Abstract
Endonuclease (Endo) IV encoded by denB of bacteriophage T4 is an enzyme that cleaves single-stranded (ss) DNA in a dC-specific manner. Also the growth of dC-substituted T4 phage and host Escherichia coli cells is inhibited by denB expression presumably because of the inhibitory effect on replication of dC-containing DNA. Recently, we have demonstrated that an efficient cleavage by Endo IV occurs exclusively at the 5′-proximal dC (dC1) within a hexameric or an extended sequence consisting of dC residues at the 5′-proximal and the 3′-proximal positions (dCs tract), in which a third dC residue within the tract affects the polarized cleavage and cleavage rate. Here we isolate and characterize two denB mutants, denB(W88R) and denB(S176N). Both mutant alleles have lost the detrimental effect on the host cell. Endo IV(W88R) shows no enzymatic activity (<0.4% of that of wild-type Endo IV). On the other hand, Endo IV(S176N) retains cleavage activity (17.5% of that of wild-type Endo IV), but has lost the polarized and restricted cleavage of a dCs tract, indicating that the Ser176 residue of Endo IV is implicated in the polarized cleavage of a dCs tract which brings about a detrimental effect on the replication of dC-containing DNA.
INTRODUCTION
Endonuclease IV (Endo IV) encoded by denB of bacteriophage T4 is implicated in restriction of deoxycytidine (dC)-containing DNA corresponding to the host Escherichia coli or its own genomic DNA in the host cells. Indeed, the synthesis of stable dC-substituted T4 (T4dC) genomic DNA requires a defective mutation of denB in addition to mutations of gene 42 (which encodes dCMP hydroxymethylase), gene 56 (dCTP/dUTPase) and either gene 46 (46/47 exonuclease) or denA (endonuclease II) (1). Under the denB−, gene 42−, gene 56−, possibly denA− background, an additional mutation in alc (unf), whose product shuts off transcription of T4dC DNA (2), results in the generation of a plaque-forming T4 phage containing dC-substituted DNA (T4dC phage) (3). Furthermore, a deficiency in denB is indispensable for the synthesis of stable T4dC genomic DNA (1) and denB expression in E. coli cells is lethal (4), even though Endo IV has little effect on the degradation of T4dC (5) or host (6–8) genomic DNA after infection of T4 phage. These results suggest that Endo IV plays a crucial role in inhibition of the replication of T4dC or host genomic DNA rather than in its degradation.
Endo IV composed of 185 amino acid residues requires Mg2+ for activity, acts only on single-stranded (ss) DNA and generates 5′ termini containing exclusively dC (4,9–12, H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). The enzyme does not cleave normal T4 genomic DNA-containing glucosylated deoxyhydroxymethylcytidine (gluc-dHMC) (4,9). Endo IV preferentially recognizes short nucleotide sequences that include 5′-dTdCdA-3′ (4,12) in a condition where a unique dC locates within a substrate oligonucleotide. Given that Endo IV is highly toxic to E. coli cells, we have synthesized the enzyme using a wheat germ cell-free protein synthesis system (13) and purified to homogeneity (4, H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). Recently, we have demonstrated that a six-base sequence comprising dC residues at the first and sixth positions (refers as a dC1–dC6 tract or a 6 dCs tract), 5′-dC1dN2dN3dN4dN5dC6-3′ (where dN represents dA, dC, dG or dT), is crucial for the efficient cleavage by Endo IV, and a third dC at the dN2dN3dN4dN5 region of a dC1–dC6 tract restricts the cleavage activity at the 5′-proximal dC1 site (restricted and polarized cleavage). A third dC residue placed within a dCs tract enhances the restricted and polarized cleavage. Furthermore, some of dCs tracts having sizes larger than six bases (extended dCs tract) also were shown to behave as a six dCs tract (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication).
Replication of the E. coli chromosome occurs bidirectionally from a single locus known as oriC (14,15), whereas bacteriophage T4 makes use of two major replication mechanisms that are regulated to occur during certain phases of the infection cycle (16,17). At early period of T4 infection, origin-dependent replication proceeds from several origins and involves initiation from R-loops (18). As infection progresses, origin-dependent replication is repressed, and recombination-dependent replication predominates (19) and involves initiation from D-loops formed by strand invasion (20). Given that R-loops, D-loops and lagging strand of replication fork include ss regions of DNA, the regions of ssDNA in these replication-specific structures may become potential substrates for Endo IV in vivo. If this is the case, Endo IV might target certain sequences in the ssDNA regions of these replication-specific structures and thereby effectively inhibit the replication. In this regard, we previously identified that short nucleotide sequences that include 5′-dT/dCdAdCdTdTdC-3′ is the most susceptible to Endo IV cleavage in vitro (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication), where the slash indicates the cleavage site by Endo IV. However, the mechanism of sequence recognition by Endo IV that is crucial for the inhibition of replication of dC-containing DNA remains unclear.
To elucidate in vivo function of Endo IV, in this study we isolated two T4dC mutants with point mutations in denB gene that result in the replacement of Trp88 of Endo IV with Arg [denB(W88R)] or that of Ser176 with Asn [denB(S176N)]. We synthesized the Endo IV enzymes encoded by the mutant denB alleles and found that the enzymatic activities of Endo IV(W88R) and Endo IV(S176N) were <0.4 and 17.5% of that of wild-type Endo IV, respectively. Neither of the two mutant alleles had a detrimental effect on the synthesis of T4dC genomic DNA nor the growth of host cells. Furthermore, the sequence 5′-dCdTdT-3′ located at the dN2dN3dN4dN5 region of a dC1–dC6 tract was shown to enhance cleavage at the dC1 site by wild-type Endo IV but not by Endo IV(S176N), indicating that the Ser176 residue is crucial for the restricted and polarized cleavage by Endo IV. These results indicate that the Ser176 residue of Endo IV implicated in the restricted and polarized cleavage at the dC1 site of a dCdTdT-containing dC1–dC6 tract is responsible for the Endo IV-mediated restriction of dC-containing DNA in E. coli cells.
