into pBTM116URA and led to the synthesis of a protein elongated by 15 aa. For LexA-peptide fusions, double-stranded oligonucleotides were inserted into pBTM116URA.
Two-Hybrid System.
Two-hybrid screenings were performed as described (17) except that a ura3− derivative of the Saccharomyces cerevisiae L40 strain together with the plasmid pBTM116URA was used, allowing counterselection against the bait plasmid in media containing 5-fluoroorotic acid (16). Two-hybrid interactions were determined based on the growth on medium lacking histidine.
Northern Blot Analysis.
Multiple tissue blots were purchased from CLONTECH. Single-stranded cDNA probes corresponding to human AF6 (nucleotides 2955–3246) (18) and EphB3 (nucleotides 423-2055) were generated, and hybridization was performed as described (3).
Transfection and Cell Lysis.
COS-1 and NIH 3T3 cells were grown in DMEM (BioWhittaker) containing 10% fetal calf serum, 2 mM l-glutamine, 100 μg/ml streptomycin, and 100 international units/ml penicillin. Cultivation and ephrine-B1 stimulation of NG108 cells stably expressing EphB2 have been described (13). Transfections of COS-1 cells were performed by using the Lipofectamine method (BRL). Forty-eight hours after transfection cells were washed twice in PBS and lysed in ice-cold lysis buffer [50 mM Hepes, pH 7.5/150 mM NaCl/10% (vol/vol) glycerol/1% Triton X-100/1.5 mM MgCl2/1 mM EGTA] freshly supplemented with 1 μg/ml leupeptin and aprotinin, 1 mM phenylmethylsulfonyl fluoride (PMSF), and 1 mM Na3VO4. Cell lysates were incubated on ice for 15 min with occasional mixing and then clarified by centrifugation for 10 min at 12,000 × g. The protein concentration was determined by using the BCA method (Pierce). NG108 and NIH 3T3 cells were lysed in RIPA buffer, which contains 20 mM Tris⋅HCl at pH 7.5, 150 mM NaCl, 1 mM EDTA, 5 mM MgCl2, 1% (vol/vol) Triton X-100, 0.25% Nonidet P-40, 0.1% SDS, 1 mM Na3VO4, 1 mM PMSF, 1 μg/ml aprotinin, and 1 μg/ml leupeptin, as described above.
Immunopreciptiation and Western Blotting.
Polyclonal EphB3 antibodies were described previously (5). Monoclonal phosphotyrosine antibodies were purchased from Santa Cruz Biotechnology. For immunoprecipitations, polyclonal AF6 antibodies or monoclonal HA antibodies (12CA5; Babco, Richmond, CA) were added to lysates and incubated on ice for 1 hr. Protein-A-Sepharose (Sigma) was added and incubation was continued for 1 hr. Beads were recovered and washed in cold lysis buffer. Precipitated proteins were analyzed by Western blotting as described (5).
In Vitro Binding Studies.
The cDNA insert of clone 31-44 encompassing the PDZ domain of AF6 was inserted into pGEX4T1 (Pharmacia) for expression in Escherichia coli. For in vitro binding studies, extracts of COS-1 cells were incubated for 1 hr at 4°C with 1 μg of either purified glutathione S-transferase (GST) or GST-clone 31-44 coupled to glutathione-Sepharose beads. Before Western blotting with EphB3 antibodies, beads were washed with cold lysis buffer.
RESULTS
Isolation of the PDZ-Domain-Containing Protein AF-6 as Interacting Partner of the Eph-Related RTKs EphB2 and EphB3.
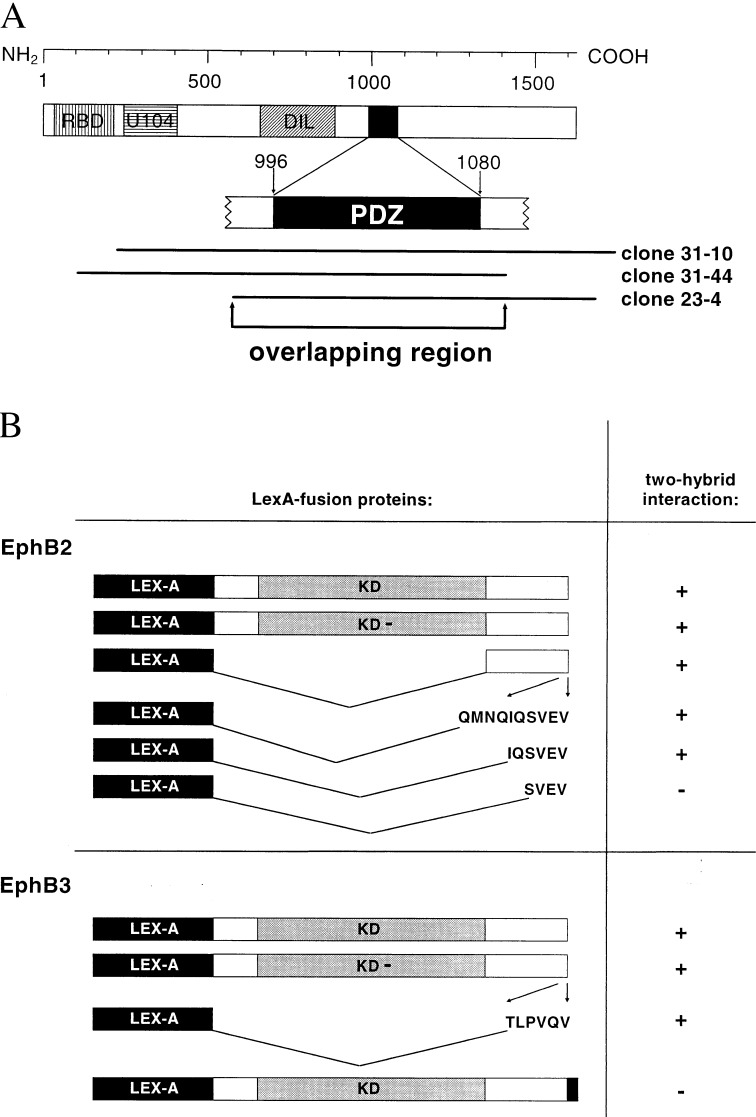
A LexA-based two-hybrid system with a size-selected murine embryonic cDNA library (15, 17) was used for a screening with the kinase-deficient cytoplasmic domain of EphB2 as bait. Three independent clones that showed specific reporter-gene activity with EphB2 as bait displayed a sequence homology of 91% to the human AF6 cDNA (18). The overlapping region of these clones encompasses the PDZ domain of AF6 (Fig. 1A). An amino acid alignment revealed a sequence identity of 100% between the human and murine PDZ domains. These data suggest that the interaction with EphB2 is mediated by the PDZ domain of AF6.
