Abstract
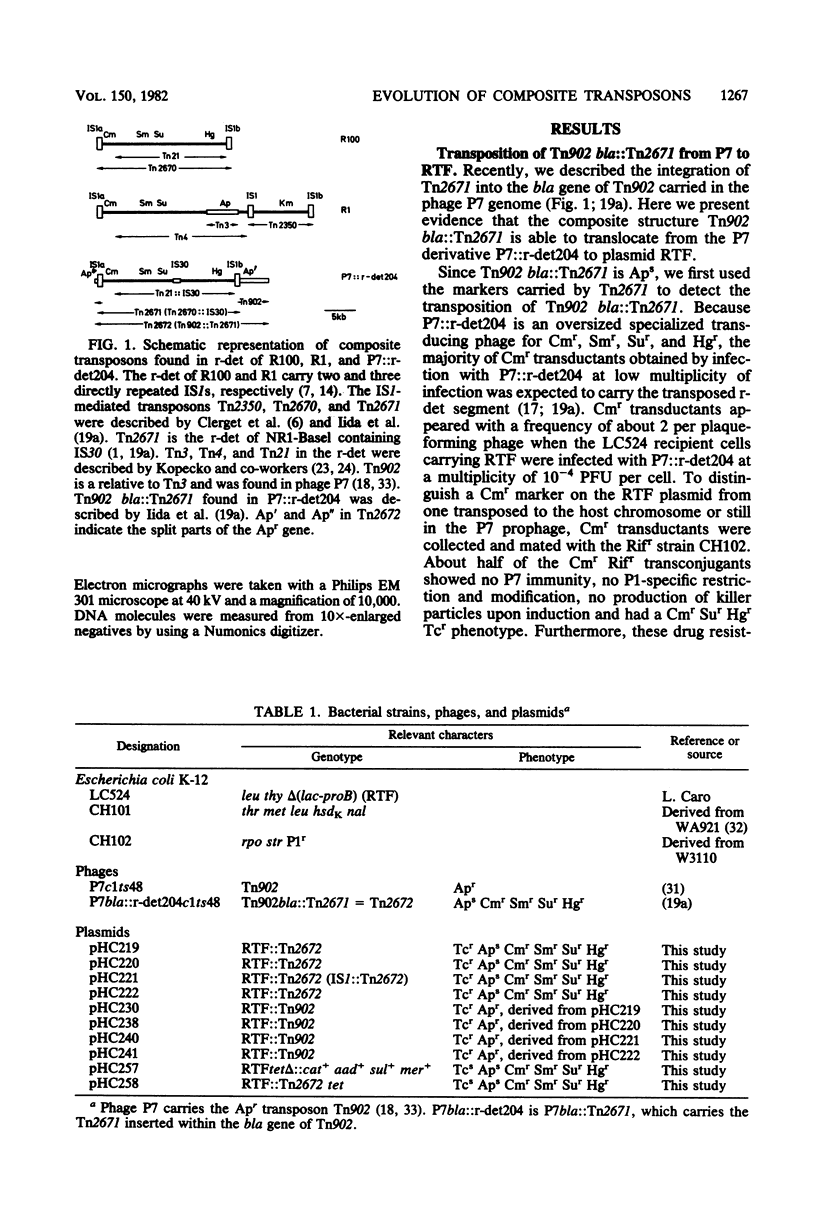
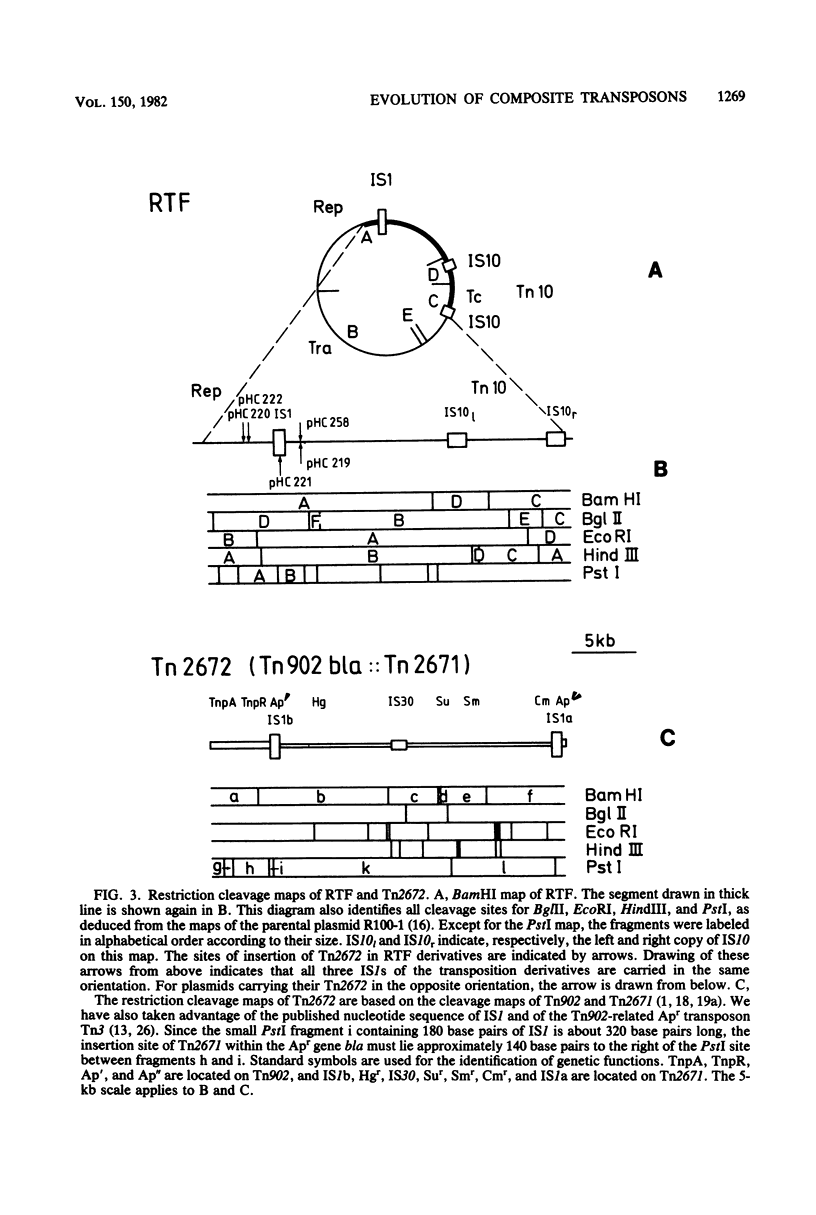
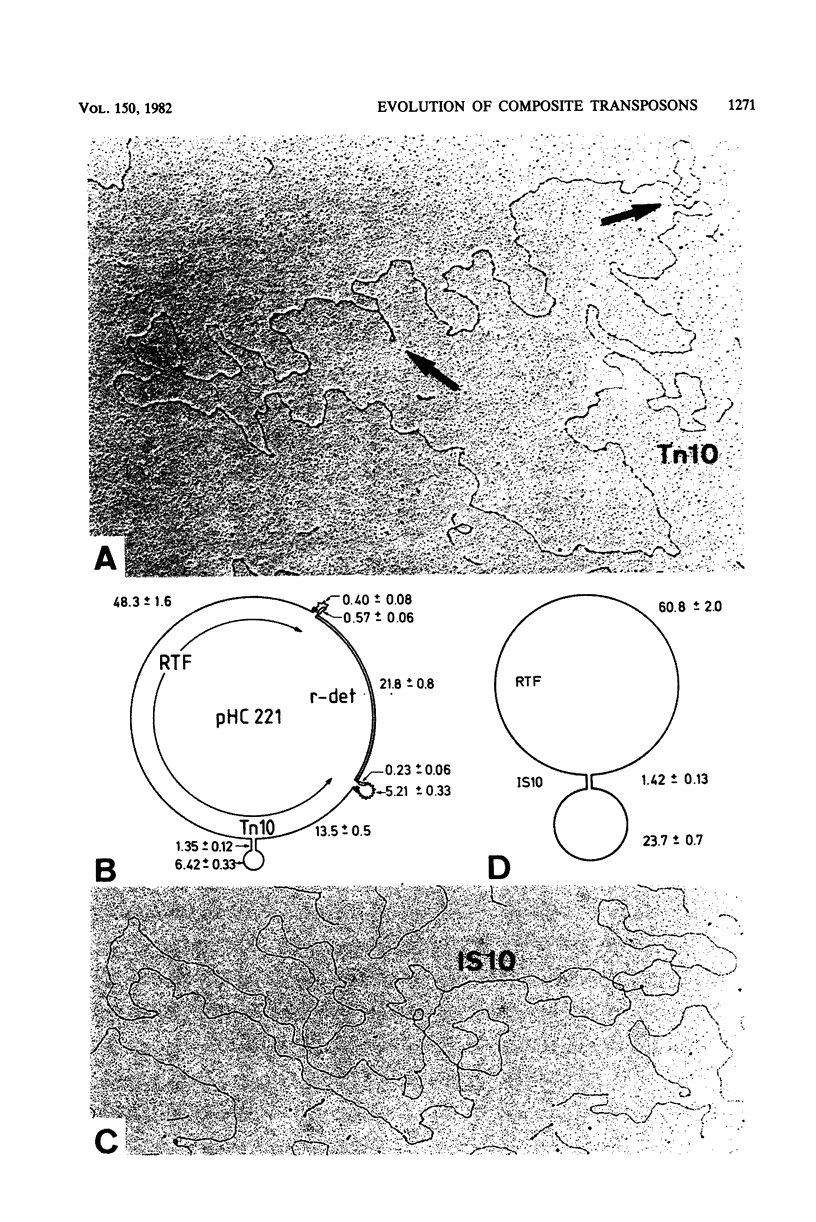
We found Tn2671 (the 23-kb long IS1-flanked r-determinant of NR1-Basel) inserted into the ampicillin resistance gene bla of the Tn3-related transposon Tn902. The resulting 28-kilobase-long composite transposon Tn2672 (= Tn902 bla::Tn2671) is stable, and it translocates as a unit into various loci including IS1 of the resistance transfer factor of R100-1. These results are discussed with respect to the evolution of R plasmids providing multiple drug resistance.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arber W., Iida S., Jütte H., Caspers P., Meyer J., Hänni C. Rearrangements of genetic material in Escherichia coli as observed on the bacteriophage P1 plasmid. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 2):1197–1208. doi: 10.1101/sqb.1979.043.01.136. [DOI] [PubMed] [Google Scholar]
- Chandler M., Allet B., Gallay E., Boy de La Tour E., Caro L. Involvement of IS1 in the dissociation of the r-determinant and RTF components of the plasmid R100.1. Mol Gen Genet. 1977 Jun 24;153(3):289–295. doi: 10.1007/BF00431594. [DOI] [PubMed] [Google Scholar]
- Chandler M., Roulet E., Silver L., Boy de la Tour E., Caro L. Tn10 mediated integration of the plasmid R100.1 into the bacterial chromosome: inverse transposition. Mol Gen Genet. 1979 May 23;173(1):23–30. doi: 10.1007/BF00267687. [DOI] [PubMed] [Google Scholar]
- Chou J., Lemaux P. G., Casadaban M. J., Cohen S. N. Transposition protein of Tn3: identification and characterisation of an essential repressor-controlled gene product. Nature. 1979 Dec 20;282(5741):801–806. doi: 10.1038/282801a0. [DOI] [PubMed] [Google Scholar]
