Abstract
CD8+ cytotoxic T lymphocytes (CTL) are strong mediators of human immunodeficiency virus type 1 (HIV-1) control, yet HIV-1 frequently mutates to escape CTL recognition. In an analysis of sequences in the Los Alamos HIV-1 database, we show that emerging CTL escape mutations were more often present at lower frequencies than the amino acid(s) that they replaced. Furthermore, epitopes that underwent escape contained amino acid sites of high variability, whereas epitopes persisting at high frequencies lacked highly variable sites. We therefore infer that escape mutations are likely to be associated with weak functional constraints on the viral protein. This was supported by an extensive analysis of one subject for whom all escape mutations within defined CTL epitopes were studied and by an analysis of all reported escape mutations of defined CTL epitopes in the HIV Immunology Database. In one of these defined epitopes, escape mutations involving the substitution of amino acids with lower database frequencies occurred, and the epitope soon reverted back to the sensitive form. We further show that this escape mutation substantially diminished viral fitness in in vitro competition assays. Coincident with the reversion in vivo, we observed the fixation of a mutation 3 amino acids C terminal to the epitope, coincident with the ablation of the corresponding CTL response. The C-terminal mutation did not restore replication fitness reduced by the escape mutation in the epitope and by itself had little effect on replication fitness. Therefore, this C-terminal mutation presumably impaired the processing and presentation of the epitope. Finally, for one persistent epitope, CTL cross-reactivity to a mutant form may have suppressed the mutant to undetected levels, whereas for two other persistent epitopes, each of two mutants showed poor cross-reactivity and appeared in the subject at later time points. Thus, a viral dynamic exists between the advantage of immune escape, peptide cross-reactivity, and the disadvantage of lost replication fitness, with the balance playing an important role in determining whether a CTL epitope will persist or decline during infection.
CD8+ cytotoxic T-lymphocyte (CTL) responses are associated with partial control of human immunodeficiency virus type 1 (HIV-1) infection (8, 36, 54) and are a selective force for mutations in the viral genome that confer escape from immune recognition (4, 5, 9, 13, 25, 26, 43, 49, 55, 60). Some CTL epitopes persist within HIV-1 over years of infection despite the maintenance of CTLs that specifically recognize them (10, 27, 33, 48, 52, 57). Thus, why escape mutations develop in some epitopes and not others remains unclear.
CTLs can effectively kill HIV-1-infected cells and inhibit viral replication in vitro (63), with high-avidity CTL responses possibly being more effective at viral clearance in vivo (2, 17). Studies of acute simian immunodeficiency virus (SIV) infection suggest that CTLs with high functional avidity can rapidly select for escape mutants (53). However, no such association was found in studies of acute HIV infection (13), and persistent recognition of epitopes by high-avidity CTLs was observed in patients with chronic HIV infection (19). The lack of evolution of some epitopes in vivo might be due to insufficient antiviral pressure (33). Indeed, some studies suggested an inverse correlation between the magnitude and breadth of CTL responses and viral load (11, 35), while other studies showed no such correlation (1, 12). Specific targeting of the Gag viral protein has been associated with the control of viral load (66), in contrast to responses that target Env and the auxiliary viral proteins (32). Targeting conferred by specific HLA alleles (21, 31, 59) and of subdominant CTL epitopes (22) have also been associated with better viral control. Further complicating this story is the fact that significant functional differences exist between CTLs targeting identical epitopes but restricted by different, albeit closely related, HLAs (40).
Recently, attention has been focused on the impact of mutations mediating immune escape on the replication fitness of the virus. The reduced fitness of CTL escape mutants of SIV (20, 23) and HIV-1 (47) has been observed in vitro. Reversion of escape mutants to their epitopic forms after infecting new hosts with different major histocompatibility complex class I alleles has been reported for both SIV and HIV-1 (23, 41), suggesting that there is often a fitness impairment associated with CTL escape mutations. In addition, a compensatory mutation has been shown to partially restore viral fitness and delay reversion of escape mutation (14). Furthermore, not all sites within an epitope are likely to be equally mutable. Sites that are more important for function should be less likely to harbor mutations that will reach appreciable frequencies. Studies of Gag and reverse transcriptase (RT) epitopes revealed that although multiple potential point mutations can ablate CTL recognition (28, 61), only oligoclonal escape mutants were observed. This suggests that fitness constraints might have limited the diversity of the observed escape mutants.
Viruses with amino acids found at higher frequencies in the Los Alamos National Laboratory (LANL) HIV sequence database (HIVDB) (38) might reasonably be hypothesized to be associated with better replication capacity. Exceptions may include those within HIV-1 peptides that have extensively been adapted following infection in a population with shared HLA alleles (39, 49; however, see reference 7). The degree of variability of amino acid sites in HIV proteins in the HIVDB has been associated with fitness constraints; i.e., amino acid sites with stronger fitness constraints may have lower variability (3, 30, 65). Furthermore, a study of V3 sequences of env has argued for site-specific amino acid frequency being a predictor of viral fitness (15).
We have therefore considered three explanations for the continued presence of certain CTL epitopes: (i) the corresponding CTLs did not impose significant immune pressure, contributing little to the control of viral infection; (ii) viral escape from these CTLs posed large fitness costs and thus little net selective advantage for the virus; and (iii) flanking site mutations contributed to the escape from recognition, possibly by preventing appropriate processing and presentation on HLA molecules. We addressed these issues in the current study, extending our analysis of one male subject comprehensively studied over the first 4 years of his HIV-1 infection (43). CTL responses were elicited in this subject against at least 25 epitopes, with continuous accumulation of escape mutations observed in 15. Two other epitopes were largely replaced by mutants, although the mutations were not confirmed to confer escape, and eight epitopes remained at high frequencies. Our studies suggest that the balance between the advantage of escaping from immune responses and the disadvantage of losing replication fitness can play an important role in determining whether a CTL epitope will persist or become lost during infection.
MATERIALS AND METHODS
Study subject and characterizations.
Clinical progression and virologic and immunologic characterizations of subject PIC1362 were reported previously (13, 43). He received no antiretroviral therapy throughout the course of the study. His HLA class I alleles, determined by sequence-specific primer molecular typing, are A*0201, A*2501, B*1801, B*5101, C*0102, and C*1203. The University of Washington institutional review board approved the study, and the subject provided written consent for participation prior to enrollment.
Here we extend the analysis of sequences of plasma virus and gamma interferon (IFN-γ) enzyme-linked immunospot (ELISPOT) assay measurements of host CTL responses to include blood samples from days 1245 and 1501 after onset of acute symptoms, in addition to the data reported previously (43). Briefly, viral sequences were derived from 5′ and 3′ half genomes from day 8 and from whole genomes from days 826 and 1245 after the onset of acute symptoms. Targeted gene fragments—gag-p17, gag-p24, pol-RT, vpr-tat, vpu-V2 (env), env-C2-V5, env-gp41, and nef—were sequenced from an additional 11 time points (days 22, 50, 76, 113, 155, 190, 344, 581, 769, 1035, and 1501), as were a gag-p24 fragment from days 414 and 491 and a gag-p17 fragment from days 491 and 667. Sequences used in this study correspond to accession numbers DQ853426 to DQ854622 in GenBank.
CTL epitopes spanning the entire HIV-1 proteome were identified by IFN-γ ELISPOT assays using peripheral blood mononuclear cells (PBMC) previously cryopreserved from 19 time points (days 8, 22, 29, 34, 50, 155, 190, 259, 304, 344, 491, 496, 680, 769, 826, 829, 1035, 1245, and 1501), again with data from the last two time points being reported for the first time here. Peptides based on HIV-1HXB2, HIV-1MN, the 2001 HIV-1 subtype B consensus (37, 43), and a subset of autologous viral sequences were used in IFN-γ ELISPOT experiments. The functional avidity of each recognized epitope and the mutant forms of these epitopes detected over time was measured by the effective concentration of peptide that elicited 50% of the peak IFN-γ response in the ELISPOT assay (EC50). Mutant peptides with EC50 values at least 10-fold higher than those of the corresponding epitopic forms were designated escape mutants.
Construction of chimeric viruses and growth competition experiments.