Figure 1.
Interaction of the PDZ domain of AF6 with the C termini of EphB2 and EphB3. (A) AF6-domain topology: ras-binding domain (RBD), homology regions to kinesin-like proteins (U104) and to class V myosins (DIL), PDZ domain (PDZ). cDNAs identified in a two-hybrid screening are shown relative to the enlarged PDZ domain. (B) Intracellular domains of EphB2 and EphB3 containing deletions or point mutations were tested in two-hybrid assays for their ability to bind to the PDZ domain of AF6. Interactions were detected by induction of HIS3 expression on medium lacking histidine. KD refers to the wild-type receptor, KD- to the kinase-deficient receptor. Sequences of C-terminal peptides are indicated. The C-terminal elongation of EphB3 by 15 aa is shown in black.
Moreover, we found that the PDZ domain of AF6 also interacts with the kinase-active intracellular portion of wild-type EphB2 (Fig. 1B). VP16 fusions of the N-terminal SH2 domain of rasGAP or the Fyn-SH2 domain were used as control. They induced reporter gene activity in combination with kinase-active EphB2 but not with the kinase-negative mutant (12), indicating that the cytoplasmic domain of EphB2 is autophosphorylated in yeast cells. Furthermore, we observed that the PDZ domain also associates with wild-type and kinase-deficient EphB3 (Fig. 1B), which seems to be because of the high homology between EphB2 and EphB3. Thus, the PDZ domain of AF6 interacts with the cytoplasmic moiety of the Eph-related kinases EphB2 and EphB3 in yeast cells independent of their kinase activity.
Six C-Terminal Residues of EphB2 and EphB3 Are Required for Binding to the PDZ Domain of AF6.
To determine the region within the cytoplasmic domains of EphB2 and EphB3 that is involved in the interaction with the PDZ domain of AF6, we analyzed a series of deletion mutants in two-hybrid assays. The PDZ-domain-mediated interaction with EphB2 and EphB3 could be confined to LexA fusions encompassing the last 6 C-terminal residues of the receptors (Fig. 1B). In contrast, a LexA fusion containing only 4 aa from the C terminus of EphB2 was not recognized by the PDZ domain. An EphB3 bait construct elongated by a C-terminal stretch of 15 aa also failed to interact with the PDZ domain of AF6 in yeast. These data lead us to propose that 6 specific residues at the C terminus of the receptors appear to be sufficient for the interaction of the PDZ domain of AF6. Furthermore, the C-terminal location of the motif seems to be critical.
The PDZ Domain of AF6 Binds to the C Termini of Different Eph-Related RTKs.
The comparison of the C-terminal hexapeptides of EphB2 and EphB3 revealed conserved valine residues at positions −1 and −3. Furthermore, they contain polar residues at position −2 and small, near neutral, residues at position −4 (Table 1). This observation suggests that the C-terminal tetrapeptide may define a critical core motif (small near neutral-V-E/Q-V) for the interaction with the PDZ domain of AF6. Since this C-terminal motif is also conserved among various Eph-related kinases, we asked whether other family members could interact with the PDZ domain of AF6 as well. To this end, we generated bait constructs encoding C-terminal hexapeptides of several Eph RTKs. As shown in Table 1, baits derived from EphA7, EphB5, and EphB6 (19–21) bound to the PDZ domain of AF6, whereas baits derived from EphA2, EphA3, EphA4, and EphA6 (19, 22–24) did not interact. The related C terminus of Nsk2/MuSK (25, 26), an RTK that does not belong to the Eph-receptor family, did not bind AF6 either. These results are in good agreement with the core motif proposed above, because hexapeptides derived from EphA7, EphB5, and EphB6 share this motif. Only EphA7 displays one amino acid substitution at position −3 (V → I), suggesting that a conservative substitution at this position does not inhibit the interaction. Peptides that do not interact with the PDZ domain of AF6 in two-hybrid assays contain at least one nonconservative substitution in the predicted core motif.
Table 1.
The C termini of five EPH-related RTKs bind to the PDZ domain of AF6
| RTK | Accession no. | Analyzed C terminus | AF6 binding |
|---|---|---|---|
| EphB2 | L36643 | IQSVEV | + |
| EphB6 | L77867 | PGSVEV | + |
| EphB5 | L23783 | LEPVEV | + |
| EphB3 | X75208 | TLPVQV | + |
| EphA3 | M83941 | NGPVPV | − |
| EphA4 | L36654 | GRMVPV | − |
| EphA7 | L36642 | GTGIQV | + |
| EphA6 | U58332 | EKGFHV | − |
| EphA2 | M59371 | TVGIPI | − |
| Nsk2 | X86445 | EGTVGV | − |
LexA peptide fusions derived from the C termini of various RTKs were coexpressed with the PDZ domain in S. cerevisiae L40.
Mutational Analysis of the PDZ-Domain-Binding Motif.
