Abstract
Using a model system, we have studied deletion formation in Bacillus subtilis. When the staphylococcal plasmids pSA2100 (7.1 kilobases) and pUB110 (4.5 kilobases) were ligated to one another at their unique XbaI sites and transformed into either rec+ or recE4 strains of B. subtilis, an intramolecular recombination event usually occurred. Two plasmids, one of 2.6 kilobases and the other of 9.0 kilobases, were consistently isolated and shown by restriction enzyme analysis to be derived by recombination occurring in the pSA2100-pUB110 cointegrate. Analysis of the sequence of the junctions of the recombinant plasmids and of the crossover regions of the parental plasmids suggested that a reciprocal, conservative, intramolecular recombination event had occurred between short 18-base-pair homologous sequences that were oriented as direct repeats and bounded by regions of dyad symmetry. Evidence is presented that the above illegitimate recombination event is biased to occur intramolecularly and that randomly chosen direct repeats of either 22 or 29 base pairs are not sufficient to support recombination. The recombination event occurs in recA1, recB2, recD3, recE5, recL16, recM13, polA59, polA13, uvr-22, uvr-13, and stb mutants of B. subtilis and does not require that the competent state be established.
Full text
PDF






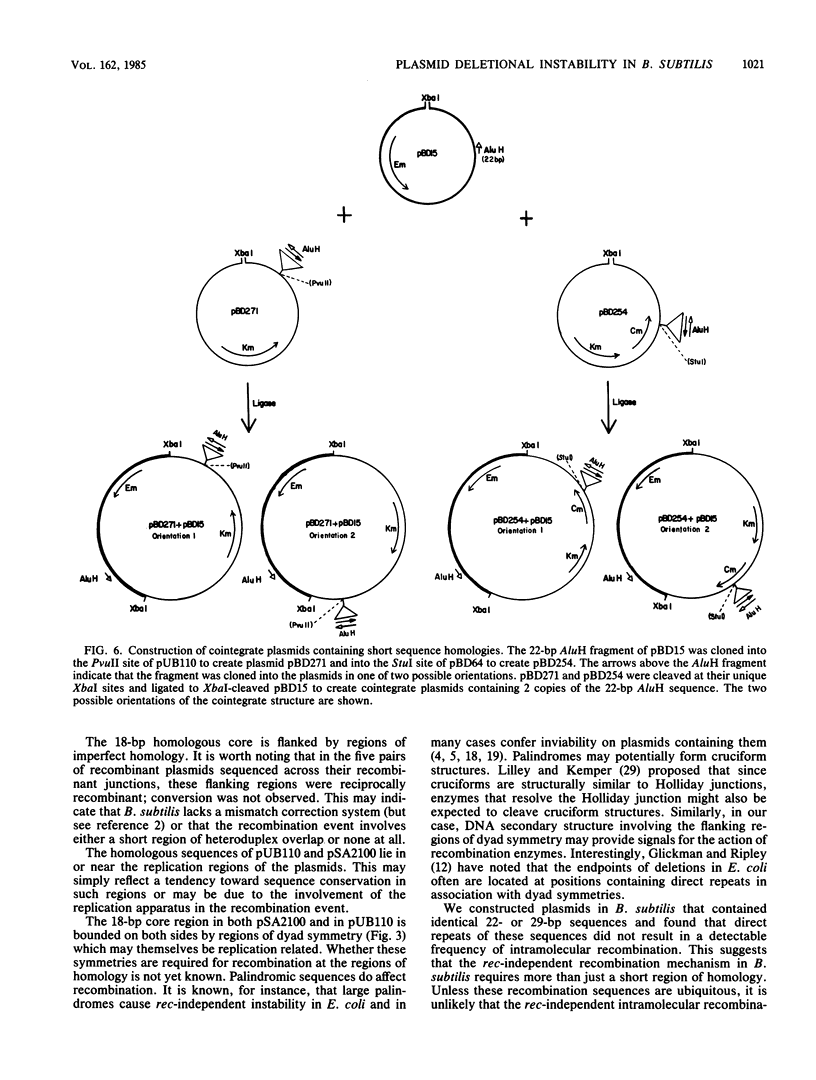


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Albertini A. M., Hofer M., Calos M. P., Miller J. H. On the formation of spontaneous deletions: the importance of short sequence homologies in the generation of large deletions. Cell. 1982 Jun;29(2):319–328. doi: 10.1016/0092-8674(82)90148-9. [DOI] [PubMed] [Google Scholar]
- Bresler S. E., Kreneva R. A., Kushev V. V. Correction of molecular heterozygotes in the course of transformation. Mol Gen Genet. 1968;102(3):257–268. doi: 10.1007/BF00385983. [DOI] [PubMed] [Google Scholar]
- Chang S., Cohen S. N. High frequency transformation of Bacillus subtilis protoplasts by plasmid DNA. Mol Gen Genet. 1979 Jan 5;168(1):111–115. doi: 10.1007/BF00267940. [DOI] [PubMed] [Google Scholar]
- Collins J. Instability of palindromic DNA in Escherichia coli. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):409–416. doi: 10.1101/sqb.1981.045.01.055. [DOI] [PubMed] [Google Scholar]
- Collins J., Volckaert G., Nevers P. Precise and nearly-precise excision of the symmetrical inverted repeats of Tn5; common features of recA-independent deletion events in Escherichia coli. Gene. 1982 Jul-Aug;19(1):139–146. doi: 10.1016/0378-1119(82)90198-6. [DOI] [PubMed] [Google Scholar]
- Contente S., Dubnau D. Characterization of plasmid transformation in Bacillus subtilis: kinetic properties and the effect of DNA conformation. Mol Gen Genet. 1979 Jan 2;167(3):251–258. doi: 10.1007/BF00267416. [DOI] [PubMed] [Google Scholar]
- Contente S., Dubnau D. Marker rescue transformation by linear plasmid DNA in Bacillus subtilis. Plasmid. 1979 Oct;2(4):555–571. doi: 10.1016/0147-619x(79)90054-4. [DOI] [PubMed] [Google Scholar]
- Doly J., Sasarman E., Anagnostopoulos C. ATP-dependent deoxyribonuclease in Bacillus subtilis and a mutant deficient in this activity. Mutat Res. 1974 Jan;22(1):15–23. doi: 10.1016/0027-5107(74)90003-7. [DOI] [PubMed] [Google Scholar]
- Dubnau D., Davidoff-Abelson R., Scher B., Cirigliano C. Fate of transforming deoxyribonucleic acid after uptake by competent Bacillus subtilis: phenotypic characterization of radiation-sensitive recombination-deficient mutants. J Bacteriol. 1973 Apr;114(1):273–286. doi: 10.1128/jb.114.1.273-286.1973. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Farabaugh P. J., Schmeissner U., Hofer M., Miller J. H. Genetic studies of the lac repressor. VII. On the molecular nature of spontaneous hotspots in the lacI gene of Escherichia coli. J Mol Biol. 1978 Dec 25;126(4):847–857. doi: 10.1016/0022-2836(78)90023-2. [DOI] [PubMed] [Google Scholar]