The p24 coding region was PCR amplified from PBMC from day 289 after the onset of acute symptoms and cloned into plasmid pNL4-3, using the yeast recombination system described previously (50). The p24 sequence from clone 298-3 from this subject, referred to as p24298-3, was identical to the nearly homogeneous viral sequence present 8 days after the onset of symptoms of acute HIV-1 infection in PIC1362. p24298-3 was inserted into plasmids pNL4-3VifA and pNL4-3VifB (45) to generate chimeric HIV-1 molecular clones. pNL4-3VifA and pNL4-3VifB have six synonymous nucleotide differences in vif to facilitate later differential recognition by hybridization. Mutant versions of p24298-3 bearing single or double mutations of E75D and V83L were generated by site-directed mutagenesis and then inserted into plasmid pNL4-3VifA. Purified plasmids were used to transfect 293 T cells. Supernatants were harvested after 72 h and used to infect U87.CD4-CXCR4 cells. Supernatants from these infections were harvested at the peak of RT activity, passed through 22-μm filters (Millipore), and used as virus stocks. Viruses were titrated on phytohemagglutinin (PHA)- and interleukin-2 (IL-2)-stimulated PBMC (56). Chimeric viruses bearing the mutant p24 were competed against virus bearing the original p24 of the patient, as well as standard SI/X4 reference HIV-1 isolates: E6 (CMU06) and A8 (92UG029), obtained from the AIDS Research and Reference Reagent Program, and C08 (TCCD08 in reference 29). Virus was added alone (mono-infection) or in pairs (dual infection) to 2 × 105 PHA-IL-2-treated PBMC at a multiplicity of infection of 0.0005 in a 48-well plate and incubated for 10 days. For dual infections, the initial ratio of the two virus strains was 1:1.
PCR and HTA.
For all virus-infected cultures, cell-associated viral DNA was extracted from lysed PBMC. For competition experiments between viruses bearing the chimeric mutant and the original p24 coding sequences, vif regions were amplified by nested PCR (R. M. Troyer, J. P. McNevin, Y. Liu, R. W. Krizan, A. Abraha, D. M. Tebit, H. Zhao, S. Avila, M. A. Lobritz, M. J. McElrath, J. I. Mullins, and E. J. Arts, unpublished data). For competition experiments between chimeric and standard SI/X4 reference virus isolates, env regions were amplified as previously described (56). Nested PCR products of vif and env were analyzed by a heteroduplex tracking assay (HTA) to determine the relative amounts of virus produced in the dual-competition experiments (56). Radiolabeled DNA probes were PCR-amplified from the regions of vif and env described above (50). Sense-strand nested primers were labeled with T4 polynucleotide kinase (Gibco BRL) and 2 μCi of [γ-32P]ATP, and probes were generated by PCR amplification followed by purification by agarose gel electrophoresis and a QIAquick gel extraction kit (QIAGEN). HTA reactions were carried out and heteroduplexes separated as described previously (16). Gels were scanned with a phosphorimager (Personal Molecular Imager FX; Bio-Rad) and bands analyzed using the Quantity One software package (Bio-Rad).
Estimation of viral fitness.
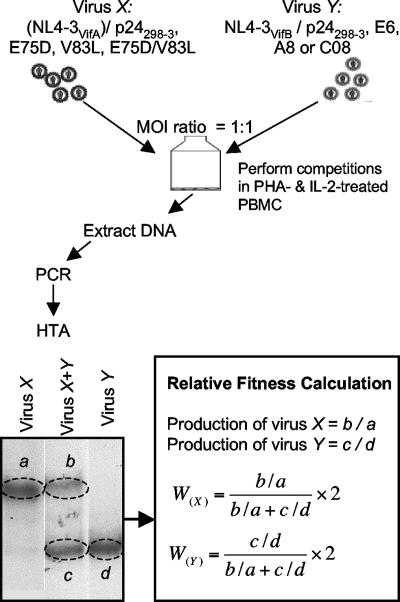
As shown in Fig. 1, the relative quantities of the two competing viruses (X and Y) produced from a dual infection were determined by HTA. To control for minor differences in probe binding efficiency, the virus production levels of dual and mono-infections were compared. Relative fitness (W) was defined as the production of an individual virus (b/a for virus X and c/d for virus Y) in a dual infection, divided by total viral production (b/a + c/d) and its initial proportion (0.5 in this study) in the inoculum (56). Therefore, the sum of the relative fitnesses of the two viruses is 2.
FIG. 1.
Schematic representation of the protocol used for fitness determination. Viruses were added to PHA- and IL-2-stimulated PBMC alone (mono-infection) or in pairs in equal amounts (dual infection), and viral production was monitored by HTA. X viruses are chimeric viruses with an original p24 (p24298-3) or p24 mutants with mutations of E75D, V83L, or E75D/V83L in plasmid pNL4-3VifA. Y viruses are chimeric viruses with p24298-3 in plasmid pNL4-3VifB or standard SI/X4 reference HIV-1 isolates (E6, A8, or C08). To control for minor differences in probe binding efficiency, the virus production levels of dual infections and mono-infections were compared. In the mono-infections, the production of virus X was a and the production of virus Y was d. In dual infections, the production of X was b/a and the production of Y was c/d. Relative fitness (W) was defined as the production of an individual virus strain (b/a or c/d) in a dual infection divided by total viral production (b/a + c/d) and its initial proportion (0.5 here) in the inoculum. The equations for the relative fitnesses of viruses X and Y are shown. MOI, multiplicity of infection.
Calculation of database frequencies of amino acids.
HIV-1 subtype B sequences, one sequence per individual, were retrieved at random from the HIVDB (38) as previously described (43). For each amino acid position within an epitope, paa, the database frequency of each amino acid in the position was counted, and its Shannon entropy score (65) was calculated using the equation −Σpaa log paa. Gaps were treated as a 21st-character state. If an amino acid was not observed at a site, we defined paa log paa as 0.
RESULTS
Evolution within CTL epitopes is associated with immunological escape.
Twenty-five CTL epitopes were identified in this subject over the first 4 years of infection (Table 1). Seventeen of them underwent evolution, and mutant forms of 15 that escaped or partially escaped CTL recognition (43) were detected. CTLs in responding to the mutant forms of the other two peptides (QAISPRTLNAW and HAPWDVNDL) had lower functional avidity, but the avidity did not reach the 10-fold lower level of that for the corresponding epitopic forms and the peptides were therefore not designated escape mutants by our criteria (43). Among the eight persistent epitopes, only one [ETINEEAAEW, p24 positions 71 to 80] developed a transient escape mutation (Fig. 2A). Thus, evolution within epitopes was strongly associated with immunological escape (P = 0.0005, Fisher's exact test).
TABLE 1.