Using an oriented peptide library technique, Songyang et al. (27) proposed optimal peptide sequences for the in vitro binding of the AF6-PDZ domain containing aliphatic or aromatic amino acids at the three C-terminal positions. However, our data indicate that C-terminal hexapeptides of Eph-related receptors containing glutamine or glutamic acid at position −2 also have the capacity to bind the PDZ domain of AF6. Thus, site-directed mutagenesis was applied to analyze whether substitutions of the aliphatic amino acids at positions −1 and −3 of the C terminus of EphB2 influences its binding properties to the PDZ domain of AF6. For this purpose we generated LexA fusions of the EphB2 C-terminal hexapeptide containing single or multiple amino acid substitutions. As shown in Table 2, substitutions of either valine at position −1 to leucine or valine at position −3 to alanine or methionine abolished the interaction in yeast cells. In contrast, a conservative substitution of valine at position −3 to leucine or isoleucine did not influence the binding capacity of the PDZ domain. Surprisingly, a nonconservative substitution of valine at position −3 to serine or threonine did not prevent the interaction, suggesting that certain single nonconservative substitutions are tolerable. In addition, a single substitution of glutamic acid at position −2 to proline did not affect binding of the PDZ domain of AF6, whereas additional mutations of either serine at position −4 to proline or both isoleucine at position −6 to asparagine and glutamine at position −5 to glycine abolished the interaction. In summary, our study suggests that binding of the PDZ domain of AF6 is not restricted to motifs found in our analysis of Eph-related receptors or in the oriented peptide library approach used by Songyang et al. (27).
Table 2.
Sequence requirements for the PDZ-mediated interaction of EphB2 and AF6
| Mutated EphB2 C terminus | AF6 binding |
|---|---|
| IQSVEV | + |
| IQSVEL | − |
| IQSLEV | + |
| IQSIEV | + |
| IQSMEV | − |
| IQSAEV | − |
| IQSSEV | + |
| IQSTEV | + |
| IQSVPV | + |
| IQPVPV | − |
| NGSVPV | − |
The C-terminal sequence of wild-type EphB2 (IQSVEV) is given in italics. Single or multiple amino acid substitutions are shown in boldface lettering. LexA peptide fusions were expressed in yeast and their ability to interact with the PDZ domain of AF6 was monitored in two-hybrid assays.
The C-Terminal Motif T/S-X-V of Eph-Related Receptors Is Not Sufficient for Binding to the PDZ Domain of AF6.
Our mutational analysis has revealed that a modified C terminus of EphB2 bearing serine or threonine at position −3 embedded in a T/S-X-V-motif can be recognized by the PDZ domain of AF6. Identical motifs were found in transmembrane proteins such as Fas, N-methyl-d-aspartate (NMDA) receptors, or Shaker-type potassium channels, which have the capacity to bind different PDZ-containing proteins (28–30). Interestingly, a two-hybrid analysis revealed that of six receptors that display a T/S-X-V motif at their C termini only mutated EphB2 had the capability to bind the PDZ domain of AF6 (Table 3).
Table 3.
Specific binding of the C termini of different transmembrane proteins to the PDZ domain of AF6
| Transmembrane protein | Accession no. | Analyzed C terminus | AF6 binding |
|---|---|---|---|
| Mutated EphB2 | — | IQSSEV | + |
| Nr2C K+ channel | P35429 | SLESEV | − |
| Nr2D K+ channel | Q03391 | SLESEV | − |
| NMDAR Kv1.4 | G57036 | AVETDV | − |
| Fas/APO-1 | X63717 | EIQSLV | − |
| NrCAM | P35331 | AMNSFV | − |
| Peptide library | — | QQAFYV | + |
| Neurexin I | M96375 | DKEYYV | + |
| Neurexin II | M96376 | DKEYYV | + |
| Neurexin III | L14851 | DKEYYV | + |
| Jagged | L38483 | RMEYIV | + |
| Ephrine B1 | P98172 | NIYYKV | − |
| Ephrine B2 | P52799 | NIYYKV | − |
| Ephrine B3 | U62755 | NIYYKV | − |
LexA peptide fusions derived from the C termini of various transmembrane proteins were tested for their ability to interact with the PDZ domain of AF6 in two-hybrid assays. NMDAR, N-methyl-d-aspartate receptor; NrCAM, neuronal cell adhesion molecule.
An additional C-terminal motif (Q-X-A-Y/F-Y/F-V) suitable for binding of the PDZ domain of AF6 was suggested by L. Cantley (personal communication) according to an analysis using an oriented peptide library (27). This peptide containing aromatic residues (Y/F) at position −3 and −2 interacted also with AF6 in our two-hybrid analysis. On the basis of this observation and the hypothesis that amino acid positions −3 and −1 are critical for binding of PDZ domains, we searched the databank for proteins with the C-terminal sequence Y-X-V. The Notch ligand Jagged (31), neurexins I, II and III (32, 33), and the transmembrane ligands of Eph-related kinases (ephrine B1, B2, and B3), which were all found to contain this motif, were tested in two-hybrid assays. Only C-terminal peptides of neurexin I-III and Jagged interacted with PDZ-AF6 (Table 3). Taken together, the analysis of two peptide motifs T/S-X-V and Y-X-V revealed that within C-terminal hexapeptides in addition to the amino acids at positions −1 and −3 other residues contribute to the selectivity of the PDZ domain.
Interaction of the PDZ Domain of AF6 and EphB3 in Vitro.