- Gass K. B., Cozzarelli N. R. Further genetic and enzymological characterization of the three Bacillus subtilis deoxyribonucleic acid polymerases. J Biol Chem. 1973 Nov 25;248(22):7688–7700. [PubMed] [Google Scholar]
- Glickman B. W., Ripley L. S. Structural intermediates of deletion mutagenesis: a role for palindromic DNA. Proc Natl Acad Sci U S A. 1984 Jan;81(2):512–516. doi: 10.1073/pnas.81.2.512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandi G., Mottes M., Sgaramella V. Specific pattern of instability of Escherichia coli HisG gene cloned in Bacillus subtilis via the Staphylococcus aureus plasmid pCS194. Plasmid. 1981 Jul;6(1):99–111. doi: 10.1016/0147-619x(81)90057-3. [DOI] [PubMed] [Google Scholar]
- Grindley N. D., Lauth M. R., Wells R. G., Wityk R. J., Salvo J. J., Reed R. R. Transposon-mediated site-specific recombination: identification of three binding sites for resolvase at the res sites of gamma delta and Tn3. Cell. 1982 Aug;30(1):19–27. doi: 10.1016/0092-8674(82)90007-1. [DOI] [PubMed] [Google Scholar]
- Gryczan T. J., Contente S., Dubnau D. Characterization of Staphylococcus aureus plasmids introduced by transformation into Bacillus subtilis. J Bacteriol. 1978 Apr;134(1):318–329. doi: 10.1128/jb.134.1.318-329.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gryczan T. J., Dubnau D. Construction and properties of chimeric plasmids in Bacillus subtilis. Proc Natl Acad Sci U S A. 1978 Mar;75(3):1428–1432. doi: 10.1073/pnas.75.3.1428. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gryczan T., Shivakumar A. G., Dubnau D. Characterization of chimeric plasmid cloning vehicles in Bacillus subtilis. J Bacteriol. 1980 Jan;141(1):246–253. doi: 10.1128/jb.141.1.246-253.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hagan C. E., Warren G. J. Lethality of palindromic DNA and its use in selection of recombinant plasmids. Gene. 1982 Jul-Aug;19(1):147–151. doi: 10.1016/0378-1119(82)90199-8. [DOI] [PubMed] [Google Scholar]
- Hagan C. E., Warren G. J. Viability of palindromic DNA is restored by deletions occurring at low but variable frequency in plasmids of Escherichia coli. Gene. 1983 Oct;24(2-3):317–326. doi: 10.1016/0378-1119(83)90092-6. [DOI] [PubMed] [Google Scholar]
- Hoch J. A., Barat M., Anagnostopoulos C. Transformation and transduction in recombination-defective mutants of Bacillus subtilis. J Bacteriol. 1967 Jun;93(6):1925–1937. doi: 10.1128/jb.93.6.1925-1937.1967. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Horinouchi S., Weisblum B. Nucleotide sequence and functional map of pC194, a plasmid that specifies inducible chloramphenicol resistance. J Bacteriol. 1982 May;150(2):815–825. doi: 10.1128/jb.150.2.815-825.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikeda H., Aoki K., Naito A. Illegitimate recombination mediated in vitro by DNA gyrase of Escherichia coli: structure of recombinant DNA molecules. Proc Natl Acad Sci U S A. 1982 Jun;79(12):3724–3728. doi: 10.1073/pnas.79.12.3724. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ikeda H., Kawasaki I., Gellert M. Mechanism of illegitimate recombination: common sites for recombination and cleavage mediated by E. coli DNA gyrase. Mol Gen Genet. 1984;196(3):546–549. doi: 10.1007/BF00436208. [DOI] [PubMed] [Google Scholar]
- Ikeda H., Moriya K., Matsumoto T. In vitro study of illegitimate recombination: involvement of DNA gyrase. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):399–408. doi: 10.1101/sqb.1981.045.01.054. [DOI] [PubMed] [Google Scholar]
