Abstract
Tousled-like kinases (TLKs) constitute a family of serine/threonine kinases conserved in plants and animals that act in a cell cycle-dependent manner. In mammals, their activity peaks during S phase, when they phosphorylate the antisilencing function protein 1 (ASF1), a histone chaperone involved in replication-dependent chromatin assembly. Here, we show that Drosophila ASF1 is also a phosphorylation target of TLK, and that the two components cooperate to control chromatin replication in vivo. By altering TLK activity through loss-of-function mutations, we show that nuclear divisions are arrested at interphase, followed by apoptosis. Overexpression of TLK alters the chromatin structure, suggesting that TLK mediates the activity of chromatin proteins. These results suggest that TLK coordinates cell cycle progression through the regulation of chromatin dynamics.
Keywords: Anti-silencing function protein 1, cell cycle, chromatin structure, histone chaperone, tousled-like kinase
DNA of eukaryotes is wrapped repetitively around histone octameres, forming nucleosomal arrays. This so-called beads-on-a-string arrangement constitutes the backbone for building higher-order chromatin structures (Luger et al. 1997). Distinct chromatin structures are assembled depending on covalent histone tail modifications (histone-code), resulting in activation or repression of gene expression. These active/silenced chromatin states are epigenetically inherited through cell divisions (Turner et al. 1992; Jenuwein and Allis 2001; Richards and Elgin 2002). During S phase, nucleosomal array, and consequently, higher-order chromatin complexes, are partially disrupted by the passage of the replication forks (Sogo et al. 1986; Gruss et al. 1993). Therefore, in order to transmit the epigenetic state of chromatin through the cell cycle, nucleosomal arrays must be duplicated faithfully during DNA replication (Enomoto and Berman 1998; Shibahara and Stillman 1999; Zhang et al. 2000).
Assembly of newly synthesized DNA onto nucleosomes is under the control of replication-dependent chromatin assembly systems (for review, see Mello and Almouzni 2001; Tyler 2002). It involves the deposition of a histone H3 and H4 tetramer onto the newly generated daughter strands, followed by incorporation of two histone H2A–H2B dimers to complete the nucleosome core particles (Smith and Stillman 1991; Lucchini et al. 2001). This process is facilitated by histone chaperones such as the chromatin assembly factor 1 (CAF-1) and by the recently identified antisilencing function protein 1 (ASF1). Additionally, ATP-dependent chromatin remodeling factors are required to produce regularly spaced arrays (for review, see Mello and Almouzni 2001; Tyler 2002).
Coupling of chromatin assembly and DNA synthesis is, in part, mediated by interactions between CAF-1 and PCNA, a loading factor for the replication machinery. This association is thought to be important for the location of the chromatin assembly factors to the sites of DNA synthesis (Shibahara and Stillman 1999; Zhang et al. 2000). Further biochemical analysis revealed that CAF-1 nucleosome assembly activity on replicating DNA is facilitated by the replication-coupling assembly factor (RCAF) complex (Tyler et al. 1999, 2001; Mello et al. 2002) composed of ASF1 and newly synthesized acetylated histones H3 and H4 (Tyler et al. 1999). Gene disruption studies of CAF-1 subunits in yeast (Tyler et al. 1999) and asf1 in yeast and flies (Sharp et al. 2001; Moshkin et al. 2002; Umehara et al. 2002) support the idea that de novo chromatin assembly is required for epigenetic inheritance of silenced chromatin. In yeast, the lack of ASF1 activity causes a substantial slowing down of the cell cycle in S phase, an effect that is enhanced by mutations in CAF-1 subunits (Le et al. 1997; Tyler et al. 1999). Both genetic and biochemical data suggest that ASF1 functions synergistically with CAF-1 in the regulation of chromatin assembly and cell divisions (Tyler et al. 1999, 2001; Sharp et al. 2001; Mello et al. 2002).
In mammals, ASF1 has been identified as a phosphorylation target of the cell cycle-regulated Tousled-like kinases (TLKs). These kinases are members of a conserved serine/threonine kinase family present in both animals and plants (Silljé and Nigg 2001). The prototype TLK in Arabidopsis thaliana was shown to be essential for proper flower and leaf development by possibly regulating cell divisions (Roe et al. 1993, 1997). Mammalian TLKs are localized in the nucleus during interphase and display maximal activity during S phase, when DNA synthesis and replication-dependent chromatin assembly occur (Silljé et al. 1999). In further studies, TLKs have also been shown to be able to phosphorylate Ser 10 of histone H3 (Li et al. 2001).
We describe the function of the single tousled-like kinase (tlk) gene of Drosophila. Analysis of tlk mutants shows that reduced tlk levels cause abnormal nuclear divisions followed by apoptotic death, and that increased tlk levels alter the chromatin structure. TLK interacts in vitro with ASF1, resulting in phosphorylation of ASF1. In vivo, TLK cooperates with ASF1 in the control of endoreplication, indicating that TLK regulates cell cycle progression through the chromatin assembly pathway.
Results
Characterization of the Drosophila tlk
The Drosophila genome contains only a single member of the Tousled-like kinase family of genes, termed tousled-like kinase (tlk). The gene is located in the region 3F of the Drosophila X chromosome (Adams et al. 2000; Fig. 1A). Comparison of the tlk cDNA sequences (LD15160, GH07910) with the corresponding genomic DNA sequences (Berkeley Drosophila Genome Project, BDGP) revealed that the gene encodes alternatively spliced transcripts. They differ in their 3′ untranslated region, resulting from the use of different polyadenylation sites. In addition, they have an alternatively spliced first exon. The predicted proteins share a common ORF of 1242 residues and differ in the N-terminal region (details in legend to Fig. 1). The structure of the gene, the deduced amino acid sequence of TLK, and a comparison with human and Arabidopsis homologs are summarized in Figure 1.
Figure 1.
Characterization of the tlk gene, tlk mutations, and tlk protein. (A) Structure of the tlk gene and position of the P-element insertion l(1)G0054. cDNA was obtained from the LD15160 clone. AE refers to the contig number of the Drosophila genome project (Adams et al. 2000). Comparison of different tlk cDNA sequences and tlk genomic sequences (Berkeley Drosophila Genome Project, BDGP) showed that tlk encodes differentially spliced transcripts. The tlk LD15160 and GH07910 transcripts differ in their 3′ untranslated region (different polyadenylation sites), and they have an alternatively spliced first exon. The predicted protein sequences encoded by tlk LD15160 and tlk GH07910 (gene CG2829) share a common ORF of 1242 residues and differ in the first putative 13 (MPVVKQRKMSRFK) and 24 amino acids (MDQFQTSLNPRKLELLESRITGVR), respectively. Database searches revealed that cDNAs LD16185 and LD09833 also encode a protein with 12 alternative N-terminal residues (MCVQKDMRRLKK) and the cDNAs LD41810 and LD16882 lack the alternatively spliced first exon. The alleles tlkΔ14 and tlkΔ40 result from an imprecise excision in which 2357 bp have been deleted (tlkΔ14) and 1143 bp have been replaced by a 22-bp sequence interval of the P-element (tlkΔ40), respectively. Nontranslated regions and coding region of the gene are shown by open and black boxes, respectively; deleted genomic DNA of the mutants is shown by brackets. PK indicates the cDNA region that encodes the tlk kinase domain; ABP (green box) indicates the position of the peptide used for antibody production. (B) Alignment of the conserved C terminus containing the Tousled kinase domain of Drosophila tlk, Arabidopsis (Atha), and human homologs (Hsap TLK1 and Hsap TLK2) using the CLUSTALW program (Thompson et al. 1994). Identical residues or conservative replacements are shaded; dots denote gaps in the alignments. The amino acid position 217 is labeled with an asterisk; A indicates that the tlkdead transgene carries a replacement of the aspartic acid residue by alanine. (C) Schematic representation of Drosophila, Arabidopsis, and human TLKs. The kinase domains (PK; yellow boxes), nuclear localization signals (red oval), leucine zipper motives (blue boxes), coiled-coil regions (green spirals), and glutamine-rich regions (green boxes) are shown. Numbers refer to positions of amino acids.