MATERIALS AND METHODS
Materials
Restriction and other enzymes for recombinant DNA technology were obtained from Takara Shuzo. T4 and T4dC genomic DNA were prepared as described previously (21). The plasmid pEUGFP was constructed as described previously (22) and pGEX-6p-1 was obtained from GE Healthcare. Escherichia coli strain KH5402-1 [ilv, thyA, thr(Am), trpE9829(Am), metE, deo, supF6(Ts)] (23) was provided by the National Institute of Genetics (Japan), and strains LE392 (supF58, supE44, hsdR514, galK2, galT22, metB1, trpR55, lacY1) and MC1061 (sup°, hsdR, mcrB, araD139, araABC-leu7679, lacX74, galU, galK, rpsL, thi) were described previously (24). λCI857 lysogen of LE392 and E. coli Bb were used as host strains for T4 phages (24). T4GT7 (3), T4dC(+) (24) and T4 amC87 (gene 42), amE51 (gene 56) and unf39x6 (alc-1) were derived from our T4 phage stocks.
Oligonucleotide substrates used in this article were purchased from Oligo Texas Co. and are listed in Table 1. A 45-base oligonucleotide, 5′-dTdAdGdAdTdTdTdAdGdTdAdAdTdTdAdTdAdTdGdCdTdCdTdTdCdAdTdTdAdGdAd AdAdAdAdTdTdAdTdTdTdGdTdGdA-3′ (=Cy5-T4A), based on the sequence of T4 DNA and labeled at its 5′ end with Cy5, was commercially synthesized (Sigma-Aldrich, USA).
Table 1.
Oligonucleotide substrates used and their abbreviated names
| Oligonucleotide | Short namea |
|---|---|
| 5′-(dC)45-3′ | [(dC)45]45/45 |
| 5′-(dC)25-3′ | [(dC)25]25/25 |
| 5′-(dA)5(dC)15(dA)5-3′ | [(dC)15]15/25 |
| 5′ -(dA)7(dC)11(dA)7-3′ | [(dC)11]11/25 |
| 5′ -(dA)9(dC)6(dA)10-3′ | [dCdCdCdCdCdC]6/25 |
| 5′ -(dA)10(dC)5(dA)10-3′ | [dCdCdCdCdC]5/25 |
| 5′ -(dA)12(dC)1(dA)12-3′ | [dC]1/25 |
| 5′-(dA)9dCdTdTdTdTdC(dA)10-3′ | [dCdTdTdTdTdC]6/25 |
| 5′-(dA)9dCdCdTdTdTdC(dA)10-3′ | [dCdCdTdTdTdC]6/25 |
| 5′-(dA)9dCdTdCdTdTdC(dA)10-3′ | [dCdTdCdTdTdC]6/25 |
| 5′-(dA)9dCdTdTdCdTdC(dA)10-3′ | [dCdTdTdCdTdC]6/25 |
| 5′-(dA)9dCdTdTdTdCdC(dA)10-3′ | [dCdTdTdTdCdC]6/25 |
aSuffix numbers indicate the nucleotide length of the target sequence (referred as a six-base tract) and total length of oligonucleotides, respectively. Suffix number is often omitted in the text. A dC1–dC6 tract is defined as a hexanucleotide comprising two dC residues at the first and sixth positions and the sequences are shown in the parentheses [ ].
Genetic manipulation of T4 phage mutants and isolation of single point mutations in T4 denB
T4 phage cross and complementation test were performed as described in a previous paper (24). Isolation of denB mutations from denB-proficient multiple mutant strain of T4 was done as described previously (24). In brief, T4GT7, which contains mutations in gene 42 (amC87), gene 56 (amE51) and alc (unf39) in addition to the deletion NB5060 (ΔrII–ΔdenB), was genetically crossed with an alc (unf39)-deficient mutant having rII- and denB-proficient genes. Genetic cross was performed on LE392 as host cells. LE392, LE392 (λCI857), MC1061 and Bb were used as host cells to identify the presence of respective mutation. After crossing on LE392, plaque formers on LE392 (λCI857) at 30°C were screened for the presence of ‘amber’ mutation(s) by spotting on Bb and LE392. Then, phage clones giving no growth on Bb host were further examined by genetic complementation tests with amC87 (gene 42), amE51 (gene 56) on Bb cells. Thus, rII- and denB-proficient phage recombinants having mutations in gene 42 (amC87) and gene 56 (amE51) besides alc (unf39), were obtained. Two such T4 clones that give plaques on MC1061 at ∼10−4, but essentially no plaque on Bb host cells were used for further study. These clones may possibly have an additional mutation derived from the original GT7 that affects endonuclease II activity (25). However, this does not affect the results reported here and therefore we do not mention this mutation later.
The plaque formers on MC1061 cells are expected to have mutation in denB in addition to the mutations in gene 42 (amC87), gene 56 (amE51) and alc (unf39). Plaques appeared on MC1061 cells were purified and the DNAs were subjected to the polymerase chain reaction (PCR) with a 5′ primer and a 3′ primer (4) to amplify the denB region. The sequence of this region was determined by the dideoxy chain termination method to identify individual points mutations of denB.
Construction of denB expression plasmids
The plasmids pBRdenBam, pBRdenB(W88R)am and pBRdenB(S176N)am were constructed as described previously (4). The denBam, denB(W88R)am and denB(S176N)am genes, in which the TAC codon for Tyr38 is replaced with a TAG nonsense codon, were constructed by PCR with a 5′ primer containing a BamHI site and a 3′ primer containing a SalI site as well as with 5′ and 3′ mutagenic primers (4). The PCR products were introduced into the BamHI–SalI sites of pBR322 (Takara Shuzo) to yield pBRdenBam, pBRdenB(W88R)am and pBRdenB(S176N)am, and the resulting plasmids were cloned in non-amber-suppressing host cells. The nucleotide sequences of the plasmid inserts were confirmed by the dideoxy chain termination method. All PCR primers were synthesized by Sigma-Aldrich. PCR was performed for 25 cycles of incubation at 98°C for 10 s, 55°C for 10 s and 72°C for 30 s with Prime STAR HS DNA polymerase (Takara Shuzo) in a Takara PCR Thermal Cycler MP.
