Abstract
The role of CD4+ T cells in the control of human immunodeficiency virus (HIV) and simian immunodeficiency virus (SIV) replication is not well understood. Even though strong HIV- and SIV-specific CD4+ T-cell responses have been detected in individuals that control viral replication, major histocompatibility complex class II (MHC-II) molecules have not been definitively linked with slow disease progression. In a cohort of 196 SIVmac239-infected Indian rhesus macaques, a group of macaques controlled viral replication to less than 1,000 viral RNA copies/ml. These elite controllers (ECs) mounted a broad SIV-specific CD4+ T-cell response. Here, we describe five macaque MHC-II alleles (Mamu-DRB*w606, -DRB*w2104, -DRB1*0306, -DRB1*1003, and -DPB1*06) that restricted six SIV-specific CD4+ T-cell epitopes in ECs and report the first association between specific MHC-II alleles and elite control. Interestingly, the macaque MHC-II alleles, Mamu-DRB1*1003 and -DRB1*0306, were enriched in this EC group (P values of 0.02 and 0.05, respectively). Additionally, Mamu-B*17-positive SIV-infected rhesus macaques that also expressed these two MHC-II alleles had significantly lower viral loads than Mamu-B*17-positive animals that did not express Mamu-DRB1*1003 and -DRB1*0306 (P value of <0.0001). The study of MHC-II alleles in macaques that control viral replication could improve our understanding of the role of CD4+ T cells in suppressing HIV/SIV replication and further our understanding of HIV vaccine design.
CD4+ T cells play a pivotal role in adaptive immunity. These cells are required for the development and maintenance of memory and cytotoxic CD8+ T cells (1, 2). In chronic viral infections, CD8+ T cells become exhausted in the absence of CD4+ T-cell help (19, 31). Thus, the ability of the immune system to initiate an effective response to a pathogen requires the active involvement of CD4+ T cells (18, 48, 50).
Previous observations have suggested that CD4+ T cells may be important in controlling lentivirus replication. Human immunodeficiency virus (HIV)-infected patients who maintained low viral loads had strong CD4+ T-cell responses (46). Similarly, in elite controller (EC) rhesus macaques, simian immunodeficiency virus (SIV)-specific CD4+ T-cell responses remained strong even after experimental, in vivo depletion of CD8+ cells and consequent viral recrudescence (15). Nonetheless, because HIV and SIV infections are characterized by a profound depletion of CD4+ T cells during the acute phase of infection (26, 33), the exact role of these cells in containing HIV or SIV replication has been difficult to define. This depletion is directed primarily to the gut-associated lymphoid tissue (4, 33, 35, 53; reviewed in references 17 and 43). Finally, HIV-specific CD4+ T cells may be preferentially targeted by the virus (3, 10, 11), further complicating the role that these cells play in control of viral replication.
While it is difficult to assign a role to CD4+ T cells because of their depletion early after infection, preservation of these cells in vaccinated humans and macaques has been correlated with a delay in the onset of AIDS-like symptoms (21, 25, 27, 32, 55). Likewise, rhesus macaques infected with live attenuated SIVmac239Δnef that induced high levels of SIV-specific CD4+ T-cell responses were protected against challenge with wild-type SIVmac239 (16). However, it is unclear whether the high numbers of SIV-specific CD4+ T cells observed after SIVmac239Δnef infection are a cause or effect of reduced viral replication. In fact, a strong, vaccine-induced CD4+ T-cell response to the SIV Env protein was correlated with accelerated disease progression in one study (49). Thus, despite increasing evidence of the importance of the CD4+ T-cell compartment in control of viral replication, the precise role of these cells remains poorly understood.
Several major histocompatibility complex class I (MHC-I) alleles are associated with control of HIV and SIV replication (6, 28, 59). In particular, the MHC-I alleles HLA-B57 and -B27 in HIV-infected patients (34, 37) and Mamu-B*17 and -B*08 in SIV-infected rhesus macaques have been correlated with control of viral replication (28, 59). Although individuals who controlled viral replication had strong HIV-specific CD4+ T-cell responses (15, 46), no clear association of particular MHC-II molecules with protection against disease progression has been demonstrated. However, an HLA class II haplotype, DRB1*13-DQB1*06, was overrepresented in individuals who maintained viral control after stopping treatment with antiretroviral drugs (29). In addition, homozygosity of a haplotype containing Mamu-DQB1*0601 was associated with faster disease progression in SIV-infected rhesus macaques (47).
In a cohort of 196 SIVmac239-infected Indian rhesus macaques, we identified sixteen ECs, most of which have controlled SIV replication to less than 1,000 viral RNA copies/ml for more than 1 year (15, 28, 59). Six of these ECs were previously used in a CD8+ T-cell depletion study (15). These ECs mounted strong SIV-specific CD4+ T-cell responses after CD8-depletion, several of which were shared among the ECs (15). The presence of these shared responses in ECs led us to hypothesize that macaque MHC-II alleles present in ECs restrict CD4+ T-cell epitopes that might be crucial for control of disease progression. We used EC 95061 and five ECs from the CD8-depletion study to map six SIV-specific CD4+ T-cell responses. We then determined the restricting MHC-II allele for each response. The entire cohort of 196 SIVmac239-infected Indian rhesus macaques was then typed for these MHC-II alleles to elucidate the association of these alleles with control of viral replication. Surprisingly, we found that two MHC-II alleles, Mamu-DRB1*1003 and -DRB1*0306, were significantly enriched in the EC group. Moreover, Mamu-B*17-positive rhesus macaques that expressed Mamu-DRB1*1003 and -DRB1*0306 had significantly lower viral loads than animals that expressed Mamu-B*17 but did not express Mamu-DRB1*1003 and -DRB1*0306. This is the first study to demonstrate an association between particular MHC-II alleles and elite control of SIV replication in SIV-infected Indian rhesus macaques.
MATERIALS AND METHODS
Animals.
SIVmac239-infected Indian rhesus macaques (Macaca mulatta) were maintained at the Wisconsin National Primate Research Center (WNPRC; University of Wisconsin—Madison, WI). Animals were cared for according to regulations set by the Guide for the Care and Use of Laboratory Animals of the National Research Council, as approved by the University of Wisconsin Institutional Animal Care and Use Committee.
Peptides.
All peptides of 15 amino acids in length were obtained through the AIDS Research and Reference reagent program, Division of AIDS, National Institute of Allergy and Infectious Diseases, National Institutes of Health (NIH): human interleukin-2 ([IL-2] item 136; Hoffman-La Roche, Inc.) as well as complete SIVmac239 peptide sets (15-mers overlapping by 11 amino acids) of Gag (item 6204), Vif (item 6205), Tat (item 6207), Pol (item 6443), Rev (item 6448), Vpr (item 6449), Vpx (item 6450), Env (item 6883), and full-length Nef (item 8762) (all from the Division of AIDS).
Cell lines and clones.
CD4+ T-cell lines were generated from whole blood peripheral blood mononuclear cells (PBMCs) separated by Ficoll density gradient centrifugation. After separation, PBMCs were depleted of CD8+ cells using anti-CD8 nonhuman primate microbeads on an AutoMACS bead separation column according to the manufacturer's protocol (Miltenyi, Auburn, CA). Depletions were 99% effective, as confirmed by surface staining with anti-CD4 antibody labeled with allophycocyanin (APC; Becton Dickinson, San Jose, CA) and anti-CD8 antibody labeled with peridinin chlorophyll protein (Becton Dickinson, San Jose, CA) (data not shown). The CD8-depleted fraction was incubated at 37°C and 5% CO2 in R15 medium with IL-7 (R15 consists of RPMI medium, 15% fetal calf serum, 1% l-glutamine, 1% antibiotic/antimycotic agent, and 50 ng/ml of IL-7) and stimulated with irradiated autologous B-lymphoblastic cell lines (BLCL) as antigen-presenting cells pulsed with SIVmac239 peptide for 3 days. R15-50 medium (R15 medium with 50 U/ml of IL-2 instead of IL-7) was added on the fourth day. On day 8, the cell line was stimulated again with irradiated autologous BLCL pulsed with SIVmac239 peptide or pooled peptides. Pool-specific or peptide-specific CD4+ T-cell lines were analyzed for peptide specificity after three rounds of restimulation. To generate SIV-specific CD4+ T-cell clones, we carried out three rounds of limiting dilution and expansion, as previously described (7, 58).
ICS.
To determine the specificity of the CD4+ T-cell lines and the MHC-II restricting alleles, intracellular cytokine stain (ICS) was used as previously described (55). Briefly, 2 × 105 BLCL or MHC-II transferent RM3 cells (5; also see below) were pulsed with 1 μM peptide for 90 min and washed three times with R10 medium (R10 consists of RPMI medium, 10% fetal calf serum, 1% l-glutamine, 1% antibiotic/antimycotic). Subsequently, the pulsed cells were added to 2 × 105 CD4+ T cells and incubated for 90 min at 37°C and 5% CO2. As negative controls, autologous BLCL or RM3 cells were pulsed with irrelevant peptide or no peptide. After 90 min of incubation, brefeldin A was added to the cells to stop the Golgi-mediated transport of proteins, and the CD4+ T cells with the BLCL or RM3 cells were incubated for five more hours. Cells were then stained with anti-CD4 antibody labeled with APC (Becton Dickinson, San Jose, CA) and anti-CD8 antibody labeled with peridinin chlorophyll protein (Becton Dickinson, San Jose, CA) and fixed with 2% paraformaldehyde (PFA) overnight at 4°C. Following overnight incubation, the cells were permeabilized with saponin and stained with anti-gamma interferon (IFN-γ) antibody labeled with fluorescein isothiocyanate (FITC; BD Biosciences, San Jose CA) and anti-tumor necrosis factor alpha antibody labeled with phycoerythrin (BD Biosciences, San Jose CA), fixed with 2% PFA, and stored at 4°C until analysis. Data were collected using a FACSCalibur flow cytometer (Becton Dickinson), and at least 50,000 events were collected. The data were analyzed using FlowJo (version 8.1.1) for Macintosh.