TLK, human TLK1 and TLK2, and the Tousled kinase of Arabidopsis (TSL) show a high degree of sequence conservation in the C-terminal serine/threonine kinase domain with 71%, 68%, and 52% sequence similarity, respectively (Fig. 1B). The catalytic domain of TLKs carries an unusual ATP-binding motif, which contains a GXGXXS (position 73–78; Fig. 1B) instead of the canonical GXGXXG motif (Hanks et al. 1988). A structural comparison of Drosophila TLK with mammalian and plant homologs is shown in Figure 1C.
tlk expression patterns and subcellular protein localization
In situ hybridization of the tlk cDNA probe to ovaries shows that the transcripts are expressed in nurse cells from where they are transported into the oocyte (data not shown). tlk transcripts remain ubiquitously distributed in the egg and throughout embryonic development (Fig. 2A). Northern blot analysis confirms that the highest amount of tlk transcript is present during the early syncytial phase of embryonic development, when rapid DNA replication and chromatin assembly occur (data not shown). Moreover, ubiquitous expression of tlk is found in proliferative tissue, the larval imaginal discs (data not shown).
Figure 2.
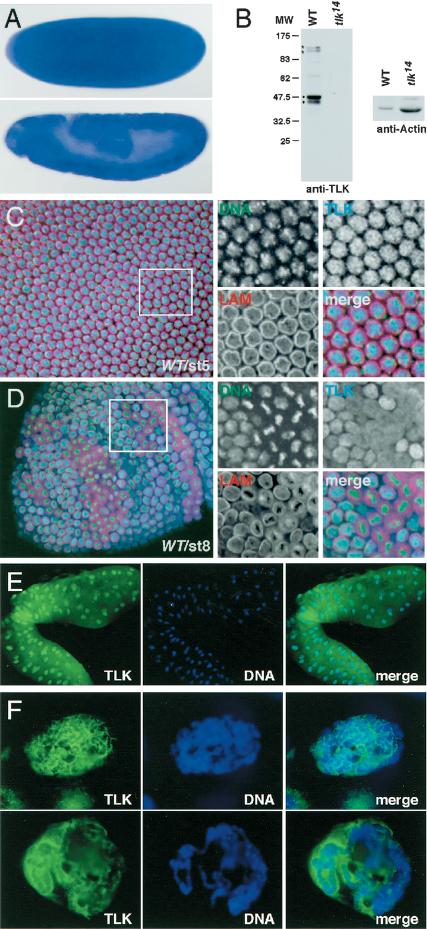
Distribution of tlk transcripts and protein. (A) In situ hybridization of whole-mount preparations from embryos (stages 3 and 9) as revealed by antisense RNA corresponding to LD15160. (B) Western blots of SDS gels loaded with crude extracts from eggs of either wild-type females (WT) or females containing a homozygous tlkΔ14 germ line (tlk14) were probed with anti-TLK antibodies; anti-actin antibodies were used to monitor the protein loading. Asterisks indicate bands corresponding to predicted sizes of TLKs, dots indicate possible degradation products (for details, see text). (C,D) Confocal images of embryos showing DNA (green; SYTOX Green), lamin (red; antilamin antibodies), or TLK (blue; anti-TLK antibodies). (C) Stage 5 wild-type embryo with synchronized nuclei at interphase; TLK is concentrated in the nuclei (see enlarged area, right). (D) Stage 8 wild-type embryo with asynchronous cell divisions. TLK is enriched in interphase nuclei. During all other stages of the division cycle, TLK is distributed diffusely and not associated with condensed chromosomes (enlarged area, right). (E) Staining of whole-mount salivary glands for TLK (green) and DNA (blue). Note that TLK is concentrated in nuclei. (F) Magnification of nuclei showing that TLK is not associated with polytene chromosomes but concentrated around them.
To examine the cellular localization of Drosophila Tousled-like kinase protein (TLK), we generated antibodies against TLK (see Materials and Methods). In Western blots prepared from crude extracts of freshly laid eggs or nuclei of embryos (Fig. 2B), anti-TLK antibodies recognize two bands of ∼135 and 125 kD, the predicted size of TLKs. In addition, the antibodies detect two smaller bands. All bands are absent from corresponding crude extracts of tlkΔ14 mutants (see below), indicating that anti-TLK antibodies are specific and have no cross-reactivity. The two smaller bands seen in the Western blots (Fig. 2B) likely represent TLK degradation products.
Early proliferation events during Drosophila embryogenesis proceed through 13 rapid and synchronous syncytial mitotic cycles. After cellularization, which occurs during nuclear division cycle 14, cells enter an extended G2 phase. Cells that continue to divide are restricted to so-called mitotic domains (Foe 1989). Immunostaining detects TLK during nuclear division cycle 10, when nuclei align at the periphery of the embryo, in a distinct cell cycle-dependent localization pattern. TLK is strongly enriched in interphase nuclei (Fig. 2C). During all other stages of mitosis, TLK is not associated with the mitotic chromosomes, but rather, diffusely distributed throughout the cells (data not shown; Fig. 2D). After cellularization, when the mitotic domains of the embryo have formed, and throughout embryonic development (Foe 1989), TLK again becomes enriched only in interphase nuclei (Fig. 2D).
The subcellular pattern of TLK during interphase suggests that the protein could be linked to chromatin. To test this hypothesis, we examined polyploid salivary glands of larvae. We found that the protein is enriched in salivary gland nuclei, in which it concentrates around the polytene chromosomes (Fig. 2E, F). However, no staining was obtained with squashed polytene chromosomes (data not shown). This observation indicates that TLK is not associated directly with DNA and may stay in only weak contact with the interphase chromatin of polytene chromosomes.
tlk is an essential gene
To assess the in vivo function of TLK, we generated mutant tlk alleles by P[lacW]-element insertion mutagenesis (Peter et al. 2002). We recovered a lethal mutation, tlkl(1)G0054, in which a P[lacW] was inserted into the first intron of the tlk gene (Fig. 1A). Homozygous tlkl(1)G0054 individuals die as larvae. This effect is due to the P[lacW] insertion into the tlk gene, because precise excision of the transposon restores full viability. We also recovered two imprecise excisions that gave rise to deletion mutants, the tlkΔ14 and tlkΔ40 alleles (Fig. 1A). As observed with the original tlkl(1)G0054 allele, the homozygous tlkΔ14 and tlkΔ40 mutant individuals die during larval stages. The fact that tlkΔ14 and tlkΔ40 mutations lack parts of the TLK coding sequence (Fig. 1A) and fail to express detectable amounts of protein as shown by anti-TLK antibody staining on Western blots of crude extracts of eggs from females containing tlkΔ14 germ-line clones (Fig. 2B) indicate that the two alleles are loss-of-function alleles and that tlkΔ14 is possibly amorphic. These data indicate that tlk wild-type activity is necessary for the viability of the organism, and that the activity of maternally derived tlk transcripts is possibly sufficient to carry the mutant individuals throughout and beyond embryogenesis (Fig. 2A).
To address the role of maternal tlk transcripts, we used the FLP-ovoD system (Chou and Perrimon 1996) to generate homozygous tlk mutant germ-line clones, which causes the loss of maternal tlk activity in embryos. Male progeny derived from the tlkl(1)G0054 mutant female germ line died during embryogenesis, whereas the female individuals, which received a paternal wild-type tlk gene copy, developed into fertile adults. This result indicates that maternally derived tlk activity has an essential function, which can be compensated for by zygotic, paternally derived tlk activity.
Corresponding experiments with tlkΔ14 and tlkΔ40 alleles resulted in embryonic lethality of both males and heterozygous females, showing that these mutants are stronger than the tlkl(1)G0054 allele. At 25°C, females with germ-line clones for both tlkΔ14 and tlkΔ40 mutations are sterile, laying a small number of eggs that collapse shortly after deposition. At 18°C, however, these females lay eggs that look normal, but a substantial portion of them appears to be shorter than wild type. tlkΔ14 mutant embryos fail to hatch, and about two-thirds die during the first nine nuclear divisions (class I embryos). The other embryos form a syncytial blastoderm and only 12.5% (of total embryos) initiate gastrulation (class II embryos). tlkΔ40 is a weaker allele, because about half of the embryos gastrulate (52% of the total embryos), and some of them even develop into hatching larvae.