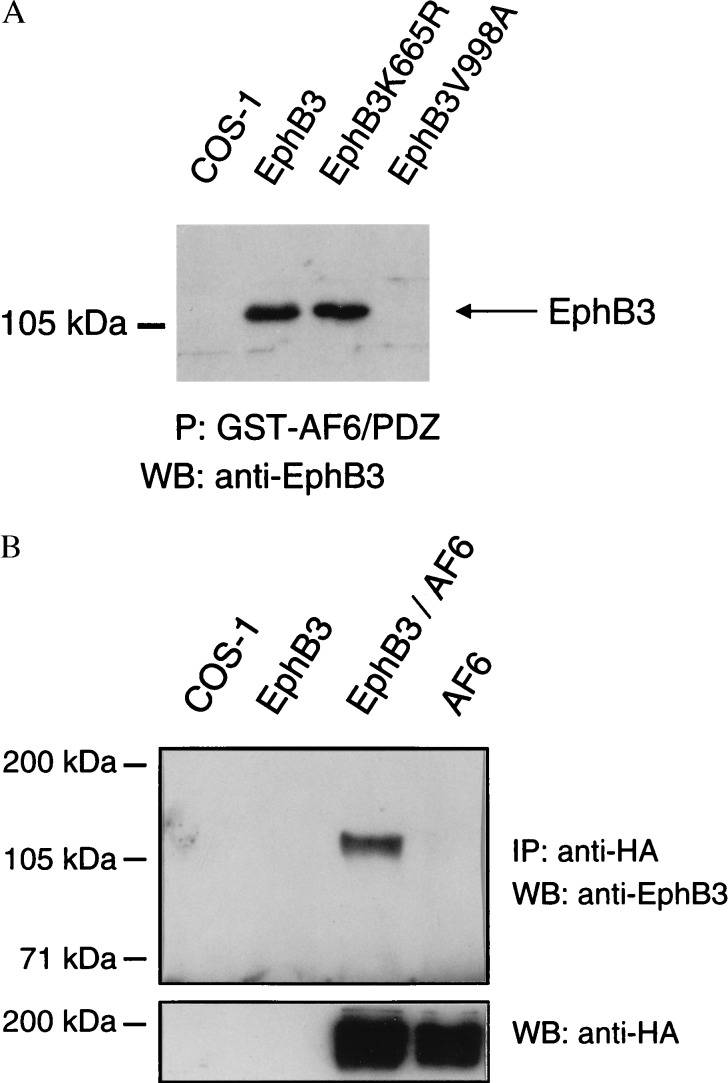
We expressed wild-type or different mutated versions of EphB3 in COS-1 cells and a GST-fusion of clone 31-44 containing the PDZ domain of AF6 (Fig. 1A) in E. coli to perform in vitro binding experiments. As shown in Fig. 2A, the AF6 fragment interacted with the wild-type receptor or kinase-dead EphB3K665R, whereas binding of the C-terminal point mutant EphB3V998A was not detectable. These data confirm the prediction based on two-hybrid assays that the PDZ domain of AF6 interacts with the C-terminal region of the Eph-related kinase EphB3 independent of its kinase activity.
Figure 2.
Association of EphB3 and the PDZ domain of AF-6 in vitro and in mammalian cells. (A) Wild-type or mutant EphB3 was expressed in COS-1 cells as indicated. Proteins that could be precipitated with the GST-AF6/PDZ fusion (P) were treated on a Western blot (WB) with EphB3 antibodies. (B) Lysates from untransfected, EphB3-transfected, EphB3- and HA-AF6-cotransfected, or HA-AF6-transfected COS-1 cells were immunoprecipitated (IP) with HA antibodies. Immunoprecipitates were probed on a Western blot (WB) with EphB3 antiserum, stripped, and reprobed with monoclonal HA antibodies.
Interaction of Full-Length AF6 with EphB3 in Mammalian Cells.
Our previous experiments have shown that overexpression of EphB3 under the transcriptional control of the cytomegalovirus enhancer-promoter causes a substantial level of receptor phosphorylation independent of ligand addition (5). Under these conditions the phosphorylation of the Y614F mutant is markedly diminished compared with the wild-type receptor. Although we cannot rule out that this mutant exhibits reduced kinase activity, we assume that Y614 is the major autophosphorylation site of EphB3 in overexpressing COS-1 cells (12).
Lysates of COS-1 cells transiently expressing HA-AF6 and EphB3 were immunoprecipitated with HA antibodies and subjected to a Western blot analysis using EphB3 antiserum to examine the interaction between EphB3 and full-length AF6 in mammalian cells. As shown in Fig. 2B, EphB3 could be coimmunoprecipitated only with HA antibodies from EphB3/HA-AF6-coexpressing cells, demonstrating that AF6 associates with EphB3 in mammalian cells.
AF6 Is a Substrate of the Kinase Activities of EphB2 and EphB3.
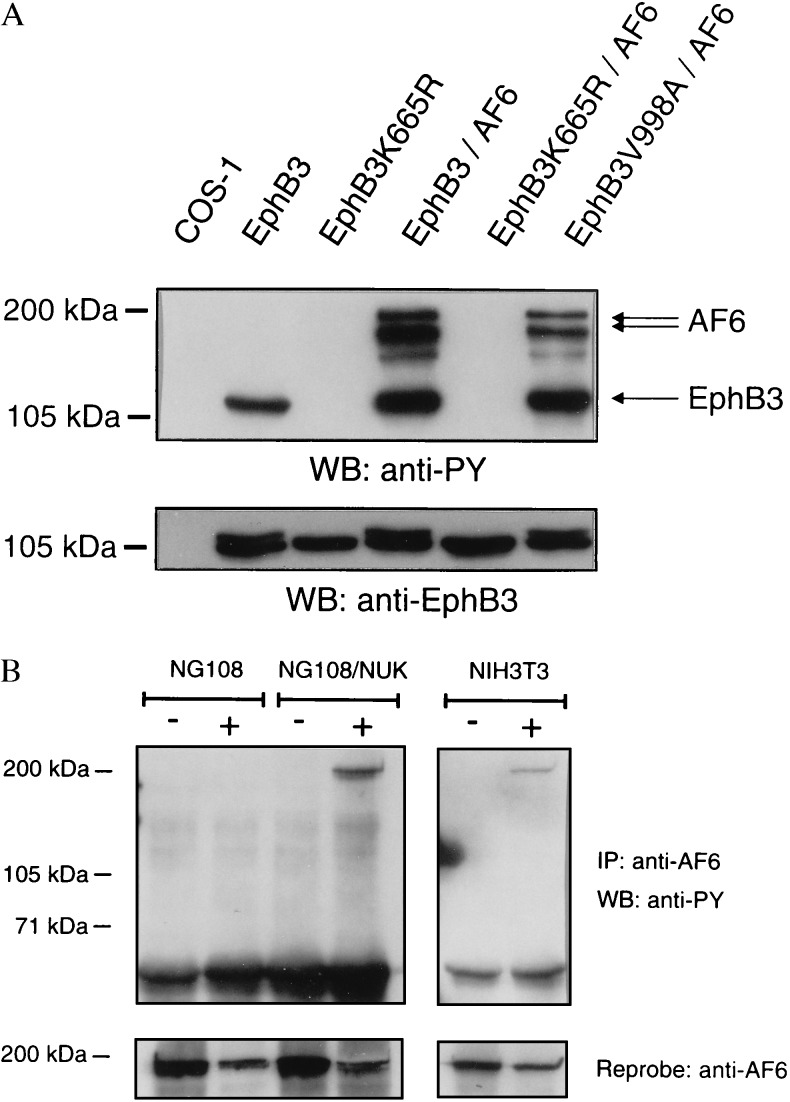
To determine whether AF6 is phosphorylated by EphB3 in mammalian cells, we cotransfected COS-1 cells with HA-AF6 in combination with either wild-type or mutated EphB3. Using phosphotyrosine-specific antibodies the Western blot analysis revealed that, in contrast to cells coexpressing kinase-dead EphB3K665R and HA-AF6, HA-AF6 is phosphorylated on tyrosine in cells coexpressing HA-AF6 and wild-type EphB3 (Fig. 3A). Interestingly, the C-terminal mutant EphB3V998A, which did not associate with the PDZ domain of AF6 in vitro (Fig. 2A) was able to phosphorylate HA-AF6 in COS-1 cells.