- Clerget M., Chandler M., Caro L. Isolation of an IS1 flanked kanamycin resistance transposon from R1drd19. Mol Gen Genet. 1980;180(1):123–127. doi: 10.1007/BF00267360. [DOI] [PubMed] [Google Scholar]
- Clerget M., Chandler M., Caro L. The structure of R1drd19: a revised physical map of the plasmid. Mol Gen Genet. 1981;181(2):183–191. doi: 10.1007/BF00268425. [DOI] [PubMed] [Google Scholar]
- Cornelis G., Saedler H. Deletions and an inversion induced by a resident IS1 of the lactose transposon Tn951. Mol Gen Genet. 1980;178(2):367–374. doi: 10.1007/BF00270486. [DOI] [PubMed] [Google Scholar]
- Foster T. J., Davis M. A., Roberts D. E., Takeshita K., Kleckner N. Genetic organization of transposon Tn10. Cell. 1981 Jan;23(1):201–213. doi: 10.1016/0092-8674(81)90285-3. [DOI] [PubMed] [Google Scholar]
- Gill R. E., Heffron F., Falkow S. Identification of the protein encoded by the transposable element Tn3 which is required for its transposition. Nature. 1979 Dec 20;282(5741):797–801. doi: 10.1038/282797a0. [DOI] [PubMed] [Google Scholar]
- Heffron F., McCarthy B. J., Ohtsubo H., Ohtsubo E. DNA sequence analysis of the transposon Tn3: three genes and three sites involved in transposition of Tn3. Cell. 1979 Dec;18(4):1153–1163. doi: 10.1016/0092-8674(79)90228-9. [DOI] [PubMed] [Google Scholar]
- Hu S., Otsubo E., Davidson N., Saedler H. Electron microscope heteroduplex studies of sequence relations among bacterial plasmids: identification and mapping of the insertion sequences IS1 and IS2 in F and R plasmids. J Bacteriol. 1975 May;122(2):764–775. doi: 10.1128/jb.122.2.764-775.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iida S. A cointegrate of the bacteriophage P1 genome and the conjugative R plasmid R100. Plasmid. 1980 May;3(3):278–290. doi: 10.1016/0147-619x(80)90041-4. [DOI] [PubMed] [Google Scholar]