Loss of tlk affects nuclear divisions and causes DNA fragmentation
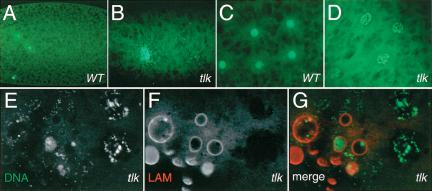
Closer examination of aged tlkΔ14 class I embryos revealed that no DNA could be detected in most of them (data not shown). Those containing DNA have a few enlarged nuclei, in which DNA forms either unstructured supernovas (Fig. 3A,B) or network-like structures (Fig. 3C,D). The latter observation reflects apoptotic nuclei as revealed by TUNEL staining (data not shown), visualization of DNA fragmentation, and nuclear envelope break-down, as detected by anti-lamin antibodies (Fig. 3E–G). The network-like DNA structures suggest defects in chromatin structure as has been reported for JIL-1 kinase (Wang et al. 2001). These results indicate that loss of maternal tlk activity impairs the early nuclear divisions of class I embryos and leads to apoptotic death.
Figure 3.
Apoptosis in tlkΔ14 class I mutant embryos. DNA staining with SYTOX Green of preblastodermal wild-type (A,C) and tlkΔ14 loss-of-function mutant embryos (B,D). Nuclei fail to replicate properly and undergo apoptosis. (E–G) DNA and lamin double staining shows DNA fragmentation and nuclear membrane breakdown in tlkΔ14 mutant embryos, both characteristics of apoptotic death. The nuclear envelope is outlined with antibodies against lamin (LAM; white in F, red in G).
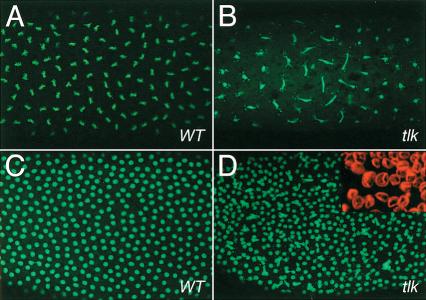
Although nuclei of class II embryos divide and migrate in a wild-type-like fashion toward the periphery, they harbor a number of distinct defects. The early nuclear divisions are asynchronous, as revealed by the appearance of both interphase and anaphase nuclei that is rarely observed in wild-type embryos (Fig. 4A,B). Furthermore, nuclei are connected by elongated DNA bridges (Fig. 4A–D), indicating incomplete chromosomal segregation. Moreover, nuclei are heterogeneous in size (Fig. 4D, inset) and the DNA of the enlarged nuclei has a uniform cloudy appearance with some strongly stained DNA spots (data not shown).
Figure 4.
Asynchronous nuclear divisons in tlkΔ14 class II mutant embryos. DNA is stained with SYTOX Green (DNA; green in A–D). The nuclear envelope is outlined with antibodies against lamin (LAM; inset, D). (A) Synchronous nuclear divisions (here: metaphase) in a preblastoderm wild-type embryo. (B) tlkΔ14 mutant embryo of younger age showing that nuclear divisions are asynchronous. Note that nuclei at the left and right margins have already divided and are separated, whereas nuclei in the central region are still in anaphase and show a defective chromosome segregation pattern (long DNA bridges). (C,D) Nuclear cycle 12 wild-type (C) and corresponding tlk mutant embryo (D). Note the loss of mitotic synchrony and uneven spacing of mutant nuclei (inset, D). The nuclear envelope is labeled with antibodies against lamin (red). tlk embryos show heterogeneity in the size of the nuclei.
The early stages of Drosophila embryogenesis undergo a rather specialized division cycle, with nuclei cycling between S and M phases only, for example, lacking the G phases in between. To find out whether TLK is also important for proliferating tissues that follow a conventional cell cycle, we performed mitotic recombination experiments to generate homozygous tlk mutant cells during imaginal disc development (Dang and Perrimon 1992). We found that imaginal disc cells homozygous for tlkΔ14 mutant are able to initially form clones of up to eight cells and then disappear from the growing tissue, whereas the wild-type twin clones continue normal growth (data not shown). These observations confirm that loss of tlk activity causes cell death.
tlk mutants show a reduced level of phosphorylated histone H3
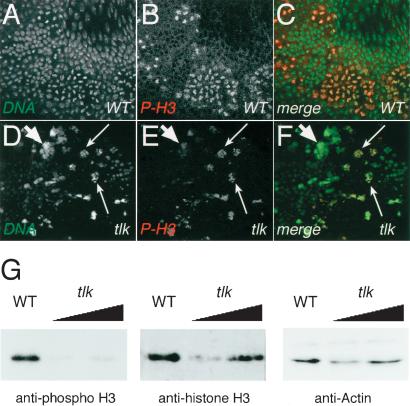
TLK1 was shown to phospholylate Ser 10 of histone H3 and to substitute for the absence of Ipl1, a kinase required for histone H3 phosphorylation in yeast (Li et al. 2001). We therefore asked whether the amount of histone H3 phosphorylation is altered in tlk mutant embryos. Figure 5A–F show that tlkΔ14 mutants have only a low number of nuclei that are stained with antibodies directed against phophorylated histone H3, a molecular marker of M-phase nuclei (Hendzel et al. 1997). This indicates that the majority of tlk mutant nuclei remain in interphase, and only a few continue mitosis (Fig. 5A–F). Reduced levels of phophorylated histone H3 in tlkΔ14 mutants could be confirmed by Western blot analysis, showing that although tlkΔ14 and wild-type embryos contain similar amounts of histone H3, the amount of phosphorylated histone H3 in tlkΔ14 mutants was reduced significantly (Fig. 5G). It is noteworthy that we failed to detect increased levels of histone H3 phosphorylation in embryos that overexpressed TLK (data not shown). Reduced levels of phosphorylated histone H3 in tlkΔ14 mutants could be an indirect consequence of fewer cells entering mitosis. In summary, these results show that the loss of tlk activity causes the nuclei to arrest in interphase. Those nuclei that are capable of entering the next division cycle fail to segregate chromosomes properly and subsequently undergo apoptosis.
Figure 5.
tlkΔ14 mutants have reduced phosphorylated histone H3 levels. DNA is stained with SYTOX Green (DNA; white in A,D; green in C,F). Nuclei in M phase are visualized by staining with anti-phospho-histone H3 antibody (P-H3; white in B,E; red in C,F). (A–C) DNA and phosphorylated histone H3 patterns in stage 8 wild-type embryos. Note P-H3 antibody stained cells within the mitotic domains. (D–F) tlkΔ14 mutant embryo stained with P-H3 antibody. Note few mitotic cells undergoing DNA fragmentation (large thin arrows). Most nuclei remain in interphase and accumulate big clumps of DNA (small fat arrows). (G) Western blots of SDS gels loaded with crude extracts from eggs that derived from either wild-type females (WT) or females containing homozygous tlkΔ14 germ lines were probed with antibodies against phosphorylated histone H3 (anti-phospho H3). Controls for loading of protein extracts; antibody staining of nonphosphorylated histone H3 (anti-histone H3) and actin (anti-Actin); increasing amounts of the protein extracts are indicated by triangles. Note that the tlkΔ14 mutants show reduced levels of phosphorylated histone H3.
Increased tlk activity alters the pattern of cell proliferation
We next asked whether increasing tlk expression also matters to dividing cells. To answer this question, we used the GAL4/UAS system (Brand and Perrimon 1993) to examine the effects of transgene-dependent tlk overexpression throughout the early embryos. Overproduction of tlk in response to some of the most active Gal4 drivers during early embryogenesis such as V32-Gal4, arm-Gal4, and en-Gal4 (see Materials and Methods) results in embryonic lethality. MS1096-driven overexpression (Milan et al. 1998) of tlk in the dorsal compartment of wing imaginal discs results in fewer and enlarged cells (data not shown). These findings are consistent with the conclusions drawn from the loss-of-function mutant analysis, showing that the level of TLK activity plays a general and provital role during the cell cycle and thereby affects proliferating tissues.
TLK interacts with ASF1 in vivo
Studies on human TLKs have shown that they are able to phosphorylate ASF1, a histone chaperone involved in chromatin assembly (Silljé and Nigg 2001). Thus, the tlk-dependent nuclear division defects in Drosophila might be mediated by ASF1 and thereby affect the chromatin assembly pathway. Therefore, we investigated whether TLK and ASF1 interact functionally in vivo by gain-of-function experiments, in which tlk and asf1 were expressed in response to the eyeless-Gal4 (ey-Gal4) driver during eye development. Figure 6A shows that overexpressing tlk or asf1 alone results in mild rough-eye phenotypes, whereas the coexpression of both genes causes a dramatic effect. Most of the derivatives of the eye-antenna imaginal disc fail to develop, resulting in a so-called pin-head phenotype (Fig. 6A). The enhancement of the phenotype due to the combined expression of asf1 and tlk strongly suggests that the products of the two genes act either in the same genetic pathway or contribute to the same process in an independent fashion.