ELISPOT assay.
An enzyme-linked immunospot (ELISPOT) assay was used to quantify IFN-γ-positive responses in PBMCs depleted in vitro of CD8+ cells, map CD4+ T-cell responses, and define the macaque MHC-II restricting alleles by using RM3 cell transferents. When fresh PBMCs were used, we performed the assay as previously described (55). Fresh CD8-depleted PBMCs were isolated to map SIV-specific CD4+ T-cell responses using several pools of 10 peptides of 15 amino acids in length, overlapping by 11 amino acids. These pools comprised most of the viral proteome. Fresh CD8-depleted PBMCs (2 × 105) were added to each well along with a 1 μM concentration of a peptide pool and incubated overnight at 37°C and 5% CO2. To map CD4+ T-cell responses, we added 2 × 103 BLCL, 1 × 103 to 2 × 103 CD4+ T cells, and a 1 μM concentration of relevant peptide per well and incubated the culture overnight at 37°C and 5% CO2. Plates were processed on day 2 as previously described (55). CD4+ T-cell lines or clones were tested for MHC-II allele restriction using autologous BLCL (as a positive control) or MHC-II-transferred RM3 cells that were pulsed with 1 μM peptide for 90 min at 37°C and 5% CO2 and washed three times with R10 medium. At least 7 × 103 peptide-pulsed BLCL or RM3 cells were added to each ELISPOT well with 1 × 103 to 2 × 103 CD4+ T cells and incubated overnight at 37°C and 5% CO2. Plates were processed on day 2 as previously described (55). As negative controls, BLCL or RM3 cells were pulsed with irrelevant peptide or no peptide. Each sample was tested at least in duplicate. The plates were read using an AID EliSpot reader and software, version 4.0 (Strassburg, Germany).
EC cDNA libraries.
To exhaustively MHC type animals and capture macaque MHC-II alleles for further assays, cDNA libraries for each EC were developed. We cultured 5 × 108 to 10 × 108 PBMCs or BLCL from each EC macaque and extracted total RNA using an RNeasy Protect Mini kit from Qiagen (Valencia, CA) according to the manufacturer's protocol. Then the mRNA was isolated from total RNA using an Oligotex mRNA midi kit from Qiagen (Valencia, CA) following the manufacturer's protocol. The extracted mRNA was used as a template to generate cDNA. The cDNA was ligated directionally into the pCMV.SPORT6 expression vector, using the Superscript Plasmid System Gateway Technology for cDNA Synthesis and Cloning from Invitrogen (Burlingame, CA) according to the manufacturer's protocol. Chemically competent Escherichia coli DH5α cells (Invitrogen, Carlsbad, CA) were transformed using the cDNA. Ampicillin-resistant colonies were plated on 150-mm plates and incubated overnight at 34°C. The following day, we added 5 ml of broth to the plate and scraped the colonies into the broth. The recombinant plasmid cDNA from these colonies was isolated and purified with a HiSpeed Plasmid Midi Kit from Qiagen (Valencia, CA). To capture macaque MHC-II genes from the cDNA libraries, we used a Gene Trapper cDNA positive selection kit from Invitrogen (Carlsbad, CA), following the manufacturer's instructions. The probes used for MHC-II capture were the following: DRA, 5′-CAACGTCCTCATCTGTTTCATCGA3-′; DRB, 5′-GCMCAGARCAAGATSCTGAGTGGW-3′; DQA, 5′-SCGCARBKTGCACTGAGAAAC-3′; DQB, 5′-TGTGCTACTWCRYCAACKGGA-3′; DPA, 5′-TCTGGCATCTGGAGGAGTTTG-3′; and DPB, 5′-GCAGCTCTTTTCATTTTGCCATCC-3′. Chemically competent E. coli DH5α cells (Invitrogen, Carlsbad, CA) were transformed with the captured MHC-II cDNA and spread onto LB agar plates (100 μg/ml ampicillin) overnight at 34°C. Colonies were single picked and grown overnight in 1 ml of CircleGrow broth with 100 μg/ml ampicillin. Plasmid cDNA was isolated the next day using a Perfectprep Plasmid 96 Vac Direct Bind kit from Eppendorf (Hamburg, Germany) according to the manufacturer's protocol. Plasmid cDNA was sequenced on an ABI 3730 DNA analyzer (Applied Biosystems, Foster City, CA) with the SP6 primer (5′-GGCCTATTTAGGTGACACTATAG-3′). More than 100 macaque MHC-II clones were identified for each cDNA library, and then clones of interest were chosen for full-length sequencing. All of the macaque MHC-II alleles sequenced from the six EC cDNA libraries are available at http://ink.primate.wisc.edu/∼watkins/supplemental.html.
RM3 transferents.
The RM3 cell line is a derivative of a human Epstein-Barr virus-transformed BLCL Raji cell line that does not express MHC-II molecules (5). To transfect a single macaque MHC-II α-chain allele and a single β-chain allele into RM3 cells, we used the cell line Nucleofector kit C and the electroporator Nucleofector I from Amaxa Inc. (Gaithersburg, MD) and followed the manufacturer's instructions. Briefly, 5 × 106 RM3 cells were washed in R10 medium and resuspended in 100 μl of Nucleofector solution C. Five micrograms of a pCMV.SPORT6 plasmid construct bearing a macaque MHC-II α-chain allele and 5 μg of a pCMV.SPORT6 plasmid construct bearing a macaque MHC-II β-chain allele were added to the cells and placed in an electroporation cuvette. The cell solution was electroporated using the Nucleofector I with the program G-16 (Amaxa Inc). Shortly after electroporation, the cells were transferred to a 12-well plate with 1.5 ml of warm (37°C) R10 medium and incubated at 37°C and 5% CO2 overnight. Following the overnight incubation, 2.5 ml of R10 medium was added, and the culture was incubated for three more days at 37°C and 5% CO2. During the fourth day of incubation, 1 × 105 transfected RM3 cells were pulsed with peptide for 90 min and washed three times, and CD4+ T-cell lines or clones were added to test in ELISPOT or ICS assays as mentioned above.
Stable RM3 transferents that express a single MHC-II α allele and a single MHC-II β allele were also generated for the alleles DRB1*1003 and DRB1*0306. The plasmids used for stable transferents were made by excising the MHC-II allele from the pCMV.SPORT6 plasmid with KpnI and NotI in NEB2 buffer. The digestion was run on an agarose gel to isolate the 1,000-bp insert, and to purify the insert a Qiagen QIAquick gel extraction kit (Valencia, CA) was used. Each MHC-II chain was ligated to a separate plasmid, pcDNA3.1(+)/Neo or pcDNA3.1/Hygro. The plasmids ligated to an MHC-II allele were used to transform chemically competent E. coli DH5α; Invitrogen, Camarillo, CA) overnight on LB agar plates plus ampicillin (100 μg/ml). The following day, one colony was picked and grown overnight in LB broth with ampicillin (100 μg/ml). The next day the plasmid constructs containing the MHC-II alleles were extracted from the chemically competent E. coli DH5α (Invitrogen, Camarillo, CA) using a HiSpeed Plasmid Midi kit from Qiagen and resuspended to a concentration of more than 1 μg/μl. The protocol for transfection of RM3 cells with the plasmid constructs pcDNA3.1(+)/Neo-pcDNA3.1/Hygro carrying an MHC-II allele was the same as described above. The transfected (pcDNA3.1(+)/Neo-pcDNA3.1/Hygro) RM3 cells underwent drug selection in R10 medium with selection markers G418 (500 μg/ml) and hygromycin B (500 μg/ml) for 1 month at 37°C in 5% CO2 before being tested.
To test the percentage of transfected RM3 cells expressing the MHC-II allele of interest, we did surface staining with 2 × 105 RM3 cells using anti-HLA-DR antibody conjugated with FITC (BD Biosciences, San Jose, CA), anti-HLA-DP antibody (Becton Dickinson, San Jose CA,), or anti-HLA-DQ antibody (Abcam Inc. Cambridge, MA); cells were incubated for 1 h at room temperature and washed twice. If the cells were stained with anti-HLA-DP or anti-HLA-DQ antibodies, a secondary mouse anti-immunoglobulin G antibody conjugated with FITC (Abcam Inc., Cambridge, CA) was added and incubated at room temperature for 1 h, followed by two washes. Finally, the transferents were fixed with 2% PFA and stored at 4°C until analysis using a FACSCalibur flow cytometer (Becton Dickinson).
MHC-II typing.
A group of 196 SIVmac239-infected rhesus macaques from the WNPRC (University of Wisconsin, Madison, WI) and the National Cancer Institute cohort were MHC-II typed for the Mamu-DRB1*0306, Mamu-DRB1*1003, Mamu-DRB*w2104, Mamu-DRB*w606, and Mamu-DPB1*06 alleles by using either group- or allele-specific amplification primers targeting unique polymorphisms within exon 2 (D. L. Fisk et al., unpublished data). Allele-level resolution was achieved and confirmed by direct DNA sequencing of the amplified PCR products.
Statistical analysis.
Statistical analyses were done as previously described (28, 59). Briefly, for analysis of the number and percentage of each allele in the cohort of 196 SIV-infected rhesus macaques shown in Table 3, we used the program PROC FREQ (SAS institute, Cary, NC). The data to calculate the enrichment of a particular MHC-II allele in the EC group was generated using PROC LOGISTIC SAS, version 9.1 (SAS institute, Cary, NC). To calculate the relative log geometric mean for the chronic phase we used PROC MIXED (SAS institute, Cary, NC). The two groups in Table 4 were independent, and the analysis was done using a non-paired t test.
TABLE 3.