The effect of denB alleles on E. coli growth was examined by growth curve analysis of E. coli KH5402-1 cells (23) harboring pBRdenBam, pBRdenB(W88R)am or pBRdenB(S176N)am. The cells were grown overnight at 42°C in LB liquid broth containing thymine and ampicillin at 2 and 100 μg/ml, respectively (LB-Thy-Amp). The resulting cells were used to inoculate (1:100 dilution) fresh LB-Thy-Amp liquid broth and were incubated with shaking (∼140 min−1) at 42 or 30°C. Samples were removed from the cultures at various times to measure the optical density at 600 nm (OD600). Given that E. coli KH5402-1 cells harbor a temperature-sensitive (Ts) allele of the supF suppressor gene, the cells transformed with pBRdenBam, pBRdenB(W88R)am or pBRdenB(S176N)am would be expected to produce the encoded Endo IV proteins when grown at 30°C but not at 42°C.
Production and purification of GST-Endo IV fusion proteins
A DNA fragment encoding a glutathione S-transferase (GST) fusion protein of Endo IV was constructed and subjected to in vitro transcription as described previously (4). The resulting mRNA was then translated in a wheat germ cell-free protein synthesis system with the use of a dialysis cup (molecular size cutoff of 12 000 Da; Daiichi Pure Chemicals) as described previously (4). The dialysis unit containing the reaction mixture was incubated for 96 h at 26°C, with the original amount of substrate mRNA being supplemented and the external solution changed every 24 h. The GST-Endo IV fusion protein was purified as described previously (4). The fusion protein was cleaved by incubation with PreScission protease (10 U/ml; GE Healthcare) for 4 h at 4°C in a glutathione-Sepharose 4B MicroSpin column (GE Healthcare). The flow-through fraction contained a protein of the predicted size for Endo IV (21.1 kDa), as revealed by SDS–polyacrylamide gel electrophoresis on a 12% gel and staining with Coomassie brilliant blue. The protein concentration of this fraction was estimated by densitometric analysis with NIH Image software and the use of trypsin inhibitor (20.1 kDa) as a standard. The Endo IV mutant proteins were obtained as described for the wild-type protein.
Kinetic analysis of Endo IV activity
Hydrolysis of oligonucleotide substrates (10 μM) was performed as described previously (4, H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). Enzyme and substrate were incubated for 30 min at 37°C in a reaction mixture (20 μl) containing 10 mM Tris–HCl (pH 8.0), 10 mM MgCl2, 1 mM dithiothreitol and bovine serum albumin (0.1 mg/ml). The reaction was stopped by the addition of 30 μl of 25 mM EDTA (pH 8.0) and 50 μl of 10% trichloroacetic acid. The resulting mixture (100 μl) was maintained on ice for 15 min and then centrifuged at 5000g for 15 min at 4°C. The amount of acid-soluble nucleotides in the supernatant fraction was quantified by measurement of absorbance at 260 nm, with molar absorption coefficients of 15 200, 7050, 12 010 and 8400 M−1 cm−1 for dA, dC, dG and dT, respectively. One unit was defined as the amount of enzyme producing 1 μmol of acid-soluble nucleotides per minute, and specific activity was defined as the enzymatic activity per milligram of enzyme. The amount of enzyme was varied such that the amount of product increased in proportion to that of the enzyme. The concentration of the substrate was also varied from 3 to 30 μM such that it spanned the Km. Oligonucleotide substrates were synthesized by Texas Genomics Japan. Kinetic parameters were determined by a least-squares fit of the data in Lineweaver–Burk plots.
Cleavage analysis of an oligonucleotide corresponding to a T4 DNA sequence by Endo IV
Hydrolysis of a 45-base oligonucleotide (Cy5–T4A, see Materials section) based on the sequence of T4 DNA and labeled at its 5′ end with Cy5 (Sigma-Aldrich) was performed as described above for the Endo IV assay at a substrate concentration of 10 μM and enzyme concentrations of either 0.5–4 μg/ml for wild-type Endo IV or 1–8 μg/ml for Endo IV(S176N). The reaction products were separated by electrophoresis on a 10% polyacrylamide gel containing 7 M urea and were visualized with a Variable Image Analyzer Typhoon 8600 (GE Healthcare).
RESULTS
Screening for single point mutations in denB
To clarify the biological significance and function of denB and the gene product, Endo IV, we screened and characterized the denB mutations and their gene products. The isolation of denB mutations was originally described in a previous paper (24). T4 mutant phages having multiple mutations in gene 42 (amC87), gene 56 (amE51) and alc (unf39) were plated on MC1061 strain as the host and plaque formers were further characterized in details (see Materials and Methods section). Thus, we identified two types of T4 phage mutants with a single point mutation in denB besides the multiple mutations, which resulted in plaque-formable T4 phages with dC-containing DNA on a restriction-minus and suppressor-negative host, such as MC1061. By sequence analysis, the first type of denB mutation revealed to have a transition mutation (U to C) in the codon Trp88 (UGG) changed to codon Arg (CGG). This mutation coincided with the denB-s19 mutation of T4dC(+) phage (24) and was named as denB(W88R). The second type of denB mutation had also a transition mutation (G to A) which resulted in a change of codon from Ser176 (AGC) to Asn (AAC), and was named as denB(S176N).