Figure 6.
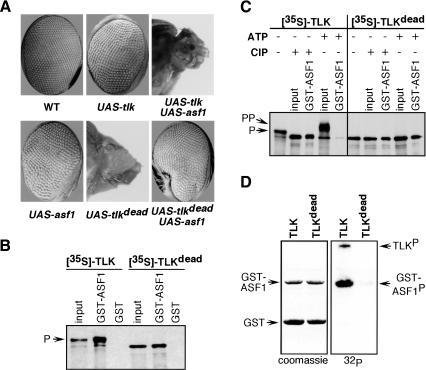
TLK acts in the same genetic pathway as ASF1 and phosphorylates it in vitro. (A) tlk interacts with asf1 in vivo. In wild-type flies, synchronized cell divisions give rise to adult eyes with regularly distributed facets. In flies overexpressing either tlk kinase or asf1 in the eye under the control of eyeless-GAL4 driver, the facets distribution is slightly altered, giving rise to a rough eye phenotype. Coexpression of both tlk and asf1 in the eye using the eyeless-GAL4 driver leads to severe developmental defects, resulting in pin-head phenotype. Overexpression of tlkdead alone in the eye with the same eyeless-GAL4 driver results in pin-head phenotype, whereas coexpression of both tlkdead and asf1 rescues the tlkdead pin-head phenotype. (B) GST pull-down assays with radiolabeled TLK and TLKdead produced by coupled in vitro transcription/translation (IVT) in the presence of [35S]methionine as detected by autoradiography. Both TLK and TLKdead bind to GST–ASF1 fusion protein, but not to GST. A total of 10% of the input mixture was loaded as control (Input). A major fraction of TLK kinase (P) migrates slower on SDS-PAGE compared with TLKdead due to autophosphorylation. (C) Stability of the interaction between TLK and ASF1 depends on TLKs autophosphorylation state. Radiolabeled TLK and TLKdead were treated with either calf intestinal alkaline phosphatase (CIP) or with ATP. CIP treatment results in a fast migrating TLK form of same mobility as TLKdead, which is pulled down by GST–ASF1. Addition of ATP results in a slower migrating form of TLK, which is presumably hyperphosphorylated (labeled PP) and does not bind GST–ASF1. Gel mobility of TLKdead and binding to GST–ASF1 protein is unchanged after treatments either with CIP or ATP. (D) Nonradiolabeled, in vitro translated TLK phosphorylates GST–ASF1 protein and itself, but not GST alone, as detected by autoradiography with 32P. TLKdead lacks phosphorylation and autophosphorylation activities because of a mutation in the catalytic domain. TLK and TLKdead proteins were not synthesized in sufficient amounts to be revealed by Coomassie staining; however, they were revealed by immunoblotting using anti-TLK antibodies (data not shown).
To further establish whether asf1 and tlk act in the same genetic pathway, we performed a complementary experiment, in which we coexpressed ASF1 and a dominant negative variant of TLK (TLKdead), which contains a single amino acid replacement (D to A, position 217 of the catalytic domain; Fig. 1B). Overexpression of tlkdead in response to ey-Gal4 had a devastating effect on eye development, leading to a pin-head phenotype, as observed in response to the combined overexpression of tlk and asf1 (see above; Fig. 6A). Interestingly, expression of asf1 in addition to tlkdead significantly suppressed the effect of tlkdead (Fig. 6A). Suppression of the tlkdead effects by coexpression of asf1 is not due to the dilution of the Gal4 driver, because coexpression of the flippase enzyme does not suppress the effects of tlkdead overexpression. These findings show that asf1 overexpression is able to inactivate TLKdead dominant negative function. Taken together, these coexpression studies suggest that in vivo ASF1 is a substrate of TLK.
TLK binds and phosphorylates ASF1 in vitro
One possibility to explain the ASF1-dependent suppression of the dominant negative effect of TLKdead is to suppose that, like in humans (Silljé and Nigg 2001), ASF1 is able to form a stable complex with the mutated kinase, and thereby suppresses its action by competition-based dilution of the active molecule.
To test this hypothesis and to further characterize the interaction between TLK and ASF1, we performed GST pull-down experiments. We found that bacterially produced GST–ASF1 fusion protein is able to pull down TLK or TLKdead synthesized and radiolabeled by in vitro transcription/translation (IVT) reaction (Fig. 6B). TLK produced in vitro migrates in two forms (slow and fast migrating; Fig. 6B,C), TLKdead migrates only as the fast moving form (Fig. 6B,C). These observations raise the possibility that in vitro synthesized TLK is phosphorylated, possibly by autophosphorylation. Figure 6D shows that in vitro synthesized TLK is active and able to phosphorylate both the GST–ASF1 fusion protein and itself. Treatment of TLK preparations with phosphatase or ATP further confirms that the fast and slow moving forms differ in phosphorylation status and provide insights into the interactions between ASF1 and TLK. We find that upon treatment with ATP, TLK migrates even slower (probably due to hyperphosphorylation) and is not pulled down by the GST–ASF1 fusion protein (Fig. 6C). We noted that ATP treatment may also alter the binding of ASF1 to hypophosphorylated TLK, as the recovery of the fast-migrating protein band seems to be reduced (Fig. 6C). This could be due to continued TLK autophosphorylation during the binding reaction or to partial ASF1 phosphorylation that may alter the interaction with TLK. In contrast, upon phosphatase treatment, TLK migrates only as the fast moving form and forms a stable complex with the GST–ASF1 fusion protein. Phosphatase or ATP treatments have no effects on TLKdead and do not alter its ability to interact with the GST–ASF1 fusion protein (Fig. 6C). These data suggest that ASF1 binds dephosphorylated (or moderately phosphorylated) TLK and dissociates upon TLK hyperphosphorylation. In addition, phosphorylation of ASF1 may regulate the interaction.
TLK and ASF1 affect chromatin structure and DNA replication
To gain further insight into the processes regulated by the TLK–ASF1 pathway during development, we have generated individuals in which UAS–asf1 and/or UAS–tlk transgenes can be ubiquitously activated. Because early acting GAL4 drivers cause embryonic lethality, we used heat-shock inducible GAL4 drivers. We found that 2-h heat-shock at any time during larval development caused UAS–asf1 or UAS–tlk-dependent lethality. We therefore decided to take advantage of the fact that the heat-shock promoter (hsp70) is leaky, and we determined the experimental conditions (25°C–30°C; see Materials and Methods) under which larvae developed into adult flies. Under these conditions, the overall growth and differentiation of larvae is not affected by expressing either asf1, tlk, or tlkdead, whereas the size of the third instar salivary glands is greatly reduced. The reason for the enhanced or even specific effect of ASF1 and TLK on salivary gland nuclei is that, in contrast to most other larval tissue, they undergo 10 rounds of endoreplication, in which DNA is replicated without daughter strands separation. Lack of endoreplication, therefore, would cause significantly smaller nuclei, and in turn, smaller salivary glands.
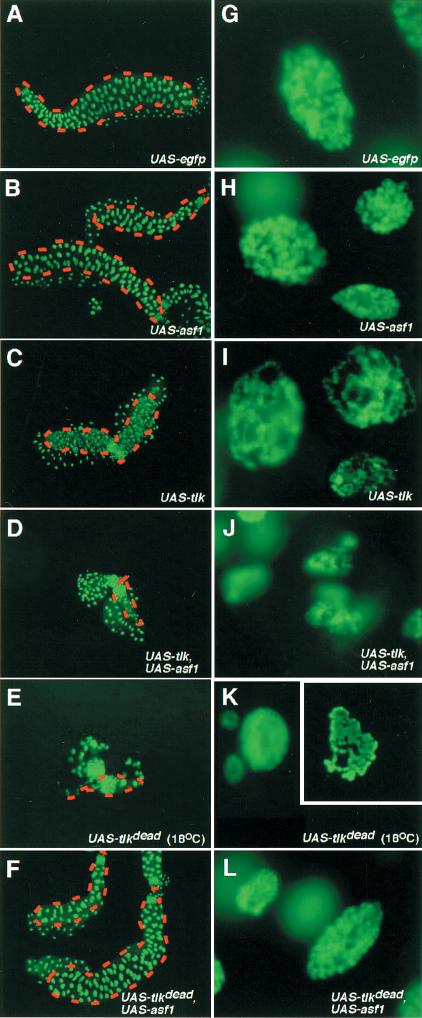
During normal salivary gland growth, endoreplication gives rise to polytene chromosomes showing a characteristic pattern of chromosomal bands. Polytene chromosomes provide a prime model to study in vivo the structure of interphase chromatin. Overexpression of ASF1 and/or TLK had severe effects on the morphology and the size of nuclei of salivary glands (Fig. 7). EGFP, which serves as an internal control for the overexpression of a protein in response to heat-shock-induced Gal4, had no such effect (Fig. 7A,G). Overexpression of asf1 results in smaller salivary glands composed of fewer cells (Fig. 7B), which contain chromosomes that are reduced in size (Fig. 7H). These findings suggest that excess of ASF1 slows down cell cycle progression or leads to cell death and that overexpression of ASF1 interferes with endoreplication. In the case of tlk overexpression, the salivary glands are also smaller, but the cell number is not significantly reduced, suggesting that the reduced size seems to be due primarily to reduced endoreplication, leading to thinner chromosomes than in wild-type and ASF-overexpressing glands (Fig. 7I; cf. Fig. 7G,H; see also Fig. 7H). In addition, tlk overexpression alters higher-order chromatin structures, as the chromosomes are grossly disrupted, losing their typical banding pattern (Fig. 7I). These observations suggest that TLK is essential for the proper assembly of higher order chromatin structures.
