Abstract
Plasmids pAD1 (37.8 megadaltons) and pAD2 (17.1 megadaltons) of Streptococcus faecalis strain DS16 have been mapped with restriction enzymes. The location of a hemolysin-bacteriocin determinant on the conjugative pAD1 plasmid was derived from analyses of transposon insertions. Electron microscope and hybridization analyses located Tn917(Em) and the streptomycin (Sm) and kanamycin (Km) resistance determinants on the nonconjugative pAD2 plasmid. It was shown previously that the erythromycin (Em) resistance associated with Tn917 is inducible and that transposition from pAD2 to pAD1 is also stimulated by exposure of cells to low concentrations of Em. Here we show that inducing concentrations of Em also increase the conjugative transfer potential of pAD1; this is possibly related to a mild and short-lived inhibitory stress placed on the cells before full induction of resistance. Selection of Em-resistant transconjugants arising from matings between DS16 and a plasmid-free recipient gave rise to transconjugants which primarily harbor stable pAD1::pAD2 cointegrates. A 30-min exposure of donors to Em (0.5 microgram/ml) before mating resulted in a severalfold increase in the number of such transconjugants. However, a small fraction (e.g., 3 of 40) of these Emr Smr Kmr transconjugants harbored pAD1::Tn917 and pAD2 molecules. Since we believe pAD2 is incapable of being mobilized by pAD1 without being covalently linked, it is likely that transfer in these cases involved cointegrates representing structural intermediates in the transposition of Tn917 from pAD2 to pAD1. It follows that such intermediates probably had two copies of Tn917 and readily resolved after transfer. (These cointegrates are different from the stable cointegrates which were shown to have only a single copy of Tn917; the latter are assumed not to be related to transposition.) Two variants with altered Tn917 transposition properties were derived. One of them transposed at an elevated frequency, whereas the other showed no detectabel transposition. In neither case was transposition influenced by Em exposure; however, both remained inducible for Em resistance.