HIV-1-specific CTL epitopes and escape mutants in PIC1362
| Epitope sequencea | HIV-1 protein (epitope position [aa])b | HLA restriction | Escape mutant sequencec | EC50 ratiod | Subtype B HIVDB frequency (%)e
|
|
|---|---|---|---|---|---|---|
| Epitope | Escape mutant | |||||
| Persistent epitopes | ||||||
| VIPMFSAL | p24 (36-43) | Cw*0102 | 84 | |||
| ETINEEAAEW | p24 (71-80) | A*2501 | ETINDEAAEW | 27 | 87 | 1 |
| FRDYVDRFYK | p24 (161-170) | B*1801 | 92 | |||
| EKEGKISKI | RT (42-50) | B*5101 | 87 | |||
| LPPVVAKEI | INT (28-36) | B*5101 | 72 | |||
| IYETYGDTW | Vpr (46-54) | A*2501 | 87 | |||
| RAIEAQQHL | gp41 (46-54) | B*5101 | 73 | |||
| YPLTFGWCF | Nef (135-143) | B*1801 | 49 | |||
| Epitopes whose frequencies decreased | ||||||
| QAISPRTLNAW | p24 (13-23) | A*2501 | 54 | |||
| NSPTRREL | p6Pol (35-42) | Cw*0102 | NSPTSPTRREL | F | 39 | 1 |
| TSPTRREL | F | 1 | ||||
| TAFTIPSI | RT (128-135) | B*5101 | TAFTIPST | >100 | 72 | 13 |
| TAFTIPSR | >100 | 1 | ||||
| TAFTIPSV | >100 | 11 | ||||
| NNETPGVRY | RT (136-144) | B*1801 | NNGTPGVRY | 22 | 5 | 0 |
| ELRQHLLRW | RT (204-212) | A*2501 | ELRQHLLKW | F | 46 | 26 |
| EAVRHFPRI | Vpr (29-37) | B*5101 | EAVRHFPRT | 43 | 33 | 7 |
| EAVRHFPRL | 18 | 3 | ||||
| HAPWDVNDL | Vpu (74-82) | Cw*0102 | 7 | |||
| PVDPRLEPW | Tat (3-11) | A*2501 | PVDPRLDPW | F | 81 | 0 |
| PVDPSLEPW | F | 9 | ||||
| PVDPKLEPW | F | 3 | ||||
| CCFHCQVC | Tat (30-37) | Cw*1203 | CCLHCQVC | F | 78 | 13 |
| CCFHCQSC | F | 0 | ||||
| AENLWVTVY | gp120 (31-39) | B*1801 | TEDLWVTVY | F | 6 | 0 |
| EEDLWVTVY | F | 0 | ||||
| AEDSWVTVY | F | 0 | ||||
| YETEVHNVW | gp120 (61-69) | B*1801 | YGTEVHNVW | 22 | 3 | 1 |
| YDTEVHNVW | 13 | 52 | ||||
| YDTEAHNVW | >83 | 8 | ||||
| NVTENFNMW | gp120 (88-96) | A*2501 | NVTEDFDMW | F | 72 | 0 |
| NVTEEFDMW | F | 0.5 | ||||
| NVTESFDMW | F | 0 | ||||
| YCAPAGFAIL | gp120 (217-226) | Cw*0102 | YCAPAGFAII | F | 60 | 2 |
| EIIGDIRQAY | gp120 (322-330) | A*2501 | QIIGDIRQAY | >200 | 1 | 0.5 |
| DIIGDIRQAH | 34 | 14 | ||||
| EIIGNIRQAH | 15.9 | 4 | ||||
| EIIGDIRQAH | F | 21 | ||||
| TLSQIVTKL | gp120 (341-349) | A*0201 | TLSKIVTKL | F | 0 | 0 |
| YSPLSLQTL | gp41 (201-209) | Cw*0102 | YSPLSLQTR | F | 2 | 6 |
| WSKSSIIGW | Nef (5-13) | A*2501 | WSKTSIIGW | F | 0.4 | 0 |
| WSKSSMIGW | F | 2 | ||||
Amino acid sequences of identified CTL epitopes are shown, with the amino acid sites that potentially experienced positive selection (43) underlined. Epitope sequences are identical to the autologous viral sequences obtained from the first time point, 8 days after the onset of acute symptoms.
HIV-1HXB2 amino acid (aa) numerations are used.
Sequences of mutant forms of identified CTL epitopes are shown, with mutated amino acids underlined.
The EC50 ratio is the EC50 of the mutant form over the EC50 of the epitope. The EC50 was measured at the time point when the epitope elicited peak CTL responses. EC50s are based on peptide titration and the generation of a dose-response curve. F, full escape (i.e., CTL recognition of the mutant form was not detected). Other mutant forms are partial (EC50 ratio ≥ 10) or not (EC50 ratio < 10) escape mutants.
Representation of the epitopes and their escape mutants in HIV-1 subtype B sequences in the LANL HIVDB (38).
FIG. 2.
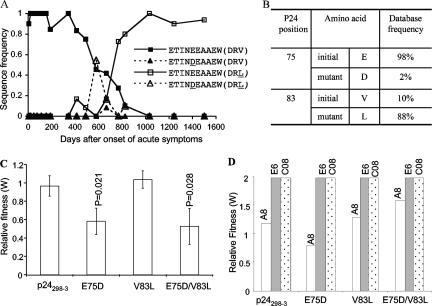
Characterization of the ETINEEAAEW p24 epitope (EW10). (A) Frequencies of occurrence of the epitope, its escape mutant, and mutants with a mutation C terminal to the epitope. Amino acids C terminal to the epitope are enclosed within parentheses. (B) Database frequencies of the initial and selected mutations of interest. (C) Relative fitnesses of chimeric viruses, all within the pNL4-3VifA backbone, containing either the original p24 coding sequence (p24298-3, the epitopic form from day 8 after the onset of symptoms of primary HIV-1 infection) or a p24 mutant: the E75D mutant (escape mutant), the V83L mutant (with a change 3 amino acids C terminal to the epitope), or the E75D V83L mutant (double mutant). (D) Relative fitnesses of p24298-3, E75D, V83L, and E75D V83L chimeric viruses, competed against laboratory strains A8, E6, and C08.
Magnitude and functional avidities of CTLs were not significantly associated with the occurrence of epitope mutations.
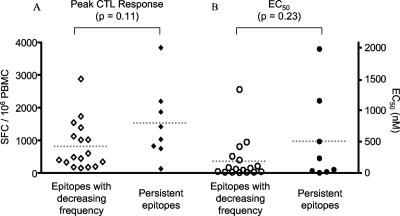
As shown in Fig. 3A, peak IFN-γ ELISPOT levels elicited by evolving epitopes trended lower but were not significantly different from those elicited by epitopes that persisted (P = 0.11, Mann-Whitney test). The functional avidities (EC50s) of the two groups were not significantly different either (Fig. 3B) (P = 0.23, Mann-Whitney test). Thus, neither the magnitude of the ELISPOT response nor the functional avidity of CTLs appeared to play a strong role in the occurrence of CTL epitope evolution.
FIG. 3.
Comparison of peak CTL responses and functional avidities between epitopes whose frequencies persisted or decreased during the infection of PIC1362. Peak CTL responses correspond to the peak levels of IFN-γ in ELISPOT assays detected over the 4-year time period examined (43). Functional avidity is represented by EC50s. Dotted horizontal lines representmean values. SFC, spot-forming cells.
Most CTL escape mutants are rare in the HIVDB.
As shown in Table 1, the epitopic forms were generally more frequently observed than the escape mutants in HIV-1 subtype B sequences in the HIVDB. To better infer the potential effect of CTL escape mutations on viral replication fitness, we compared the HIV-1 subtype B database frequencies of amino acids at positions within their epitopic and CTL escape mutant forms. Of the 15 epitopes that accumulated escape mutations in this subject, 7 differed from the subtype B consensus sequence. Unlike other epitopes that developed escape mutations via amino acid substitutions, the NSPTRREL epitope of p6Pol (positions 35 to 42) developed escape mutations by the insertion of three amino acids, Ser, Pro, and Thr, between positions 4 and 5 (43). The NSPTRREL epitope was not included in the following comparisons. Of the 24 amino acid sites associated with escape that were analyzed (43), 13 were located within seven epitopes with sequences different from the subtype B consensus sequence. Therefore, this analysis based on database frequency is not significantly biased by epitopes being exclusively “consensus-like.”
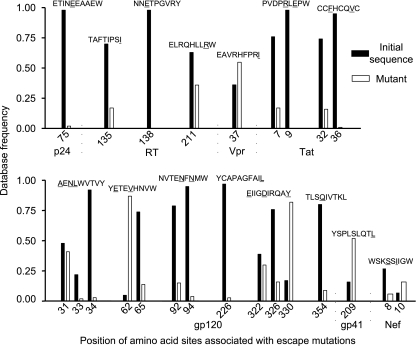
As shown in Fig. 4, the mutant form containing 19 of the 24 amino acid sites that determined escape occurred at a lower frequency in the database. In 14 cases, the amino acid in the initial epitopic form was found at a database frequency of >50%, whereas the sum of the frequencies of all the observed mutant forms was less than half the frequency of the initial form. Overall, database frequencies of the mutant-form amino acids were significantly lower than the frequencies of the epitopic-form amino acids (P = 0.0015, Wilcoxon signed-rank test). We therefore inferred that most of the CTL escape mutations in our study incurred fitness costs, and we made the logical assumption that in the absence of other selective pressures, HIV-1 is more likely to conserve sequences that improve replication efficiency.
FIG. 4.
Database frequencies of amino acids at sites associated with CTL escape mutations in epitopic or mutant forms. When multiple mutations were observed for a site, the database frequency of the mutant forms is shown, corresponding to the sum of the frequency of each mutation. Wild-type epitopes are presented above the columns. Underlined amino acids indicate sites at which mutations associated with CTL escape were demonstrated.
Fitness cost associated with an escape mutation within a p24 epitope.