Particular MHC class II alleles were significantly enriched in an EC cohorta
| Animalb | Presence of the indicated MHC-II allelec
|
|||||
|---|---|---|---|---|---|---|
| Mamu-DRB1*0306 | Mamu-DRB1*1003 | Mamu-DPB1*06 | Mamu-DRB*w606 | Mamu-DRB*w2104 | Mamu-B*17d | |
| 00032 | − | − | + | − | − | − |
| 02019 | − | − | − | − | − | − |
| 98057 | − | − | − | − | − | − |
| 99x032 | − | − | + | − | − | − |
| C59Z | − | − | − | + | + | − |
| 00078 | + | + | − | − | − | − |
| 01064 | + | + | − | − | − | − |
| 95061 | + | + | + | + | + | + |
| 95071 | + | + | + | − | − | + |
| 96112 | + | + | − | − | − | + |
| 98016 | + | + | − | − | − | + |
| AJ11 | + | + | + | − | − | + |
| 95096 | − | − | + | − | − | + |
| 98014 | − | − | + | − | − | + |
| 99C051 | − | − | − | − | − | + |
| 96C114 | − | − | − | − | − | + |
| Frequency in EC (%)e | 44 | 44 | 44 | 13 | 13 | 69d |
| Frequency in whole cohort (%) | 23 | 20 | 36 | 11 | 10 | 19d |
ECs were defined as animals that maintained plasma virus concentrations below 1,000 viral RNA copies/ml during the chronic phase. Most animals were infected for more than 1 year. Animals 00032 and 02019 were part of an ongoing study and had been infected for less than 1 year (28).
Animals 95061, 95096, 96112, 98014, 99C051, and 96C114 were vaccinated before challenge; however, only vaccines not associated with control were included (59). Animals 96C114 and C59Z were housed at the National Cancer Institute.
A total of 196 SIV-infected animals were genotyped using sequence-specific PCR (D. L. Fisk et al., unpublished data). Additionally, animals 00078, 01064, 95061, 95071, 95096, 98016 and AJ11 were genotyped using cDNA libraries.
Mamu-B*17 has been shown to be significantly enriched in ECs, as previously published (59).
Bold numbers represent a significant enrichment for the EC group (n = 16). The enrichment was determined using PROC LOGISTIC in SAS, version 9.1 (SAS Institute, Cary, NC). For Mamu-DRB1*0306, P = 0.05; for Mamu-DRB1*1003, P = 0.02.
TABLE 4.
Difference in SIV viral loads between Indian rhesus macaques that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 and animals that expressed Mamu-B*17 but did not express Mamu-DRB1*1003 or -DRB1*0306
| Allele(s) expressed by group | No. of animals in group (no. of ECs) | Mean log viral load | SD | t value | P value |
|---|---|---|---|---|---|
| Mamu-B*17, -DRB1*1003, -DRB1*0306 | 13 (5) | 3.91 | 1.83 | ||
| Mamu-B*17 (not Mamu-DRB1*1003 or -DRB1*0306) | 28 (4) | 5.05 | 1.43 | 9.38 | <0.0001 |
Microsatellite analysis.
Multiplex PCR assays were performed using previously described primers designed to amplify loci in the macaque MHC region (42, 56). PCR setup, reaction conditions, and the fragment analysis procedure were described in Wiseman et al. (56). Identification of Mamu-B*17 by microsatellite analysis was described by Wojcechowskyj et al. (57). Microsatellite identification of Mamu-DRB1*1003/*0306 was described in Penedo et al. (42). The identification of the MHC-II-associated locus D05-104184 in the rhesus genomic MHC sequence was done by Daza-Vamenta et al. (8) using ETANDEM (EMBOSS suite of software) (44). The locus was amplified using a 5′ hexachlorofluorescein-labeled forward primer (5′-CCGGCAGGAAAATATTCTGAG-3′) and a reverse primer (5′-AGGACGGCAGTTTGAGTTTG-3′).
Nucleotide sequence accession numbers.
The following macaque MHC-II alleles were identified and named according to Klein et al. (23) and Robinson et al. (45), and sequences have been deposited in the EMBL database (accession numbers in parentheses): Mamu-DPA1*0201 (EF204945), -DPA1*0202 (EF204948), -DPA1*0203 (EF204950), -DPA1*0204 (EF204947), -DPA1*0205 (EF362455), -DPA1*0601 (EF204949), -DPA1*0701 (EF204946), and -DPB1*17 (EF544137). Full-length sequences have been determined for the first time for the following macaque MHC-II alleles: Mamu-DQB1*0601 (EF426708), -DQB1*0602 (EF490967), -DQB1*0605 (EF362447), -DQB1*1801 (EF362442), -DQB1*1804 (EF362443), -DQB1*1809 (EF362445), -DQB1*1811 (EF362446), -DPB1*01 (EF362434), -DPB1*04 (EF362435), -DPB1*06 (EF490966), -DPB1*10 (EF362436), -DQA1*0105 (EU008936), -DQA1*05 (EF362438), -DQA1*06 (EF362439), -DQA1*09 (EF362440), -DQA1*2302 (EF362441), -DRB1*0309 (EF362437), and -DRB*w602 (EF544139).
RESULTS
CD4+ T-cell responses in SIV-infected EC macaques.
HIV- and SIV-infected ECs have broad and strong virus-specific CD4+ T-cell responses (15, 46), suggesting that these cells might play an important role in the containment of virus replication. After in vivo depletion of CD8+ cells in ECs, several SIV-specific CD4+ T-cell responses expanded vigorously (15). Interestingly, some of these responses were shared among the ECs used in this previous study (15). To analyze these CD4+ T-cell responses further, we used five of the ECs initially enrolled in the CD8+ cell depletion study as well as rhesus macaque 95061, which is an EC that has maintained viral loads lower than 50 copies/ml for more than 5 years (7, 59). Using PBMCs from animal 95061 (depleted in vitro of CD8+ cells), we mapped IFN-γ-positive responses directed against particular SIV peptide pools. We found that the majority of CD4+ T-cell responses in this animal were directed against Gag-derived peptides (Fig. 1A). Indeed, Gag has been shown to be a highly immunogenic protein that elicits strong CD4+-specific T-cell responses in HIV- and SIV-infected patients (15, 20).
FIG. 1.
SIV-specific CD4+ T-cell responses in EC 95061. CD8-depleted PBMCs from animal 95061 were added to ELISPOT wells with SIVmac239 peptide pools comprising 10 peptides of 15 amino acids in length overlapping by 11 amino acids spanning most of the viral proteome. An asterisk (*) indicates CD4+ T-cell responses subsequently mapped in panel B (A). Several positive peptide pools from animal 95061 were mapped to peptides of 15 amino acids in length (B). The IFN-γ production in panel B was normalized to the IFN-γ secretion of each SIV-specific CD4+ T-cell response when stimulated with the peptide pool. Peptide sequences above each mapped pool in panel B represent the peptide(s) that elicited the highest IFN-γ secretion within each pool. SFC, spot-forming cells.
The Gag161-211 E pool (SIV capsid) elicited the strongest response using in vitro CD8-depleted PBMCs from animal 95061. Interestingly, this pool also elicited a positive response using in vitro CD8-depleted PBMCs from all the other EC macaques studied (Fig. 1A and Table 1). The Mamu-A*01-restricted Gag181-189 CM9 epitope (where CM is an epitope comprised of 9 residues) is located within the Gag161-211 E pool. However, the response elicited by the Gag161-211 E pool was not CD8+ T-cell derived because PBMCs were in vitro depleted of CD8+ cells before each assay (data not shown). The Gag81-131 C pool (SIV matrix) elicited another response from in vitro CD8-depleted PBMCs and was shared by the ECs (Fig. 1A and Table 1). Animal 95061 also had several SIV-specific CD4+ T-cell responses outside Gag. These responses were directed against the Vpx1-43 A, Rev1-51 A, and Nef126-176 D pools (Fig. 1A). Interestingly, the Vpx1-43 A pool elicited a positive response from CD8-depleted PBMCs in five of six macaques in the EC cohort (Table 1), and the Nef126-176 D pool elicited a positive response from CD8-depleted PBMCs in all of the ECs studied (Table 1). PBMCs depleted in vitro of CD8+ cells from all the ECs responded to the Rev1-51 A pool (Table 1), and this response was an immunodominant response in one EC (15).
TABLE 1.
SIVmac239-specific CD4+ T-cell responses to five peptide pools in ECs
| Animal | WPIa | CD4+ T-cell response to the indicated peptideb
|
||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Gag81-131 Cc
|
Gag161-211 Ed
|
Vpx1-43 A
|
Rev1-51 A
|
Nef126-176 D
|
||||||||
| Pool | TE15/ KT15 | Pool | CD15 | QA15 | Pool | VE15 | Pool | ET15/ RY15 | Pool | RI15 | ||
| 95096 | 355 | + | + | + | − | − | − | − | + | + | + | + |
| AJ11 | 227 | + | − | + | − | + | + | + | + | + | + | + |
| 00078 | 155 | − (+)e | − | + | − | − | + | + | + | + | + | − |
| 95071 | 123 | + | − | + | − | + | + | + | + | + | + | − |
| 98016 | 186 | + | − | + | + | + | + | + | + | − | + | + |
| 95061 | 380 | + | + | + | + | + | + | + | + | + | + | + |
WPI, number of weeks after SIV infection when SIV-specific CD4+ T-cell responses were tested in each animal.
Pools of SIV peptides mapped in Fig. 1A and B were tested on CD8-depleted PBMCs from each EC. TE15/KT15, peptides Gag97-111 TE15/Gag101-115 KT15; CD15, peptide Gag181-195 CD15; QA15, peptide Gag197-211 QA15; VE15, peptide Vpx29-43 VE15; ET15/RY15, peptides Rev9-23 ET15/Rev13-27 RY15; RI15, peptide Nef138-152 RI15.
Gag81-131 C pool is located in the SIV matrix protein.
Gag161-211 E pool is located within the SIV capsid protein.
A CD4+ T-cell response (+) was detected during the study of Friedrich et al. (15).
Mapping and allele restriction of epitopes recognized by CD4+ T cells in ECs.