Full text
PDF


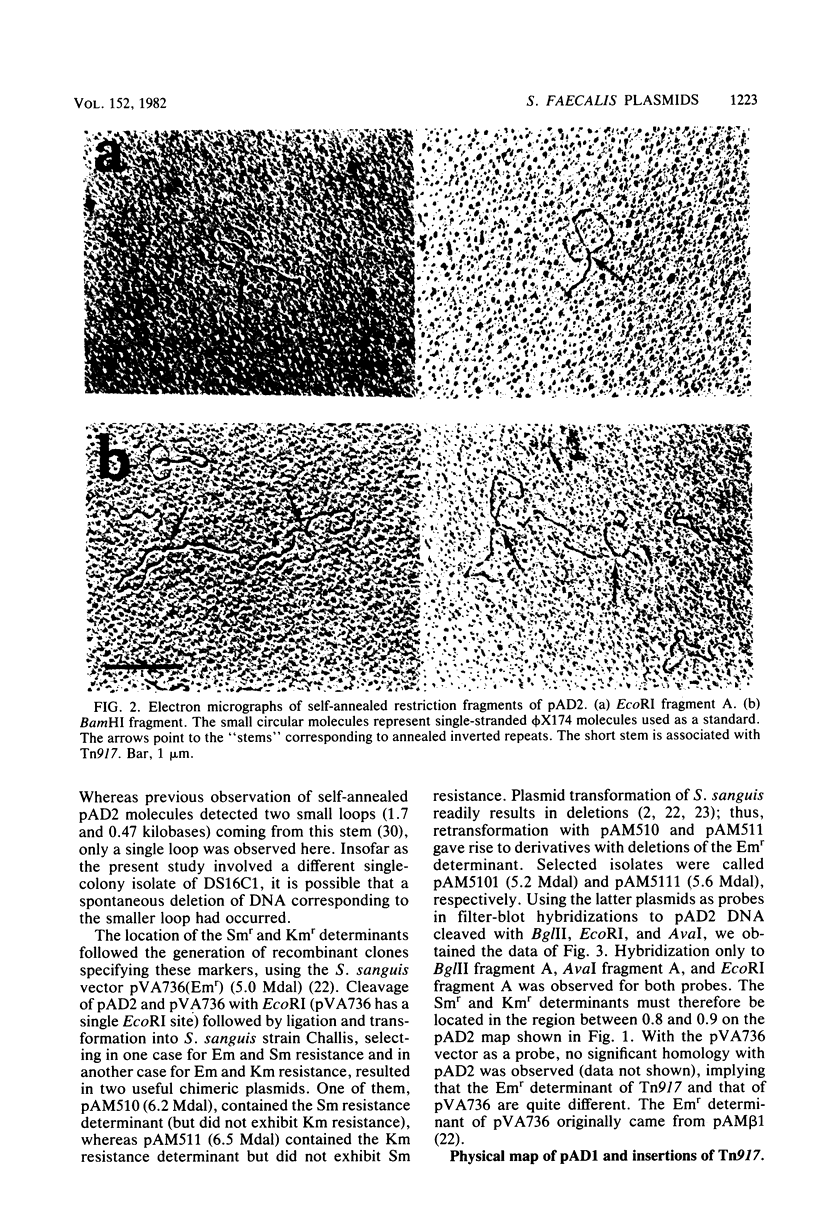


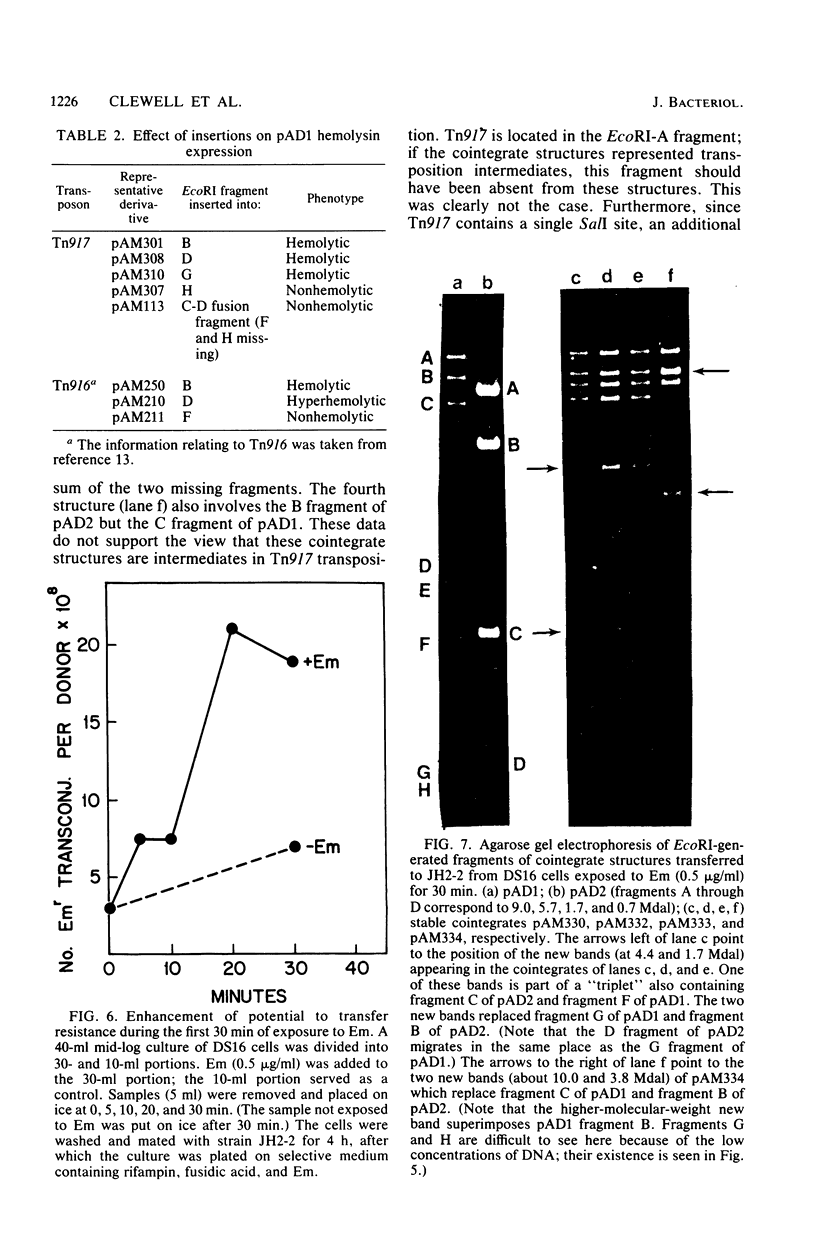

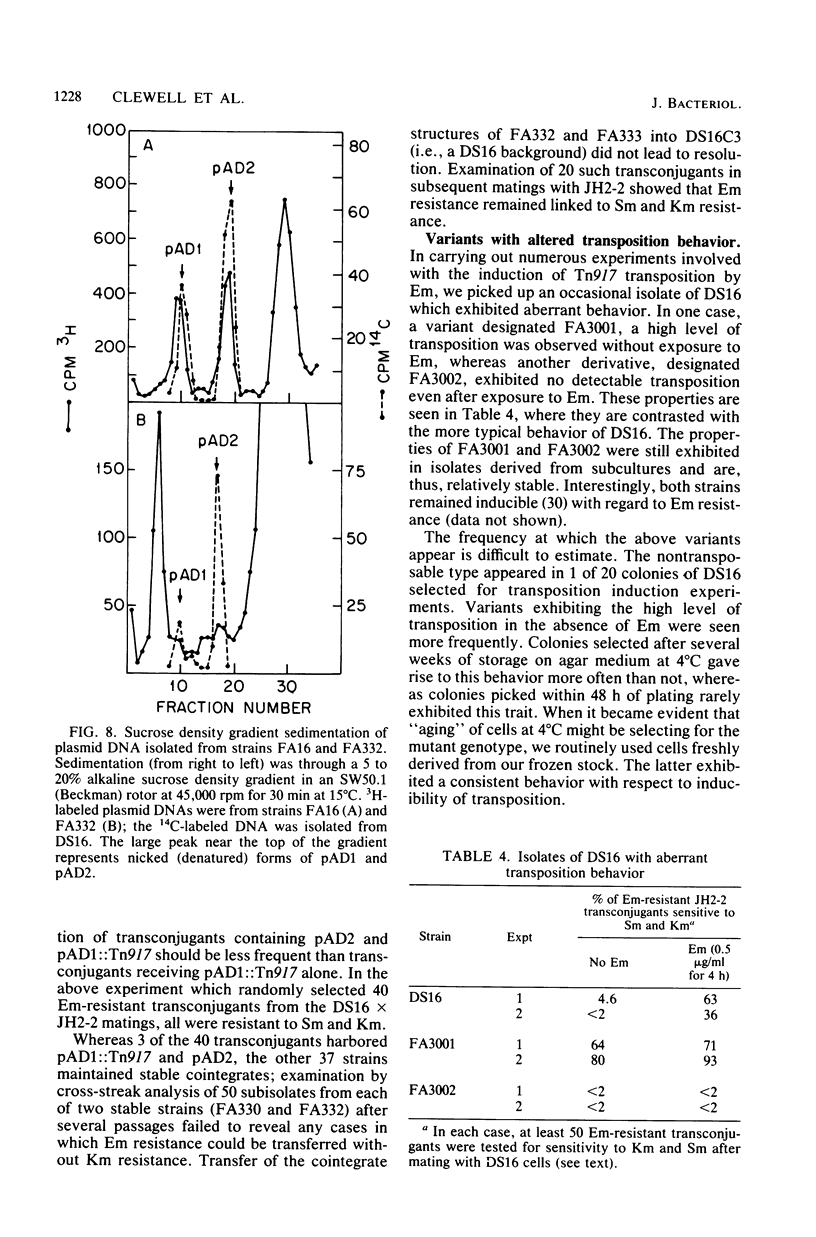


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arthur A., Sherratt D. Dissection of the transposition process: a transposon-encoded site-specific recombination system. Mol Gen Genet. 1979 Oct 1;175(3):267–274. doi: 10.1007/BF00397226. [DOI] [PubMed] [Google Scholar]
- Behnke D. Plasmid transformation of Streptococcus sanguis (Challis) occurs by circular and linear molecules. Mol Gen Genet. 1981;182(3):490–497. doi: 10.1007/BF00293940. [DOI] [PubMed] [Google Scholar]
- Clark A. J., Warren G. J. Conjugal transmission of plasmids. Annu Rev Genet. 1979;13:99–125. doi: 10.1146/annurev.ge.13.120179.000531. [DOI] [PubMed] [Google Scholar]
- Clewell D. B., Helinski D. R. Properties of a supercoiled deoxyribonucleic acid-protein relaxation complex and strand specificity of the relaxation event. Biochemistry. 1970 Oct 27;9(22):4428–4440. doi: 10.1021/bi00824a026. [DOI] [PubMed] [Google Scholar]
- Clewell D. B. Plasmids, drug resistance, and gene transfer in the genus Streptococcus. Microbiol Rev. 1981 Sep;45(3):409–436. doi: 10.1128/mr.45.3.409-436.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clewell D. B., Yagi Y., Dunny G. M., Schultz S. K. Characterization of three plasmid deoxyribonucleic acid molecules in a strain of Streptococcus faecalis: identification of a plasmid determining erythromycin resistance. J Bacteriol. 1974 Jan;117(1):283–289. doi: 10.1128/jb.117.1.283-289.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crisona N. J., Clark A. J. Increase in conjugational transmission frequency of nonconjugative plasmids. Science. 1977 Apr 8;196(4286):186–187. doi: 10.1126/science.322280. [DOI] [PubMed] [Google Scholar]