Figure 3.
Phosphorylation of full-length AF6 depends on the catalytic activity of the receptor. (A) COS-1 cells were cotransfected with various mutants of EphB3 and HA-AF6. Cell lysates were probed on a Western blot with phosphotyrosine antibodies. AF6 is recognized as multiple bands presumably due to degradation of the overexpressed protein. The EphB3 and HA-AF6 content of the lysates was compared on separate blots using HA (not shown) and EphB3 antisera (Lower). (B) NIH 3T3, NG108, or NG108 cells stably expressing EphB2 (NG108/NUK) were challenged with 2 μg/ml clustered ephrine-B1 Fc (+) or human Fc antibodies (Sigma) for 30 min (−). Subsequently, cells were scraped off, washed twice in PBS, and lysed in RIPA buffer. Cell lysates were immunoprecipitated (IP) with a polyclonal AF6 antiserum and probed on a Western blot (WB) using monoclonal phosphotyrosine antibodies (PY99, Santa Cruz Biotechnology), stripped, and reprobed with AF6 antibodies.
In addition to the experiment with COS-1 cells showing constitutive activation of EphB2, we analyzed NIH 3T3 cells that exhibited endogenous expression of AF-6 and EphB3 (6). In this case ephrine-B1 stimulation is required for receptor activation. Lysates of nonstimulated and ephrine-B1-stimulated NIH 3T3 cells were immunoprecipitated with an AF6 antiserum and probed on a Western blot with phosphotyrosine antibodies. Fig. 3B demonstrates that phosphorylation of AF6 depends on ephrine-B1 stimulation in NIH 3T3 cells.
In addition, NG108 cells stably transfected with EphB2 (13) were used to investigate whether AF6 is also a substrate for EphB2. The Western blot in Fig. 3B illustrates that AF6 is phosphorylated by EphB2 in a ligand-dependent manner.
The Interaction with Full-Length AF6 Correlates to the Kinase Activity of EphB3.
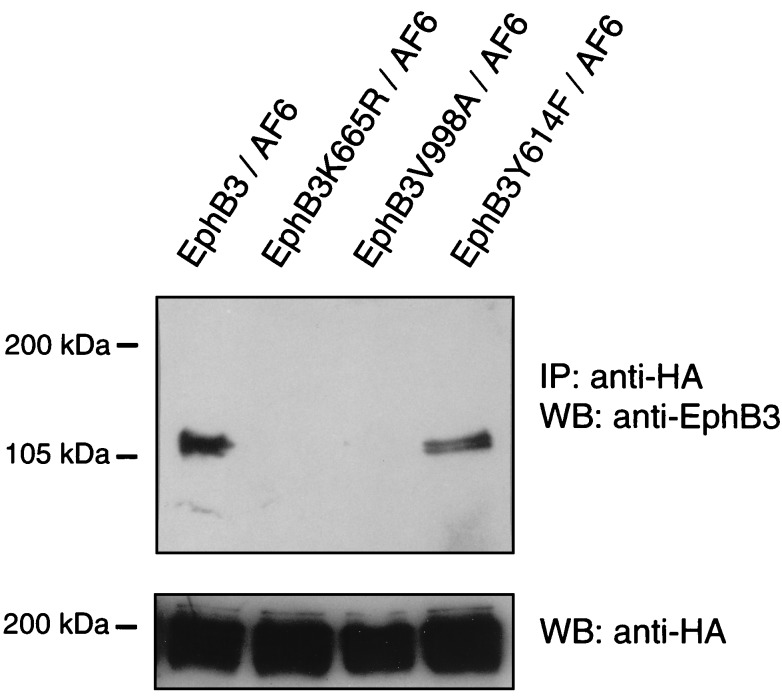
Because the C-terminal mutant EphB3V998A was able to phosphorylate HA-AF6, we analyzed the interaction in COS-1 cells in more detail using various mutants of EphB3. In agreement with results obtained from in vitro binding assays, wild-type EphB3 interacted strongly with full-length AF6, whereas the C-terminal mutant EphB3V998A did not (Fig. 4). Moreover, we analyzed whether the interaction of full-length AF6 with EphB3 in COS-1 cells depends on the catalytic activity of EphB3. Surprisingly, in anti-HA immunoprecipitates prepared from lysates of COS-1 cells coexpressing HA-AF6 and the kinase-dead mutant EphB3K665R we could not detect the kinase-deficient EphB3 (Fig. 4). These data indicate that the interaction of EphB3 with full-length AF6 in mammalian cells depends on the activation status of the receptor. Thus, the binding properties of the full-length protein differ clearly from the isolated PDZ domain of AF6 in vitro and from results obtained in the two-hybrid system.
Figure 4.