To map the responses directed against the Gag81-131 C, Gag161-211 E, Vpx1-43 A, Rev1-51 A, and Nef126-176 D pools, we generated CD4+ T-cell lines specific for each pool from animal 95061. The Gag81-131 C-specific CD4+ T-cell line had the strongest IFN-γ response against two overlapping peptides Gag97-111 TE15 and Gag101-115 KT15 (Fig. 1B and Table 2). It has been reported previously that this 19-amino-acid sequence was targeted by a CD4+ T-cell response (54), but the restricting allele was not described. IFN-γ secretion was also seen in vitro using CD8-depleted PBMCs stimulated with these two overlapping peptides in another EC, animal 95096 (Table 1). Mapping HIV- and SIV-specific CD4+ T-cell epitopes has historically been difficult because these epitopes often cannot be narrowed to a single optimal peptide (39). The epitope contained in Gag97-111 TE15 and Gag101-115 KT15 could not be mapped to a single peptide. However, we mapped the epitope to two peptides, Gag102-111 QE10 and Gag103-112 IT10 (Fig. 2A and Table 2). These two peptides span the region where Gag97-111 TE15 and Gag101-115 KT15 overlap (except for the threonine in peptide Gag103-112 IT10).
TABLE 2.
SIVmac239-specific CD4+ T-cell epitope localization, mapping, and restricting MHC-II allele
| SIV protein | Pool position (aa)a | Pool mapping (sequence)b | Mapped epitope (sequence) | MHC-II restricting allele |
|---|---|---|---|---|
| Gag | Matrix (81-131) | Gag97-111 TE15 (TEEAKQIVQRHLVVE); Gag101-115 KT15 (KQIVQRHLVVETGTT) | Gag102-111 QE10 (QIVQRHLVVE); Gag103-112 IT10 (IVQRHLVVET | Mamu-DRB*w606 |
| Capsid (161-211) | Gag181-195 CD15 (CTPYDINQMLNCVGD) | Gag184-193 YV10 (YDINQMLNCV) | Mamu-DRB*w2104 | |
| Capsid (161-211) | Gag197-211 QA15 (QAAMQIIRDIINEEA) | Gag200-210 ME11 (MQIIRDIINEE) | Mamu-DRB1*0306 | |
| Vpx | Vpx (1-43) | Vpx29-43 VE15 (VEEINREAVNHLPRE) | Vpx31-40 EL10 (EINREAVNHL); Vpx32-41 IP10 (INREAVNHLP) | Mamu-DRB1*1003 |
| Rev | Rev (1-51) | Rev9-23 ET15 (ELRKRLRLIHLLHQT); Rev13-27 RY15 (RLRLIHLLHQTINPY) | Rev13-23 RT11 (RLRLIHLLHQT) | Mamu-DPB1*06 |
| Nef | Nef (126-176) | Nef138-152 RI15 (RHRILDIYLEKEEGI) | Nef141-150 IE10 (ILDIYLEKEE); Nef142-151 LG10 (LDIYLEKEEG) | Mamu-DRB*w606 |
SIVmac239 amino acid (aa) region that elicited a SIV-specific CD4+ T-cell response using peptide pools of 10 peptides of 15 amino acids in length overlapping by 11 amino acids.
Pool mapping indicates the peptide (15 amino acids in length) that was found to elicit the highest IFN-γ secretion from SIV pool-specific CD4+ T cells (Fig. 1B).
FIG. 2.
Mapping of SIV-specific CD4+ T-cell responses. Peptides of 10 or 11 amino acids in length were used to map each SIV-specific CD4+ T-cell line or clone. Gag97-111 TE15/Gag101-115 KT15 was mapped with peptides of 10 amino acids in length (A), Gag181-195 CD15 was mapped using peptides of 10 amino acids in length (B), Gag197-211 QA15 was mapped using peptides of 11 amino acids in length (C), Vpx29-43 VE15 was mapped with peptides of 10 amino acids in length (D), Rev9-23 ET15/Rev13-27 RY15 was mapped with peptides of 11 amino acids in length (E), and Nef138-152 RI15 was mapped using peptides of 10 amino acids in length (F). The IFN-γ production was normalized for each SIV-specific CD4+ T-cell response when stimulated with the relevant peptide, shown in bold, and autologous BLCL.
Few MHC-II alleles that restrict SIV-specific CD4+ T-cell epitopes have been previously described (12, 24). Using the MHC-II alleles cloned from a cDNA library derived from animal 95061, we transfected RM3 cells with these MHC-II alleles to analyze the restriction of the identified CD4+ T-cell responses. (The MHC-II alleles expressed by animal 95061 and the rest of the ECs are available at http://ink.primate.wisc.edu/∼watkins/supplemental.html.) CD4+ T cells specific for Gag97-111 TE15/Gag101-115 KT15 (an epitope comprised of two peptides) responded to relevant peptide-pulsed RM3 cells expressing Mamu-DRB*w606, but this response was not elicited using other macaque MHC-II alleles (Fig. 3A, B). Mamu-DRB*w606 pulsed with an irrelevant peptide did not induce secretion of IFN-γ from CD4+ T cells specific for the Gag97-111 TE15/Gag101-115 KT15 epitope (data not shown).
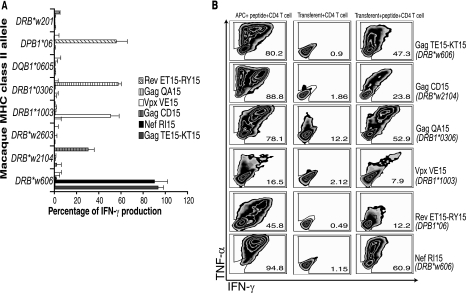
FIG. 3.
Identification of the macaque MHC-II restricting allele of SIV-specific CD4+ T-cell responses using RM3 transferents. RM3 cell transferents expressing a single macaque MHC-II allele (one α-chain allele and one β-chain allele) were tested with SIV-specific CD4+ T-cell responses. Shown are the ELISPOT analyses of eight macaque MHC-II alleles tested with SIV-specific CD4+ T cells (A). ICS analyses of SIV-specific CD4+ T-cells responses stimulated with RM3 cells expressing the relevant restricting MHC-II allele found in panel A (B). The following DNA constructs encoding MHC-II α-chain alleles were used to transfect RM3 cells with their corresponding β-chain allele: Mamu-DRA1*01041 with all of the DNA constructs encoding a DRB chain (Mamu-DRB*w201, -DRB1*0306, -DRB1*1003, -DRB*w2603, -DRB*w2104, and -DRB*w606), Mamu-DQA1*09 with Mamu-DQB1*0605, and Mamu-DPA1*0202 with Mamu-DPB1*06. The IFN-γ secretion shown in panel A was normalized for each SIV-specific CD4+ T-cell response when stimulated with homologous BLCL and pulsed with relevant peptide. APC+peptide+CD4 T cell, autologous BLCL pulsed with a relevant peptide plus specific CD4+ T cells; Transferent+CD4 T cell, SIV-specific CD4+ T cells stimulated with RM3 cells expressing the restricting MHC-II allele specific for the response and pulsed without peptide; Transferent+peptide+CD4 T cell, SIV-specific CD4+ T cells stimulated with RM3 cells expressing the restricting MHC-II allele specific for the response and pulsed with a relevant peptide.
Gag161-211 E-pool-specific CD4+ T-cell lines responded to two nonoverlapping peptides, Gag181-195 CD15 and Gag197-211 QA15 (Fig. 1B and Table 2), located in the capsid protein. Although CD8-depleted lymphocytes from the six ECs included in this study were reactive to the Gag161-211 E pool (Table 1), the mapping of the responses to this pool in these animals showed that not all had CD8-depleted lymphocytes responding to both peptides, Gag181-195 CD15 and Gag197-211 QA15. CD8-depleted lymphocytes from only two ECs responded to the Gag181-195 CD15 peptide, while CD8-depleted lymphocytes from four ECs responded to stimulation with the Gag197-211 QA15 peptide (Table 1). A sequence of 20 amino acids containing Gag181-195 CD15 has been reported previously to be the target of a CD4+ T-cell response (54) and Mills et al. (38) reported a CD4+ T-cell line that responded to a sequence of 30 amino acids containing Gag197-211 QA15. None of these studies reported the MHC-II restricting allele of either epitope. We mapped the Gag181-195 CD15-specific CD4+ T-cell response to a single peptide, Gag184-193 YV10 (Fig. 2B and Table 2). Because Gag181-195 CD15 contains the Mamu-A*01 epitope Gag181-189 CM9, we also tested Gag181-195 CD15-specific CD4+ T cells for reactivity against the peptide Gag181-189 CM9. The IFN-γ secretion of the CD4+ T-cell response stimulated with the peptide Gag181-189 CM9 was not higher than background levels (data not shown). Gag181-195 CD15-specific CD4+ T cells responded to stimulation with RM3 cells transfected with Mamu-DRB*w2104 and pulsed with peptide Gag181-195 CD15 (Fig. 3A and B and Table 2); however, the same allele pulsed with an irrelevant peptide did not induce IFN-γ secretion from Gag181-195 CD15-specific CD4+ T cells (data not shown). RM3 cells transfected with other MHC-II alleles and pulsed with peptide Gag181-195 CD15 did not elicit IFN-γ production from Gag181-195 CD15-specific CD4+ T cells (Fig. 3A). The Gag197-211 QA15-specific CD4+ T-cell response was mapped to a peptide of 11 amino acids in length, Gag200-210 ME11 (Fig. 2C and Table 2). Gag197-211 QA15-specific CD4+ T cells responded to RM3 cells expressing Mamu-DRB1*0306 and pulsed with the Gag197-211 QA15 peptide (Fig. 3A and B and Table 2); other alleles did not induce IFN-γ production from Gag197-211 QA15 (Fig. 3A).