- Crisona N. J., Nowak J. A., Nagaishi H., Clark A. J. Transposon-mediated conjugational transmission of nonconjugative plasmids. J Bacteriol. 1980 May;142(2):701–713. doi: 10.1128/jb.142.2.701-713.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunny G. M., Brown B. L., Clewell D. B. Induced cell aggregation and mating in Streptococcus faecalis: evidence for a bacterial sex pheromone. Proc Natl Acad Sci U S A. 1978 Jul;75(7):3479–3483. doi: 10.1073/pnas.75.7.3479. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunny G. M., Clewell D. B. Transmissible toxin (hemolysin) plasmid in Streptococcus faecalis and its mobilization of a noninfectious drug resistance plasmid. J Bacteriol. 1975 Nov;124(2):784–790. doi: 10.1128/jb.124.2.784-790.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunny G. M., Craig R. A., Carron R. L., Clewell D. B. Plasmid transfer in Streptococcus faecalis: production of multiple sex pheromones by recipients. Plasmid. 1979 Jul;2(3):454–465. doi: 10.1016/0147-619x(79)90029-5. [DOI] [PubMed] [Google Scholar]
- Franke A. E., Clewell D. B. Evidence for a chromosome-borne resistance transposon (Tn916) in Streptococcus faecalis that is capable of "conjugal" transfer in the absence of a conjugative plasmid. J Bacteriol. 1981 Jan;145(1):494–502. doi: 10.1128/jb.145.1.494-502.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Franke A. E., Clewell D. B. Evidence for conjugal transfer of a Streptococcus faecalis transposon (Tn916) from a chromosomal site in the absence of plasmid DNA. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):77–80. doi: 10.1101/sqb.1981.045.01.014. [DOI] [PubMed] [Google Scholar]
- Gill R., Heffron F., Dougan G., Falkow S. Analysis of sequences transposed by complementation of two classes of transposition-deficient mutants of Tn3. J Bacteriol. 1978 Nov;136(2):742–756. doi: 10.1128/jb.136.2.742-756.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goebel W., Lindenmaier W., Pfeifer F., Schrempf H., Schelle B. Transposition and insertion of intact, deleted and enlarged ampicillin transposon Tn3 from mini-R1 (Rsc) plasmids into transfer factors. Mol Gen Genet. 1977 Nov 29;157(2):119–129. doi: 10.1007/BF00267389. [DOI] [PubMed] [Google Scholar]
- Guyer M. S. The gamma delta sequence of F is an insertion sequence. J Mol Biol. 1978 Dec 15;126(3):347–365. doi: 10.1016/0022-2836(78)90045-1. [DOI] [PubMed] [Google Scholar]