Detrimental effect of wild-type and mutant-type denB alleles on E. coli growth
As described in a previous paper (4), attempts to clone the wild-type denB gene in several different expression vectors, including pET21a under strict control of the T7 promoter, were unsuccessful, presumably because of a highly toxic effect of the denB product (Endo IV) on the host E. coli cells. We therefore examined the effect of denB expression on E. coli by using KH5402-1 (supF6Ts) cells. The E. coli strain has a temperature-sensitive allele of the supF suppressor gene (23) with a plasmid containing an amber mutant allele of denB, in which the TAC codon for Tyr38 is mutated to a TAG nonsense codon. The E. coli KH5402-1 cells were used to examine the effects of T4 phage-encoded denB (4) and ndd (26) genes on the growth of host cells, and a significant growth defect was observed when KH5402-1 cells transformed with a plasmid containing an amber mutant allele of denB (4) or ndd (26) gene were cultured at 30°C.
To know whether the denB(W88R) and denB(S176N) alleles allow the growth of E. coli cells, we transformed KH5402-1 cells with a plasmid-containing denB mutant alleles in which an amber mutation had been introduced at the codon for Tyr38 (TAC). Cells cultured at 30°C would thus be expected to produce the full-length Endo IV mutant enzymes, whereas those cultured at 42°C would not. We previously showed that culture at 30°C of KH5402-1 cells transformed with pBRdenBam reduced colony-forming ability compared with that apparent at 42°C, and most (99%) of the original colonies that formed at 30°C did not give rise to colonies in the presence of the selection drug (ampicillin) at 42°C (4). These results indicated that most colonies that formed at 30°C consisted of cells lacking pBRdenBam and that most cells that expressed wild-type Endo IV were not viable. Here we examined the effects of denB expression on the growth of KH5402-1 cells transformed with pBRdenBam. KH5402-1 cells transformed with pBRdenBam revealed essentially no growth at 30°C, but normal growth at 42°C, and KH5402-1 cells transformed with pBR322 grow normally both at 30 and 42°C (Figure 1A). In contrast, the growth at 42 or 30°C of KH5402-1 cells transformed with pBR322 containing amber mutant forms of denB(W88R) or denB(S176N) alleles did not differ substantially from that of those transformed with pBR322 alone (Figure 1B). These results suggested that E. coli cells expressing the intact Endo IV mutant enzymes, Endo IV(W88R) and Endo IV(S176N), were viable and that, in contrast to wild-type Endo IV, these mutant enzymes do not show detrimental effect on the host cells.
Figure 1.
Effect of denB(W88R) and denB(S176N) alleles on E. coli growth. Cultures of KH5402-1 cells harboring either pBR322 or pBRdenBam (A) or pBRW88Ram or pBRS176Nam (B) were grown overnight in LB-Thy-Amp liquid broth at 42°C, and the cells were then used to inoculate fresh LB-Thy-Amp liquid broth and incubated at 42°C or 30°C. Samples were removed at the indicated times for determination of OD600. Data are means of values from two independent experiments.
Enzymatic activities of Endo IV(W88R) and Endo IV(S176N)
To examine the Endo IV activity encoded by these denB mutant alleles, we synthesized wild-type and mutant Endo IV as GST fusion proteins by in vitro translation with a wheat germ cell-free protein synthesis system (4). The amounts of purified Endo IV(W88R) and Endo IV(S176N) obtained were similar to that of wild-type Endo IV. We previously developed the acid-solubility assay for Endo IV activity (4). The acid-solubility assay is based on the endonucleolytic activity of Endo IV to generate acid-soluble products from an acid-insoluble oligonucleotide substrate. All of the oligonucleotide substrates were so designed as to be acid-insoluble before Endo IV cleavage and to generate acid-soluble products after the cleavage. The acid-solubility assay was successfully applied to the kinetic analysis of Endo IV with oligonucleotide substrates consisting of two stretches of appropriate length of oligo(dA) attached to both the 5′ and 3′ termini of varied dC-containing sequence (4, H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). All of the oligonucleotide substrates used in this study were identical to those used in the previous study (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication).
The denB product of T4dC phages would not be expected to possess enzymatic activity, because Endo IV catalyzes endonucleolytic cleavage of dC-containing ssDNA. To confirm this expectation, we examined the enzymatic activities of Endo IV(W88R), Endo IV(S176N) and wild-type Endo IV with 45-base 5′-(dC)45-3′ ([(dC)45]45/45), 25-base 5′-(dC)25-3′ ([(dC)25]25/25) and various oligonucleotides as substrates listed in Table 1. The activity of wild-type Endo IV decreased as the length of dC tract decreased from 15 to 5, and its enzymatic activity with [dC]1/25 did not differ from that with [dCdCdCdCdC]5/25 (Figure 2). As expected, Endo IV(W88R) showed essentially no Endo IV activity, which was <0.4% of that of wild-type Endo IV with [(dC)25]25/25. In contrast, Endo IV(S176N) showed a substantial level of Endo IV activity, which was 17.5% of that of wild-type Endo IV with [(dC)25]25/25 and equal to that of wild-type Endo IV with [dCdCdCdCdC]5/25, even though the denB(S176N) allele allowed the synthesis of T4dC genomic DNA and the growth of host cells. In addition, the activity of wild-type Endo IV increased markedly (by a factor of 20) as the length of dC tract increased from five to six, whereas that of Endo IV(S176N) did not (by a factor of 2). These results indicated that the S176N mutation might affect the sequence preference of Endo IV.
Figure 2.
Effect of dC tract length on enzymatic activities of wild-type and mutant (S176N or W88R) forms of Endo IV. Enzymatic activity was determined by measurement of the amount of acid-soluble nucleotides released from the substrate (10 μM). All substrates with the exception of [(dC)45]45/45 contained 25 nt. The specific activity of the wild-type (WT) enzyme with the [(dC)25]25/25 substrate was ∼8.0 U/mg. Relative activity was calculated by dividing the enzymatic activity of each Endo IV enzyme observed with each substrate by that apparent with the wild-type enzyme and [(dC)25]25/25 as substrate. Data are means of values from two independent experiments.
