Abstract
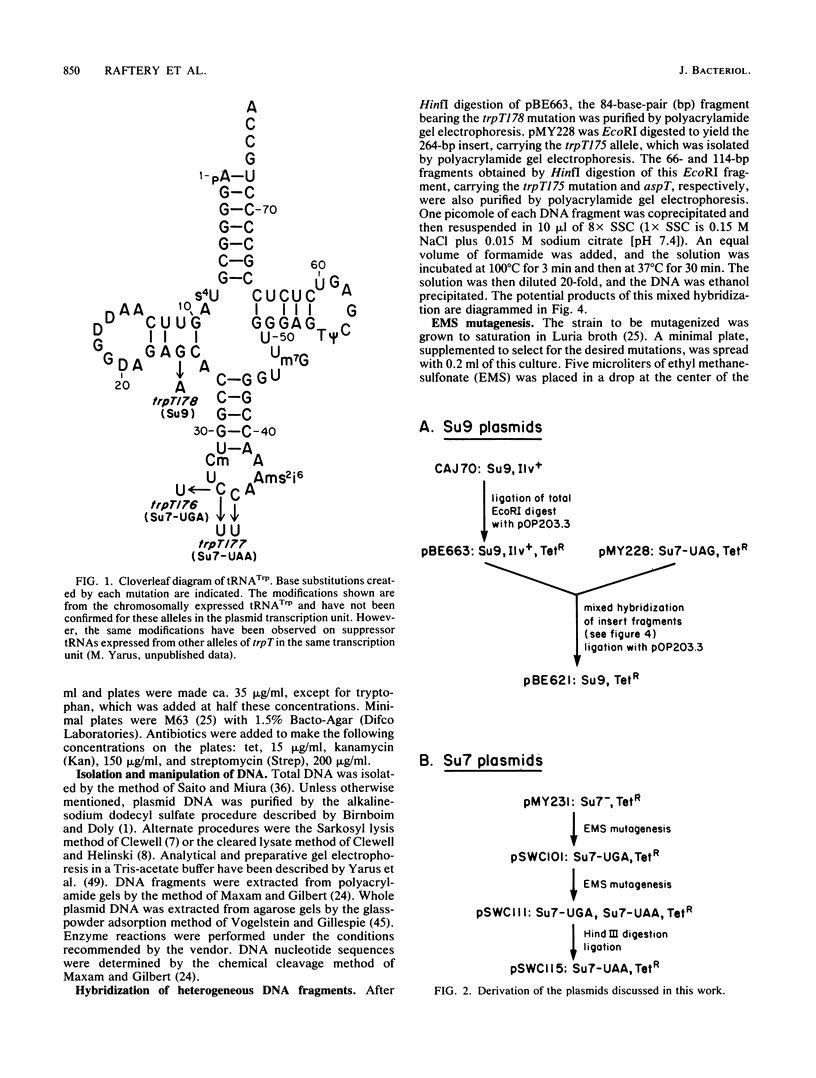
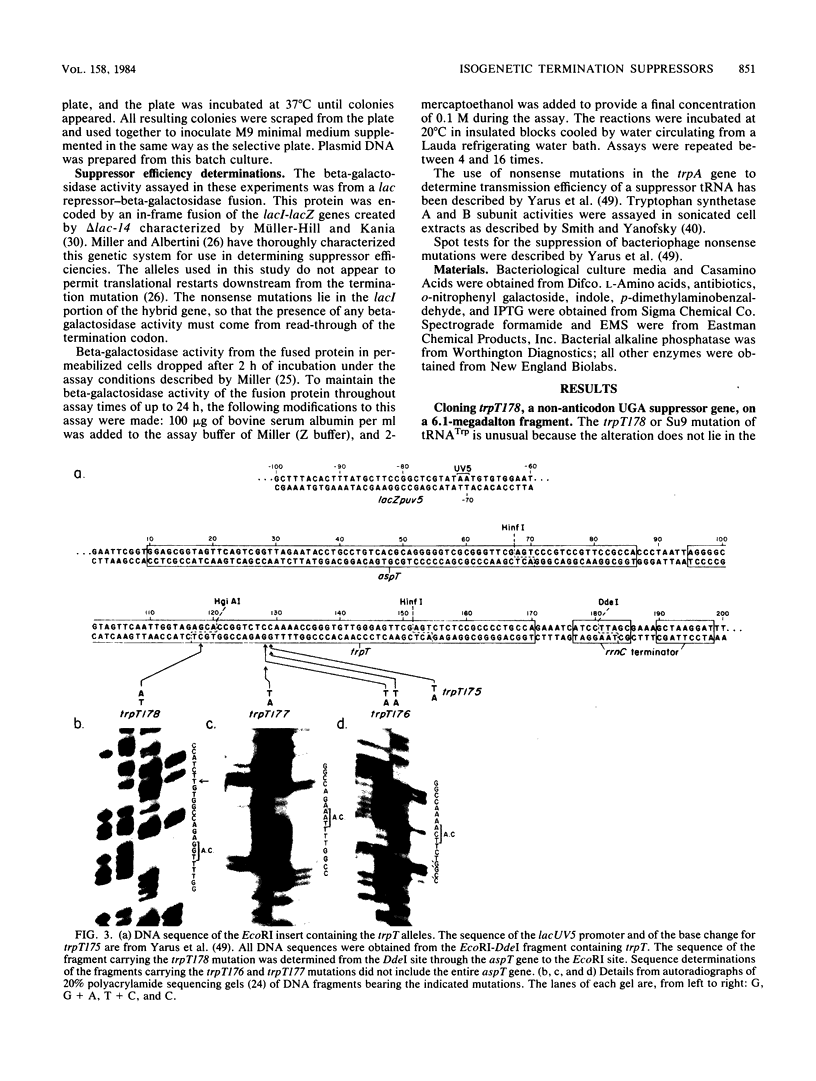
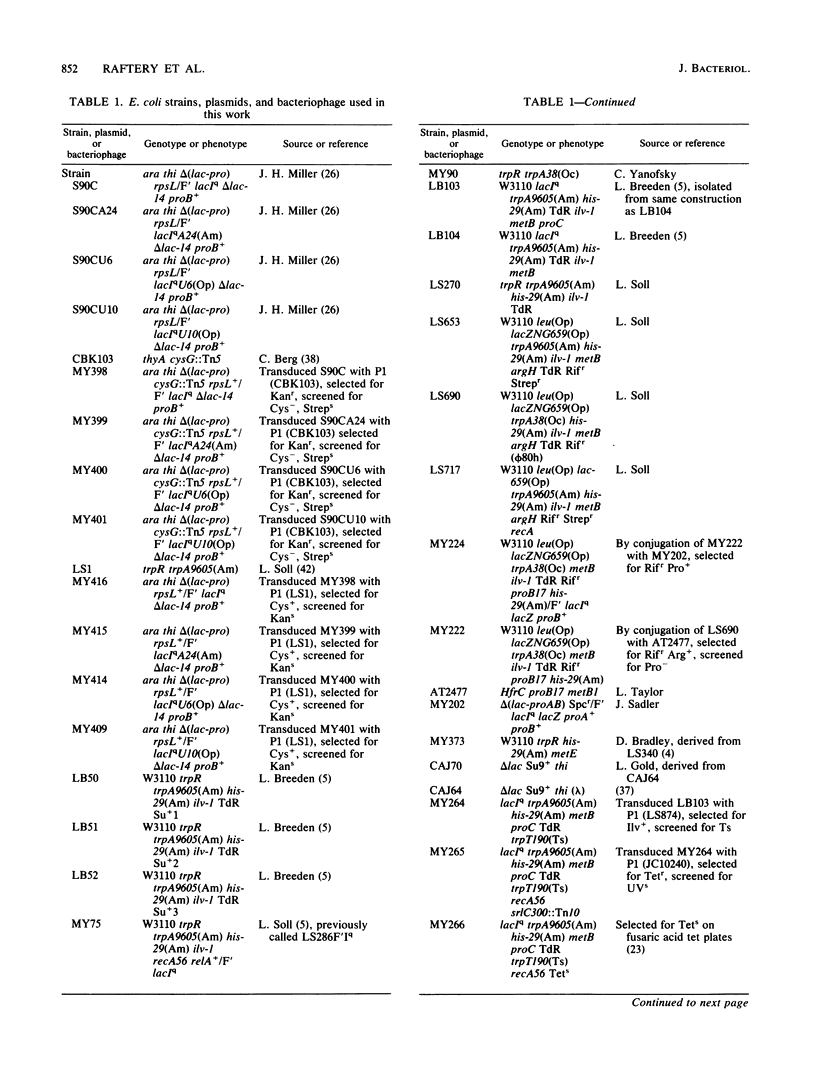
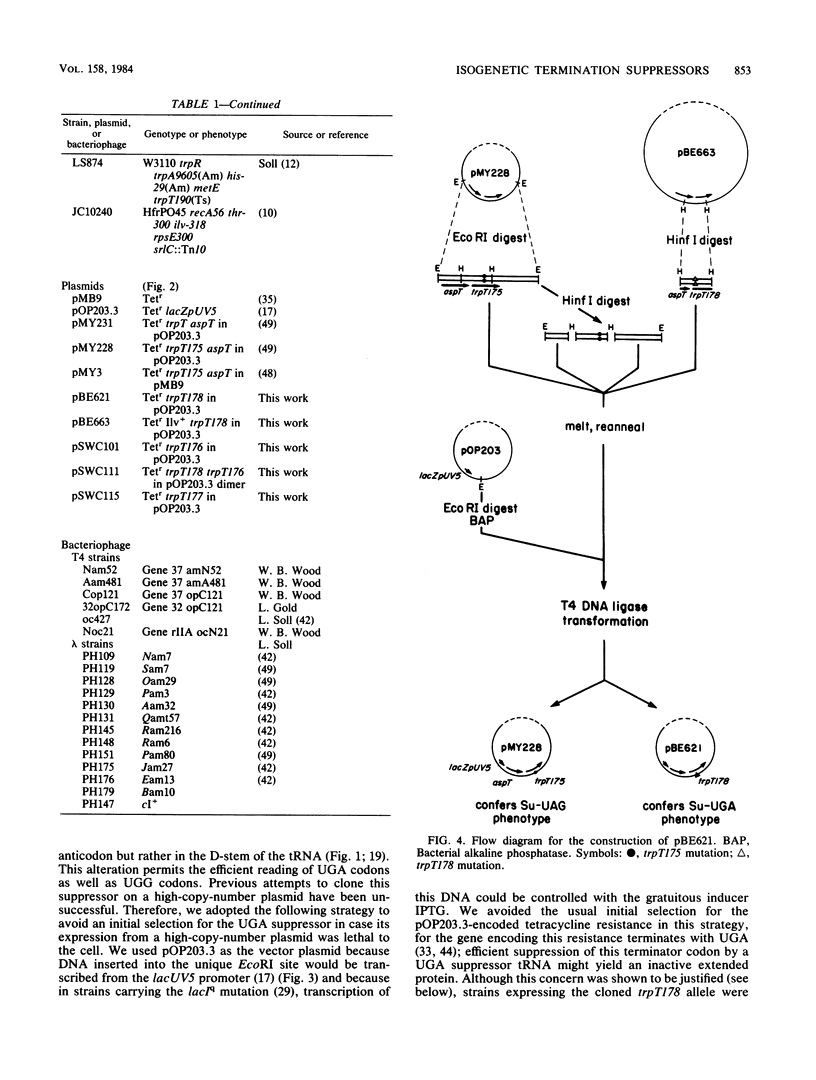
We have cloned an isogenetic set of UAG, UAA, and UGA suppressors. These include the Su7 -UAG, Su7 -UAA, and Su7 -UGA suppressors derived from base substitutions in the anticodon of Escherichia coli tRNATrp and also Su9 , a UGA suppressor derived from a base substitution in the D-arm of the same tRNA. These genes are cloned on high-copy-number plasmids under lac promoter control. The construction of the Su7 -UAG plasmid and the wild-type trpT plasmid have been previously described ( Yarus , et al., Proc. Natl. Acad. Sci. U.S.A. 77:5092-5097, 1980). Su7 -UAA ( trpT177 ) is a weak suppressor which recognizes both UAA and UAG nonsense codons and probably inserts glutamine. Su7 -UGA ( trpT176 ) is a strong UGA suppressor which may insert tryptophan. Su9 ( trpT178 ) is a moderately strong UGA suppressor which also recognizes UGG (Trp) codons, and it inserts tryptophan. The construction of these plasmids is detailed within. Data on the DNA sequences of these trpT alleles and on amino acid specificity of the suppressors are presented. The efficiency of the cloned suppressors at certain nonsense mutations has been measured and is discussed with respect to the context of these codons.
Full text
PDF


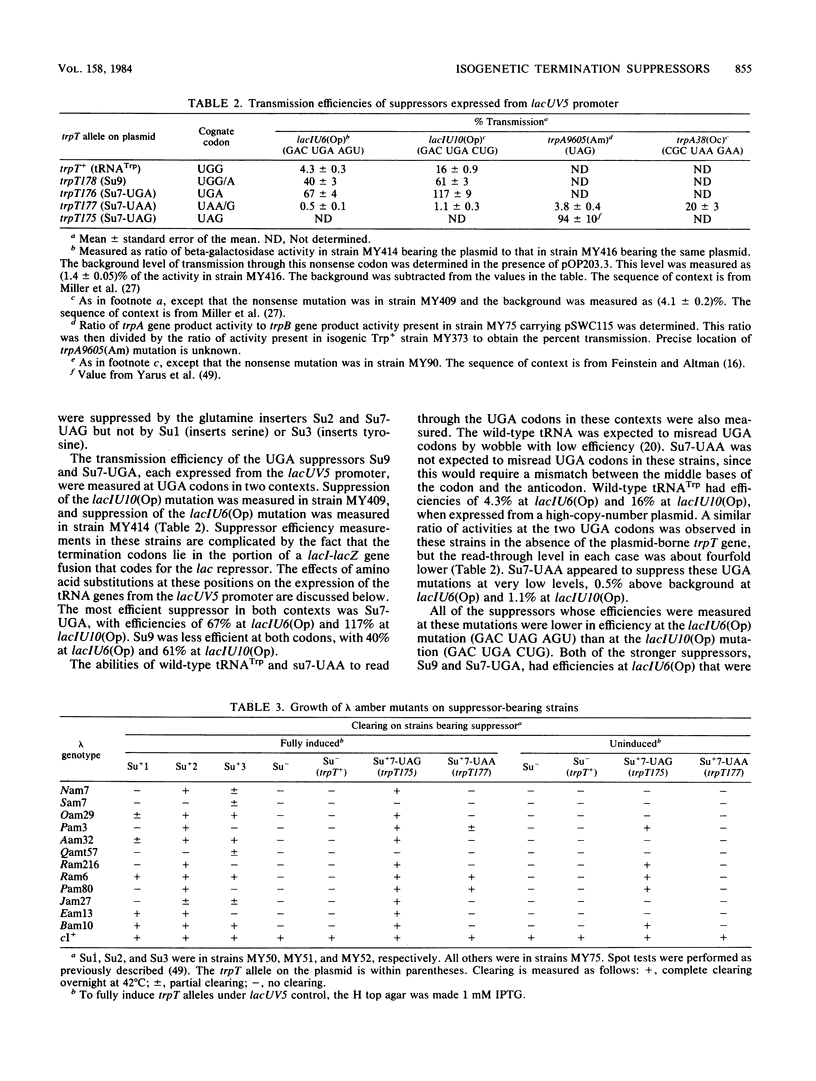




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bossi L. Context effects: translation of UAG codon by suppressor tRNA is affected by the sequence following UAG in the message. J Mol Biol. 1983 Feb 15;164(1):73–87. doi: 10.1016/0022-2836(83)90088-8. [DOI] [PubMed] [Google Scholar]