Figure 7.
Overexpression of the TLK–ASF1 pathway affects chromatin structure and endoreplication in salivary glands. (A,G) Salivary glands from larvae overexpressing egfp at 25°C under Heat-Shock-gal4 (HS-gal4) driver show normal cell number (128 per lobe) and morphology of polytene chromosomes. (B,H) Overexpression of asf1 results in smaller salivary glands with only half of the normal cell number (upper salivary gland). Occasionally, some of the salivary glands have normal size and almost normal cell number (lower salivary gland). Polytene chromosomes appear thinner, indicating that DNA is underreplicated. (C,I) Overexpression of tlk produces small salivary glands with nearly normal cell number, but with chromosomes that are extremely thin and unstructured, showing global changes in higher-order chromatin structure. (D,J) Coexpression of both tlk and asf1 gives rise to tiny salivary glands with few cells (∼1/10 of the normal) and tiny chromosomes. (E,K) Expression of the dominant negative tlkdead allele almost completely abolishes cell proliferation in salivary glands, even at 18°C. In these glands, chromosomes appear to be underreplicated. Chromosomes from few preparations that could be obtained show a banded pattern (K, inset). (F,L) Coexpressing asf1 with tlkdead partially rescues tlkdead effects on cell cycle progression and DNA replication. Note that G–L (with the exception of the inset in K) are presented at the same magnification.
Coexpressing asf1 and tlk together aggravates the salivary-gland mutant phenotypes as compared with overexpression of either asf1 or tlk only. Under these conditions, salivary glands are composed of only a few cells (Fig. 7D) that contain strongly underreplicated DNA (Fig. 7J). As observed with the eye (see above), overexpression of tlkdead affects salivary gland development severely, that is, we could only recover individuals with salivary glands by growing the larvae at 18°C. These glands are composed of only a few cells, all with underreplicated polytene chromosomes. This finding shows that the presence of tlkdead strongly inhibits endoreplication and interferes with cell divisions or cell viability (Fig. 7E,K). The effect of tlkdead overexpression can be suppressed by coexpression of asf1 (Fig. 7F,L), confirming an interaction between ASF1 and TLK. Despite a strong effect on chromatin structure and endoreplication in salivary glands, larvae overexpressing either asf1 and/or tlk at 30°C (hsp-driver) are able to develop into adults that, however, lack a number of the large thoracic bristles (macrochaetae). Also, the sex ratio of the flies hatching in these overexpression studies was skewed toward females, indicating that males are more sensitive to the altered levels of tlk and asf1 (data not shown). Altogether, these data imply that the TLK–ASF1 pathway is important for both DNA replication and chromatin assembly, suggesting a mechanism by which TLK controls cell cycle progression.
Discussion
The tlk gene encodes a conserved serine/threonine kinase with homologs in plants and animals. In humans, the TLKs are activated during the S phase of the cell cycle and were shown to phosphorylate at least two targets, ASF1 (Silljé et al. 1999; Silljé and Nigg 2001) and histone H3 (Li et al. 2001). ASF1 acts as a histone chaperone and participates in replication-dependent chromatin assembly (Tyler et al. 1999). Phosphorylated histone H3 is an important factor in the condensation of chromatin during mitosis (Wei et al. 1999). These discoveries revealed a possible link between the TLK family members and regulation of chromatin condensation and/or the assembly of chromatin during cell cycle progression. More recently, TLK activity has been shown to be down-regulated upon exposure to DNA-damaging ionizing radiation during S phase, a response depending on ATM and Chk1 kinases (Groth et al. 2003).
We show that Drosophila tlk is essential for nuclear divisions and cell viability, and required for the maintenance of the higher-order chromatin structure of interphase chromosomes. We also demonstrate that Drosophila TLK phophorylates ASF1, and that both proteins cooperate in the control of chromatin replication in vivo, suggesting a mechanism that links cell proliferation and a TLK-dependent chromatin assembly pathway.
TLK is essential for nuclear divisions and cell viability
The Drosophila tlk gene is transcribed throughout development in both somatic and germ-line tissues. Its activity is essential for survival as required for normal nuclear divisions in proliferating tissues. Loss of tlk causes an arrest of cell cycle progression at interphase and subsequently leads to apoptotic death.
Nuclear division defects are already detected in early embryos that lack tlk. Normally, early Drosophila embryos develop in a syncytium, in which the nuclei proceed through rapid nuclear divisions undergoing S and M transitions only. When embryos lack TLK activity, nuclear cleavages stop. In both Drosophila and human cells, TLK is enriched in S-phase nuclei and reaches its maximal activity during S phase (Silljé et al. 1999). These observations suggest that TLK activity is required for proper DNA replication and/or proper chromatin assembly. The elongated DNA masses (Fig. 4), however, imply that the most prominent effects occur during anaphase. Therefore, we cannot exclude the possibility that TLK affects not only the replication and chromatin assembly during interphase, but also other steps in the cell cycle (Fig. 8). In humans, TLK also phosphorylates histone H3 (Li et al. 2001), a step thought to be important for progression through mitosis. Although our results support a role of Drosophila TLK in histone H3 phosphorylation (Fig. 5G), they do not demonstrate that histone H3 is a direct phosphorylation target of TLK.
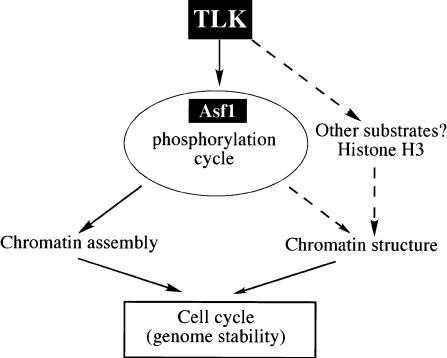
Figure 8.
Possible control over cell cycle progression by TLK via chromatin. Altering TLK expression affects cell cycle progression most likely via Asf1. Altered Asf1 phosphorylation could affect chromatin assembly in S phase and/or during DNA repair. TLK may also act on chromatin through histone H3 (Li et al. 2001), and have additional functions via as yet unidentified substrates.
Chromatin assembly factor ASF1 is a phosphorylation target of TLK
In support of a role of TLK in controlling the assembly and/or condensation of chromatin, we found that, as in humans (Silljé and Nigg 2001), Drosophila chromatin assembly factor ASF1 is phosphorylated by TLK. However, unlike in humans, in which TLKs interact only transiently with ASF1a and ASF1b (Silljé and Nigg 2001), we found that Drosophila TLK and ASF1 proteins form a stable complex that is disrupted upon TLK hyperphosphorylation. Interestingly, in human, TLK kinase is hyperphosphorylated during S phase, a time point which coincides with maximal TLK activity. Dephosphorylated forms of TLK are found throughout the rest of the cell cycle (Silljé et al. 1999). In light of these observations, we suggest that Drosophila ASF1 stably binds dephosphorylated TLK prior to initiation of DNA replication. During S phase, the initiation of DNA replication probably results in the activation of TLK concomitant to its autophosphorylation and the subsequent release of phosphorylated ASF1 from the complex. Phosphorylated ASF1 may then be used in de novo chromatin assembly coupled to DNA replication. A dynamic mode of the TLK–ASF1 interaction may also explain the rather weak association of TLK with polytene chromosomes and the absence of TLK-associated chromosomal bands on squashed chromosomes (Fig. 2E,F).