Sequence preferences of Endo IV and Endo IV(S176N)
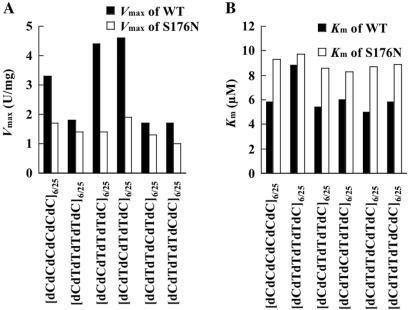
To examine a difference in the enzymatic activity with [dCdCdCdCdCdC]6/25 between wild-type Endo IV and Endo IV(S176N) in more detail, we compared their kinetic parameters for the hydrolysis of this substrate (Figure 3). The Vmax of Endo IV(S176N) with [dCdCdCdCdCdC]6/25 was reduced by a factor of 1.9 and its Km was increased by a factor of 1.6 compared with the corresponding values of wild-type Endo IV, indicating that the denB(S176N) mutation resulted in both a lower hydrolysis rate and a lower substrate affinity.
Figure 3.
Analysis and comparison of the sequence preferences of wild-type and S176N mutant forms of Endo IV. The Vmax (A) and Km (B) of the wild-type (WT) and S176N mutant forms of Endo IV were determined with series of 25-base oligonucleotides [dCdYdYdYdYdC]6/25 (where dY represents dT or dC) and shown by black columns (WT) and white columns (S176N) in the figure. Data are means of values from two independent experiments.
Since the activity of Endo IV(S176N) did not markedly increase compared with that of wild-type Endo IV as the length of dC tract increased from five to six (Figure 2), the denB(S176N) mutation might affect recognition of 5′-dC1dN2dN3dN4dN5dC6-3′ (dC1–dC6) tract by Endo IV (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). To examine this possibility, we compared the kinetic parameters of wild-type Endo IV and Endo IV(S176N) for the hydrolysis of [dCdTdTdTdTdC]6/25, [dCdCdTdTdTdC]6/25, [dCdTdCdTdTdC]6/25, [dCdTdTdCdTdC]6/25 and [dCdTdTdTdCdC]6/25 oligonucleotides (Figure 3). The Km and Vmax of Endo IV(S176N) with [dCdTdTdTdTdC]6/25 were similar to those of wild-type Endo IV with this substrate (Figure 3), indicating that the denB(S176N) mutation does not affect recognition of the dC1 and dC6 residues of a dC1–dC6 tract by Endo IV. In contrast, the Vmax values of wild-type Endo IV with [dCdCdTdTdTdC]6/25 and [dCdTdCdTdTdC] 6/25, both of which contain 5′-dCdTdT-3′ at the dN2dN3dN4dN5 region, were ∼2.5-fold greater than its Vmax with [dCdTdTdTdTdC]6/25 (Figure 3A), whereas the Vmax values of Endo IV(S176N) with all the substrates tested were similar to each other (Figure 3A), indicating that the denB(S176N) mutation disrupts recognition of a 5′-dCdTdT-3′ sequence located at the dN2dN3dN4dN5 region of a dC1–dC6 tract by Endo IV. In addition, the Km values of wild-type Endo IV with [dCdCdTdTdTdC]6/25, [dCdTdCdTdTdC]6/25, [dCdTdTdCdTdC]6/25 and [dCdTdTdTdCdC]6/25, all of which contain a dC residue at the dN2dN3dN4dN5 region, were ∼1.6-fold lower than its Km with [dCdTdTdTdTdC]6/25 (Figure 3B), whereas the Km values of Endo IV(S176N) with all the substrates tested were substantially identical (Figure 3B), indicating that Endo IV(S176N) markedly reduces the affinity to the third dC residue within a dC1–dC6 tract. Thus, the mutant enzyme is unable to recognize a 5′-dCdTdT-3′ sequence within a dC1–dC6 tract and did not show enhanced cleavage at the same site observed by wild-type Endo IV.
Endo IV(S176N) is not able to cleave at the 5′ proximal dC (dC1) in a dC1–dC6 tract
We have previously demonstrated that a 5′-dCdTdT-3′ sequence placed at the dN2dN3dN4dN5 region of a dC1–dC6 tract directs the cleavage activity of Endo IV at the dC1 site (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). Since Endo IV(S176N) did not recognize 5′-dCdTdT-3′ element within a dC1–dC6 tract, the denB(S176N) mutation might affect the restricted and polarized cleavage at the dC1 site by Endo IV. To examine this possibility, we compared the cleavage patterns of a Cy5-labeled 45-base oligonucleotide corresponding to a T4 DNA sequence containing a 5′-dC1dT2dC3dT4dT5dC6-3′ tract (Cy5–T4A) by wild-type Endo IV and Endo IV(S176N) (Figure 4). The maximal concentrations of wild-type Endo IV and Endo IV(S176N) used in these assays were 4 and 8 μg/ml, respectively, and the cleavage products were separated by electrophoresis on a polyacrylamide gel containing 7 M urea. The substrate contains three dC residues at positions 20, 22 and 25, which correspond to the dC1, dC3 and dC6 positions, respectively, of the 5′-dC1dT2dC3dT4dT5dC6-3′ tract. Wild-type Endo IV (Figure 4A) cleaved exclusively at the dC1 site in dG19–dC20–dT21 (marked c) while Endo IV(S176N) (Figure 4B) cleaved most efficiently at the dC6 site in dT24–dC25–dA26 (marked a) and less efficiently at the dC3 site in dT21–dC22–dT23 (marked b), indicating that the denB(S176N) mutation disrupts the restricted and polarized cleavage at the dC1 site (marked c) of the 5′-dC1dT2dC3dT4dT5dC6-3′ tract by Endo IV. These results suggest that Ser176 of Endo IV recognizing the third dC residues included 5′-dCdTdT-3′ within a dC1–dC6 tract is crucial for the restricted and polarized cleavage at the dC1 site by Endo IV. In addition, although Endo IV(S176N) has lost the ability to restrict the cleavage site to the dC1 site, it retains an ability to cleave dC-specific manner and a 5′-dTdCdA-3′ preference that is characteristic of wild-type Endo IV (4).
