Abstract
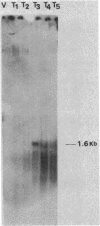
The DNA sequence of the structural gene for glucose dehydrogenase (EC 1.1.1.47) of Bacillus subtilis was determined and comprises 780 base pairs. The subunit molecular weight of glucose dehydrogenase as deduced from the nucleotide sequence is 28,196, which agrees well with the subunit molecular weight of 31,500 as determined from sodium dodecyl sulfate-polyacrylamide gel electrophoresis. The sequence of the 49 amino acids at the NH2 terminus of glucose dehydrogenase purified from sporulating B. subtilis cells matched the amino acid sequence derived from the DNA sequence. Glucose dehydrogenase was purified from an Escherichia coli strain harboring pEF1, a plasmid that contains the B. subtilis gene encoding glucose dehydrogenase. This enzyme has the identical amino acid sequence at the NH2 terminus as the B. subtilis enzyme. A putative ribosome-binding site, 5'-AGGAGG-3', which is complementary to the 3' end of the 16S rRNA of B. subtilis, was found 6 base pairs preceding the translational start codon of the structural gene of glucose dehydrogenase. No known promoterlike DNA sequences that are recognized by B. subtilis RNA polymerases were present immediately preceding the translational start site of the glucose dehydrogenase structural gene. The glucose dehydrogenase gene was found to be under sporulation control at the trancriptional level. A transcript of 1.6 kilobases hybridized to a DNA fragment within the structural gene of glucose dehydrogenase. This transcript was synthesized 3 h after the cessation of vegetative growth concomitant to the appearance of glucose dehydrogenase.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bradford M. M. A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem. 1976 May 7;72:248–254. doi: 10.1016/0003-2697(76)90527-3. [DOI] [PubMed] [Google Scholar]
- Chaudhry G. R., Halpern Y. S., Saunders C., Vasantha N., Schmidt B. J., Freese E. Mapping of the glucose dehydrogenase gene in Bacillus subtilis. J Bacteriol. 1984 Nov;160(2):607–611. doi: 10.1128/jb.160.2.607-611.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferrari E., Henner D. J., Hoch J. A. Isolation of Bacillus subtilis genes from a charon 4A library. J Bacteriol. 1981 Apr;146(1):430–432. doi: 10.1128/jb.146.1.430-432.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ferrari F. A., Trach K., Hoch J. A. Sequence analysis of the spo0B locus reveals a polycistronic transcription unit. J Bacteriol. 1985 Feb;161(2):556–562. doi: 10.1128/jb.161.2.556-562.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fort P., Errington J. Nucleotide sequence and complementation analysis of a polycistronic sporulation operon, spoVA, in Bacillus subtilis. J Gen Microbiol. 1985 May;131(5):1091–1105. doi: 10.1099/00221287-131-5-1091. [DOI] [PubMed] [Google Scholar]
- Fujita Y., Ramaley R., Freese E. Location and properties of glucose dehydrogenase in sporulating cells and spores of Bacillus subtilis. J Bacteriol. 1977 Oct;132(1):282–293. doi: 10.1128/jb.132.1.282-293.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hunkapiller M. W., Hood L. E. Direct microsequence analysis of polypeptides using an improved sequenator, a nonprotein carrier (polybrene), and high pressure liquid chromatography. Biochemistry. 1978 May 30;17(11):2124–2133. doi: 10.1021/bi00604a016. [DOI] [PubMed] [Google Scholar]
- Jörnvall H., von Bahr-Lindström H., Jany K. D., Ulmer W., Fröschle M. Extended superfamily of short alcohol-polyol-sugar dehydrogenases: structural similarities between glucose and ribitol dehydrogenases. FEBS Lett. 1984 Jan 9;165(2):190–196. doi: 10.1016/0014-5793(84)80167-2. [DOI] [PubMed] [Google Scholar]
- Lepesant-Kejzlarová J., Lepesant J. A., Walle J., Billault A., Dedonder R. Revision of the linkage map of Bacillus subtilis 168: indications for circularity of the chromosome. J Bacteriol. 1975 Mar;121(3):823–834. doi: 10.1128/jb.121.3.823-834.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Losick R., Pero J. Cascades of Sigma factors. Cell. 1981 Sep;25(3):582–584. doi: 10.1016/0092-8674(81)90164-1. [DOI] [PubMed] [Google Scholar]
