Abstract
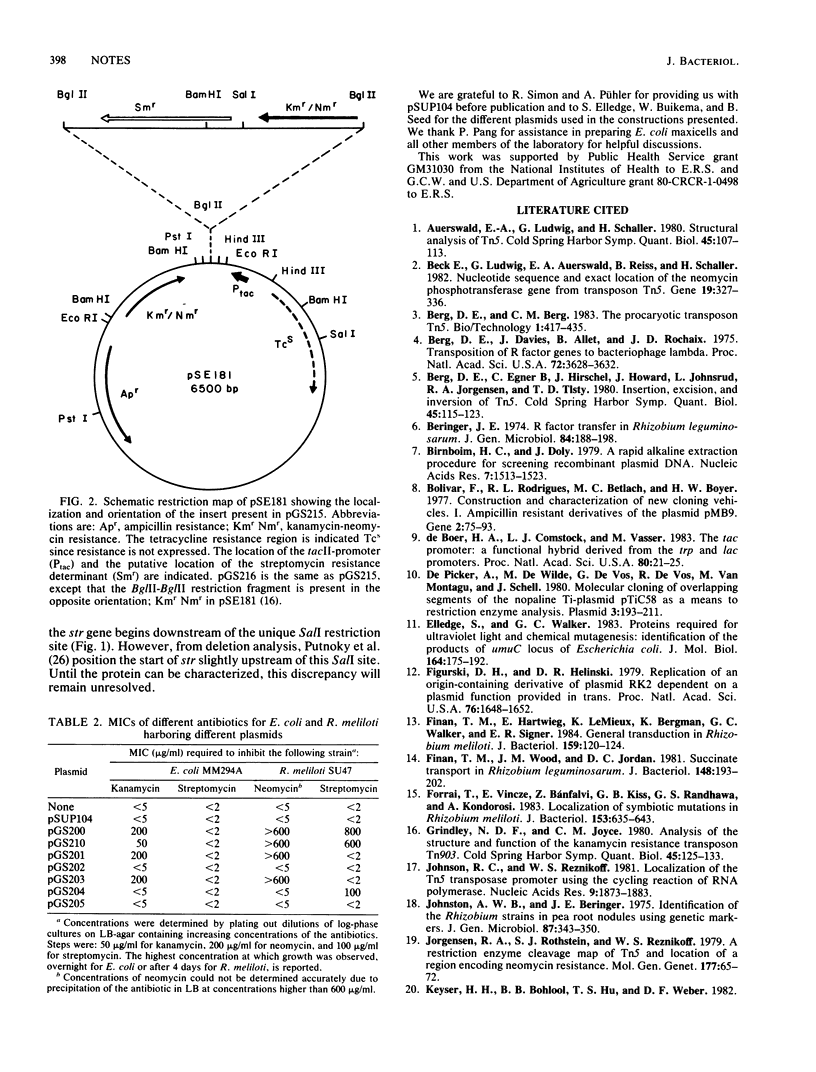
Transposon Tn5 encodes streptomycin resistance in addition to kanamycin-neomycin resistance. This resistance was not detectable in Escherichia coli but was efficiently expressed in Rhizobium meliloti and certain other strains. By analysis of cloned Tn5 restriction endonuclease fragments, the streptomycin resistance (str) gene was located in the right-hand side of the central region as the transposon is conventionally drawn. Transcription of str appeared to originate at pL, the promoter for the neo gene (neomycin phosphotransferase type II). Expression of streptomycin resistance in E. coli was obtained after cloning of the neo-str region downstream of a strong E. coli promoter. A construct in which PL was deleted also showed differential expression of streptomycin resistance.
Full text
PDF



Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Auerswald E. A., Ludwig G., Schaller H. Structural analysis of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):107–113. doi: 10.1101/sqb.1981.045.01.019. [DOI] [PubMed] [Google Scholar]
- Beck E., Ludwig G., Auerswald E. A., Reiss B., Schaller H. Nucleotide sequence and exact localization of the neomycin phosphotransferase gene from transposon Tn5. Gene. 1982 Oct;19(3):327–336. doi: 10.1016/0378-1119(82)90023-3. [DOI] [PubMed] [Google Scholar]
- Berg D. E., Davies J., Allet B., Rochaix J. D. Transposition of R factor genes to bacteriophage lambda. Proc Natl Acad Sci U S A. 1975 Sep;72(9):3628–3632. doi: 10.1073/pnas.72.9.3628. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Berg D. E., Egner C., Hirschel B. J., Howard J., Johnsrud L., Jorgensen R. A., Tlsty T. D. Insertion, excision, and inversion of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):115–123. doi: 10.1101/sqb.1981.045.01.020. [DOI] [PubMed] [Google Scholar]
- Beringer J. E. R factor transfer in Rhizobium leguminosarum. J Gen Microbiol. 1974 Sep;84(1):188–198. doi: 10.1099/00221287-84-1-188. [DOI] [PubMed] [Google Scholar]
- Birnboim H. C., Doly J. A rapid alkaline extraction procedure for screening recombinant plasmid DNA. Nucleic Acids Res. 1979 Nov 24;7(6):1513–1523. doi: 10.1093/nar/7.6.1513. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bolivar F., Rodriguez R. L., Betlach M. C., Boyer H. W. Construction and characterization of new cloning vehicles. I. Ampicillin-resistant derivatives of the plasmid pMB9. Gene. 1977;2(2):75–93. doi: 10.1016/0378-1119(77)90074-9. [DOI] [PubMed] [Google Scholar]
- Depicker A., De Wilde M., De Vos G., De Vos R., Van Montagu M., Schell J. Molecular cloning of overlapping segments of the nopaline Ti-plasmid pTiC58 as a means to restriction endonuclease mapping. Plasmid. 1980 Mar;3(2):193–211. doi: 10.1016/0147-619x(80)90109-2. [DOI] [PubMed] [Google Scholar]
- Elledge S. J., Walker G. C. Proteins required for ultraviolet light and chemical mutagenesis. Identification of the products of the umuC locus of Escherichia coli. J Mol Biol. 1983 Feb 25;164(2):175–192. doi: 10.1016/0022-2836(83)90074-8. [DOI] [PubMed] [Google Scholar]
- Figurski D. H., Helinski D. R. Replication of an origin-containing derivative of plasmid RK2 dependent on a plasmid function provided in trans. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1648–1652. doi: 10.1073/pnas.76.4.1648. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finan T. M., Hartweig E., LeMieux K., Bergman K., Walker G. C., Signer E. R. General transduction in Rhizobium meliloti. J Bacteriol. 1984 Jul;159(1):120–124. doi: 10.1128/jb.159.1.120-124.1984. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Finan T. M., Wood J. M., Jordan D. C. Succinate transport in Rhizobium leguminosarum. J Bacteriol. 1981 Oct;148(1):193–202. doi: 10.1128/jb.148.1.193-202.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Forrai T., Vincze E., Bánfalvi Z., Kiss G. B., Randhawa G. S., Kondorosi A. Localization of symbiotic mutations in Rhizobium meliloti. J Bacteriol. 1983 Feb;153(2):635–643. doi: 10.1128/jb.153.2.635-643.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grindley N. D., Joyce C. M. Analysis of the structure and function of the kanamycin-resistance transposon Tn903. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):125–133. doi: 10.1101/sqb.1981.045.01.021. [DOI] [PubMed] [Google Scholar]