- Iida S., Arber W. Multiple physical differences in the genome structure of functionally related bacteriophages P1 and P7. Mol Gen Genet. 1979 Jun 20;173(3):249–261. doi: 10.1007/BF00268635. [DOI] [PubMed] [Google Scholar]
- Iida S., Arber W. On the role of IS1 in the formation of hybrids between the bacteriophage P1 and the R plasmid NR1. Mol Gen Genet. 1980 Jan;177(2):261–270. doi: 10.1007/BF00267437. [DOI] [PubMed] [Google Scholar]
- Iida S., Arber W. Plaque forming specialized transducing phage P1: isolation of P1CmSmSu, a precursor of P1Cm. Mol Gen Genet. 1977 Jun 24;153(3):259–269. doi: 10.1007/BF00431591. [DOI] [PubMed] [Google Scholar]
- Iida S., Hänni C., Echarti C., Arber W. Is the IS1-flanked r-determinant of the R plasmid NR1 a transposon? J Gen Microbiol. 1981 Oct;126(2):413–425. doi: 10.1099/00221287-126-2-413. [DOI] [PubMed] [Google Scholar]
- Iida S., Meyer J., Arber W. Cointegrates between bacteriophage P1 DNA and plasmid pBR322 derivatives suggest molecular mechanisms for P1-mediated transduction of small plasmids. Mol Gen Genet. 1981;184(1):1–10. doi: 10.1007/BF00271186. [DOI] [PubMed] [Google Scholar]
- Iida S., Meyer J., Arber W. Genesis and natural history of IS-mediated transposons. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):27–43. doi: 10.1101/sqb.1981.045.01.006. [DOI] [PubMed] [Google Scholar]
- Kleckner N., Chan R. K., Tye B. K., Botstein D. Mutagenesis by insertion of a drug-resistance element carrying an inverted repetition. J Mol Biol. 1975 Oct 5;97(4):561–575. doi: 10.1016/s0022-2836(75)80059-3. [DOI] [PubMed] [Google Scholar]