- McLaughlin J. R., Murray C. L., Rabinowitz J. C. Unique features in the ribosome binding site sequence of the gram-positive Staphylococcus aureus beta-lactamase gene. J Biol Chem. 1981 Nov 10;256(21):11283–11291. [PubMed] [Google Scholar]
- Messing J., Vieira J. A new pair of M13 vectors for selecting either DNA strand of double-digest restriction fragments. Gene. 1982 Oct;19(3):269–276. doi: 10.1016/0378-1119(82)90016-6. [DOI] [PubMed] [Google Scholar]
- Piggot P. J., Coote J. G. Genetic aspects of bacterial endospore formation. Bacteriol Rev. 1976 Dec;40(4):908–962. doi: 10.1128/br.40.4.908-962.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Piggot P. J., Curtis C. A., de Lencastre H. Use of integrational plasmid vectors to demonstrate the polycistronic nature of a transcriptional unit (spoIIA) required for sporulation of Bacillus subtilis. J Gen Microbiol. 1984 Aug;130(8):2123–2136. doi: 10.1099/00221287-130-8-2123. [DOI] [PubMed] [Google Scholar]
- Prasad C., Diesterhaft M., Freese E. Initiation of spore germination in glycolytic mutants of Bacillus subtilis. J Bacteriol. 1972 Apr;110(1):321–328. doi: 10.1128/jb.110.1.321-328.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Price C. W., Doi R. H. Genetic mapping of rpoD implicates the major sigma factor of Bacillus subtilis RNA polymerase in sporulation initiation. Mol Gen Genet. 1985;201(1):88–95. doi: 10.1007/BF00397991. [DOI] [PubMed] [Google Scholar]
- Ramaley R. F., Vasantha N. Glycerol protection and purification of Bacillus subtilis glucose dehydrogenase. J Biol Chem. 1983 Oct 25;258(20):12558–12565. [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Setlow B., Setlow P. Levels of oxidized and reduced pyridine nucleotides in dormant spores and during growth, sporulation, and spore germination of Bacillus megaterium. J Bacteriol. 1977 Feb;129(2):857–865. doi: 10.1128/jb.129.2.857-865.1977. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shine J., Dalgarno L. The 3'-terminal sequence of Escherichia coli 16S ribosomal RNA: complementarity to nonsense triplets and ribosome binding sites. Proc Natl Acad Sci U S A. 1974 Apr;71(4):1342–1346. doi: 10.1073/pnas.71.4.1342. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Silverman S. J., Rose M., Botstein D., Fink G. R. Regulation of HIS4-lacZ fusions in Saccharomyces cerevisiae. Mol Cell Biol. 1982 Oct;2(10):1212–1219. doi: 10.1128/mcb.2.10.1212. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tinoco I., Jr, Borer P. N., Dengler B., Levin M. D., Uhlenbeck O. C., Crothers D. M., Bralla J. Improved estimation of secondary structure in ribonucleic acids. Nat New Biol. 1973 Nov 14;246(150):40–41. doi: 10.1038/newbio246040a0. [DOI] [PubMed] [Google Scholar]
- Vasantha N., Freese E. Enzyme changes during Bacillus subtilis sporulation caused by deprivation of guanine nucleotides. J Bacteriol. 1980 Dec;144(3):1119–1125. doi: 10.1128/jb.144.3.1119-1125.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vasantha N., Uratani B., Ramaley R. F., Freese E. Isolation of a developmental gene of Bacillus subtilis and its expression in Escherichia coli. Proc Natl Acad Sci U S A. 1983 Feb;80(3):785–789. doi: 10.1073/pnas.80.3.785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wax R., Freese E. Initiation of the germination of Bacillus subtilis spores by a combination of compounds in place of L-alanine. J Bacteriol. 1968 Feb;95(2):433–438. doi: 10.1128/jb.95.2.433-438.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wittmann-Liebold B., Graffunder H., Kohls H. A device coupled to a modified sequenator for the automated conversion of anilinothiazolinones into PTH amino acids. Anal Biochem. 1976 Oct;75(2):621–633. doi: 10.1016/0003-2697(76)90117-2. [DOI] [PubMed] [Google Scholar]
- Zimmerman C. L., Appella E., Pisano J. J. Rapid analysis of amino acid phenylthiohydantoins by high-performance liquid chromatography. Anal Biochem. 1977 Feb;77(2):569–573. doi: 10.1016/0003-2697(77)90276-7. [DOI] [PubMed] [Google Scholar]