Figure 4.
Comparison of cleavage patterns of oligonucleotide containing the 5′-dCdTdCdTdTdC-3′ corresponding to a T4 DNA sequence by wild-type and S176N mutant forms of Endo IV. A 45-base oligonucleotide (Cy5–T4A) based on the sequence of T4 DNA and labeled at its 5′ end with Cy5 was used as the substrate (10 μM) for assay of the activity of wild-type (0.5, 1.0, 2.0 or 4.0 μg/ml) (A) or S176N mutant (1.0, 2.0, 4.0 or 8.0 μg/ml) (B) forms of Endo IV. The reaction products were separated by electrophoresis on a 10% polyacrylamide gel containing 7 M urea and were visualized with an image analyzer. Lane (−) represents a reaction mixture incubated in the absence of enzyme. Lane M represents a mixture of oligonucleotides labeled at their 5′ ends with Cy5 and with sequences identical to those of residues 1 to 19, 1 to 21 or 1 to 24 of the substrate. Cleavage sites of the substrate are indicated by arrows, and dC sites in dG19–dC20–dT21, dT21–dC22–dT23 and dT24–dC25–dA26 are indicated by a, b and c, which correspond to the dC1, dC3 and dC6 sites of the 5′-dC1dT2dC3dT4dT5dC6-3′ tract, respectively.
DISCUSSION
Role of Ser176 in the sequence preference of Endo IV
Only limited information on the sequence preference of Endo IV has been available (4,12), and the small number of Endo IV-related proteins in the genome sequence databases has restricted the amount of insight provided by such proteins into the mechanism of sequence recognition by Endo IV. Recently, we have found that a 5′-dC1dN2dN3dN4dN5dC6-3′ (dC1–dC6) tract is crucial for the efficient cleavage by wild-type Endo IV (Figure 5A) (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication). A third dC residue located at the dN2dN3dN4dN5 region of a dC1–dC6 tract improves the affinity of Endo IV to substrates, and a 5′-dCdTdT-3′ sequence at the dN2dN3dN4dN5 region restricts and enhances the Endo IV cleavage at the dC1 site (Figures 4A and 5a) (H. Ohshima, N. Hirano and H. Takahashi, submitted for publication).
Figure 5.
Schematic representation of the dCs tract model for restricted dC-specific cleavage by Endo IV and the role of Ser176. (A) A dCs tract is shown as a horizontal line with two closed boxes (dC6 and dC1) from the 3′ to 5′ direction. A downward arrow indicates the point where an endonucleolytic cleavage by Endo IV occurs and a rightward arrow indicates the element enhancing and restricting the enzymatic activity (Vmax and Km) of Endo IV. Endo IV binds to both dC1 and dC6 residues, and especially the Ser176 residue of the enzyme contributes to recognition of a dC residue located at the dN2dN3dN4dN5 region of a dCs tract and leads the enzyme to exhibit cytotoxicity and recognize the 5′-dCdTdT-3′ trinucleotide element enhancing the cleavage activity at the dC1 site. This interaction is required for the restricted and polarized cleavage at the dC1 site by Endo IV. (B) Replacement of Ser176 with Asn disrupts the interaction between Endo IV and a dC residue located at the dN2dN3dN4dN5 region, resulting in the losses of restricted and polarized cleavage at the dC1 site and cytotoxicity.
In this report, we have shown that the affinity of Endo IV(S176N) to the third dC residue located at the dN2dN3dN4dN5 region is markedly reduced and therefore a 5′-dCdTdT-3′ sequence at the dN2dN3dN4dN5 region does not either restrict or enhance the cleavage activity of Endo IV(S176N) at the dC1 site (Figures 4B and 5b), whereas Endo IV(S176N) retains an ability to cleave dC-specific manner and a 5′-dTdCdA-3′ preference (Figure 4B). These results suggest that the Ser176 residue that contributes to recognition of the third dC residue located at the dN2dN3dN4dN5 region of a dC1–dC6 tract is crucial for enhancement of the cleavage activity of Endo IV with a dCdTdT-containing dC1–dC6 tract and the restricted and polarized cleavage at the dC1 site by Endo IV. In addition, because the Ser176 residue markedly increases the hydrolysis rate of Endo IV with a dCdTdT-containing dC1–dC6 tract, Ser176 may contribute to accommodation of the cleavage site (dC1 site) to the active site of Endo IV enzyme, resulting in the efficient cleavage at the dC1 site by Endo IV.
Possible mechanism of Endo IV action in vivo
A deficiency in denB suppresses the lack or arrest of T4dC genomic DNA synthesis in a gene 56− (1,27,28) or gene 42−, gene 46−, gene 56− (1) background, respectively, and T4dC phage mutants with a denB+ background cease synthesizing their genomic DNA within 30 min after infection (1). In addition, several denB point mutations that allow stable T4dC genomic DNA synthesis in the gene 56− background result in a complete loss of Endo IV activity (confirmed in vitro) (29). Moreover, denB expression in E. coli cells is lethal (4). Together, these various observations so far suggest that a lack of Endo IV activity is indispensable for the replication of dC-containing DNA.
We have now identified the denB(W88R) and denB(S176N) alleles that allowed plaque-forming T4dC phage in the gene 42−, gene 56−, alc−, possibly denA− background and did not exhibit the detrimental effect on host cell growth, suggesting that these denB alleles do not inhibit the replication of both T4dC and host genomic DNAs. Indeed, Endo IV(W88R) exhibited essentially no Endo IV activity (<0.4% of that of wild-type Endo IV), consistent with the suggested indispensability of a deficiency of denB for the replication of dC-containing DNA. However, Endo IV(S176N) possessed a substantial level of Endo IV activity (17.5% of that of wild-type Endo IV) and a 5′-dTdCdA-3′ preference. The comparison of the enzymatic activities of wild-type Endo IV and Endo IV(S176N) indicated that the denB(S176N) mutation did not enhance the cleavage activity of Endo IV with a dCdTdT-containing dC1–dC6 tract and lost the restricted and polarized cleavage at the dC1 site, which might be responsible for the loss of the ability of Endo IV to restrict dC-containing DNA in the host cells.