- Heffron F., Bedinger P., Champoux J. J., Falkow S. Deletions affecting the transposition of an antibiotic resistance gene. Proc Natl Acad Sci U S A. 1977 Feb;74(2):702–706. doi: 10.1073/pnas.74.2.702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heffron F., McCarthy B. J., Ohtsubo H., Ohtsubo E. DNA sequence analysis of the transposon Tn3: three genes and three sites involved in transposition of Tn3. Cell. 1979 Dec;18(4):1153–1163. doi: 10.1016/0092-8674(79)90228-9. [DOI] [PubMed] [Google Scholar]
- Horinouchi S., Weisblum B. Posttranscriptional modification of mRNA conformation: mechanism that regulates erythromycin-induced resistance. Proc Natl Acad Sci U S A. 1980 Dec;77(12):7079–7083. doi: 10.1073/pnas.77.12.7079. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacob A. E., Hobbs S. J. Conjugal transfer of plasmid-borne multiple antibiotic resistance in Streptococcus faecalis var. zymogenes. J Bacteriol. 1974 Feb;117(2):360–372. doi: 10.1128/jb.117.2.360-372.1974. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kopecko D. J., Cohen S. N. Site specific recA--independent recombination between bacterial plasmids: involvement of palindromes at the recombinational loci. Proc Natl Acad Sci U S A. 1975 Apr;72(4):1373–1377. doi: 10.1073/pnas.72.4.1373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macrina F. L., Jones K. R., Wood P. H. Chimeric streptococcal plasmids and their use as molecular cloning vehicles in Streptococcus sanguis (Challis). J Bacteriol. 1980 Sep;143(3):1425–1435. doi: 10.1128/jb.143.3.1425-1435.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Macrina F. L., Tobian J. A., Jones K. R., Evans R. P. Molecular cloning in the Streptococci. Basic Life Sci. 1982;19:195–210. doi: 10.1007/978-1-4684-4142-0_17. [DOI] [PubMed] [Google Scholar]
- Meyer R., Boch G., Shapiro J. Transposition of DNA inserted into deletions of the Tn5 kanamycin resistance element. Mol Gen Genet. 1979 Mar 9;171(1):7–13. doi: 10.1007/BF00274009. [DOI] [PubMed] [Google Scholar]