EphB3 binding to full-length AF6 depends on the catalytic activity of the receptor. COS-1 cells were cotransfected with various mutants of EphB3 and HA-AF6. Cell lysates were immunoprecipitated (IP) with HA antibodies and probed on a Western blot (WB) using EphB3 antisera, stripped, and reprobed with monoclonal HA antibodies.
The binding properties of full-length AF6 could be regulated either by the autophosphorylation of the receptor or by the EphB3-mediated phosphorylation of AF6. Since we found recently that Y614 in the EphB3 RTK serves as major autophosphorylation and multidocking site for SH2-domain-mediated interactions (12), we wondered whether this residue could be involved in the association with AF6. However, coimmunoprecipitation experiments showed that substitution of Y614 to phenylalanine did not abolish the binding capacity of EphB3 to AF6 in mammalian cells (Fig. 4).
EphB3 and AF6 Show Similar Patterns of Expression in Human Tissues.
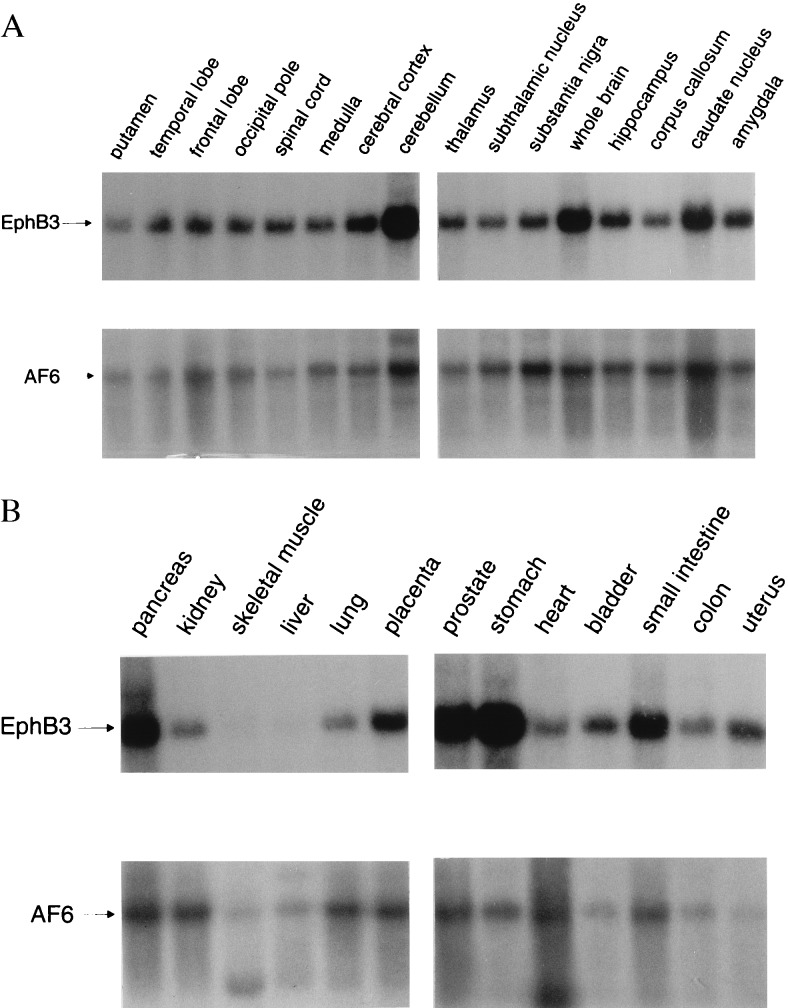
Previous studies have demonstrated a broad distribution of EphB2, EphB5, EphB6, and EphA7 in adult tissues, with most frequent expression in brain (19–21). Since the detailed expression patterns of EphB3 and AF6 were unknown, a Northern blot analysis using a broad spectrum of adult tissues was performed. While EphB3 is transcribed as an mRNA of 4.6 kb, the AF6 probe detected transcripts of 8 kb in all tissues analyzed and, at a lower frequency of expression, a message of 5 kb in certain tissues such as heart and skeletal muscle (Fig. 5). This analysis of adult tissues also reveals the overlapping mRNA expression of AF6 and different members of the family of Eph-related receptors, which have the capability to interact with the PDZ domain of AF6 as shown above. Interestingly, the dissection of human brain, a tissue with most abundant expression of Eph-related receptors, revealed coexpression of EphB3 and AF6 in all functional domains analyzed.
Figure 5.
Northern blot analysis of different neuronal (A) and nonneuronal (B) human tissues. Blots were probed with a 291-bp fragment of the AF6 cDNA encompassing the PDZ domain under high-stringency conditions. After exposure blots were stripped and reprobed with an EphB3-specific cDNA probe under the same conditions.
DISCUSSION
Binding Properties of the PDZ Domain of AF6.
Using the two-hybrid system and in vitro binding studies, we could demonstrate that the PDZ domain of AF6 binds to the Eph-related RTKs EphB3 and EphB2 independent of their catalytic activity. The interaction involves 6 amino acids at the C terminus of the receptors in a sequence-specific fashion. In addition, we describe the association of the PDZ domain with the C-terminal hexapeptides derived from the Eph-related RTKs EphA7, EphB5, and EphB6. The Eph-related kinases that bind to the PDZ domain of AF6 share a C-terminal core motif (small near neutral-V/I-E/Q-V) that seems to play a major role for the interaction.
Recently, it has been reported that PDZ domains can be grouped into two subsets on the basis of their binding properties: PDZ domains of group 1 select peptides with an E-S/T-X-V/I consensus motif, whereas PDZ domains of group 2, including AF6-PDZ, select peptides containing hydrophobic or aromatic amino acids at positions −1, −2, and −3 (27). In this report we found that binding of the PDZ domain of AF6 is not restricted to peptides containing hydrophobic or aromatic amino acids at positions −2 and −3: C-terminal hexapeptides of Eph-related receptors containing glutamine or glutamic acid at position −2 also have the capacity to bind the PDZ domain of AF6. In addition, an amino acid substitution at position −3 of EphB2 generating a S/T-X-V motif did not abolish binding. Thus, we asked whether the PDZ domain of AF6 could also recognize peptides that bind PDZ domains belonging to group 1. However, S/T-X-V-containing hexapeptides derived from Kv1.4 NMDA receptor, Nr2C and Nr2D subunits of Shaker-type potassium channels, as well as from FAP that associate with PDZ domains of group 1 (28–30, 34) did not interact with AF6-PDZ. Since mutated EphB2 and the analyzed potassium channels share the C-terminal S-E-V motif, the interaction is clearly influenced by amino acids at positions −4, −5, and −6. Thus, AF6-PDZ mediates highly specific interactions and does not crossreact with C termini of proteins recognized by PDZ domains of group 1.