A similar mechanism has been suggested in yeast for the interactions between the RAD53 checkpoint kinase and ASF1 (Emili et al. 2001). In this case, activation of RAD53 by hyperphosphorylation induced by DNA damage results in the release of ASF1 from the complex. Upon release, ASF1 is thought to function in chromatin assembly on repaired DNA (Emili et al. 2001). However, the yeast ASF1 protein does not seem to be phosphorylated by the RAD53 kinase. In addition, there is no homolog of TLK in yeast, suggesting that the phosphorylation of ASF1, as observed in both humans and fly, is likely to be specific for higher eukaryotes.
It is interesting to note that the C-terminal part of the yeast ASF1 protein is enriched in acidic residues relative to its higher eukaryotic counterparts. This raises the possibility that the phosphorylation of ASF1 in Drosophila and humans adds negative charges to the C-terminal tail of ASF1 to allow fly or human ASF1 to attain a structure similar to that of yeast ASF1. Although, in yeast, an ASF1 protein lacking the polyanionic stretch is sufficient to complement asf1-growth defects and establishment of telomeric silencing in vivo, we believe that the negatively charged protein may enhance the deposition of histones. In support of this hypothesis, it was found that the yeast ASF1 protein lacking the polyanionic tail displayed markedly reduced binding to histones H3/H4 (Umehara et al. 2002).
TLK and chromatin assembly
Overexpression of TLK appears to block endoreplication of polytene chromosomes in the salivary glands, further substantiating the role of TLK in DNA replication. By overexpression of both genes, we were able to show that TLK cooperates with ASF1 in this process, arguing that the TLK-dependent effects on endoreplication are mediated through components of the chromatin assembly pathway (Fig. 7). ASF1 in yeast has been found to be important for histone gene regulation (Sutton et al. 2001). In this process, ASF1 cooperates directly with the histone regulation protein (HIR). Studies in which the HIR was up-regulated show that excess of the protein leads to the repression of histone gene transcription, followed by a block of DNA synthesis and an arrest of the cell cycle in S phase (Sharp et al. 2001; Nelson et al. 2002). In addition, the Xenopus laevis homolog of HIR, called HIRA, is able to assemble nucleosomes on naked DNA, suggesting a more general role for HIR proteins in chromatin organization (Ray-Gallet et al. 2002). These findings raise the possibility that TLK affects endoreplication by regulating DNA replication through the ASF1–HIR pathway, a proposal that will need to be addressed by future experiments.
TLK is not directly associated with distinct sites on polytene chromosomes, but TLK overexpression significantly disrupts their higher-order chromatin structure (Fig. 7I). Thus, TLK may participate in chromatin assembly and/or condensation by, for example, regulating or modifying chromatin proteins (Fig. 8). Several mutations have been reported that affect the higher-order chromatin structure of polytene chromosomes. Whereas mutations in iswi and nurf301, two genes encoding proteins involved in nucleosome sliding, affect mostly male X chromosome structure, mutations in the jil-1 kinase and Su(var)2–10 disrupt the chromatin structure of all polytene chromosomes (Deuring et al. 2000; Hari et al. 2001; Li et al. 2001; Wang et al. 2001; Badenhorst et al. 2002). Within such a scenario, TLK could exert its function in the maintenance of higher-order chromatin structures by modulating the activity of ASF1 or some other chromatin protein during chromatin assembly. Possible mechanisms by which altering TLK activity leads to a disruption of chromatin and cell cycle arrest are summarized in Figure 8.
Although further experiments will be required to determine exactly how TLK-dependent phosphorylation affects cell cycle progression, our findings suggest the existence of a crucial link between cell cycle progression and chromatin structure. First, we show that the lack of TLK activity leads to an arrest of cell cycle progression at interphase, and as consequence, to reduced levels of phosphorylated histone H3. Second, the fact that TLK seems to interact with chromatin, and that the only two known substrates of TLK are proteins involved in chromatin structure (histone H3 in humans and ASF1 in both humans and Drosophila) suggests that TLK activity may be targeted primarily to chromatin proteins. Finally, a closer examination of the chromosomes following TLK overexpression demonstrates that an alteration of TLK activity causes a global change in chromatin structure. These findings raise the intriguing possibility that the lack of proper chromatin structure may be the cause of the cell cycle arrest, and that there might be a specific cell cycle checkpoint determining whether the chromatin is properly assembled. In support of this hypothesis, a recent study shows that treating tissue-culture cells with chromatin-modifying drugs activates the ATM kinase, a protein whose activity is thought to lead to the cell cycle arrest following exposure to DNA-damaging ionizing radiation (Bakkenist and Kastan 2003). In addition, altering in vivo chromatin assembly by expressing a dominant negative mutant of the largest CAF-1 subunit induces S-phase arrest through activation of DNA-damage checkpoint pathway (Ye et al. 2003). It is interesting, then, to speculate that the cell cycle arrest following the alteration of chromatin structure due to TLK misregulation may occur through the activation of the DNA damage checkpoint pathway.
Materials and methods
Isolation and characterization of tlk alleles
Maintenance and handling of fly stocks and crosses were done according to standard techniques (Peter et al. 2002). l(1)G0054 is a lethal PlacW insertion in the first intron of tlk (Peter et al. 2002) that was used for subsequent studies. Isolation of new alleles by P-element excision was carried out with jump-starter females [genotype w tlkl(1)G0054/w FM6; TM2 Ubx Δ2–3/+] that were crossed with Y/FM6 males. P-element excisions were scored in female progeny (white-eye phenotype) and crossed with Y/FM7 GFP males, so that progeny can be distinguished on the basis of the visible GFP marker. Of 1250 independent excision events, we established 36 homozygous lethal strains that were balanced over FM7 GFP. The lethal phase of each tlk allele was determined with hemizygous males, genomic DNA surrounding the original l(1)G0054 insertion site was amplified by PCR and the DNA fragments were sequenced (Peter et al. 2002). Further detailed analysis was done with two deletion alleles, tlkΔ14 and tlkΔ40 (see text and Fig. 1).
Generation of germ-line clones
The alleles tlkl(1)G0054, tlkΔ14, and tlkΔ40 were each recombined onto a chromosome carrying P(w+ FRT)101. Homozygous cell clones were generated by mitotic recombination using the FRT/FLP system (Dang and Perrimon 1992); germ-line clones were generated using the FLP-ovoD system (Chou and Perrimon 1996). Virgin females of the genotype w, tlk, FRT101/FM7 GFP were mated with w ovoD1 v FRT101/Y; HS-Flp males. Flies were transferred to fresh food once every 24 h, and F1 larvae were heat shocked (37°C; 2 h) during the second and third instar larval stages to induce mitotic recombination events in proliferating germ cells. Embryos derived from a tlk homozygous female germ line were examined at 18°C and 25°C.
Antibody production, immunocytochemistry, and microscopy
Rabbit anti-TLK antibodies were generated against the peptide KSQIVGNQKTIDQHKCHIAKC located within the TLK N-terminal domain (position 585–606; Eurogentec). The position of the peptide within the ORF of the TLK coding sequence is indicated in Figure 1A. The cysteine in position 16 of the peptide was modified C(t-Bu) to block the coupling in internal positions. Both antiserum and affinity-purified antibodies (Thiopropyl Sepharose 6b, Amersham Pharmacia) were tested with Western blots of nuclear extracts of wild-type and tlk mutant embryos (see Fig. 2B) and by immunostaining of such embryos. Immunoreactivity on Western blots was detected with extracts of wild-type but not of tlk mutant embryos, indicating that the antibodies were specific for TLK. They were used in a 1:1000 dilution. Inmunostainings of wild-type and en-Gal4/UAS–tlk mutant embryos (showing overexpression of TLK in an ectopic stripe pattern) with antibodies against Tousled are fully competed by incubation of the antibody with the peptide antigen at concentrations below 10 μg/mL. We also used mouse anti-Lamin (1:200), rabbit anti-phospho-histone H3 (1:100; Upstate Biotechnology) as well as Cy3- or Alexa 488-, Alexa 543-, and Cy5- or Alexa 633-conjugated secondary antibodies as recommended by the manufacturer (Molecular Probes). For nuclear staining, samples were treated with RNAse A (Sigma; 400 μg/mL; 20 min) and incubated with 0.1 μM SYTOX Green (Molecular Probes; 2 h). Fixation of Drosophila embryos was in 4% paraformaldehyde. In general, embryos were incubated with the primary antibodies (PBS; overnight at 4°C), and then with the secondary antibodies and SYTOX Green. Embryos were mounted in VECTASHIELD mounting medium (Vectorlabs) and viewed by confocal microscopy. Digoxigenin-labeled antisense LD15160 RNA was used for whole-mount preparations of ovaries and embryos (Hauptmann and Gerster 1996). Immunostaining of whole-mounted salivary glands or squashed polytene chromosomes were done using a 1:200 dilution of anti-TLK antibody (Platero et al. 1995; Hari et al. 2001). DNA was stained with 1 μg/mL of DAPI.