SIV-specific CD4+ T cells from ECs preferentially targeted Gag (Fig. 1A, animal 95061) (15). Nevertheless, there were three non-Gag peptide pools that elicited IFN-γ production in PBMCs depleted in vitro of CD8+ cells from 95061 (Fig. 1A). CD4+ T-cell lines specific for the Vpx1-43 A pool were generated and used to map this CD4+ T-cell response to peptide Vpx29-43 VE15 (Fig. 1B and Table 2). The Vpx29-43 VE15-specific CD4+ T-cell response was further mapped to two peptides, Vpx31-40 EL10 and Vpx32-41 IP10 (Fig. 2D and Table 2). Interestingly, five of the six ECs had detectable IFN-γ-secreting responses directed against the Vpx29-43 VE15 peptide (Table 1). Vpx29-43 VE15-specific CD4+ T cells were restricted by RM3 cells expressing the MHC-II allele Mamu-DRB1*1003, and this response was not elicited using other MHC-II alleles (Fig. 3A and B and Table 2).
Rev1-51 A-pool-specific CD4+ T cells responded to two overlapping peptides, Rev9-23 ET15 and Rev13-27 RY15 (Fig. 1B and Table 2). CD8-depleted PBMCs from five of the ECs used in this study responded to these two peptides (Table 1). Previously, Dzuris et al. (12) reported a CD4+ T-cell response against these two peptides, Rev9-23 ET15 and Rev13-27 RY15. This response was restricted by Mamu-DRB*w201. Vogel et al. (54) also reported that the Rev9-23 ET15 and Rev13-27 RY15 peptides elicited IFN-γ secretion from SIV-specific CD4+ T cells. Using Rev9-23 ET15/Rev13-27 RY15-specific CD4+ T cells, we mapped this response to the peptide Rev13-23 RT11 (Fig. 2E and Table 2). The sequence of the peptide Rev13-23 RT11 corresponded to the amino acids that overlap between peptides Rev9-23 ET15 and Rev13-27 RY15. CD4+ T cells specific for Rev9-23 ET15/Rev13-27 RY15 were restricted by Mamu-DPB1*06 (Fig. 3A and B and Table 2). Although, Dzuris et al. (12) showed that the restricting allele for the epitope within peptides Rev9-23 ET15 and Rev13-27 RY15 was Mamu-DRB*w201, this allele did not elicit IFN-γ release from our Rev9-23 ET15/Rev13-27 RY15-specific CD4+ T cells (Fig. 3A). This region of Rev could contain two different CD4+ T-cell epitopes restricted by two different MHC-II alleles.
The peptide Nef138-152 RI15 elicited the strongest IFN-γ secretion by CD4+ T cells specific for the Nef126-176 D pool (Fig. 1B and Table 2). Vogel et al. (54) previously reported that this peptide induced IFN-γ secretion by CD4+ T cells in SIV-infected macaques; however, the restricting allele was not reported. PBMCs depleted in vitro of CD8+ cells from four ECs responded to the Nef138-152 RI15 peptide (Table 1). Using Nef138-152 RI15-specific CD4+ T cells, we mapped this response to two peptides, Nef141-150 IE10 and Nef142-151 LG10 (Fig. 2F and Table 2). Nef138-152 RI15-specific CD4+ T cells responded only to relevant peptide-pulsed RM3 cells expressing Mamu-DRB*w606, and other MHC-II alleles did not elicit the secretion of IFN-γ (Fig. 3A and B). Nef138-152 RI15-specific CD4+ T cells were not stimulated by Mamu-DRB*w606 pulsed with an irrelevant peptide (data not shown).
Five of the CD4+ T-cell responses described above (summarized in Table 2) were restricted by an MHC-II DR allele, and an MHC-II DP allele restricted one of them. The responses restricted by the MHC-II DR alleles were tested on RM3 cells transfected with either Mamu-DRA1*01041 or -DRA1*01024. The response restricted by the MHC-II DP allele was tested on RM3 cells transfected with either DPA1*0202 or DPA1*0601. The different MHC-II α-chains did not affect the IFN-γ secretion from SIV-specific CD4+ T-cell responses (data not shown).
Certain MHC-II alleles are enriched in ECs.
Although individuals who control viral replication have strong HIV-specific CD4+ T-cell responses (15, 46), no clear association of particular MHC-II molecules with protection against disease progression has been demonstrated. However, evidence suggests that MHC-II alleles might be important in the control of retroviral replication (29).
To define the association between MHC-II alleles and SIV control, we genotyped 196 SIVmac239-infected Indian rhesus macaques from the WNPRC and the National Cancer Institute for each of the restricting MHC-II alleles described above. Regression analysis was done using PROC LOGISTIC SAS, version 9.1 (SAS Institute, Cary, NC), as previously described (28, 59), to determine if the frequency of these MHC-II alleles was enriched in the EC group. Table 3 shows that the MHC-II alleles Mamu-DPB1*06, -DRB*w606, and -DRB*w2104 were no more frequent in the EC cohort than they were in the entire infected cohort. On the other hand, the MHC-II alleles Mamu-DRB1*0306 and -DRB1*1003 were significantly enriched in the EC cohort (n = 16) compared to the entire cohort of SIV-infected rhesus macaques (n = 196). Both of these alleles were present at a frequency of 44% in the EC cohort but were present only at a frequency of less than 23% in the entire SIV-infected cohort (P values of 0.05 and 0.02 for Mamu-DRB1*0306 and Mamu-DRB1*1003, respectively) (Table 3). Previously de Groot et al. (9) showed that these two MHC-II alleles segregate together along with Mamu-DRA1*01041 as part of the same genetic configuration.
To determine whether the MHC-II alleles described above contributed to control of SIV replication, we used the program PROC MIXED to estimate the relative log geometric mean of plasma viremia for the chronic phase (>10 weeks) in all of the 196 SIV-infected animals. The EC status, vaccine status (only vaccines not associated with control were included) (59), and all genotyped MHC-II alleles were included as covariates in the model. Unlike the macaque MHC-I alleles Mamu-B*08 and Mamu-B*17, the presence of any of the five MHC-II alleles did not affect viral load in either the acute phase (data not shown) or the chronic phase of infection (http://ink.primate.wisc.edu/∼watkins/supplemental.html).
Interestingly, five of the seven ECs that expressed Mamu-DRB1*0306 and Mamu-DRB1*1003 also expressed Mamu-B*17 (Table 3). Thus, we compared the viral load of animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 (n = 13) with those that expressed Mamu-B*17 but did not express Mamu-DRB1*0306 or Mamu-DRB1*1003 (n = 28) in the 196 SIV-infected rhesus macaque cohort. Surprisingly, animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 had significantly lower viral loads than animals that expressed Mamu-B*17 but did not express Mamu-DRB1*1003 or -DRB1*0306 (P = <0.0001) (Table 4).
The effect of Mamu-DRB1*1003 and -DRB1*0306 on viral loads in SIV-infected animals that also expressed the Mamu-B*17 allele could possibly be due to a linked gene carried along with these three alleles in a single haplotype. To analyze whether Mamu-B*17, -DRB1*1003, and -DRB1*0306 were always inherited in the same haplotype, we used microsatellite markers previously described for Indian rhesus macaques (42, 56, 57). We used seven of 13 animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 from the 196 SIV-infected cohort to analyze the linkage among these alleles (Table 5) (the microsatellite markers used for this analysis are available at http://ink.primate.wisc.edu/∼watkins/supplemental.html). To increase the number of animals, we included two animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 that were infected with different strains of SIV (Table 5). We found that Mamu-B*17 and the two MHC-II DR alleles (Mamu-DRB1*1003 and Mamu-DRB1*0306) were inherited as one haplotype in six animals (Table 5; also see http://ink.primate.wisc.edu/∼watkins/supplemental.html), and three animals inherited Mamu-B*17 separately from the Mamu-DRB1*1003/-DRB1*0306 configuration (Table 5; also see http://ink.primate.wisc.edu/∼watkins/supplemental.html). Thus, although Mamu-B*17 can be inherited separately from the alleles Mamu-DRB1*1003 and -DRB1*0306, we cannot exclude the possibility that the effect of lower viral loads in animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 was due to another gene carried as part of a haplotype that contains these three alleles.
TABLE 5.
Inheritance of Mamu-B*17, -DRB1*1003, and -DRB1*0306 alleles
| Animala | Presence of alleleb
|
Linkagec | ECd | ||
|---|---|---|---|---|---|
| Mamu-B*17 | Mamu-DRB1*1003 | Mamu-DRB1*0306 | |||
| 98016 | + | + | + | N | Y |
| 99006 | + | + | + | N | Y |
| 00026 | + | + | + | N | N |
| 95071 | + | + | + | Y | Y |
| 95061 | + | + | + | Y | Y |
| 98017 | + | + | + | Y | N |
| 96112 | + | + | + | Y | Y |
| 96072 | + | + | + | Y | N |
| 99026 | + | + | + | Y | N |
Animals were infected with SIVmac239 and were part of the 196 SIV-infected cohort except for animals 99006 and 00026. These two animals were infected with different strains of SIV. Animal 99006 was infected with a variant of SIVmac239 that had three immunodominant epitopes mutated (14). Animal 00026 was infected with SIVmac251 (the clone SIVmac239 was derived subsequently from this strain).
All animals were typed for the alleles Mamu-B*17, -DRB1*1003, and -DRB1*0306 using sequence-specific PCR, and results were confirmed with microsatellites. These alleles were also typed in animals 98016, 95071, and 95061 using cDNA libraries.
Linkage indicates whether Mamu-B*17 was in the same chromosome with Mamu-DRB1*1003 and -DRB1*0306. Y, yes; N, no.
Y, yes; N, no.
