Abstract
A deletion mutation of the gene for protein S (tps), a development-specific protein of Myxococcus xanthus, was constructed. No significant differences in the process of fruiting body formation or the yield of myxospores were observed between mutant and wild-type cells. On the other hand, when the tps gene was deleted together with a 2.0-kilobase sequence including the ops gene immediately upstream of the tps gene, fruiting body formation was substantially delayed, and the yield of myxospores was reduced. These results indicate that protein S is not essential for differentiation of M. xanthus, whereas a gene product(s) coded from the sequence upstream of the tps gene appears to be required for normal fruiting body formation.
Full text
PDF
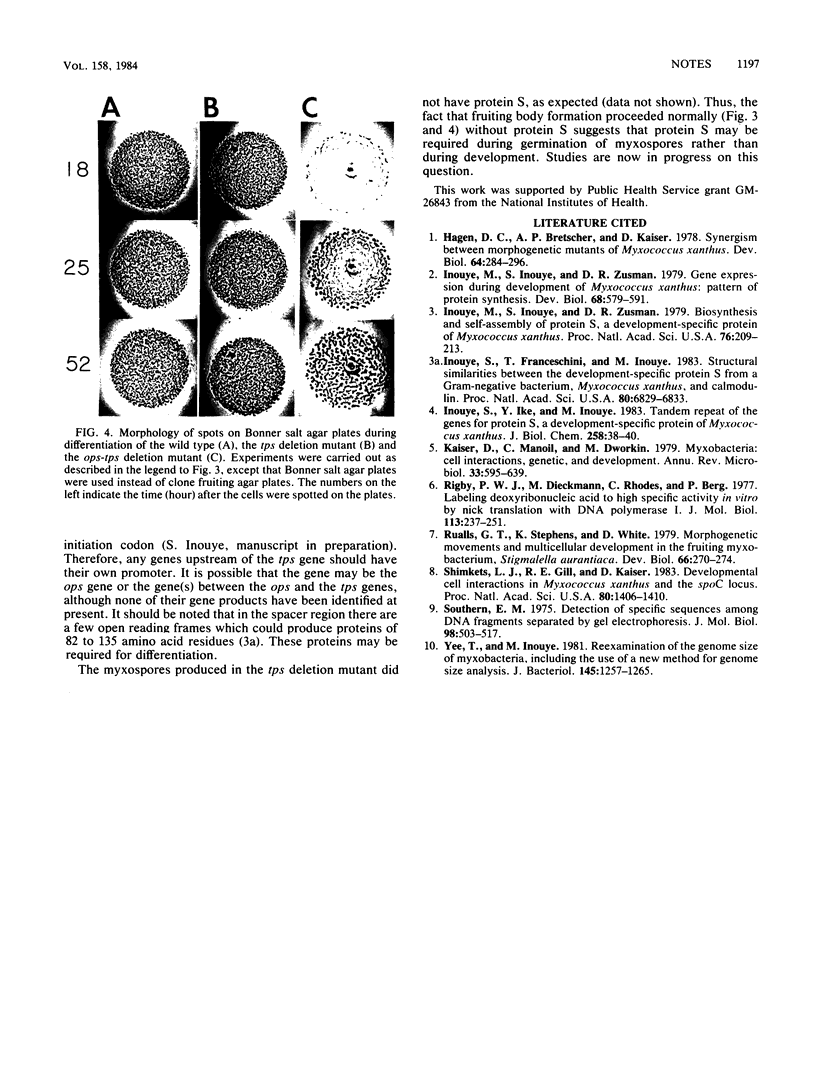
Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Hagen D. C., Bretscher A. P., Kaiser D. Synergism between morphogenetic mutants of Myxococcus xanthus. Dev Biol. 1978 Jun;64(2):284–296. doi: 10.1016/0012-1606(78)90079-9. [DOI] [PubMed] [Google Scholar]
- Inouye M., Inouye S., Zusman D. R. Biosynthesis and self-assembly of protein S, a development-specific protein of Myxococcus xanthus. Proc Natl Acad Sci U S A. 1979 Jan;76(1):209–213. doi: 10.1073/pnas.76.1.209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye M., Inouye S., Zusman D. R. Gene expression during development of Myxococcus xanthus: pattern of protein synthesis. Dev Biol. 1979 Feb;68(2):579–591. doi: 10.1016/0012-1606(79)90228-8. [DOI] [PubMed] [Google Scholar]
- Inouye S., Franceschini T., Inouye M. Structural similarities between the development-specific protein S from a gram-negative bacterium, Myxococcus xanthus, and calmodulin. Proc Natl Acad Sci U S A. 1983 Nov;80(22):6829–6833. doi: 10.1073/pnas.80.22.6829. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inouye S., Ike Y., Inouye M. Tandem repeat of the genes for protein S, a development-specific protein of Myxococcus xanthus. J Biol Chem. 1983 Jan 10;258(1):38–40. [PubMed] [Google Scholar]
- Kaiser D., Manoil C., Dworkin M. Myxobacteria: cell interactions, genetics, and development. Annu Rev Microbiol. 1979;33:595–639. doi: 10.1146/annurev.mi.33.100179.003115. [DOI] [PubMed] [Google Scholar]
- Qualls G. T., Stephens K., White D. Morphogenetic movements and multicellular development in the fruiting Myxobacterium, Stigmatella aurantiaca. Dev Biol. 1978 Sep;66(1):270–274. doi: 10.1016/0012-1606(78)90291-9. [DOI] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Shimkets L. J., Gill R. E., Kaiser D. Developmental cell interactions in Myxococcus xanthus and the spoC locus. Proc Natl Acad Sci U S A. 1983 Mar;80(5):1406–1410. doi: 10.1073/pnas.80.5.1406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Yee T., Inouye M. Reexamination of the genome size of myxobacteria, including the use of a new method for genome size analysis. J Bacteriol. 1981 Mar;145(3):1257–1265. doi: 10.1128/jb.145.3.1257-1265.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]