Overexpression of TLK
The UAS/GAL4 system was used as decribed (Brand and Perrimon 1993). The UAS–tlk transgene was constructed by inserting the NotI–Asp718 fragment of LD15160, containing the full-length ORF of the tlk cDNA, into the pUAST vector. The UAS–tlkdead transgene contains a replacement of the aspartic acid residue in position 217 of catalytic domain by alanine (Fig. 1). The UAS–asf1 line was described previously (Moshkin et al. 2002). The UAS–tlk, UAS–tlkdead, and UAS–asf1 lines were crossed with the different Gal4 drivers described in the text. The heat-shock-dependent promoter (hsp70) was used to express the Gal4-dependent UAS–asf1 and/or UAS–tlk under conditions that have been described (Moshkin et al. 2002). For details on the driver lines used, see FlyBase (http://flybase.bio.indiana.edu).
GST pull-down
TLK and TLKdead were synthesized using rabbit reticulocyte in vitro transcription-translation system (IVT) in the presence of [35S]methionine, as described by the manufacturer (Promega). GST–ASF1 fusion protein was expressed and purified from bacteria (Moshkin et al. 2002). GST pull-down experiments with TLK or TLKdead were performed for 1–2 h at 4°C in binding buffer (20 mM HEPES-KOH at pH 7.6, 150 mM KCl, 2.5 mM MgCl2, 10% glycerol, 1% Triton X-100, 1 mM DTT, and 10 mg/mL BSA) with GST–ASF1 or GST (negative control) coupled to the Glutathione Sepharose 4B beads (Amersham). Beads were washed three times with 10-bed volumes of buffer (20 mM HEPES-KOH at pH 7.6; 150 mM KCl; 2.5 mM MgCl2; 1% Triton X-100; 1 mM DTT). Proteins were separated on 6% SDS-PAGE. Bound TLK or TLKdead were revealed by autoradiography.
TLK autophosphorylation and phosphatase treatments
For autophosphorylation studies, [35S]methionine radiolabeled IVT-produced TLK or TLKdead were incubated for 30 min at 30°C in TLK kinase buffer (50 mM Tris-HCl at pH 7.5; 10 mM MgCl2; 1 mM EGTA; 0.5 mM DTT; 5 mM NaF) in the presence of 10 μm ATP (Silljé et al. 1999). Dephosphorylation was performed with 10 U of calf intestinal alkaline phosphatase in supplied buffer (NEB) for 2 h at 37°C (Silljé et al. 1999). Reactions were subsequently diluted in GST pull-down binding buffer and incubated with GST–ASF1 coupled to beads for 1–2 h at 4°C. After washes, proteins were separated on 6% SDS-PAGE and revealed by autoradiography.
In vitro kinase assays
TLK–GST–ASF1 or TLKdead–GST–ASF1 complexes obtained after GST pull-downs with nonlabeled IVT-produced TLK or TLKdead were incubated for 30 min at 30°C in TLK kinase buffer in the presence of 10 μm ATP, 2 μCi of [γ-32P]ATP (Hartmann Analytic). The reactions were stopped by boiling in Laemmli SDS-PAGE loading buffer. GST alone was tested as negative control. To monitor the amount of complexes, proteins were stained with Coomassie after separation on 10% SDS-PAGE. TLK and TLKdead were detected by immunoblotting with anti-TLK antibodies (data not shown). 32P incorporation was revealed by autoradiography.
Acknowledgments
We thank T. Häder for initial work on tlk and U. Schäfer for providing the tlk P-element. We thank R. Maeda for comments on the manuscript, and D. Pauli, M. Zeidler, and R. Pflanz for discussions. We also thank J. Wuest for scanning EM, and P. Spierer for support. Special thanks to Olga Derkatch. We acknowledge E. Favre, G. Dowe, G. Faustino, and H. Taubert for technical assistance. Work was supported by grants from the Swiss National Foundation, the state of Geneva (F.K.), and the Max-Planck Society (H.J.).
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked “advertisement” in accordance with 18 USC section 1734 solely to indicate this fact.
Article and publication are at http://www.genesdev.org/cgi/doi/10.1101/gad.276703.
Corresponding authors.
References
- Adams M.D., Celniker, S.E., Holt, R.A., Evans, C.A., Gocayne, J.D., Amanatides, P.G., Scherer, S.E., Li, P.W., Hoskins, R.A., Galle, R.F., et al. 2000. The genome sequence of Drosophila melanogaster. Science 287: 2185-2195. [DOI] [PubMed] [Google Scholar]
- Badenhorst P., Voas, M., Rebay, I., and Wu, C. 2002. Biological functions of the ISWI chromatin remodeling complex NURF. Genes & Dev. 16: 3186-3198. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bakkenist C.J. and Kastan, M.B. 2003. DNA damage activates ATM through intermolecular autophosphorylation and dimer dissociation. Nature 421: 499-506. [DOI] [PubMed] [Google Scholar]
- Brand A.H. and Perrimon, N. 1993. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 118: 401-415. [DOI] [PubMed] [Google Scholar]
- Chou T.B. and Perrimon, N. 1996. The autosomal FLP-DFS technique for generating germline mosaics in Drosophila melanogaster. Genetics 144: 1673-1679. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dang D.T. and Perrimon, N. 1992. Use of a yeast site-specific recombinase to generate embryonic mosaics in Drosophila. Dev. Genet. 13: 367-375. [DOI] [PubMed] [Google Scholar]
- Deuring R., Fanti, L., Armstrong, J.A., Sarte, M., Papoulas, O., Prestel, M., Daubresse, G., Verardo, M., Moseley, S.L., Berloco, M., et al. 2000. The ISWI chromatin-remodeling protein is required for gene expression and the maintenance of higher order chromatin structure in vivo. Mol. Cell 5: 355-365. [DOI] [PubMed] [Google Scholar]
- Emili A., Schieltz, D.M., Yates III, J.R., and Hartwell, L.H. 2001. Dynamic interaction of DNA damage checkpoint protein Rad53 with chromatin assembly factor Asf1. Mol. Cell 7: 13-20. [DOI] [PubMed] [Google Scholar]
- Enomoto S. and Berman, J. 1998. Chromatin assembly factor I contributes to the maintenance, but not the re-establishment, of silencing at the yeast silent mating loci. Genes & Dev. 12: 219-232. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Foe V.E. 1989. Mitotic domains reveal early commitment of cells in Drosophila embryos. Development 107: 1-22. [PubMed] [Google Scholar]
- Groth A., Lukas, J., Nigg, E.A., Silljé, H.H.W., Wernstedt, C., Bartek, J., and Hansen, K. 2003. Human Tousled like kinases are targeted by a ATM- and Chk1-dependent DNA damage checkpoint. EMBO J. 22: 1676-1687. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gruss C., Wu, J., Koller, T., and Sogo, J.M. 1993. Disruption of the nucleosomes at the replication fork. EMBO J. 12: 4533-4545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hanks S.K., Quinn, A.M., and Hunter, T. 1988. The protein kinase family: Conserved features and deduced phylogeny of the catalytic domains. Science 241: 42-52. [DOI] [PubMed] [Google Scholar]
- Hari K.L., Cook, K.R., and Karpen, G.H. 2001. The Drosophila Su(var)2–10 locus regulates chromosome structure and function and encodes a member of the PIAS protein family. Genes & Dev. 15: 1334-1348. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hauptmann G. and Gerster, T. 1996. Multicolor whole mount in situ hybridisation to Drosophila embryos. Genes & Dev. Evol. 206: 292-295. [DOI] [PubMed] [Google Scholar]
- Hendzel M.J., Wei, Y., Mancini, M.A., VanHooser, A., Ranalli, T., Brinkley, B.R., BazettJones, D.P., and Allis, C.D. 1997. Mitosis-specific phosphorylation of histone H3 initiates primarily within pericentromeric heterochromatin during G2 and spreads in an ordered fashion coincident with mitotic chromosome condensation. Chromosoma 106: 348-360. [DOI] [PubMed] [Google Scholar]