DISCUSSION
Two macaque MHC-II DR alleles were enriched in a group of SIV-infected ECs that have controlled viral replication for longer than 1 year. For the first time, we show that a particular combination of MHC-I and -II alleles may be important for control of SIV replication. Unlike the lower viral loads seen with certain human MHC-I and macaque MHC-I alleles in HIV and SIV infections (34, 37, 59), particular MHC-II alleles were not associated with lower viral loads (http://ink.primate.wisc.edu/∼watkins/supplemental.html). However, our data suggest that CD4+ T-cell responses might have an important role in the control of SIV replication because Indian rhesus macaques that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 (n = 13) had significantly lower viral loads than animals that expressed Mamu-B*17 but did not express Mamu-DRB1*1003 or -DRB1*0306 (n = 28; P < 0.0001). These results suggest that although CD4+ T cells alone may not be capable of controlling SIV replication, the presence of certain CD4+ T-cell responses in concert with CD8+ T-cell responses might have an important impact on control of SIV replication. It is probably not surprising that MHC-II alleles by themselves were not directly associated with control of viral replication. Evidence from in vivo CD8 depletion studies (15, 30) and analysis of viral escape (13, 40) had implicated CD8+ cytotoxic T cells as critical factors in the control of viral replication. However, the viral loads in animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 were lower than those found in animals that expressed Mamu-B*17 but did not express Mamu-DRB1*1003 or -DRB1*0306, which suggests that CD8+ cytotoxic T cells may be more efficacious at controlling viral replication when adequate CD4+ T-cell help is present. We found that these two MHC-II alleles restricted two different SIV-specific CD4+ T-cell responses, Vpx29-43 VE15 (Mamu-DRB1*1003) and Gag197-211 QA15 (Mamu-DRB1*0306).
We also analyzed whether Mamu-B*17 was inherited along with Mamu-DRB1*1003 and -DRB1*0306 as part of the same haplotype. We found that in some cases these three alleles belonged to the same haplotype, and in other cases Mamu-B*17 was inherited separately from Mamu-DRB1*1003 and -DRB1*0306 (Table 5; also see http://ink.primate.wisc.edu/∼watkins/supplemental.html). We cannot completely exclude the possibility that the effect of the lower viral loads seen in animals that expressed Mamu-B*17, -DRB1*1003, and -DRB1*0306 was due to an effect of another gene or genes inherited as part of this haplotype. To completely exclude this possibility, we would need to compare viral loads from animals that have these alleles in the same haplotype with viral loads of animals that inherited these alleles in different chromosomes. Unfortunately, we currently do not have a large enough number of animals that inherited Mamu-B*17 separately from Mamu-DRB1*1003 and -DRB1*0306 to do a suitable comparison.
Previously de Groot et al. (9) showed that Mamu-DRB1*1003 and -DRB1*0306 belong to a configuration that is inherited as a unit; however, the P values for these two MHC-II alleles (for Mamu-DRB1*0306, P = 0.05; for Mamu-DRB1*1003, P = 0.02) in the enrichment analysis was different (Table 3). This difference was observed because Mamu-DRB1*0306 can be part of a different configuration including Mamu-DRB1*1007 (9). This may have been the case for seven animals that were positive for Mamu-DRB1*0306 and not for Mamu-DRB1*1003 (data not shown). It is also possible that these two alleles were not always inherited as a complete unit (9). This may have been the case for two animals that were positive for Mamu-DRB1*1003 but not for Mamu-DRB1*0306 in the 196 SIV-infected cohort (data not shown).
Few SIV-specific CD4+ T-cell responses have been reported to date (12, 24, 38, 54), and there is even less information about the MHC-II alleles that restrict SIV epitopes recognized by Indian rhesus macaque CD4+ T cells (12, 24). In this study, we mapped several SIV-specific CD4+ T-cell epitopes (Fig. 2 and Table 2) and reported their MHC-II restricting alleles (Fig. 3 and Table 2). Interestingly, we found that one of the identified epitopes (Gag181-195 CD15) overlapped entirely with the Mamu-A*01-restricted CD8+ T-cell epitope, Gag181-189 CM9. The linkage of CD4+ T-cell epitopes with CD8+ T-cell epitopes has previously been observed in HIV (60), in a HIV murine model (51), and in infection with Chlamydia trachomatis (22) and Yersinia enterocolitica (36). It is possible that as our knowledge of SIV-specific CD4+ T-cell epitopes increases, more SIV-specific CD4+ and CD8+ T-cell epitopes will be found to overlap with each other. However, whether there is any advantage to having clustered CD4+ T-cell and CD8+ T-cell epitopes is not yet known. It is possible that the clustering of epitopes has more to do with the ability of the cellular machinery to process those regions better than others (22).
Some of the ECs in this study had CD4+ T-cell responses against the same region of the virus yet expressed different MHC-II alleles. CD8-depleted lymphocytes from animal 95096 responded to Gag97-111 TE15/Gag101-115 KT15 and Nef138-152 RI15 peptides (Table 1). However, this animal was negative for Mamu-DRB*w606 (http://ink.primate.wisc.edu/∼watkins/supplemental.html). CD8-depleted lymphocytes from the ECs AJ11 and 98016 were reactive against Nef138-152 RI15 (Table 1) although these animals were also negative for Mamu-DRB*w606 (http://ink.primate.wisc.edu/∼watkins/supplemental.html). Similarly, CD8-depleted lymphocytes from animal 98016 responded to the peptide Gag181-195 CD15 (Table 1); however, this animal was negative Mamu-DRB*w2104 (http://ink.primate.wisc.edu/∼watkins/supplemental.html). Thus, it is likely that more than one MHC-II allele might restrict these peptides. Several MHC-II molecules can present the same peptide to CD4+ T cells (41, 52). On the other hand, it is plausible that within long peptides, several different CD4+ T-cell responses might be elicited. We found that the CD4+ T-cell epitope Rev9-23 ET15/Rev13-27 RY15 was restricted by Mamu-DPB1*06 in macaque 95061 and not by Mamu-DRB*w201, as has been previously suggested (12). This apparent disparity could be explained by the existence of two different CD4+ T-cell epitopes that were located in this same SIV region but may differ in a couple of amino acids and consequently were restricted by two different MHC-II alleles.
In this study, we describe SIV-specific CD4+ T-cell responses from ECs. It would also be of interest to examine if these CD4+ T-cell responses were present in rhesus macaques that cannot control viral replication to low levels (<1,000 viral RNA copies/ml). However, due to the highly pathogenic nature of SIVmac239 used to infect our cohort of Indian rhesus macaques, most of the SIV-specific memory CD4+ T cells are depleted during infection (17, 26). It is, therefore, exceedingly difficult to detect SIV-specific CD4+ T-cell responses in animals that do not control viral replication during the chronic phase of infection.
In conclusion, we have defined the restricting alleles for six different SIVmac239-specific CD4+ T-cell epitopes. We also found that the macaque MHC-II alleles Mamu-DRB1*1003 and -DRB1*0306 were enriched in our EC group. These two MHC-II alleles by themselves were not correlated with lower SIV loads. However, rhesus macaques that expressed the alleles Mamu-DRB1*1003, -DRB1*0306, and Mamu-B*17 had lower SIV loads than animals that expressed Mamu-B*17 but did not express Mamu-DRB1*1003 or -DRB1*0306. The study of ECs with particular MHC-II alleles could improve our comprehension of how some individuals are able to control SIV and HIV replication and further our understanding of HIV vaccine design.
Acknowledgments
We thank the animal care staff at the WNPRC for housing and caring for the Indian rhesus macaque colony. We also thank R. Paul Johnson for kindly providing us with Mamu-DPB1*06 and Mamu-DPB1*10 stable transferents and Matija Peterlin for supplying us with the RM3 cell line. We gratefully acknowledge David H. O'Connor, Jonah B. Sacha, Laura E. Valentine, and Lara Vojnov for helpful discussions.
This research was supported by NIH grants R24-RR016038, R24-RR015371, R01-AI049120, and R01-AI052056. Part of this work was made possible by grant P51-RR000167 from the National Center for Research Resources, a component of NIH, to the WNPRC at the University of Wisconsin—Madison. This work was also supported in part with federal funds from the National Cancer Institute, NIH, under contract NO1-CO-12400 and by the Intramural Research Program of the NIH through the Center for Cancer Research, National Cancer Institute. This research was conducted in part at a facility constructed with support from Research Facilities Improvement grants RR15459-01 and RR020141-01 (WNPRC).
The contents of this publication are solely the responsibility of the authors and do not necessarily represent the official views or policies of the National Center for Research Resources, NIH, or the Department of Health and Human Services, nor does mention of trade names, commercial products, or organizations imply endorsement by the U.S. government.
We declare that we have no competing financial interests.
Footnotes
Published ahead of print on 7 November 2007.