Replication of E. coli genome initiates from a single origin, oriC (14,15), whereas bacteriophage T4 makes use of two major mechanisms of replication initiation, which involve R-loops at several origins, oriA, oriE, oriF and oriG, and D-loops in the process of recombination-dependent replication (16–20). However, because dCdTdT-containing dC1–dC6 tracts are not found abundantly in the proximal regions of any of T4 and E. coli origins, Endo IV may not specifically target the initiation-specific structures of replication at these origins but target various ssDNA regions of replicating DNA, such as those involved in R-loops, D-loops and lagging strand of replication fork. In fact, no sequence consisting of more than six consecutive dC residues crucial for the efficient cleavage by Endo IV is found in T4 genomic DNA (168 903 bp), whereas dCdTdT-containing dC1–dC6 tracts being the enhanced target sequence by Endo IV are found frequently in T4 genomic DNA with an average frequency of 1 per 938 bp. Thus, recognition of the sequences that include a dCdTdT-containing dC1–dC6 hexanucleotide tract in T4 genomic DNA by Endo IV may be crucial for the effective inhibition of T4dC genomic DNA synthesis.
The mechanism of restriction of dC-containing DNA in the host cells by Endo IV remains elusive. The denB(S176N) mutation reduced the enzymatic activity with the oligo(dC) substrates and the Vmax/Km values with the substrates containing a dCdTdT-containing dC1–dC6 tract only by a factor of ∼5 (Figures 2 and 3), raising a question why the denB(S176N) allele allows both T4 and E. coli replication-apparatuses to replicate their dC-containing genomic DNAs. A five-fold reduction of the Endo IV activity in vivo may simply make both T4 and E. coli replication-apparatuses possible to replicate dC-containing DNA due to contribution of the endogenous repair systems. Alternatively, if the cleavage at the dC1 site of a dCdTdT-containing dC1–dC6 tract is critical for the inhibition of both replications by T4 and E. coli replication-apparatuses, a loss of the restricted cleavage at the dC1 site of the tract may greatly reduce the accumulation of damage to both replicating DNAs in vivo. In vivo studies of the effects of denB mutant alleles on E. coli growth, which retain the restricted cleavage at the dC1 site of the tract but reduce the enzymatic activity, may provide insight into the mechanism of Endo IV-mediated restriction of dC-containing DNA in E. coli cells.
ACKNOWLEDGEMENTS
We thank Y. Yada, Y. Masutani and T. Kurahashi for technical support, N. Arai and H. Mitsusawa for helpful discussion. This work was supported in part by Grant-in-Aid for Scientific Research, the 21st Century COE Program and the High-Tech Research Project from the Ministry of Education, Culture, Sports, Science and Technology of Japan. Funding to pay the Open Access publication charges for this article was provided by High-Tech Research Project for private universities: matching fund from MEXT.
Conflict of interest statement. None declared.
Footnotes
The authors wish it to be known that, in their opinion, the first two authors should be regarded as joint First Authors.
REFERENCES
- 1.Carlson K, Overvatn A. Bacteriophage T4 endonucleases II and IV, oppositely affected by dCMP hydroxymethylase activity, have different roles in the degradation and in the RNA polymerase-dependent replication of T4 cytosine-containing DNA. Genetics. 1986;114:669–685. doi: 10.1093/genetics/114.3.669. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Snyder L, Gold L, Kutter E. A gene of bacteriophage T4 whose product prevents true late transcription on cytosine-containing T4 DNA. Proc. Natl Acad. Sci. USA. 1976;73:3098–3102. doi: 10.1073/pnas.73.9.3098. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Wilson GG, Young KY, Edlin GJ, Konigsberg W. High-frequency generalized transduction by bacteriophage T4. Nature. 1979;280:80–82. doi: 10.1038/280080a0. [DOI] [PubMed] [Google Scholar]
- 4.Hirano N, Ohshima H, Takahashi H. Biochemical analysis of the substrate specificity and sequence preference of endonuclease IV from bacteriophage T4, a dC-specific endonuclease implicated in restriction of dC-substituted T4 DNA synthesis. Nucleic Acids Res. 2006a;34:4743–4751. doi: 10.1093/nar/gkl553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Carlson K, Wiberg JS. In vivo cleavage of cytosine-containing bacteriophage T4 DNA to genetically distinct, discretely sized fragments. J. Virol. 1983;48:18–30. doi: 10.1128/jvi.48.1.18-30.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Warner HR, Snustad DP, Jorgensen SE, Koerner JF. Isolation of bacteriophage T4 mutants defective in the ability to degrade host deoxyribonucleic acid. J. Virol. 1970;5:700–708. doi: 10.1128/jvi.5.6.700-708.1970. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hercules K, Munro JL, Mendelsohn S, Wiberg JS. Mutants in a nonessential gene of bacteriophage T4 which are defective in the degradation of Escherichia coli deoxyribonucleic acid. J. Virol. 1971;7:95–105. doi: 10.1128/jvi.7.1.95-105.1971. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Souther A, Bruner R, Elliott J. Degradation of Escherichia coli chromosome after infection by bacteriophage T4: role of bacteriophage gene D2a. J. Virol. 1972;10:979–984. doi: 10.1128/jvi.10.5.979-984.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Sadowski PD, Hurwitz J. Enzymatic breakage of deoxyribonucleic acid. II. Purification and properties of endonuclease IV from T4 phage-infected Escherichia coli. J. Biol. Chem. 1969;244:6192–6198. [PubMed] [Google Scholar]