The frequency of p24 epitope EW10 (ETINEEAAEW) decreased to ∼50% of the viral population around day 600 (Fig. 2A) as the ETINDEAAEW escape mutant emerged. (Underlined letters indicate mutated residues throughout.) However, the mutant did not persist and was replaced by the original epitopic form within another 200 days. Examining the database frequencies of the mutant and epitopic forms, we found that glutamic acid (E) in the epitopic form was present in 98% of subtype B sequences (Fig. 2B). Therefore, the short-lived appearance of the mutant form may have been due to a high fitness cost imposed by the E75D mutation. Interestingly, a mutation flanking this epitope, V83L, was present, and its occurrence reached 100% by day 1035 as the EW10 epitopic sequence again became nearly fixed in the viral population (Fig. 2A). Given that mutations flanking CTL epitopes can impair proteolytic processing and lead to escape (4, 18, 64), we hypothesized that the V83L mutation might mediate CTL escape without compensating for the impaired viral replication fitness conferred by the E75D mutation.
To address the above hypothesis, we assessed the fitness impact of both the E75D and V83L mutations in growth competition assays with PBMC. Recombinant viruses were generated with the p24 coding sequence reflecting the early consensus sequence in PIC1362. Synonymous nucleotide changes were introduced into the vif gene to allow distinction between strains by DNA hybridization (Fig. 1); these synonymous differences had no impact on viral fitness (Fig. 2C). We then introduced amino acid changes into the p24 coding sequence corresponding to E75D, V83L, and E75D/V83L. We found that V83L had no effect or conferred slightly higher fitness (Fig. 2C); however, E75D- and E75D/V83L-containing viruses were significantly less fit (P = 0.021 and P = 0.028, respectively). Thus, the E75D CTL escape mutation conferred a significant negative impact on viral fitness that was not reversed by the V83L mutation. Since the CTL response targeting this epitope also declined following the V83L mutation becoming dominant (data not shown), the data are consistent with the V83L mutation conferring a processing defect, allowing the E75D mutation to revert back to the epitopic form without further recognition by CTLs.
When chimeric viruses containing the original or mutant p24 EW10 were competed against SI/X4 reference HIV-1 isolates (Fig. 2D), only the E75D virus had lower fitness, and only to reference strain A8. All p24 chimeric viruses out-competed HIV-1 isolates E6 and C08. Furthermore, we could not detect differences in replication kinetics between the viruses containing the original and mutant p24s in mono-infections (data not shown). Thus, the decreases in fitness due to these CTL escape mutations were subtle, requiring competition against autologous viruses to discern significant fitness differences.
Sites of CTL escape mutations are associated with high variability.
To explore the relationship between fitness constraints conferred by amino acid changes within a CTL epitope, we calculated the Shannon entropy of each amino acid from its level of conservation in the HIVDB; lower Shannon entropies indicate lower variability and stronger hypothesized fitness constraints. We defined the entropy of an epitope as the mean of the entropies of each amino acid site in the epitope. For example, for p24 epitope VIPMFSAL, the entropies of the individual amino acid sites are 0.035, 0.018, 0, 0, 0, 0.240, 0.018, and 0, respectively; thus, the entropy of the epitope is given as 0.039.
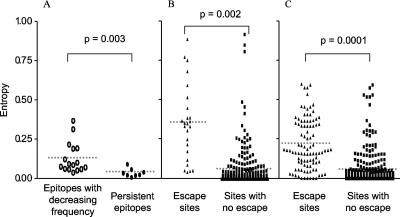
As shown in Fig. 5A, evolving epitopes within PIC1362 had significantly higher entropies (P = 0.003, Mann-Whitney test) than persistent epitopes. Across all epitopes, sites associated with CTL escape had significantly higher entropies than sites not associated with CTL escape (P = 0.002, Mann-Whitney test) (Fig. 5B). This association was also found when we compared the Shannon entropies of all 42 CTL epitopes defined in the HIV immunology database (34) for which escape mutations have been documented (Fig. 5C) (P < 0.001, Mann-Whitney test). Two of the epitopes found within the database were recognized in PIC1362, both of which developed escape mutations.
FIG. 5.
Comparison of entropies of epitopes with decreasing frequencies and epitopes that persisted in PIC1362 (A), sites associated and not associated with CTL escape in PIC1362 (B), and sites associated and not associated with CTL escape mutations in the HIV immunology database (C). Dotted horizontal lines represent mean values.
Analysis of mutations at sites of high entropy in persistent epitopes.
The upper three quartiles of the sites associated with CTL escape mutations had entropies greater than 0.17. However, only 3 of the 73 amino acid sites (4%) encompassing the eight persistent epitopes, each in different epitopes, had entropies greater than 0.17. These results suggest that escape mutations are more likely to occur within sites of high variability and infer relatively low fitness costs. By contrast, variations at sites that impart stringent functional constraints are less likely to occur, so that such epitopes, persisting as initially detected, remain at high frequencies.
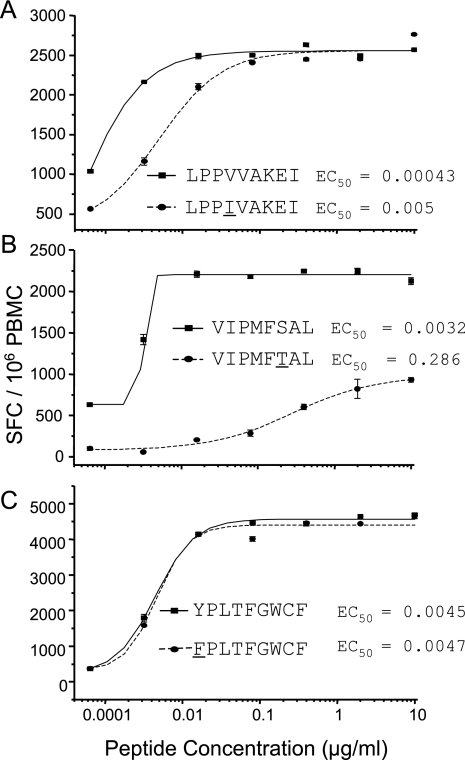
The three persistent epitopes containing sites with entropies greater than 0.17 were VIPMFSAL in p24, LPPVVAKEI in IN, and YPLTFGWCF in Nef. The underlined amino acids had database entropies of 0.24, 0.25, and 0.34, respectively. However, the frequencies of all three of the 8- to 9-amino-acid epitopes remained over 75% in this subject throughout the course of study, and each epitopic sequence represented the database consensus sequence, with each amino acid being found in the HIVDB at frequencies over 70%. We then assessed whether certain mutations at these higher entropy sites would be able to mediate escape by examining CTL responses in this subject against peptides corresponding to the mutants most frequently found in the database (38). The LPPIVAKEI mutant was detected in the subject in 1/10 clones from day 826, and the VIPMFTAL mutant was detected at days 1245 and 1501 in 1/10 and 2/16 clones, respectively. The EC50s of the LPPIVAKEI and VIPMFTAL mutants were 11- and 88-fold higher, respectively, than the EC50s of the corresponding epitopic peptides (Fig. 6A and B). Thus, these two mutant peptides may have the capacity to mediate low levels of escape and emerge over time (43). The FPLTFGWCF Nef mutant peptide, however, was not detected in this subject, and the EC50 that we determined was similar to that of the epitopic form (Fig. 6C). Hence, this mutation would have been unlikely to mediate CTL escape.
FIG. 6.
CTL responses to persistent epitopes containing sites of high entropy and their most frequent mutants found in the HIVDB. (A) LPPVVAKEI IN epitope and the LPPIVAKEI mutant. (B) VIPMFSAL p24 epitope and the VIPMFTAL mutant. (C) YPLTFGWCF Nef epitope and the FPLTFGWCF mutant.
DISCUSSION
We found two main patterns of CTL epitope evolution during the first 4 years of one male subject's HIV-1 infection: within two-thirds of the detected epitopes, escape mutants emerged to replace the susceptible epitope in the virus population over time, while for other epitopes, little genetic variation was observed and the initial sequence remained dominant. In our study subject, PIC1362, as well as in the HIVDB (34), amino acid sites associated with CTL escape mutations were more variable than those not associated with escape. Conversely, persistent epitopes lacked sites that were highly varied in the HIVDB. In three cases, persistent epitopes had amino acid sites of some variability; however, the cross-reactivities of CTL responses to common mutant forms were noted to different degrees. This cross-reactivity may have prevented the emergence of the mutant.