- Iordănescu S. Recombinant plasmid obtained from two different, compatible staphylococcal plasmids. J Bacteriol. 1975 Nov;124(2):597–601. doi: 10.1128/jb.124.2.597-601.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jones I. M., Primrose S. B., Ehrlich S. D. Recombination between short direct repeats in a recA host. Mol Gen Genet. 1982;188(3):486–489. doi: 10.1007/BF00330053. [DOI] [PubMed] [Google Scholar]
- Lacey R. W., Chopra I. Genetic studies of a multi-resistant strain of Staphylococcus aureus. J Med Microbiol. 1974 May;7(2):285–297. doi: 10.1099/00222615-7-2-285. [DOI] [PubMed] [Google Scholar]
- Lilley D. M., Kemper B. Cruciform-resolvase interactions in supercoiled DNA. Cell. 1984 Feb;36(2):413–422. doi: 10.1016/0092-8674(84)90234-4. [DOI] [PubMed] [Google Scholar]
- Lopez P., Espinosa M., Greenberg B., Lacks S. A. Generation of deletions in pneumococcal mal genes cloned in Bacillus subtilis. Proc Natl Acad Sci U S A. 1984 Aug;81(16):5189–5193. doi: 10.1073/pnas.81.16.5189. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marvo S. L., King S. R., Jaskunas S. R. Role of short regions of homology in intermolecular illegitimate recombination events. Proc Natl Acad Sci U S A. 1983 May;80(9):2452–2456. doi: 10.1073/pnas.80.9.2452. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Naito A., Naito S., Ikeda H. Homology is not required for recombination mediated by DNA gyrase of Escherichia coli. Mol Gen Genet. 1984;193(2):238–243. doi: 10.1007/BF00330674. [DOI] [PubMed] [Google Scholar]
- Novick R. P., Projan S. J., Rosenblum W., Edelman I. Staphylococcal plasmid cointegrates are formed by host- and phage-mediated general rec systems that act on short regions of homology. Mol Gen Genet. 1984;195(1-2):374–377. doi: 10.1007/BF00332777. [DOI] [PubMed] [Google Scholar]
- Okubo S., Romig W. R. Impaired transformability of Bacillus subtilis mutant sensitive to mitomycin C and ultraviolet radiation. J Mol Biol. 1966 Feb;15(2):440–454. doi: 10.1016/s0022-2836(66)80120-1. [DOI] [PubMed] [Google Scholar]
- Ostroff G. R., Pène J. J. Molecular cloning with bifunctional plasmid vectors in Bacillus subtilis: isolation of a spontaneous mutant of Bacillus subtilis with enhanced transformability for Escherichia coli-propagated chimeric plasmid DNA. J Bacteriol. 1983 Nov;156(2):934–936. doi: 10.1128/jb.156.2.934-936.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Primrose S. B., Ehrlich S. D. Isolation of plasmid deletion Mutants and study of their instability. Plasmid. 1981 Sep;6(2):193–201. doi: 10.1016/0147-619x(81)90066-4. [DOI] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Scheer-Abramowitz J., Gryczan T. J., Dubnau D. Origin and mode of replication of plasmids pE194 and pUB110. Plasmid. 1981 Jul;6(1):67–77. doi: 10.1016/0147-619x(81)90054-8. [DOI] [PubMed] [Google Scholar]
- Searashi T., Strauss B. Relation of the repair of damage induced by a monofunctional alkylating agent to the repair of damage induced by ultraviolet light in Bacillus subtilis. Biochem Biophys Res Commun. 1965 Sep 22;20(6):680–687. doi: 10.1016/0006-291x(65)90069-0. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Tanaka T. recE4-Independent recombination between homologous deoxyribonucleic acid segments of Bacillus subtilis plasmids. J Bacteriol. 1979 Sep;139(3):775–782. doi: 10.1128/jb.139.3.775-782.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Uhlén M., Flock J. I., Philipson L. RecE independent deletions of recombinant plasmids in Bacillus subtilis. Plasmid. 1981 Mar;5(2):161–169. doi: 10.1016/0147-619x(81)90017-2. [DOI] [PubMed] [Google Scholar]