- Shapiro J. A. Molecular model for the transposition and replication of bacteriophage Mu and other transposable elements. Proc Natl Acad Sci U S A. 1979 Apr;76(4):1933–1937. doi: 10.1073/pnas.76.4.1933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sharp P. A., Hsu M. T., Otsubo E., Davidson N. Electron microscope heteroduplex studies of sequence relations among plasmids of Escherichia coli. I. Structure of F-prime factors. J Mol Biol. 1972 Nov 14;71(2):471–497. doi: 10.1016/0022-2836(72)90363-4. [DOI] [PubMed] [Google Scholar]
- Sherratt D., Arthur A., Burke M. Transposon-specified, site-specific recombination systems. Cold Spring Harb Symp Quant Biol. 1981;45(Pt 1):275–281. doi: 10.1101/sqb.1981.045.01.040. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Tomich P. K., An F. Y., Clewell D. B. A transposon (Tn917) in Streptococcus faecalis that exhibits enhanced transposition during induction of drug resistance. Cold Spring Harb Symp Quant Biol. 1979;43(Pt 2):1217–1221. doi: 10.1101/sqb.1979.043.01.138. [DOI] [PubMed] [Google Scholar]
- Tomich P. K., An F. Y., Clewell D. B. Properties of erythromycin-inducible transposon Tn917 in Streptococcus faecalis. J Bacteriol. 1980 Mar;141(3):1366–1374. doi: 10.1128/jb.141.3.1366-1374.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tomich P. K., An F. Y., Damle S. P., Clewell D. B. Plasmid-related transmissibility and multiple drug resistance in Streptococcus faecalis subsp. zymogenes strain DS16. Antimicrob Agents Chemother. 1979 Jun;15(6):828–830. doi: 10.1128/aac.15.6.828. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wahl G. M., Stern M., Stark G. R. Efficient transfer of large DNA fragments from agarose gels to diazobenzyloxymethyl-paper and rapid hybridization by using dextran sulfate. Proc Natl Acad Sci U S A. 1979 Aug;76(8):3683–3687. doi: 10.1073/pnas.76.8.3683. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yagi Y., Clewell D. B. Plasmid-determined tetracycline resistance in Streptococcus faecalis: tandemly repeated resistance determinants in amplified forms of pAMalpha1 DNA. J Mol Biol. 1976 Apr 15;102(3):583–600. doi: 10.1016/0022-2836(76)90336-3. [DOI] [PubMed] [Google Scholar]