- Bossi L., Ruth J. R. The influence of codon context on genetic code translation. Nature. 1980 Jul 10;286(5769):123–127. doi: 10.1038/286123a0. [DOI] [PubMed] [Google Scholar]
- Bradley D., Park J. V., Soll L. TRNA2Gln Su+2 mutants that increase amber suppression. J Bacteriol. 1981 Feb;145(2):704–712. doi: 10.1128/jb.145.2.704-712.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Breeden L., Yarus M., Cline S. A cloned suppressor tRNA gene relaxes stringent control. Mol Gen Genet. 1980;179(1):125–133. doi: 10.1007/BF00268454. [DOI] [PubMed] [Google Scholar]
- Buckingham R. H., Kurland C. G. Codon specificity of UGA suppressor tRNATrp from Escherichia coli. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5496–5498. doi: 10.1073/pnas.74.12.5496. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clewell D. B., Helinski D. R. Supercoiled circular DNA-protein complex in Escherichia coli: purification and induced conversion to an opern circular DNA form. Proc Natl Acad Sci U S A. 1969 Apr;62(4):1159–1166. doi: 10.1073/pnas.62.4.1159. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clewell D. B. Nature of Col E 1 plasmid replication in Escherichia coli in the presence of the chloramphenicol. J Bacteriol. 1972 May;110(2):667–676. doi: 10.1128/jb.110.2.667-676.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crick F. H. Codon--anticodon pairing: the wobble hypothesis. J Mol Biol. 1966 Aug;19(2):548–555. doi: 10.1016/s0022-2836(66)80022-0. [DOI] [PubMed] [Google Scholar]