Immune pressure from CTLs is an important driving force for the evolution of HIV-1, and the accumulation of escape mutants leads to a loss of epitopes in the individual's viral population (3, 9, 43). The persistence of some epitopes has been hypothesized to be due to insufficient antiviral pressure from the corresponding CTL responses (33). However, we did not find persistent epitopes to be associated with lower-magnitude CD8+ T-cell responses or lower functional avidities as measured by IFN-γ ELISPOT assays. Others have also reported that IFN-γ production (42, 58) and the functional avidity (6, 62) of IFN-γ-secreting CTLs may not reflect their antiviral effect (46). Therefore, further studies are required to evaluate the antiviral effect of evolving and persistent epitopes.
Although CTL responses positively select for escape mutants, they may exact fitness costs on viral replication, as shown here for the EW10 epitope. It is interesting to note that most (79%) escape mutations that we observed involved substitutions of amino acids with less database representation. This suggests that database frequency, a surrogate marker of survival in the population, and viral fitness are strongly linked. For escape mutants to accumulate, the selective advantage to HIV-1 must outweigh the replication fitness cost. Thus, it is more likely to detect escape mutations at sites of high variability, where substitutions are less likely to exact a substantial fitness cost. It should be noted, however, that as we have shown here, the fitness costs of amino acid substitutions may be subtle and detectable only when competed against autologous virus strains, as opposed to reference laboratory isolates.
HIV-1 uses complex mechanisms to evade CTL responses against conserved epitopes. As shown here for the analysis of the EW10 epitope, the escape mutation, which had a distinct fitness disadvantage in the context of the autologous protein, was short-lived. Simultaneously, a C-terminal mutation was positively selected for and became fixed in the viral population, followed by the epitope escape mutation reverting back to the susceptible form. We infer that, as has been shown for other epitopes (4, 18, 64), the C-terminal mutation impaired epitope processing without incurring a further fitness cost. Consistent with this observation, the CTL response against this epitope was lost over the following year, despite reversion to the susceptible form. It is interesting to note that the HIVDB frequency of the protective amino acid substitution 83L is quite high, suggesting that this epitope might be shielded from recognition in most circulating viruses.
Three persistent epitopes, the YPLTFGWCF, VIPMFSAL, and LPPVVAKEI epitopes, had amino acid sites of considerable variability. For the YPLTFGWCF epitope, however, the most frequent mutations in the database were not detected in this subject, and we found that the EC50 of the mutant was similar to that of the epitopic form. Hence, such a mutant is unlikely to be able to mediate CTL escape and consequently is unable to accumulate to a detectable level. In contrast, the VIPMFSAL and LPPVVAKEI peptides, the most frequent mutants found in the database, were detected in the subject at later time points, and the EC50s of these mutants exhibited over 10-fold increases. Therefore, these mutants correspond to escape mutants by our definition. Taking all three epitopes into account, the suppressive impact on mutation is negatively correlated with the level of cross-reactivity and the likelihood of emergence of the escape mutant.
It has also been shown that the occurrence and accumulation of CTL escape mutants can significantly be affected by stochastic events (44). Thus, compared to other epitopes in this subject, it might take longer for the two last-mentioned epitopes to be replaced by escape mutants. In addition, it should be noted that our definition of an escape mutant, based on changes in its EC50, is arbitrary, not based on functional testing. An in vitro study (6) has shown that there is an avidity threshold for specific CTLs, below which the ability to kill infected cells rapidly drops to none and above which killing efficiency rapidly plateaus. Therefore, for different epitopes, differences in EC50s do not necessarily reflect differences in the capacity of the epitopes to elicit antiviral CTL responses. The differences in avidity threshold between epitopes also suggest that for some specific CTLs, epitopic variants with small changes in avidity near the threshold could completely ablate antiviral activity but that for other CTLs, even a large change in avidity further from the threshold could have little effect. Therefore, although the most frequent mutants found in the database for the VIPMFSAL and LPPVVAKEI epitopes may be true CTL escape mutants, we could not rule out the possibility that the avidities of the epitopic forms were far from the threshold, and thus, the mutants could not escape CTL responses.
Finally, we found the LPPVVAKEI epitope to elicit a class I B51-restricted CD8+ CTL response, resulting in the emergence of LPPIVAKEI as a partial escape mutation. However, in another study, the LPPVVAKEI peptide was found to mediate escape from B51-restricted immune responses against the LPPIVAKEI peptide (39). This Ile is relatively frequent in the database (25%), suggesting that this partial escape is associated with only a weak impact on fitness.
In summary, we provide evidence that CTL epitope evolution is determined by the balance between the benefit of immune escape and the disadvantage of subsequent replication fitness cost. Therefore, our results support approaches to design HIV vaccines that can elicit potent antiviral CTLs that restrict HIV replication (24, 51) and emphasize the importance of targeting epitopes with strong fitness constraints.
Acknowledgments
We thank Peter B. Gilbert for statistical assistance and Indira Genowati for technical assistance.
This work was supported by grants from the U.S. Public Health Services, including AI57005 for the Seattle Primary Infection Program, R37-AI47734, AI27757 for the University of Washington Center for AIDS Research, and M01-RR-00037 for leukapheresis.
Footnotes
Published ahead of print on 29 August 2007.
REFERENCES
- 1.Addo, M. M., X. G. Yu, A. Rathod, D. Cohen, R. L. Eldridge, D. Strick, M. N. Johnston, C. Corcoran, A. G. Wurcel, C. A. Fitzpatrick, M. E. Feeney, W. R. Rodriguez, N. Basgoz, R. Draenert, D. R. Stone, C. Brander, P. J. R. Goulder, E. S. Rosenberg, M. Altfeld, and B. D. Walker. 2003. Comprehensive epitope analysis of human immunodeficiency virus type 1 (HIV-1)-specific T-cell responses directed against the entire expressed HIV-1 genome demonstrate broadly directed responses, but no correlation to viral load. J. Virol. 77:2081-2092. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Alexander-Miller, M. A., K. C. Parker, T. Tsukui, C. D. Pendleton, J. E. Coligan, and J. A. Berzofsky. 1996. Molecular analysis of presentation by HLA-A2.1 of a promiscuously binding V3 loop peptide from the HIV-envelope protein to human cytotoxic T lymphocytes. Int. Immunol. 8:641-649. [DOI] [PubMed] [Google Scholar]