- Kopecko D. J., Brevet J., Cohen S. N. Involvement of multiple translocating DNA segments and recombinational hotspots in the structural evolution of bacterial plasmids. J Mol Biol. 1976 Dec;108(2):333–360. doi: 10.1016/s0022-2836(76)80124-6. [DOI] [PubMed] [Google Scholar]
- Mise K., Arber W. Plaque-forming transducing bacteriophage P1 derivatives and their behaviour in lysogenic conditions. Virology. 1976 Jan;69(1):191–205. doi: 10.1016/0042-6822(76)90206-3. [DOI] [PubMed] [Google Scholar]
- Ohtsubo H., Ohtsubo E. Nucleotide sequence of an insertion element, IS1. Proc Natl Acad Sci U S A. 1978 Feb;75(2):615–619. doi: 10.1073/pnas.75.2.615. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reif H. J. Genetic evidence for absence of transposition functions from the internal part of Tn981 a relative of Tn9. Mol Gen Genet. 1980;177(4):667–674. doi: 10.1007/BF00272678. [DOI] [PubMed] [Google Scholar]
- Rosner J. L., Guyer M. S. Transposition of IS1-lambdaBIO-IS1 from a bacteriophage lambda derivative carrying the IS1-cat-IS1 transposon (Tn9). Mol Gen Genet. 1980 Apr;178(1):111–120. doi: 10.1007/BF00267219. [DOI] [PubMed] [Google Scholar]
- Wood W. B. Host specificity of DNA produced by Escherichia coli: bacterial mutations affecting the restriction and modification of DNA. J Mol Biol. 1966 Mar;16(1):118–133. doi: 10.1016/s0022-2836(66)80267-x. [DOI] [PubMed] [Google Scholar]
- Yun T., Vapnek D. Electron microscopic analysis of bacteriophages P1, P1Cm, and P7. Determination of genome sizes, sequence homology, and location of antibiotic-resistance determinants. Virology. 1977 Mar;77(1):376–385. doi: 10.1016/0042-6822(77)90434-2. [DOI] [PubMed] [Google Scholar]