- Csonka L. N., Clark A. J. Construction of an Hfr strain useful for transferring recA mutations between Escherichia coli strains. J Bacteriol. 1980 Jul;143(1):529–530. doi: 10.1128/jb.143.1.529-530.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dagert M., Ehrlich S. D. Prolonged incubation in calcium chloride improves the competence of Escherichia coli cells. Gene. 1979 May;6(1):23–28. doi: 10.1016/0378-1119(79)90082-9. [DOI] [PubMed] [Google Scholar]
- Eisenberg S. P., Soll L., Yarus M. The purification and sequence of a temperature-sensitive tryptophan tRNA. J Biol Chem. 1979 Jun 25;254(12):5562–5566. [PubMed] [Google Scholar]
- Eisenberg S. P., Yarus M. The structure and aminoacylation of a temperature-sensitive tRNATrp (Escherichia coli). J Biol Chem. 1980 Feb 10;255(3):1128–1137. [PubMed] [Google Scholar]
- Engelberg-Kulka H. UGA suppression by normal tRNA Trp in Escherichia coli: codon context effects. Nucleic Acids Res. 1981 Feb 25;9(4):983–991. doi: 10.1093/nar/9.4.983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinstein S. I., Altman S. Coding properties of an ochre-suppressing derivative of Escherichia coli tRNAITyr. J Mol Biol. 1977 May 25;112(3):453–470. doi: 10.1016/s0022-2836(77)80192-7. [DOI] [PubMed] [Google Scholar]
- Feinstein S. I., Altman S. Context effects on nonsense codon suppression in Escherichia coli. Genetics. 1978 Feb;88(2):201–219. doi: 10.1093/genetics/88.2.201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fuller F. A family of cloning vectors containing the lacUV5 promoter. Gene. 1982 Jul-Aug;19(1):43–54. doi: 10.1016/0378-1119(82)90187-1. [DOI] [PubMed] [Google Scholar]