- 3.Allen, T. M., M. Altfeld, S. C. Geer, E. T. Kalife, C. Moore, K. M. O'Sullivan, I. DeSouza, M. E. Feeney, R. L. Eldridge, E. L. Maier, D. E. Kaufmann, M. P. Lahaie, L. Reyor, G. Tanzi, M. N. Johnston, C. Brander, R. Draenert, J. K. Rockstroh, H. Jessen, E. S. Rosenberg, S. A. Mallal, and B. D. Walker. 2005. Selective escape from CD8+ T-cell responses represents a major driving force of human immunodeficiency virus type 1 (HIV-1) sequence diversity and reveals constraints on HIV-1 evolution. J. Virol. 79:13239-13249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Allen, T. M., M. Altfeld, X. G. Yu, K. M. O'Sullivan, M. Lichterfeld, S. Le Gall, M. John, B. R. Mothe, P. K. Lee, E. T. Kalife, D. E. Cohen, K. A. Freedberg, D. A. Strick, M. N. Johnston, A. Sette, E. S. Rosenberg, S. A. Mallal, P. J. R. Goulder, C. Brander, and B. D. Walker. 2004. Selection, transmission, and reversion of an antigen-processing cytotoxic T-lymphocyte escape mutation in human immunodeficiency virus type 1 infection. J. Virol. 78:7069-7078. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Allen, T. M., X. G. Yu, E. T. Kalife, L. L. Reyor, M. Lichterfeld, M. John, M. Cheng, R. L. Allgaier, S. Mui, N. Frahm, G. Alter, N. V. Brown, M. N. Johnston, E. S. Rosenberg, S. A. Mallal, C. Brander, B. D. Walker, and M. Altfeld. 2005. De novo generation of escape variant-specific CD8+ T-cell responses following cytotoxic T-lymphocyte escape in chronic human immunodeficiency virus type 1 infection. J. Virol. 79:12952-12960. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bennett, M. S., H. L. Ng, M. Dagarag, A. Ali, and O. O. Yang. 2007. Epitope-dependent avidity thresholds for cytotoxic T-lymphocyte clearance of virus-infected cells. J. Virol. 81:4973-4980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bhattacharya, T., M. Daniels, D. Heckerman, B. Foley, N. Frahm, C. Kadie, J. Carlson, K. Yusim, B. McMahon, B. Gaschen, S. Mallal, J. I. Mullins, D. C. Nickle, J. Herbeck, C. Rousseau, G. H. Learn, T. Miura, C. Brander, B. Walker, and B. Korber. 2007. Founder effects in the assessment of HIV polymorphisms and HLA allele associations. Science 315:1583-1586. [DOI] [PubMed] [Google Scholar]
- 8.Borrow, P., H. Lewick, B. H. Hahn, G. M. Shaw, and M. B. A. Oldstone. 1994. Virus-specific CD8+ cytotoxic T-lymphocyte activity associated with control of viremia in primary human immunodeficiency virus type 1 infection. J. Virol. 68:6103-6110. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Borrow, P., H. Lewicki, X. Wei, M. S. Horwitz, N. Peffer, H. Meyers, J. A. Nelson, J. E. Gairin, B. H. Hahn, M. B. Oldstone, and G. M. Shaw. 1997. Antiviral pressure exerted by HIV-1-specific cytotoxic T lymphocytes (CTLs) during primary infection demonstrated by rapid selection of CTL escape virus. Nat. Med. 3:205-211. [DOI] [PubMed] [Google Scholar]
- 10.Brander, C., K. E. Hartman, A. K. Trocha, N. G. Jones, R. P. Johnson, B. Korber, P. Wentworth, S. P. Buchbinder, S. Wolinsky, B. D. Walker, and S. A. Kalams. 1998. Lack of strong immune selection pressure by the immunodominant, HLA-A*0201-restricted cytotoxic T lymphocyte response in chronic human immunodeficiency virus-1 infection. J. Clin. Investig. 101:2559-2566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Buseyne, F., J. Le Chenadec, B. Corre, F. Porrot, M. Burgard, C. Rouzioux, S. Blanche, M. J. Mayaux, and Y. Riviere. 2002. Inverse correlation between memory gag-specific cytotoxic T lymphocytes and viral replication in human immunodeficiency virus-infected children. J. Infect. Dis. 186:1589-1596. [DOI] [PubMed] [Google Scholar]
- 12.Cao, J., J. McNevin, S. Holte, L. Fink, L. Corey, and M. J. McElrath. 2003. Comprehensive analysis of human immunodeficiency virus type 1 (HIV-1)-specific gamma interferon-secreting CD8+ T cells in primary HIV-1 infection. J. Virol. 77:6867-6878. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Cao, J., J. McNevin, U. Malhotra, and M. J. McElrath. 2003. Evolution of CD8+ T cell immunity and viral escape following acute HIV-1 infection. J. Immunol. 171:3837-3846. [DOI] [PubMed] [Google Scholar]
- 14.Crawford, H., J. G. Prado, A. Leslie, S. Hué, I. Honeyborne, S. Reddy, M. van der Stok, Z. Mncube, C. Brander, C. Rousseau, J. I. Mullins, R. Kaslow, P. Goepfert, S. Allen, E. Hunter, J. Mulenga, P. Kiepiela, B. D. Walker, and P. J. R. Goulder. 2007. Compensatory mutation partially restores fitness and delays reversion of escape mutation within the immunodominant HLA-B*5703-restricted gag epitope in chronic human immunodeficiency virus type 1 infection. J. Virol. 81:8346-8351. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.da Silva, J. 2006. Site-specific amino acid frequency, fitness and the mutational landscape model of adaptation in human immunodeficiency virus type 1. Genetics 174:1689-1694. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Delwart, E. L., H. W. Sheppard, B. D. Walker, J. Goudsmit, and J. I. Mullins. 1994. Human immunodeficiency virus type 1 evolution in vivo tracked by DNA heteroduplex mobility assays. J. Virol. 68:6672-6683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Derby, M., M. Alexander-Miller, R. Tse, and J. Berzofsky. 2001. High-avidity CTL exploit two complementary mechanisms to provide better protection against viral infection than low-avidity CTL. J. Immunol. 166:1690-1697. [DOI] [PubMed] [Google Scholar]
- 18.Draenert, R., S. Le Gall, K. J. Pfafferott, A. J. Leslie, P. Chetty, C. Brander, E. C. Holmes, S. C. Chang, M. E. Feeney, M. M. Addo, L. Ruiz, D. Ramduth, P. Jeena, M. Altfeld, S. Thomas, Y. Tang, C. L. Verrill, C. Dixon, J. G. Prado, P. Kiepiela, J. Martinez-Picado, B. D. Walker, and P. J. Goulder. 2004. Immune selection for altered antigen processing leads to cytotoxic T lymphocyte escape in chronic HIV-1 infection. J. Exp. Med. 199:905-915. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Draenert, R., C. L. Verrill, Y. Tang, T. M. Allen, A. G. Wurcel, M. Boczanowski, A. Lechner, A. Y. Kim, T. Suscovich, N. V. Brown, M. M. Addo, and B. D. Walker. 2004. Persistent recognition of autologous virus by high-avidity CD8 T cells in chronic, progressive human immunodeficiency virus type 1 infection. J. Virol. 78:630-641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Fernandez, C. S., I. Stratov, R. De Rose, K. Walsh, C. J. Dale, M. Z. Smith, M. B. Agy, S.-L. Hu, K. Krebs, D. I. Watkins, D. H. O'Connor, M. P. Davenport, and S. J. Kent. 2005. Rapid viral escape at an immunodominant simian-human immunodeficiency virus cytotoxic T-lymphocyte epitope exacts a dramatic fitness cost. J. Virol. 79:5721-5731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Frahm, N., S. Adams, P. Kiepiela, C. H. Linde, H. S. Hewitt, M. Lichterfeld, K. Sango, N. V. Brown, E. Pae, A. G. Wurcel, M. Altfeld, M. E. Feeney, T. M. Allen, T. Roach, M. A. St. John, E. S. Daar, E. Rosenberg, B. Korber, F. Marincola, B. D. Walker, P. J. R. Goulder, and C. Brander. 2005. HLA-B63 presents HLA-B57/B58-restricted cytotoxic T-lymphocyte epitopes and is associated with low human immunodeficiency virus load. J. Virol. 79:10218-10225. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Frahm, N., P. Kiepiela, S. Adams, C. H. Linde, H. S. Hewitt, K. Sango, M. E. Feeney, M. M. Addo, M. Lichterfeld, M. P. Lahaie, E. Pae, A. G. Wurcel, T. Roach, M. A. St. John, M. Altfeld, F. M. Marincola, C. Moore, S. Mallal, M. Carrington, D. Heckerman, T. M. Allen, J. I. Mullins, B. T. Korber, P. J. Goulder, B. D. Walker, and C. Brander. 2006. Control of human immunodeficiency virus replication by cytotoxic T lymphocytes targeting subdominant epitopes. Nat. Immunol. 7:173-178. [DOI] [PubMed] [Google Scholar]
