Abstract
Loss of tumor suppressor gene expression via mutations plays a critical role in cancer development, particularly when occurring in heterozygous cells. These so-called “second step” mutational events are often large in size and arise most often from chromosome loss, mitotic recombination, or interstitial deletion. An open question in cancer research is whether different chromosomes are equally susceptible to formation of large mutations, or alternatively if the unique sequence of each chromosome will lead to chromosome specific mutational spectra. To address this question, the spectra of second step mutations were determined for chromosomes 8 and 11 in Aprt and Tk mutants, respectively, isolated from primary kidney clones heterozygous for both loci. The results showed that the spectra of large mutational events were essentially the same. This observation suggests that internal and external cellular environments provide the driving force for large autosomal mutational events, and that chromosome structure per se is the substrate upon which these forces act.
INTRODUCTION
Tumor suppressor gene inactivation plays a causal role in cancer development and progression. A critical change occurs when cells heterozygous for a given tumor suppressor gene lose expression of the functional allele [1]. Although small intragenic events such as base-pair substitutions, frame shifts, and epigenetic silencing can lead to loss of tumor suppressor gene expression, most second step events that eliminate the remaining functional allele are large in size. Large autosomal mutations generally fall into one of three categories: mitotic recombination, chromosome loss, or interstitial deletion [2].
External and internal factors that trigger large autosomal mutations have been reported. Best documented are genotoxin exposures that trigger large mutational events via DNA damage or mitotic spindle damage [3-5]. Deficiencies for DNA repair genes such as p53 and Mlh1 [6,7] can also increase the frequency and alter the spectra of large autosomal mutations. Moreover, distinct autosomal mutational spectra have been observed in different mouse tissues suggesting that cell type and/or cell location play important roles for influencing formation of large mutations [8]. While these studies offer insight into extracellular and intracellular determinants that trigger large mutations, and in many cases the types of mutations that occur (i.e., mutational signatures), they do not explain if chromosome structure and/or sequence play a role. All chromosomes are alike in that they contain a centromere, two telomeres, and common repetitive sequences. They are different, however, in primary sequence, DNA methylation patterns, and distribution of heterochromatic and euchromatic regions [9-11]. A basic issue that has not been addressed with regards to formation of large autosomal mutations is whether different chromosomes have similar propensities to undergo specific types of mutational events because of their common structures, or alternatively if differences in primary sequence and chromatin organization play the dominant role. More simply stated, the issue is whether each chromosome has a distinct mutational spectrum. Addressing this issue will help distinguish environmental and genetic effects from chromosomal effects when examining mutational spectra in different cancers.
To compare the spectra of mutations for different chromosomes, we created clonal short-term mouse kidney cultures that were heterozygous for the Aprt and Tk loci on chromosomes 8 and 11, respectively. By selecting for Aprt and Tk mutant cells in parallel cultures, this approach provided a model system to examine large mutations affecting two different chromosomes. The advantage of using these target loci for a comparative analysis of mutational spectra is that both are located near the telomere of their respective chromosomes, which eliminates having to consider locus position when analyzing the results. The results show that the spectra for large autosomal mutations were essentially the same for mouse chromosomes 8 and 11.
MATERIALS AND METHODS
Isolation of kidney cell lines
C57BL/6 mice heterozygous for both Aprt [12] and Tk [13] were bred with DBA/2 mice to obtain B6D2F1 hybrids that were also heterozygous for both loci. Minced kidneys from four heterozygous hybrid mice were digested with a collagenase solution (Liberase Blendzyme 3, Roche Diagnostics, Indianapolis) and approximately 3% of each digested kidney was plated into a total of five 100 mm dishes in DMEM medium supplemented with 15% heat-inactivated fetal bovine serum. Three weeks later, primary clones were isolated with a cloning ring, trypsinized, and plated in individual wells within 24 well plates. When confluent, the primary clones were expanded sequentially to three wells, a 25 cm2 flask, and finally a 75cm2 flask. Cells from the confluent 75cm2 flasks were then used for the measurement of mutant frequencies, as described below.