- Hirsh D., Gold L. Translation of the UGA triplet in vitro by tryptophan transfer RNA's. J Mol Biol. 1971 Jun 14;58(2):459–468. doi: 10.1016/0022-2836(71)90363-9. [DOI] [PubMed] [Google Scholar]
- Hirsh D. Tryptophan transfer RNA as the UGA suppressor. J Mol Biol. 1971 Jun 14;58(2):439–458. doi: 10.1016/0022-2836(71)90362-7. [DOI] [PubMed] [Google Scholar]
- Knowlton R. G., Soll L., Yarus M. Dual specificity of su+ 7 tRNA. Evidence for translational discrimination. J Mol Biol. 1980 Jun 5;139(4):705–720. doi: 10.1016/0022-2836(80)90056-x. [DOI] [PubMed] [Google Scholar]
- LENNOX E. S. Transduction of linked genetic characters of the host by bacteriophage P1. Virology. 1955 Jul;1(2):190–206. doi: 10.1016/0042-6822(55)90016-7. [DOI] [PubMed] [Google Scholar]
- Maloy S. R., Nunn W. D. Selection for loss of tetracycline resistance by Escherichia coli. J Bacteriol. 1981 Feb;145(2):1110–1111. doi: 10.1128/jb.145.2.1110-1111.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Miller J. H., Albertini A. M. Effects of surrounding sequence on the suppression of nonsense codons. J Mol Biol. 1983 Feb 15;164(1):59–71. doi: 10.1016/0022-2836(83)90087-6. [DOI] [PubMed] [Google Scholar]
- Miller J. H., Coulondre C., Farabaugh P. J. Correlation of nonsense sites in the lacI gene with specific codons in the nucleotide sequence. Nature. 1978 Aug 24;274(5673):770–775. doi: 10.1038/274770a0. [DOI] [PubMed] [Google Scholar]