- Jenuwein T. and Allis, C.D. 2001. Translating the histone code. Science 293: 1074-1080. [DOI] [PubMed] [Google Scholar]
- Le S., Davis, C., Konopka, J.B., and Sternglanz, R. 1997. Two new S-phase-specific genes from Saccharomyces cerevisiae. Yeast 13: 1029-1042. [DOI] [PubMed] [Google Scholar]
- Li Y., DeFatta, R., Anthony, C., Sunavala, G., and De Benedetti, A. 2001. A translationally regulated Tousled kinase phosphorylates histone H3 and confers radioresistance when overexpressed. Oncogene 20: 726-738. [DOI] [PubMed] [Google Scholar]
- Lucchini R., Wellinger, R.E., and Sogo, J.M. 2001. Nucleosome positioning at the replication fork. EMBO J. 20: 7294-7302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luger K., Mader, A.W., Richmond, R.K., Sargent, D.F., and Richmond, T.J. 1997. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389: 251-260. [DOI] [PubMed] [Google Scholar]
- Mello J.A. and Almouzni, G. 2001. The ins and outs of nucleosome assembly. Curr. Opin. Genet. Dev. 11: 136-141. [DOI] [PubMed] [Google Scholar]
- Mello J.A., Silljé, H.H., Roche, D.M., Kirschner, D.B., Nigg, E.A., and Almouzni, G. 2002. Human Asf1 and CAF-1 interact and synergize in a repair-coupled nucleosome assembly pathway. EMBO Rep. 3: 329-334. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Milan M., Diaz-Benjumea, F.J., and Cohen, S.M. 1998. Beadex encodes an LMO protein that regulates Apterous LIM-homeodomain activity in Drosophila wing development: A model for LMO oncogene function. Genes & Dev. 12: 2912-2920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moshkin Y.M., Armstrong, J.A., Maeda, R.K., Tamkun, J.W., Verrijzer, P., Kennison, J.A., and Karch, F. 2002. Histone chaperone ASF1 cooperates with the Brahma chromatin-remodelling machinery. Genes & Dev. 16: 2621-2626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nelson D.M., Ye, X., Hall, C., Santos, H., Ma, T., Kao, G.D., Yen, T.J., Harper, J.W., and Adams, P.D. 2002. Coupling of DNA synthesis and histone synthesis in S phase independent of cyclin/cdk2 activity. Mol. Cell. Biol. 22: 7459-7472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Peter A., Schöttler, P., Werner, M., Beinert, N., Dowe, G., Burkert, P., Mourkioti, F., Dentzer, L., He, Y., Deak, P., et al.. 2002. Mapping and identification of essential gene functions on the X chromosome of Drosophila. EMBO Rep. 3: 34-38. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Platero J.S., Hartnett, T., and Eissenberg, J.C. 1995. Functional analysis of the chromo domain of HP1. EMBO J. 14: 3977-3986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ray-Gallet D., Quivy, J.P., Scamps, C., Martini, E.M., Lipinski, M., and Almouzni, G. 2002. HIRA is critical for a nucleosome assembly pathway independent of DNA synthesis. Mol. Cell 9: 1091-1100. [DOI] [PubMed] [Google Scholar]
- Richards E.J. and Elgin, S.C. 2002. Epigenetic codes for heterochromatin formation and silencing: Rounding up the usual suspects. Cell 108: 489-500. [DOI] [PubMed] [Google Scholar]
- Roe J.L., Rivin, C.J., Sessions, R.A., Feldmann, K.A., and Zambryski, P.C. 1993. The Tousled gene in A. thaliana encodes a protein kinase homolog that is required for leaf and flower development. Cell 75: 939-950. [DOI] [PubMed] [Google Scholar]
- Roe J.L., Nemhauser, J.L., and Zambryski, P.C. 1997. TOUSLED participates in apical tissue formation during gynoecium development in Arabidopsis. Plant Cell 9: 335-353. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp J.A., Fouts, E.T., Krawitz, D.C., and Kaufman, P.D. 2001. Yeast histone deposition protein Asf1p requires Hir proteins and PCNA for heterochromatic silencing. Curr. Biol. 11: 463-473. [DOI] [PubMed] [Google Scholar]
- Shibahara K. and Stillman, B. 1999. Replication-dependent marking of DNA by PCNA facilitates CAF-1-coupled inheritance of chromatin. Cell 96: 575-585. [DOI] [PubMed] [Google Scholar]
- Silljé H.H. and Nigg, E.A. 2001. Identification of human Asf1 chromatin assembly factors as substrates of Tousled-like kinases. Curr. Biol. 11: 1068-1073. [DOI] [PubMed] [Google Scholar]
- Silljé H.H., Takahashi, K., Tanaka, K., Van Houwe, G., and Nigg, E.A. 1999. Mammalian homologues of the plant Tousled gene code for cell-cycle-regulated kinases with maximal activities linked to ongoing DNA replication. EMBO J. 18: 5691-5702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Smith S. and Stillman, B. 1991. Stepwise assembly of chromatin during DNA replication in vitro. EMBO J. 10: 971-980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sogo J.M., Stahl, H., Koller, T., and Knippers, R. 1986. Structure of replicating simian virus 40 minichromosomes. The replication fork, core histone segregation and terminal structures. J. Mol. Biol. 189: 189-204. [DOI] [PubMed] [Google Scholar]
- Sutton A., Bucaria, J., Osley, M.A., and Sternglanz, R. 2001. Yeast ASF1 protein is required for cell cycle regulation of histone gene transcription. Genetics 158: 587-596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Thompson J.D., Higgins, D.G., and Gibson, T.J. 1994. CLUSTAL W: Improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res. 22: 4673-4680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Turner B.M., Birley, A.J., and Lavender, J. 1992. Histone H4 isoforms acetylated at specific lysine residues define individual chromosomes and chromatin domains in Drosophila polytene nuclei. Cell 69: 375-384. [DOI] [PubMed] [Google Scholar]
- Tyler J.K. 2002. Chromatin assembly. Eur. J. Biochem. 269: 2268-2274. [DOI] [PubMed] [Google Scholar]
- Tyler J.K., Adams, C.R., Chen, S.R., Kobayashi, R., Kamakaka, R.T., and Kadonaga, J.T. 1999. The RCAF complex mediates chromatin assembly during DNA replication and repair. Nature 402: 555-560. [DOI] [PubMed] [Google Scholar]
- Tyler J.K., Collins, K.A., Prasad-Sinha, J., Amiott, E., Bulger, M., Harte, P.J., Kobayashi, R., and Kadonaga, J.T. 2001. Interaction between the Drosophila CAF-1 and ASF1 chromatin assembly factors. Mol. Cell. Biol. 21: 6574-6584. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Umehara T., Chimura, T., Ichikawa, N., and Horikoshi, M. 2002. Polyanionic stretch-deleted histone chaperone cia1/Asf1p is functional both in vivo and in vitro. Genes Cells 7: 59-73. [DOI] [PubMed] [Google Scholar]
- Wang Y., Zhang, W., Jin, Y., Johansen, J., and Johansen, K.M. 2001. The JIL-1 tandem kinase mediates histone H3 phosphorylation and is required for maintenance of chromatin structure in Drosophila. Cell 105: 433-443. [DOI] [PubMed] [Google Scholar]
- Wei Y., Yu, L.L., Bowen, J., Gorovsky, M.A., and Allis, C.D. 1999. Phosphorylation of histone H3 is required for proper chromosome condensation and segregation. Cell 97: 99-109. [DOI] [PubMed] [Google Scholar]
- Ye X.F., Franco, A.A., Santos, H., Nelson, D.M., Kaufman, P.D., and Adams, P.D. 2003. Defective S phase chromatin assembly causes DNA damage, activation of the S phase checkpoint, and S phase arrest. Mol. Cell 11: 341-351. [DOI] [PubMed] [Google Scholar]
- Zhang Z., Shibahara, K., and Stillman, B. 2000. PCNA connects DNA replication to epigenetic inheritance in yeast. Nature 408: 221-225. [DOI] [PubMed] [Google Scholar]