Songyang et al. (27) suggested a consensus peptide (Q-X-A-Y/F-Y/F-V) for the binding to the PDZ domain of AF6, placing it into group 2 of PDZ domains. Using a hexapeptide derived from this consensus motif (QQAFYV), we could confirm this prediction in yeast cells. Moreover, we have shown binding of the PDZ domain of AF6 to C-terminal peptides corresponding to the Notch ligand Jagged and neurexins I, II, and III, which share the consensus motif for association with PDZ domains of group 2. Taken together, our data suggest that within C-terminal hexapeptides each amino acid contributes to the selectivity for specific PDZ domains.
Binding of EphB3 to Full-Length AF6.
In vitro binding studies confirmed our results from two-hybrid assays: The PDZ domain of AF6 binds to both wild-type EphB3 and kinase-deficient EphB3K665R, indicating that this interaction does not depend on the catalytic activity of the receptor. In contrast, the C-terminal mutated receptor EphB3V998A did not bind to the separated PDZ domain. However, full-length AF6 differs clearly in its binding behavior, because coimmunoprecipitations revealed that its association to EphB3 in mammalian cells is strictly dependent on the catalytic activity of the receptor: In contrast to the wild-type receptor, the kinase-dead mutant EphB3K665R did not interact with full-length AF6 in these experiments. Moreover, we could demonstrate that in NIH 3T3 and NG108 cells endogenous AF6 is phosphorylated specifically by EphB3 and EphB2 in a ligand-dependent fashion. Interestingly, in COS-1 cells AF-6 is also phosphorylated by the C-terminal mutant EphB3V998A, which does not bind to the PDZ domain. Thus, phosphorylation of AF6 does not depend on the PDZ-domain-mediated interaction.
The diverging binding properties of full-length AF6 compared with the PDZ domain alone may be due to conformational hindrance, because the PDZ domain as element of the full-length protein might be masked, requiring a two-step mechanism for binding: A conformational change of AF6 could be a prerequisite to release the PDZ domain for its interaction with EphB3. This unmasking of the PDZ domain could be achieved as follows: EphB3-mediated phosphorylation of AF6 that is independent of the PDZ-domain-mediated interaction could induce a structural change of the AF6 protein followed by unmasking of the PDZ domain. This hypothesis could be tested by mapping and subsequent mutation of the phosphorylation sites in AF6. However, we cannot rule out that minor phosphorylation sites of EphB3 could contribute to the observed interaction. To answer this question all autophosphorylation sites of the receptor need to mapped and mutated.
Recently, it has been shown that AF6 binds to activated ras in vitro (35). The interaction occurs at the N terminus of the AF6 protein and is mediated by a sequence motif that is conserved among various proteins (36). Since activated EphB3 interacts with rasGAP (12), AF6 might locate ras to EphB3-bound rasGAP, resulting in a modulation of ras-mediated signaling. We found that the PDZ domain of AF6 is able to interact with a variety of transmembrane proteins. Therefore, it could function as a scaffold protein that locates receptors to specific regions within the cytoplasmic membrane or links them to defined signal transduction pathways.
Acknowledgments
We are grateful to Stanley Fields and Paul Bartel for providing the plasmid pBTM116. The plasmid pVP16 and the two-hybrid cDNA library were generous gifts from Stanley Hollenberg. Yeast strain L40 was kindly provided by Rolf Sternglanz and the ura− derivative, by Stanley Hollenberg. The Georg-Speyer-Haus is supported by the Bundesgesundheitsministerium and the Hessisches Ministerium für Wissenschaft und Kunst. This work was further supported by grants from the Deutsche Forschungsgemeinschaft (RU 242/11-1, STR 336/5-1) and the Sonderforschungsbereich 474.
Footnotes
This paper was submitted directly (Track II) to the Proceedings Office.
Abbrevations: RTK, receptor tyrosine kinase; GST, glutathione S-transferase; HA, influenza virus hemagglutinin epitope.