- Miller J. H., Coulondre C., Hofer M., Schmeissner U., Sommer H., Schmitz A., Lu P. Genetic studies of the lac repressor. IX. Generation of altered proteins by the suppression of nonsence mutations. J Mol Biol. 1979 Jun 25;131(2):191–222. doi: 10.1016/0022-2836(79)90073-1. [DOI] [PubMed] [Google Scholar]
- Müller-Hill B., Crapo L., Gilbert W. Mutants that make more lac repressor. Proc Natl Acad Sci U S A. 1968 Apr;59(4):1259–1264. doi: 10.1073/pnas.59.4.1259. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Müller-Hill B., Kania J. Lac repressor can be fused to beta-galactosidase. Nature. 1974 Jun 7;249(457):561–563. doi: 10.1038/249561a0. [DOI] [PubMed] [Google Scholar]
- Neidhardt F. C., Bloch P. L., Smith D. F. Culture medium for enterobacteria. J Bacteriol. 1974 Sep;119(3):736–747. doi: 10.1128/jb.119.3.736-747.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pannekoek H., Maat J., van den Berg E., Noordermeer I. Structure of a promotor on plasmid pMB9 derived from plasmid pSC101. Nucleic Acids Res. 1980 Apr 11;8(7):1535–1550. doi: 10.1093/nar/8.7.1535. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peden K. W. Revised sequence of the tetracycline-resistance gene of pBR322. Gene. 1983 May-Jun;22(2-3):277–280. doi: 10.1016/0378-1119(83)90112-9. [DOI] [PubMed] [Google Scholar]
- Ravetch J. V., Ohsumi M., Model P., Vovis G. F., Fischhoff D., Zinder N. D. Organization of a hybrid between phage f1 and plasmid pSC101. Proc Natl Acad Sci U S A. 1979 May;76(5):2195–2198. doi: 10.1073/pnas.76.5.2195. [DOI] [PMC free article] [PubMed] [Google Scholar]