- 23.Friedrich, T. C., E. J. Dodds, L. J. Yant, L. Vojnov, R. Rudersdorf, C. Cullen, D. T. Evans, R. C. Desrosiers, B. R. Mothe, J. Sidney, A. Sette, K. Kunstman, S. Wolinsky, M. Piatak, J. Lifson, A. L. Hughes, N. Wilson, D. H. O'Connor, and D. I. Watkins. 2004. Reversion of CTL escape-variant immunodeficiency viruses in vivo. Nat. Med. 10:275-281. [DOI] [PubMed] [Google Scholar]
- 24.Garber, D. A., and M. B. Feinberg. 2003. AIDS vaccine development: the long and winding road. AIDS Rev. 5:131-139. [PubMed] [Google Scholar]
- 25.Geels, M. J., M. Cornelissen, H. Schuitemaker, K. Anderson, D. Kwa, J. Maas, J. T. Dekker, E. Baan, F. Zorgdrager, R. van den Burg, M. van Beelen, V. V. Lukashov, T.-M. Fu, W. A. Paxton, L. van der Hoek, S. A. Dubey, J. W. Shiver, and J. Goudsmit. 2003. Identification of sequential viral escape mutants associated with altered T-cell responses in a human immunodeficiency virus type 1-infected individual. J. Virol. 77:12430-12440. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Goulder, P. J., R. E. Phillips, R. A. Colbert, S. McAdam, G. Ogg, M. A. Nowak, P. Giangrande, G. Luzzi, B. Morgan, A. Edwards, A. J. McMichael, and S. Rowland-Jones. 1997. Late escape from an immunodominant cytotoxic T-lymphocyte response associated with progression to AIDS. Nat. Med. 3:212-217. [DOI] [PubMed] [Google Scholar]
- 27.Hay, C. M., D. J. Ruhl, N. O. Basgoz, C. C. Wilson, J. M. Billingsley, M. P. DePasquale, R. T. D'Aquila, S. M. Wolinsky, J. M. Crawford, D. C. Montefiori, and B. D. Walker. 1999. Lack of viral escape and defective in vivo activation of human immunodeficiency virus type 1-specific cytotoxic T lymphocytes in rapidly progressive infection. J. Virol. 73:5509-5519. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jamieson, B. D., O. O. Yang, L. Hultin, M. A. Hausner, P. Hultin, J. Matud, K. Kunstman, S. Killian, J. Altman, K. Kommander, B. Korber, J. Giorgi, and S. Wolinsky. 2003. Epitope escape mutation and decay of human immunodeficiency virus type 1-specific CTL responses. J. Immunol. 171:5372-5379. [DOI] [PubMed] [Google Scholar]
- 29.Johnston, E. R., L. S. Zijenah, S. Mutetwa, R. Kantor, C. Kittinunvorakoon, and D. A. Katzenstein. 2003. High frequency of syncytium-inducing and CXCR4-tropic viruses among human immunodeficiency virus type 1 subtype C-infected patients receiving antiretroviral treatment. J. Virol. 77:7682-7688. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Jones, N. A., X. Wei, D. R. Flower, M. Wong, F. Michor, M. S. Saag, B. H. Hahn, M. A. Nowak, G. M. Shaw, and P. Borrow. 2004. Determinants of human immunodeficiency virus type 1 escape from the primary CD8+ cytotoxic T lymphocyte response. J. Exp. Med. 200:1243-1256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Kiepiela, P., A. J. Leslie, I. Honeyborne, D. Ramduth, C. Thobakgale, S. Chetty, P. Rathnavalu, C. Moore, K. J. Pfafferott, L. Hilton, P. Zimbwa, S. Moore, T. Allen, C. Brander, M. M. Addo, M. Altfeld, I. James, S. Mallal, M. Bunce, L. D. Barber, J. Szinger, C. Day, P. Klenerman, J. Mullins, B. Korber, H. M. Coovadia, B. D. Walker, and P. J. Goulder. 2004. Dominant influence of HLA-B in mediating the potential co-evolution of HIV and HLA. Nature 432:769-775. [DOI] [PubMed] [Google Scholar]
- 32.Kiepiela, P., K. Ngumbela, C. Thobakgale, D. Ramduth, I. Honeyborne, E. Moodley, S. Reddy, C. de Pierres, Z. Mncube, N. Mkhwanazi, K. Bishop, M. van der Stok, K. Nair, N. Khan, H. Crawford, R. Payne, A. Leslie, J. Frater, N. McCarthy, C. Brander, G. H. Learn, D. C. Nickle, C. Rousseau, H. Coovadia, J. I. Mullins, D. Heckerman, B. D. Walker, and P. Goulder. 2006. CD8+ T cell responses to different HIV proteins have discordant associations with viral load. Nat. Med. 13:46-53. [DOI] [PubMed] [Google Scholar]
- 33.Koibuchi, T., T. M. Allen, M. Lichterfeld, S. K. Mui, K. M. O'Sullivan, A. Trocha, S. A. Kalams, R. P. Johnson, and B. D. Walker. 2005. Limited sequence evolution within persistently targeted CD8 epitopes in chronic human immunodeficiency virus type 1 infection. J. Virol. 79:8171-8181. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Korber, B. T. M., C. Brander, B. F. Haynes, R. Koup, J. P. Moore, B. D. Walker, and D. I. Watkins (ed.). 2005. HIV molecular immunology 2005. LA-UR 06-0036. Theoretical Biology and Biophysics, Los Alamos National Laboratory, Los Alamos, NM.
- 35.Kostense, S., G. S. Ogg, E. H. Manting, G. Gillespie, J. Joling, K. Vandenberghe, E. Z. Veenhof, D. van Baarle, S. Jurriaans, M. R. Klein, and F. Miedema. 2001. High viral burden in the presence of major HIV-specific CD8+ T cell expansions: evidence for impaired CTL effector function. Eur. J. Immunol. 31:677-686. [DOI] [PubMed] [Google Scholar]
- 36.Koup, R. A., J. T. Safrit, Y. Cao, C. A. Andrews, G. McLeod, W. Borkowsky, C. Farthing, and D. D. Ho. 1994. Temporal association of cellular immune responses with the initial control of viremia in primary human immunodeficiency virus type 1 syndrome. J. Virol. 68:4650-4655. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Kuiken, C., B. Foley, E. Freed, B. Hahn, B. Korber, P. Marx, F. McCutchan, J. W. Mellors, and S. Wolinksy (ed.). 2002. HIV sequence compendium 2002. LA-UR 03-3564. Theoretical Biology and Biophysics Group, Los Alamos National Laboratory, Los Alamos, NM.
- 38.Leitner, T., B. Foley, B. Hahn, P. Marx, F. McCutchan, J. W. Mellors, S. Wolinsky, and B. Korber (ed.). 2006. HIV sequence compendium 2005. Theoretical Biology and Biophysics Group, Los Alamos National Laboratory, Los Alamos, NM.
- 39.Leslie, A., D. Kavanagh, I. Honeyborne, K. Pfafferott, C. Edwards, T. Pillay, L. Hilton, C. Thobakgale, D. Ramduth, R. Draenert, S. Le Gall, G. Luzzi, A. Edwards, C. Brander, A. K. Sewell, S. Moore, J. Mullins, C. Moore, S. Mallal, N. Bhardwaj, K. Yusim, R. Phillips, P. Klenerman, B. Korber, P. Kiepiela, B. Walker, and P. Goulder. 2005. Transmission and accumulation of CTL escape variants drive negative associations between HIV polymorphisms and HLA. J. Exp. Med. 201:891-902. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Leslie, A., D. A. Price, P. Mkhize, K. Bishop, A. Rathod, C. Day, H. Crawford, I. Honeyborne, T. E. Asher, G. Luzzi, A. Edwards, C. M. Rousseau, J. I. Mullins, G. Tudor-Williams, V. Novelli, C. Brander, D. C. Douek, P. Kiepiela, B. D. Walker, and P. J. Goulder. 2006. Differential selection pressure exerted on HIV by CTL targeting identical epitopes but restricted by distinct HLA alleles from the same HLA supertype. J. Immunol. 177:4699-4708. [DOI] [PubMed] [Google Scholar]
- 41.Leslie, A. J., K. J. Pfafferott, P. Chetty, R. Draenert, M. M. Addo, M. Feeney, Y. Tang, E. C. Holmes, T. Allen, J. G. Prado, M. Altfeld, C. Brander, C. Dixon, D. Ramduth, P. Jeena, S. A. Thomas, A. St. John, T. A. Roach, B. Kupfer, G. Luzzi, A. Edwards, G. Taylor, H. Lyall, G. Tudor-Williams, V. Novelli, J. Martinez-Picado, P. Kiepiela, B. D. Walker, and P. J. Goulder. 2004. HIV evolution: CTL escape mutation and reversion after transmission. Nat. Med. 10:282-289. [DOI] [PubMed] [Google Scholar]
- 42.Lim, D. G., K. Bieganowska Bourcier, G. J. Freeman, and D. A. Hafler. 2000. Examination of CD8+ T cell function in humans using MHC class I tetramers: similar cytotoxicity but variable proliferation and cytokine production among different clonal CD8+ T cells specific to a single viral epitope. J. Immunol. 165:6214-6220. [DOI] [PubMed] [Google Scholar]