References
- 1.van der Geer P, Hunter T, Lindberg R A. Annu Rev Cell Biol. 1994;10:251–337. doi: 10.1146/annurev.cb.10.110194.001343. [DOI] [PubMed] [Google Scholar]
- 2.Pawson T. Nature (London) 1995;373:573–580. doi: 10.1038/373573a0. [DOI] [PubMed] [Google Scholar]
- 3.Böhme B, Holtrich U, Wolf G, Luzius H, Grzeschik K H, Strebhardt K, Rübsamen-Waigmann H. Oncogene. 1993;8:2857–2862. [PubMed] [Google Scholar]
- 4.Eph Nomenclature Committee. Cell. 1997;90:403–404. doi: 10.1016/s0092-8674(00)80500-0. [DOI] [PubMed] [Google Scholar]
- 5.Böhme B, VandenBos T, Cerretti D P, Park L S, Holtrich U, Rübsamen-Waigmann H, Strebhardt K. J Biol Chem. 1996;271:24747–24752. doi: 10.1074/jbc.271.40.24747. [DOI] [PubMed] [Google Scholar]
- 6.Gale N W, Holland S J, Valenzuela D M, Flenniken A, Pan L, Henkemeyer M, Strebhardt K, Hirai H, Wilkinson D G, Pawson T, Davis S, Yancopoulos G D. Neuron. 1996;17:9–19. doi: 10.1016/s0896-6273(00)80276-7. [DOI] [PubMed] [Google Scholar]
- 7.Orioli D, Henkemeyer M, Lemke G, Klein R, Pawson T. EMBO J. 1996;15:6035–6049. [PMC free article] [PubMed] [Google Scholar]
- 8.Pandey A, Lazar D F, Saltiel A, Dixit V M. J Biol Chem. 1994;269:30154–30157. [PubMed] [Google Scholar]
- 9.Pandey A, Duan H, Dixit V M. J Biol Chem. 1995;270:19201–19204. doi: 10.1074/jbc.270.33.19201. [DOI] [PubMed] [Google Scholar]
- 10.Stein E, Cerretti D P, Daniel T O. J Biol Chem. 1996;271:23588–23593. doi: 10.1074/jbc.271.38.23588. [DOI] [PubMed] [Google Scholar]
- 11.Ellis C, Kasmi F, Ganju P, Walls E, Panayotou G, Reith A. Oncogene. 1996;12:1727–1736. [PubMed] [Google Scholar]
- 12.Hock B, Böhme B, Karn T, Feller S, Rübsamen-Waigmann H, Strebhardt K. Oncogene. 1998;17:255–260. doi: 10.1038/sj.onc.1201907. [DOI] [PubMed] [Google Scholar]
- 13.Holland S J, Gale N W, Gish G D, Roth R A, Songyang Z, Cantley L C, Henkemeyer M, Yancopoulos G D, Pawson T. EMBO J. 1997;16:3877–3888. doi: 10.1093/emboj/16.13.3877. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Mizushima S, Nagata S. Nucleic Acids Res. 1990;18:5322. doi: 10.1093/nar/18.17.5322. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Hollenberg S M, Sternglanz R, Cheng P F, Weintraub H. Mol Cell Biol. 1995;15:3813–1822. doi: 10.1128/mcb.15.7.3813. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Karn T, Hock B, Holtrich U, Adamski M, Strebhardt K, Rübsamen-Waigmann H. Virology. 1998;246:45–52. doi: 10.1006/viro.1998.9157. [DOI] [PubMed] [Google Scholar]
- 17.Vojtek A B, Hollenberg S M, Cooper J A. Cell. 1993;74:205–214. doi: 10.1016/0092-8674(93)90307-c. [DOI] [PubMed] [Google Scholar]
- 18.Prasad R, Gu Y, Alder H, Nakamura T, Canaani O, Saito H, Huebner K, Gale R P, Nowell P C, Kuriyama K, Miyazaki Y, Croce C M, Canaani E. Cancer Res. 1993;53:5624–5628. [PubMed] [Google Scholar]
- 19.Fox G M, Holst P L, Chute H T, Lindberg R A, Janssen A M, Basu R, Welcher A W. Oncogene. 1995;10:897–905. [PubMed] [Google Scholar]
- 20.Sajjadi F G, Pasquale E B. Oncogene. 1993;8:1808–1813. [PubMed] [Google Scholar]
- 21.Gurniak C B, Berg L. Oncogene. 1996;13:777–786. [PubMed] [Google Scholar]
- 22.Lindberg R A, Hunter T. Mol Cell Biol. 1990;10:6316–6324. doi: 10.1128/mcb.10.12.6316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Wicks I P, Wilkinson D, Salvaris E, Boyd A W. Proc Natl Acad Sci USA. 1992;89:1611–1615. doi: 10.1073/pnas.89.5.1611. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Maisonpierre P C, Barrezueta N X, Yancopoulos G D. Oncogene. 1993;8:3277–3288. [PubMed] [Google Scholar]
- 25.Ganju P, Walls E, Brennan J, Reith A D. Oncogene. 1995;11:281–290. [PubMed] [Google Scholar]
- 26.Valenzuela D M, Stitt T N, DiStefano P S, Rojas E, Mattsson K, Compton D L, Nunez L, Park J S, Stark J L, Gies D R. Neuron. 1995;15:573–584. doi: 10.1016/0896-6273(95)90146-9. [DOI] [PubMed] [Google Scholar]
- 27.Songyang Z, Fanning A S, Fu C, Xu J, Marfatia S M, Chishti A H, Crompton A, Chan A C, Anderson J M, Cantley L C. Science. 1997;275:73–77. doi: 10.1126/science.275.5296.73. [DOI] [PubMed] [Google Scholar]
- 28.Sato T, Irie S, Kitada S, Reed J C. Science. 1996;268:411–414. doi: 10.1126/science.7536343. [DOI] [PubMed] [Google Scholar]
- 29.Kim E, Niethammer M, Rothschild A, Jan Y N, Sheng M. Nature (London) 1995;378:85–88. doi: 10.1038/378085a0. [DOI] [PubMed] [Google Scholar]
- 30.Niethammer M, Kim E, Sheng M. J Neurosci. 1995;16:2157–2163. doi: 10.1523/JNEUROSCI.16-07-02157.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Lindsell C E, Shawber C J, Boulter J, Weinmaster G D. Cell. 1995;80:907–917. doi: 10.1016/0092-8674(95)90294-5. [DOI] [PubMed] [Google Scholar]
- 32.Ushkaryov Y A, Petrenko A G, Geppert M, Südhof T C. Science. 1992;257:50–56. doi: 10.1126/science.1621094. [DOI] [PubMed] [Google Scholar]
- 33.Ushkaryov Y A, Südhof T C. Proc Natl Acad Sci USA. 1993;90:6410–6414. doi: 10.1073/pnas.90.14.6410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Kornau H-C, Schenker L T, Kennedy M B, Seeburg P H. Science. 1995;269:1737–1740. doi: 10.1126/science.7569905. [DOI] [PubMed] [Google Scholar]
- 35.Kuriyama M, Harada N, Kuroda S, Yamamoto T, Nakafuku M, Iwamatsu A, Prasad R, Croce C, Canaani E, Kaibuchi K. J Biol Chem. 1995;271:607–610. doi: 10.1074/jbc.271.2.607. [DOI] [PubMed] [Google Scholar]
- 36.Ponting C P, Benjamin D R. Trends Biochem Sci. 1996;21:422–424. doi: 10.1016/s0968-0004(96)30038-8. [DOI] [PubMed] [Google Scholar]