- Johnson R. C., Reznikoff W. S. Localization of the Tn5 transposase promoter using the cycling reaction of RNA polymerase. Nucleic Acids Res. 1981 Apr 24;9(8):1873–1883. doi: 10.1093/nar/9.8.1873. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnston A. W., Beringer J. E. Identification of the rhizobium strains in pea root nodules using genetic markers. J Gen Microbiol. 1975 Apr;87(2):343–350. doi: 10.1099/00221287-87-2-343. [DOI] [PubMed] [Google Scholar]
- Jorgensen R. A., Rothstein S. J., Reznikoff W. S. A restriction enzyme cleavage map of Tn5 and location of a region encoding neomycin resistance. Mol Gen Genet. 1979;177(1):65–72. doi: 10.1007/BF00267254. [DOI] [PubMed] [Google Scholar]
- Meade H. M., Long S. R., Ruvkun G. B., Brown S. E., Ausubel F. M. Physical and genetic characterization of symbiotic and auxotrophic mutants of Rhizobium meliloti induced by transposon Tn5 mutagenesis. J Bacteriol. 1982 Jan;149(1):114–122. doi: 10.1128/jb.149.1.114-122.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyer R. J., Shapiro J. A. Genetic organization of the broad-host-range IncP-1 plasmid R751. J Bacteriol. 1980 Sep;143(3):1362–1373. doi: 10.1128/jb.143.3.1362-1373.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Putnoky P., Kiss G. B., Ott I., Kondorosi A. Tn5 carries a streptomycin resistance determinant downstream from the kanamycin resistance gene. Mol Gen Genet. 1983;191(2):288–294. doi: 10.1007/BF00334828. [DOI] [PubMed] [Google Scholar]
- Reznikoff W. S. Tn5 transposition and its regulation. Cell. 1982 Dec;31(2 Pt 1):307–308. doi: 10.1016/0092-8674(82)90123-4. [DOI] [PubMed] [Google Scholar]
- Rothstein S. J., Jorgensen R. A., Postle K., Reznikoff W. S. The inverted repeats of Tn5 are functionally different. Cell. 1980 Mar;19(3):795–805. doi: 10.1016/s0092-8674(80)80055-9. [DOI] [PubMed] [Google Scholar]
- Rothstein S. J., Jorgensen R. A., Yin J. C., Yong-di Z., Johnson R. C., Reznikoff W. S. Genetic organization of Tn5. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):99–105. doi: 10.1101/sqb.1981.045.01.018. [DOI] [PubMed] [Google Scholar]
- Sancar A., Wharton R. P., Seltzer S., Kacinski B. M., Clarke N. D., Rupp W. D. Identification of the uvrA gene product. J Mol Biol. 1981 May 5;148(1):45–62. doi: 10.1016/0022-2836(81)90234-5. [DOI] [PubMed] [Google Scholar]
- Sutcliffe J. G. Complete nucleotide sequence of the Escherichia coli plasmid pBR322. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 1):77–90. doi: 10.1101/sqb.1979.043.01.013. [DOI] [PubMed] [Google Scholar]
- Thuring R. W., Sanders J. P., Borst P. A freeze-squeeze method for recovering long DNA from agarose gels. Anal Biochem. 1975 May 26;66(1):213–220. doi: 10.1016/0003-2697(75)90739-3. [DOI] [PubMed] [Google Scholar]
- Ward J. M., Grinsted J. Analysis of the Inc P-1 group plasmids R906 and R751 and their relationship to RP1. Plasmid. 1982 Nov;8(3):244–252. doi: 10.1016/0147-619x(82)90062-2. [DOI] [PubMed] [Google Scholar]
- de Boer H. A., Comstock L. J., Vasser M. The tac promoter: a functional hybrid derived from the trp and lac promoters. Proc Natl Acad Sci U S A. 1983 Jan;80(1):21–25. doi: 10.1073/pnas.80.1.21. [DOI] [PMC free article] [PubMed] [Google Scholar]