- 10.Ling V. Partial digestion of 32P-fd DNA with T4 endonuclease IV. FEBS Lett. 1971;19:50–54. doi: 10.1016/0014-5793(71)80602-6. [DOI] [PubMed] [Google Scholar]
- 11.Sadowski PD, Bakyta I. T4 endonuclease IV. Improved purification procedure and resolution from T4 endonuclease III. J. Biol. Chem. 1972;247:405–412. [PubMed] [Google Scholar]
- 12.Bernardi A, Maat J, de Waard A, Bernardi G. Preparation and specificity of endonuclease IV induced by bacteriophage T4. Eur. J. Biochem. 1976;66:175–179. doi: 10.1111/j.1432-1033.1976.tb10437.x. [DOI] [PubMed] [Google Scholar]
- 13.Madin K, Sawasaki T, Ogasawara T, Endo Y. A highly efficient and robust cell-free protein synthesis system prepared from wheat embryos: plants apparently contain a suicide system directed at ribosomes. Proc. Natl Acad. Sci. USA. 2000;97:559–564. doi: 10.1073/pnas.97.2.559. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Bird RE, Louarn J, Martuscelli J, Caro L. Origin and sequence of chromosome replication in Escherichia coli. J. Mol. Biol. 1972;70:549–566. doi: 10.1016/0022-2836(72)90559-1. [DOI] [PubMed] [Google Scholar]
- 15.Prescott DM, Kuempel PL. Bidirectional replication of the chromosome in Escherichia coli. Proc. Natl Acad. Sci. USA. 1972;69:2842–2845. doi: 10.1073/pnas.69.10.2842. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Kreuzer KN, Morrical SW. Initiation of DNA replication. In: Karam JD, editor. Molecular Biology of Bacteriophage T4. Washington, DC: American Society for Microbiology; 1994. pp. 28–42. [Google Scholar]
- 17.Mosig G, Colowick N, Gruidl ME, Chang A, Harvey AJ. Multiple initiation mechanisms adapt phage T4 DNA replication to physiological changes during T4's development. FEMS Microbiol. Rev. 1995;17:83–98. doi: 10.1111/j.1574-6976.1995.tb00190.x. [DOI] [PubMed] [Google Scholar]
- 18.Kinch KC, Kreuzer KN. RNA-DNA hybrid formation at a bacteriophage T4 replication origin. J. Mol. Biol. 1997;14:915–926. doi: 10.1006/jmbi.1996.0844. [DOI] [PubMed] [Google Scholar]
- 19.Kreuzer KN. Recombination-dependent DNA replication in phage T4. Trends Biochem. Sci. 2000;25:165–173. doi: 10.1016/s0968-0004(00)01559-0. [DOI] [PubMed] [Google Scholar]
- 20.Formosa T, Alberts BM. DNA synthesis dependent on genetic recombination: characterization of a reaction catalyzed by purified bacteriophage T4 proteins. Cell. 1986;47:793–806. doi: 10.1016/0092-8674(86)90522-2. [DOI] [PubMed] [Google Scholar]
- 21.Takahashi H, Shimizu M, Saito H, Ikeda Y. Studies of viable T4 bacteriophage containing cytosine-substituted DNA (T4dC phage). II. Cleavage of T4dC DNA by endonuclease SalI and BamHI. Mol. Gen. Genet. 1979;168:49–53. doi: 10.1007/BF00267932. [DOI] [PubMed] [Google Scholar]
- 22.Hirano N, Sawasaki T, Tozawa Y, Endo Y, Takai K. Tolerance for random recombination of domains in prokaryotic and eukaryotic translation systems: limited interdomain misfolding in a eukaryotic translation system. Proteins. 2006b;64:343–354. doi: 10.1002/prot.21008. [DOI] [PubMed] [Google Scholar]
- 23.Kimura M, Yura T, Nagata T. Isolation and characterization of Escherichia coli dnaA amber mutants. J. Bacteriol. 1980;144:649–655. doi: 10.1128/jb.144.2.649-655.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Takahashi H, Saito H, Ikeda Y. Viable T4 bacteriophage containing cytosine substituted DNA (T4dC phage). I. Behavior towards the restriction-modification systems of Escherichia coli and derivation of a new T4 phage strain (T4dC) having the complete T4 genome. J. Gen. Appl. Microbiol. 1978;24:297–306. [Google Scholar]
- 25.Carlson K, Raleigh EA, Hattman S. Restriction and modification. In: Karam JD, editor. Molecular Biology of Bacteriophage T4. Washington, DC: American Society for Microbiology Press; 1994. pp. 369–381. [Google Scholar]
- 26.Bouet JY, Campo NJ, Krisch HM, Louarn JM. The effects on Escherichia coli of expression of the cloned bacteriophage T4 nucleoid disruption (ndd) gene. Mol. Microbiol. 1996;20:519–528. doi: 10.1046/j.1365-2958.1996.5411067.x. [DOI] [PubMed] [Google Scholar]
- 27.Bruner R, Souther A, Suggs S. Stability of cytosine-containing deoxyribonucleic acid after infection by certain T4 rII-D deletion mutants. J. Virol. 1972;10:88–92. doi: 10.1128/jvi.10.1.88-92.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kutter E, Beug A, Sluss R, Jensen L, Bradley D. The production of undegraded cytosine-containing DNA by bacteriophage T4 in the absence of dCTPase and endonucleases II and IV, and its effects on T4-directed protein synthesis. J. Mol. Biol. 1975;99:591–607. doi: 10.1016/s0022-2836(75)80174-4. [DOI] [PubMed] [Google Scholar]
- 29.Vetter D, Sadowski PD. Point mutants in the D2a region of bacteriophage T4 fail to induce T4 endonuclease IV. J. Virol. 1974;14:207–213. doi: 10.1128/jvi.14.2.207-213.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]