REFERENCES
- 1.Altfeld, M., and E. S. Rosenberg. 2000. The role of CD4(+) T helper cells in the cytotoxic T lymphocyte response to HIV-1. Curr. Opin. Immunol. 12375-380. [DOI] [PubMed] [Google Scholar]
- 2.Bevan, M. J. 2004. Helping the CD8(+) T-cell response. Nat. Rev. Immunol. 4595-602. [DOI] [PubMed] [Google Scholar]
- 3.Brenchley, J. M., L. E. Ruff, J. P. Casazza, R. A. Koup, D. A. Price, and D. C. Douek. 2006. Preferential infection shortens the life span of human immunodeficiency virus-specific CD4+ T cells in vivo. J. Virol. 806801-6809. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Brenchley, J. M., T. W. Schacker, L. E. Ruff, D. A. Price, J. H. Taylor, G. J. Beilman, P. L. Nguyen, A. Khoruts, M. Larson, A. T. Haase, and D. C. Douek. 2004. CD4+ T cell depletion during all stages of HIV disease occurs predominantly in the gastrointestinal tract. J. Exp. Med. 200749-759. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Calman, A. F., and B. M. Peterlin. 1987. Mutant human B cell lines deficient in class II major histocompatibility complex transcription. J. Immunol. 1392489-2495. [PubMed] [Google Scholar]
- 6.Carrington, M., and S. J. O'Brien. 2003. The influence of HLA genotype on AIDS. Annu. Rev. Med. 54535-551. [DOI] [PubMed] [Google Scholar]
- 7.Chung, C., W. Lee, J. T. Loffredo, B. Burwitz, T. C. Friedrich, J. P. Giraldo Vela, G. Napoe, E. G. Rakasz, N. A. Wilson, D. B. Allison, and D. I. Watkins. 2007. Not all cytokine-producing CD8+ T cells suppress simian immunodeficiency virus replication. J. Virol. 811517-1523. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Daza-Vamenta, R., G. Glusman, L. Rowen, B. Guthrie, and D. E. Geraghty. 2004. Genetic divergence of the rhesus macaque major histocompatibility complex. Genome Res. 141501-1515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.de Groot, N., G. G. Doxiadis, N. G. De Groot, N. Otting, C. Heijmans, A. J. Rouweler, and R. E. Bontrop. 2004. Genetic makeup of the DR region in rhesus macaques: gene content, transcripts, and pseudogenes. J. Immunol. 1726152-6157. [DOI] [PubMed] [Google Scholar]
- 10.Demoustier, A., B. Gubler, O. Lambotte, M. G. de Goer, C. Wallon, C. Goujard, J. F. Delfraissy, and Y. Taoufik. 2002. In patients on prolonged HAART, a significant pool of HIV-infected CD4 T cells are HIV specific. AIDS 161749-1754. [DOI] [PubMed] [Google Scholar]
- 11.Douek, D. C., J. M. Brenchley, M. R. Betts, D. R. Ambrozak, B. J. Hill, Y. Okamoto, J. P. Casazza, J. Kuruppu, K. Kunstman, S. Wolinsky, Z. Grossman, M. Dybul, A. Oxenius, D. A. Price, M. Connors, and R. A. Koup. 2002. HIV preferentially infects HIV-specific CD4+ T cells. Nature 41795-98. [DOI] [PubMed] [Google Scholar]
- 12.Dzuris, J. L., J. Sidney, H. Horton, R. Correa, D. Carter, R. W. Chesnut, D. I. Watkins, and A. Sette. 2001. Molecular determinants of peptide binding to two common rhesus macaque major histocompatibility complex class II molecules. J. Virol. 7510958-10968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Feeney, M. E., Y. Tang, K. A. Roosevelt, A. J. Leslie, K. McIntosh, N. Karthas, B. D. Walker, and P. J. Goulder. 2004. Immune escape precedes breakthrough human immunodeficiency virus type 1 viremia and broadening of the cytotoxic T-lymphocyte response in an HLA-B27-positive long-term-nonprogressing child. J. Virol. 788927-8930. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Friedrich, T. C., E. J. Dodds, L. J. Yant, L. Vojnov, R. Rudersdorf, C. Cullen, D. T. Evans, R. C. Desrosiers, B. R. Mothe, J. Sidney, A. Sette, K. Kunstman, S. Wolinsky, M. Piatak, J. Lifson, A. L. Hughes, N. Wilson, D. H. O'Connor, and D. I. Watkins. 2004. Reversion of CTL escape-variant immunodeficiency viruses in vivo. Nat. Med. 10275-281. [DOI] [PubMed] [Google Scholar]
- 15.Friedrich, T. C., L. E. Valentine, L. J. Yant, E. G. Rakasz, S. M. Piaskowski, J. R. Furlott, K. L. Weisgrau, B. Burwitz, G. E. May, E. J. Leon, T. Soma, G. Napoe, S. V. Capuano III, N. A. Wilson, and D. I. Watkins. 2007. Subdominant CD8+ T-cell responses are involved in durable control of AIDS virus replication. J. Virol. 813465-3476. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gauduin, M. C., Y. Yu, A. Barabasz, A. Carville, M. Piatak, J. D. Lifson, R. C. Desrosiers, and R. P. Johnson. 2006. Induction of a virus-specific effector-memory CD4+ T cell response by attenuated SIV infection. J. Exp. Med. 2032661-2672. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Haase, A. T. 2005. Perils at mucosal front lines for HIV and SIV and their hosts. Nat. Rev. Immunol. 5783-792. [DOI] [PubMed] [Google Scholar]
- 18.Janssen, E. M., E. E. Lemmens, T. Wolfe, U. Christen, M. G. von Herrath, and S. P. Schoenberger. 2003. CD4+ T cells are required for secondary expansion and memory in CD8+ T lymphocytes. Nature 421852-856. [DOI] [PubMed] [Google Scholar]
- 19.Kalams, S. A., and B. D. Walker. 1998. The critical need for CD4 help in maintaining effective cytotoxic T lymphocyte responses. J. Exp. Med. 1882199-2204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Kaufmann, D. E., P. M. Bailey, J. Sidney, B. Wagner, P. J. Norris, M. N. Johnston, L. A. Cosimi, M. M. Addo, M. Lichterfeld, M. Altfeld, N. Frahm, C. Brander, A. Sette, B. D. Walker, and E. S. Rosenberg. 2004. Comprehensive analysis of human immunodeficiency virus type 1-specific CD4 responses reveals marked immunodominance of Gag and Nef and the presence of broadly recognized peptides. J. Virol. 784463-4477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kawada, M., T. Tsukamoto, H. Yamamoto, A. Takeda, H. Igarashi, D. I. Watkins, and T. Matano. 2007. Long-term control of simian immunodeficiency virus replication with central memory CD4+ T-cell preservation after nonsterile protection by a cytotoxic T-lymphocyte-based vaccine. J. Virol. 815202-5211. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Kim, S. K., and R. DeMars. 2001. Epitope clusters in the major outer membrane protein of Chlamydia trachomatis. Curr. Opin. Immunol. 13429-436. [DOI] [PubMed] [Google Scholar]
- 23.Klein, J., R. E. Bontrop, R. L. Dawkins, H. A. Erlich, U. B. Gyllensten, E. R. Heise, P. P. Jones, P. Parham, E. K. Wakeland, and D. I. Watkins. 1990. Nomenclature for the major histocompatibility complexes of different species: a proposal. Immunogenetics 31217-219. [DOI] [PubMed] [Google Scholar]
- 24.Lekutis, C., and N. L. Letvin. 1997. HIV-1 envelope-specific CD4+ T helper cells from simian/human immunodeficiency virus-infected rhesus monkeys recognize epitopes restricted by MHC class II DRB1*0406 and DRB*W201 molecules. J. Immunol. 1592049-2057. [PubMed] [Google Scholar]
- 25.Letvin, N. L., J. R. Mascola, Y. Sun, D. A. Gorgone, A. P. Buzby, L. Xu, Z. Y. Yang, B. Chakrabarti, S. S. Rao, J. E. Schmitz, D. C. Montefiori, B. R. Barker, F. L. Bookstein, and G. J. Nabel. 2006. Preserved CD4+ central memory T cells and survival in vaccinated SIV-challenged monkeys. Science 3121530-1533. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Li, Q., L. Duan, J. D. Estes, Z. M. Ma, T. Rourke, Y. Wang, C. Reilly, J. Carlis, C. J. Miller, and A. T. Haase. 2005. Peak SIV replication in resting memory CD4+ T cells depletes gut lamina propria CD4+ T cells. Nature 4341148-1152. [DOI] [PubMed] [Google Scholar]
- 27.Lichterfeld, M., D. E. Kaufmann, X. G. Yu, S. K. Mui, M. M. Addo, M. N. Johnston, D. Cohen, G. K. Robbins, E. Pae, G. Alter, A. Wurcel, D. Stone, E. S. Rosenberg, B. D. Walker, and M. Altfeld. 2004. Loss of HIV-1-specific CD8+ T cell proliferation after acute HIV-1 infection and restoration by vaccine-induced HIV-1-specific CD4+ T cells. J. Exp. Med. 200701-712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Loffredo, J. T., J. Maxwell, Y. Qi, C. E. Glidden, G. J. Borchardt, T. Soma, A. T. Bean, D. R. Beal, N. A. Wilson, W. M. Rehrauer, J. D. Lifson, M. Carrington, and D. I. Watkins. 2007. Mamu-B*08-positive macaques control simian immunodeficiency virus replication. J. Virol. 818827-8832. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Malhotra, U., S. Holte, S. Dutta, M. M. Berrey, E. Delpit, D. M. Koelle, A. Sette, L. Corey, and M. J. McElrath. 2001. Role for HLA class II molecules in HIV-1 suppression and cellular immunity following antiretroviral treatment. J. Clin. Investig. 107505-517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Matano, T., R. Shibata, C. Siemon, M. Connors, H. C. Lane, and M. A. Martin. 1998. Administration of an anti-CD8 monoclonal antibody interferes with the clearance of chimeric simian/human immunodeficiency virus during primary infections of rhesus macaques. J. Virol. 72164-169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Matloubian, M., R. J. Concepcion, and R. Ahmed. 1994. CD4+ T cells are required to sustain CD8+ cytotoxic T-cell responses during chronic viral infection. J. Virol. 688056-8063. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Mattapallil, J. J., D. C. Douek, A. Buckler-White, D. Montefiori, N. L. Letvin, G. J. Nabel, and M. Roederer. 2006. Vaccination preserves CD4 memory T cells during acute simian immunodeficiency virus challenge. J. Exp. Med. 2031533-1541. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Mattapallil, J. J., D. C. Douek, B. Hill, Y. Nishimura, M. Martin, and M. Roederer. 2005. Massive infection and loss of memory CD4+ T cells in multiple tissues during acute SIV infection. Nature 4341093-1097. [DOI] [PubMed] [Google Scholar]