- SAITO H., MIURA K. I. PREPARATION OF TRANSFORMING DEOXYRIBONUCLEIC ACID BY PHENOL TREATMENT. Biochim Biophys Acta. 1963 Aug 20;72:619–629. [PubMed] [Google Scholar]
- Sambrook J. F., Fan D. P., Brenner S. A strong suppressor specific for UGA. Nature. 1967 Apr 29;214(5087):452–453. doi: 10.1038/214452a0. [DOI] [PubMed] [Google Scholar]
- Shaw K. J., Berg C. M. Escherichia coli K-12 auxotrophs induced by insertion of the transposable element Tn5. Genetics. 1979 Jul;92(3):741–747. doi: 10.1093/genetics/92.3.741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Singhal R. P., Roberts E. F., Vakharia V. N. Structure of transfer RNAs: listing of 150 additional sequences. Prog Nucleic Acid Res Mol Biol. 1983;28:211–249. doi: 10.1016/s0079-6603(08)60088-4. [DOI] [PubMed] [Google Scholar]
- Soll L., Berg P. Recessive lethal nonsense suppressor in Escherichia coli which inserts glutamine. Nature. 1969 Sep 27;223(5213):1340–1342. doi: 10.1038/2231340a0. [DOI] [PubMed] [Google Scholar]
- Soll L., Berg P. Recessive lethals: a new class of nonsense suppressors in Escherichia coli. Proc Natl Acad Sci U S A. 1969 Jun;63(2):392–399. doi: 10.1073/pnas.63.2.392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soll L. Mutational alterations of tryptophan-specific transfer RNA that generate translation suppressors of the UAA, UAG and UGA nonsense codons. J Mol Biol. 1974 Jun 25;86(2):233–243. doi: 10.1016/0022-2836(74)90015-1. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Vogelstein B., Gillespie D. Preparative and analytical purification of DNA from agarose. Proc Natl Acad Sci U S A. 1979 Feb;76(2):615–619. doi: 10.1073/pnas.76.