- 43.Liu, Y., J. McNevin, J. Cao, H. Zhao, I. Genowati, K. Wong, S. McLaughlin, M. D. McSweyn, K. Diem, C. E. Stevens, J. Maenza, H. He, D. C. Nickle, D. Shriner, S. E. Holte, A. C. Collier, L. Corey, M. J. McElrath, and J. I. Mullins. 2006. Selection on the human immunodeficiency virus type 1 proteome following primary infection. J. Virol. 80:9519-9529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 44.Liu, Y., J. I. Mullins, and J. E. Mittler. 2006. Waiting times for the appearance of cytotoxic T-lymphocyte escape mutants in chronic HIV-1 infection. Virology 347:140-146. [DOI] [PubMed] [Google Scholar]
- 45.Lobritz, M. A., A. J. Marozsan, R. M. Troyer, and E. J. Arts. 2007. Natural variation in the V3 crown of human immunodeficiency virus type 1 affects replicative fitness and entry inhibitor sensitivity. J. Virol. 81:8258-8269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Loffredo, J. T., B. J. Burwitz, E. G. Rakasz, S. P. Spencer, J. J. Stephany, J. P. G. Vela, S. R. Martin, J. Reed, S. M. Piaskowski, J. Furlott, K. L. Weisgrau, D. S. Rodrigues, T. Soma, G. Napoé, T. C. Friedrich, N. A. Wilson, E. G. Kallas, and D. I. Watkins. 2007. The antiviral efficacy of simian immunodeficiency virus-specific CD8+ T cells is unrelated to epitope specificity and is abrogated by viral escape. J. Virol. 81:2624-2634. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Martinez-Picado, J., J. G. Prado, E. E. Fry, K. Pfafferott, A. Leslie, S. Chetty, C. Thobakgale, I. Honeyborne, H. Crawford, P. Matthews, T. Pillay, C. Rousseau, J. I. Mullins, C. Brander, B. D. Walker, D. I. Stuart, P. Kiepiela, and P. Goulder. 2006. Fitness cost of escape mutation in p24 Gag in association with control of human immunodeficiency virus type 1. J. Virol. 80:3617-3623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Meyerhans, A., G. Dadaglio, J. P. Vartanian, P. Langlade-Demoyen, R. Frank, B. Asjo, F. Plata, and S. Wain-Hobson. 1991. In vivo persistence of a HIV-1-encoded HLA-B27-restricted cytotoxic T lymphocyte epitope despite specific in vitro reactivity. Eur. J. Immunol. 21:2637-2640. [DOI] [PubMed] [Google Scholar]
- 49.Moore, C. B., M. John, I. R. James, F. T. Christiansen, C. S. Witt, and S. A. Mallal. 2002. Evidence of HIV-1 adaptation to HLA-restricted immune responses at a population level. Science 296:1439-1443. [DOI] [PubMed] [Google Scholar]
- 50.Moore, D. M., E. J. Arts, Y. Gao, and A. J. Marozsan. 2005. A yeast recombination-based cloning system to produce chimeric HIV-1 viruses and express HIV-1 genes. Methods Mol. Biol. 304:369-385. [DOI] [PubMed] [Google Scholar]
- 51.Mullins, J. I. 1997. Curtailing the AIDS pandemic. Science 276:1955-1957. (Letter.) [DOI] [PubMed] [Google Scholar]
- 52.Nietfield, W., M. Bauer, M. Fevrier, R. Maier, B. Holzwarth, R. Frank, B. Maier, Y. Riviere, and A. Meyerhans. 1995. Sequence constraints and recognition by CTL of an HLA-B27-restricted HIV-1 gag epitope. J. Immunol. 154:2189-2197. [PubMed] [Google Scholar]
- 53.O'Connor, D. H., T. M. Allen, T. U. Vogel, P. Jing, I. P. DeSouza, E. Dodds, E. J. Dunphy, C. Melsaether, B. Mothe, H. Yamamoto, H. Horton, N. Wilson, A. L. Hughes, and D. I. Watkins. 2002. Acute phase cytotoxic T lymphocyte escape is a hallmark of simian immunodeficiency virus infection. Nat. Med. 8:493-499. [DOI] [PubMed] [Google Scholar]
- 54.Ogg, G. S., X. Jin, S. Bonhoeffer, P. R. Dunbar, M. A. Nowak, S. Monard, J. P. Segal, Y. Cao, S. L. Rowland-Jones, V. Cerundolo, A. Hurley, M. Markowitz, D. D. Ho, D. F. Nixon, and A. J. McMichael. 1998. Quantitation of HIV-1-specific T lymphocytes and plasma viral load of viral RNA. Science 279:2103-2106. [DOI] [PubMed] [Google Scholar]
- 55.Price, D. A., P. J. Goulder, P. Klenerman, A. K. Sewell, P. J. Easterbrook, M. Troop, C. R. Bangham, and R. E. Phillips. 1997. Positive selection of HIV-1 cytotoxic T lymphocyte escape variants during primary infection. Proc. Natl. Acad. Sci. USA 94:1890-1895. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Quiñones-Mateu, M. E., S. C. Ball, A. J. Marozsan, V. S. Torre, J. L. Albright, G. Vanham, G. van der Groen, R. L. Colebunders, and E. J. Arts. 2000. A dual infection/competition assay shows a correlation between ex vivo human immunodeficiency virus type 1 fitness and disease progression. J. Virol. 74:9222-9233. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Safrit, J. T., C. A. Andrews, T. Zhu, D. D. Ho, and R. A. Koup. 1994. Characterization of human immunodeficiency virus type 1-specific cytotoxic T lymphocyte clones isolated during acute seroconversion: recognition of autologous virus sequences within a conserved immunodominant epitope. J. Exp. Med. 179:463-472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Snyder, J. E., W. J. Bowers, A. M. Livingstone, F. E. Lee, H. J. Federoff, and T. R. Mosmann. 2003. Measuring the frequency of mouse and human cytotoxic T cells by the Lysispot assay: independent regulation of cytokine secretion and short-term killing. Nat. Med. 9:231-235. [DOI] [PubMed] [Google Scholar]
- 59.Trachtenberg, E., B. Korber, C. Sollars, T. B. Kepler, P. T. Hraber, E. Hayes, R. Funkhouser, M. Fugate, J. Theiler, Y. S. Hsu, K. Kunstman, S. Wu, J. Phair, H. Erlich, and S. Wolinsky. 2003. Advantage of rare HLA supertype in HIV disease progression. Nat. Med. 9:928-935. [DOI] [PubMed] [Google Scholar]
- 60.Wolinsky, S. M., B. T. Korber, A. U. Neumann, M. Daniels, K. J. Kunstman, A. J. Whetsell, M. R. Furtado, Y. Cao, D. D. Ho, and J. T. Safrit. 1996. Adaptive evolution of human immunodeficiency virus-type 1 during the natural course of infection. Science 272:537-542. [DOI] [PubMed] [Google Scholar]
- 61.Yang, O. O., P. T. Sarkis, A. Ali, J. D. Harlow, C. Brander, S. A. Kalams, and B. D. Walker. 2003. Determinant of HIV-1 mutational escape from cytotoxic T lymphocytes. J. Exp. Med. 197:1365-1375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Yang, O. O., P. T. Sarkis, A. Trocha, S. A. Kalams, R. P. Johnson, and B. D. Walker. 2003. Impacts of avidity and specificity on the antiviral efficiency of HIV-1-specific CTL. J. Immunol. 171:3718-3724. [DOI] [PubMed] [Google Scholar]
- 63.Yang, O. O., and B. D. Walker. 1997. CD8+ cells in human immunodeficiency virus type I pathogenesis: cytolytic and noncytolytic inhibition of viral replication. Adv. Immunol. 66:273-311. [DOI] [PubMed] [Google Scholar]
- 64.Yokomaku, Y., H. Miura, H. Tomiyama, A. Kawana-Tachikawa, M. Takiguchi, A. Kojima, Y. Nagai, A. Iwamoto, Z. Matsuda, and K. Ariyoshi. 2004. Impaired processing and presentation of cytotoxic-T-lymphocyte (CTL) epitopes are major escape mechanisms from CTL immune pressure in human immunodeficiency virus type 1 infection. J. Virol. 78:1324-1332. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 65.Yusim, K., C. Kesmir, B. Gaschen, M. M. Addo, M. Altfeld, S. Brunak, A. Chigaev, V. Detours, and B. T. Korber. 2002. Clustering patterns of cytotoxic T-lymphocyte epitopes in human immunodeficiency virus type 1 (HIV-1) proteins reveal imprints of immune evasion on HIV-1 global variation. J. Virol. 76:8757-8768. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Zuñiga, R., A. Lucchetti, P. Galvan, S. Sanchez, C. Sanchez, A. Hernandez, H. Sanchez, N. Frahm, C. H. Linde, H. S. Hewitt, W. Hildebrand, M. Altfeld, T. M. Allen, B. D. Walker, B. T. Korber, T. Leitner, J. Sanchez, and C. Brander. 2006. Relative dominance of Gag p24-specific cytotoxic T lymphocytes is associated with human immunodeficiency virus control. J. Virol. 80:3122-3125. [DOI] [PMC free article] [PubMed] [Google Scholar]