- 34.McNeil, A. J., P. L. Yap, S. M. Gore, R. P. Brettle, M. McColl, R. Wyld, S. Davidson, R. Weightman, A. M. Richardson, and J. R. Robertson. 1996. Association of HLA types A1-B8-DR3 and B27 with rapid and slow progression of HIV disease. QJM 89177-185. [DOI] [PubMed] [Google Scholar]
- 35.Mehandru, S., M. A. Poles, K. Tenner-Racz, A. Horowitz, A. Hurley, C. Hogan, D. Boden, P. Racz, and M. Markowitz. 2004. Primary HIV-1 infection is associated with preferential depletion of CD4+ T lymphocytes from effector sites in the gastrointestinal tract. J. Exp. Med. 200761-770. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Mertz, A. K., P. Wu, T. Sturniolo, D. Stoll, M. Rudwaleit, R. Lauster, J. Braun, and J. Sieper. 2000. Multispecific CD4+ T cell response to a single 12-mer epitope of the immunodominant heat-shock protein 60 of Yersinia enterocolitica in Yersinia-triggered reactive arthritis: overlap with the B27-restricted CD8 epitope, functional properties, and epitope presentation by multiple DR alleles. J. Immunol. 1641529-1537. [DOI] [PubMed] [Google Scholar]
- 37.Migueles, S. A., M. S. Sabbaghian, W. L. Shupert, M. P. Bettinotti, F. M. Marincola, L. Martino, C. W. Hallahan, S. M. Selig, D. Schwartz, J. Sullivan, and M. Connors. 2000. HLA B*5701 is highly associated with restriction of virus replication in a subgroup of HIV-infected long-term nonprogressors. Proc. Natl. Acad. Sci. USA 972709-2714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mills, K. H., A. L. Barnard, M. Williams, M. Page, C. Ling, E. J. Stott, P. Silvera, F. Taffs, A. S. Kingsman, S. E. Adams, et al. 1991. Vaccine-induced CD4+ T cells against the simian immunodeficiency virus Gag protein. Epitope specificity and relevance to protective immunity. J. Immunol. 1473560-3567. [PubMed] [Google Scholar]
- 39.Norris, P. J., H. F. Moffett, C. Brander, T. M. Allen, K. M. O'Sullivan, L. A. Cosimi, D. E. Kaufmann, B. D. Walker, and E. S. Rosenberg. 2004. Fine specificity and cross-clade reactivity of HIV type 1 Gag-specific CD4+ T cells. AIDS Res. Hum. Retrovir. 20315-325. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.O'Connor, D. H., B. R. Mothe, J. T. Weinfurter, S. Fuenger, W. M. Rehrauer, P. Jing, R. R. Rudersdorf, M. E. Liebl, K. Krebs, J. Vasquez, E. Dodds, J. Loffredo, S. Martin, A. B. McDermott, T. M. Allen, C. Wang, G. G. Doxiadis, D. C. Montefiori, A. Hughes, D. R. Burton, D. B. Allison, S. M. Wolinsky, R. Bontrop, L. J. Picker, and D. I. Watkins. 2003. Major histocompatibility complex class I alleles associated with slow simian immunodeficiency virus disease progression bind epitopes recognized by dominant acute-phase cytotoxic T-lymphocyte responses. J. Virol. 779029-9040. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.O'Sullivan, D., T. Arrhenius, J. Sidney, M. F. Del Guercio, M. Albertson, M. Wall, C. Oseroff, S. Southwood, S. M. Colon, F. C. Gaeta, et al. 1991. On the interaction of promiscuous antigenic peptides with different DR alleles: identification of common structural motifs. J. Immunol. 1472663-2669. [PubMed] [Google Scholar]
- 42.Penedo, M. C., R. E. Bontrop, C. M. Heijmans, N. Otting, R. Noort, A. J. Rouweler, N. de Groot, N. G. de Groot, T. Ward, and G. G. Doxiadis. 2005. Microsatellite typing of the rhesus macaque MHC region. Immunogenetics 57198-209. [DOI] [PubMed] [Google Scholar]
- 43.Picker, L. J., and D. I. Watkins. 2005. HIV pathogenesis: the first cut is the deepest. Nat. Immunol. 6430-432. [DOI] [PubMed] [Google Scholar]
- 44.Rice, P., I. Longden, and A. Bleasby. 2000. EMBOSS: the European molecular biology open software suite. Trends Genet. 16276-277. [DOI] [PubMed] [Google Scholar]
- 45.Robinson, J., M. J. Waller, P. Parham, N. de Groot, R. Bontrop, L. J. Kennedy, P. Stoehr, and S. G. E. Marsh. 2003. IMGT/HLA and IMGT/MHC: sequence databases for the study of the major histocompatibility complex. Nucleic Acids Res. 31311-314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Rosenberg, E. S., J. M. Billingsley, A. M. Caliendo, S. L. Boswell, P. E. Sax, S. A. Kalams, and B. D. Walker. 1997. Vigorous HIV-1-specific CD4+ T cell responses associated with control of viremia. Science 2781447-1450. [DOI] [PubMed] [Google Scholar]
- 47.Sauermann, U., C. Stahl-Hennig, N. Stolte, T. Muhl, M. Krawczak, M. Spring, D. Fuchs, F. J. Kaup, G. Hunsmann, and S. Sopper. 2000. Homozygosity for a conserved MHC class II DQ-DRB haplotype is associated with rapid disease progression in simian immunodeficiency virus-infected macaques: results from a prospective study. J. Infect. Dis. 182716-724. [DOI] [PubMed] [Google Scholar]
- 48.Shedlock, D. J., and H. Shen. 2003. Requirement for CD4 T cell help in generating functional CD8 T cell memory. Science 300337-339. [DOI] [PubMed] [Google Scholar]
- 49.Staprans, S. I., A. P. Barry, G. Silvestri, J. T. Safrit, N. Kozyr, B. Sumpter, H. Nguyen, H. McClure, D. Montefiori, J. I. Cohen, and M. B. Feinberg. 2004. Enhanced SIV replication and accelerated progression to AIDS in macaques primed to mount a CD4 T cell response to the SIV envelope protein. Proc. Natl. Acad. Sci. USA 10113026-13031. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Sun, J. C., M. A. Williams, and M. J. Bevan. 2004. CD4+ T cells are required for the maintenance, not programming, of memory CD8+ T cells after acute infection. Nat. Immunol. 5927-933. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 51.Surman, S., T. D. Lockey, K. S. Slobod, B. Jones, J. M. Riberdy, S. W. White, P. C. Doherty, and J. L. Hurwitz. 2001. Localization of CD4+ T cell epitope hotspots to exposed strands of HIV envelope glycoprotein suggests structural influences on antigen processing. Proc. Natl. Acad. Sci. USA 984587-4592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.van der Burg, S. H., K. M. Kwappenberg, A. Geluk, M. van der Kruk, O. Pontesilli, E. Hovenkamp, K. L. Franken, K. E. van Meijgaarden, J. W. Drijfhout, T. H. Ottenhoff, C. J. Melief, and R. Offringa. 1999. Identification of a conserved universal Th epitope in HIV-1 reverse transcriptase that is processed and presented to HIV-specific CD4+ T cells by at least four unrelated HLA-DR molecules. J. Immunol. 162152-160. [PubMed] [Google Scholar]
- 53.Veazey, R. S., M. DeMaria, L. V. Chalifoux, D. E. Shvetz, D. R. Pauley, H. L. Knight, M. Rosenzweig, R. P. Johnson, R. C. Desrosiers, and A. A. Lackner. 1998. Gastrointestinal tract as a major site of CD4+ T cell depletion and viral replication in SIV infection. Science 280427-431. [DOI] [PubMed] [Google Scholar]
- 54.Vogel, T. U., H. Horton, D. H. Fuller, D. K. Carter, K. Vielhuber, D. H. O'Connor, T. Shipley, J. Fuller, G. Sutter, V. Erfle, N. Wilson, L. J. Picker, and D. I. Watkins. 2002. Differences between T cell epitopes recognized after immunization and after infection. J. Immunol. 1694511-4521. [DOI] [PubMed] [Google Scholar]
- 55.Wilson, N. A., J. Reed, G. S. Napoe, S. Piaskowski, A. Szymanski, J. Furlott, E. J. Gonzalez, L. J. Yant, N. J. Maness, G. E. May, T. Soma, M. R. Reynolds, E. Rakasz, R. Rudersdorf, A. B. McDermott, D. H. O'Connor, T. C. Friedrich, D. B. Allison, A. Patki, L. J. Picker, D. R. Burton, J. Lin, L. Huang, D. Patel, G. Heindecker, J. Fan, M. Citron, M. Horton, F. Wang, X. Liang, J. W. Shiver, D. R. Casimiro, and D. I. Watkins. 2006. Vaccine-induced cellular immune responses reduce plasma viral concentrations after repeated low-dose challenge with pathogenic simian immunodeficiency virus SIVmac239. J. Virol. 805875-5885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 56.Wiseman, R. W., J. A. Wojcechowskyj, J. M. Greene, A. J. Blasky, T. Gopon, T. Soma, T. C. Friedrich, S. L. O'Connor, and D. H. O'Connor. 2007. Simian immunodeficiency virus SIVmac239 infection of major histocompatibility complex-identical cynomolgus macaques from Mauritius. J. Virol. 81349-361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Wojcechowskyj, J. A., L. J. Yant, R. W. Wiseman, S. L. O'Connor, and D. H. O'Connor. 2007. Control of simian immunodeficiency virus SIVmac239 is not predicted by inheritance of Mamu-B*17-containing haplotypes. J. Virol. 81406-410. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Yang, O. O., P. T. Sarkis, A. Trocha, S. A. Kalams, R. P. Johnson, and B. D. Walker. 2003. Impacts of avidity and specificity on the antiviral efficiency of HIV-1-specific CTL. J. Immunol. 1713718-3724. [DOI] [PubMed] [Google Scholar]
- 59.Yant, L. J., T. C. Friedrich, R. C. Johnson, G. E. May, N. J. Maness, A. M. Enz, J. D. Lifson, D. H. O'Connor, M. Carrington, and D. I. Watkins. 2006. The high-frequency major histocompatibility complex class I allele Mamu-B*17 is associated with control of simian immunodeficiency virus SIVmac239 replication. J. Virol. 805074-5077. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 60.Zhan, X., J. L. Hurwitz, S. A. Brown, and K. S. Slobod. 2007. HIV-1 envelope T cell epitope “hotspots” among mice and humans and among CD4+ and CD8+ T cell subpopulations. AIDS Res. Hum. Retrovir. 23471-476. [DOI] [PubMed] [Google Scholar]