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiner A. M., Weber K. Natural read-through at the UGA termination signal of Q-beta coat protein cistron. Nat New Biol. 1971 Sep 15;234(50):206–209. doi: 10.1038/newbio234206a0. [DOI] [PubMed] [Google Scholar]
- Yaniv M., Folk W. R., Berg P., Soll L. A single mutational modification of a tryptophan-specific transfer RNA permits aminoacylation by glutamine and translation of the codon UAG. J Mol Biol. 1974 Jun 25;86(2):245–260. doi: 10.1016/0022-2836(74)90016-3. [DOI] [PubMed] [Google Scholar]
- Yarus M. Isolation and properties of a plasmid which expresses the E. coli Su+7 amber suppressor tRNA gene. Mol Gen Genet. 1979 Mar 5;170(3):291–298. doi: 10.1007/BF00267062. [DOI] [PubMed] [Google Scholar]
- Yarus M., McMillan C., 3rd, Cline S., Bradley D., Snyder M. Construction of a composite tRNA gene by anticodon loop transplant. Proc Natl Acad Sci U S A. 1980 Sep;77(9):5092–5096. doi: 10.1073/pnas.77.9.5092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yates J. L., Gette W. R., Furth M. E., Nomura M. Effects of ribosomal mutations on the read-through of a chain termination signal: studies on the synthesis of bacteriophage lambda O gene protein in vitro. Proc Natl Acad Sci U S A. 1977 Feb;74(2):689–693. doi: 10.1073/pnas.74.2.689. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Young R. A. Transcription termination in the Escherichia coli ribosomal RNA operon rrnC. J Biol Chem. 1979 Dec 25;254(24):12725–12731. [PubMed] [Google Scholar]