Determination of mutant frequencies
Spontaneous Aprt and Tk mutant frequencies were determined for 17 expanded primary clones isolated from 4 mice heterozygous for both selectable loci. To determine the mutant frequencies, 1.3 × 106 cells were plated into thirteen 100 mm dishes (105 cells per dish). Aprt mutants were selected with 80 μg/ml 2,6-diamopurine (DAP) (4 dishes) or 4 μg/ml 2-fluoroadenine (FA) (4 dishes); Tk mutants were selected with 15 μg/ml trifluorothymidine (TFT) (5 dishes). All selection agents were obtained from Sigma (St. Louis, MO). Cloning efficiency dishes in non-selective medium were also set up; the cloning efficiencies ranged between 10 and 37% for the expanded primary kidney clones. Mutant clones were identified after 3 weeks and quantified by direct visualization under the microscope or staining with 0.5% crystal violet. Mutant clones that were observed under the microscope were expanded as above for the parental primary clones until a sufficient number of cells were obtained for a molecular analysis (see below). Mutant frequencies were determined by dividing the number of resistant cells that grew in a given selection medium by the total number of cells plated, with an adjustment for the cloning efficiency.
Molecular analysis
DNA preparations from expanded mutant clones were examined for retention or loss of heterozygosity (LOH) for polymorphic microsatellite loci (Invitrogen, Carlsbad, CA) on chromosome 8 for Aprt mutants or chromosome 11 for Tk mutants. The LOH analysis was conducted at the Plant-Microbe Genomics Facility at Ohio State University. That facility uses an ABI Prism 3700 DNA analyser to separate fluorescently labelled PCR products.
Statistical analysis
Mutant frequencies in different selection regimens (DAP vs. FA) were examined using the empirical logistic transformation [14,15] followed by weighted-least-squares. Chi-squared tests were used to compare FA, DAP, and TFT mutational spectra.
RESULTS
Mutant frequencies and initial screen for loss or retention of the expressed allele
Aprt mutant frequencies were determined by selection with 2,6-diamopurine (DAP) and separately by selection with 2-fluroadenine (FA). In all but one case, the Aprt mutant frequencies were higher for DAP-selected cells than for FA-selected cells (Table 1, clone 759K1f), The average difference was 4.6 times higher mutant frequencies when DAP was used for selection (p < 0.001, 95% CI: 2.54 – 8.29 times higher). FA has a tighter affinity for the Aprt enzyme than DAP [16], which could lead to selection against mutants containing intragenic events (silencing or base-pair changes) with FA selection that would have survived DAP selection. However, a molecular analysis for loss or retention of the wild type Aprt allele in DAP-selected clones versus FA-selected clones showed no significant difference. In this analysis, 23 of 51 (45.1%) DAP resistant clones and 19 of 55 FA (34.5%) resistant clones were found to have retained heterozygosity for Aprt (i.e. both the wild type and knockout alleles were retained) (p = 0.36) demonstrating that FA selection did not preferentially eliminate small intragenic events.
Table 1.
Mutant Frequencies.
| Clone #a | DAPb | FAc | TFTd |
|---|---|---|---|
| 735K1j | 24.0 × 10−4 | 4.3 × 10−4 | 4.8 × 10−4 |
| 740K4e | 17.1 × 10−4 | 2.0 × 10−4 | 2.1 × 10−4 |
| 740K3g | 3.0 × 10−4 | 1.1 × 10−4 | 1.1 × 10−4 |
| 740K2e | 128.3 × 10−4 | 7.3 × 10−4 | 4.0 × 10−4 |
| 740K1d | 1.3 × 10−4 | 1.5 × 10−4 | 1.6 × 10−4 |
| 740K1b | 4.4 × 10−4 | 1.0 × 10−4 | 13.5 × 10−4 |
| 740K1f | 2.5 × 10−4 | < 0.3 × 10−4 | 4.9 × 10−4 |
| 878K4e | 10.6 × 10−4 | 1.2 × 10−4 | 0.5 × 10−4 |
| 878K2a | 4.0 × 10−4 | 0.3 × 10−4 | 0.4 × 10−4 |
| 878K1e | 2.7 × 10−4 | 0.7 × 10−4 | 0.8 × 10−4 |
| 878K3a | 6.8 × 10−4 | 1.2 × 10−4 | 1.9 × 10−4 |
| 878K3g | 4.7 × 10−4 | 2.4 × 10−4 | 6.3 × 10−4 |
| 878K2c | 5.8 × 10−4 | 0.2 × 10−4 | 0.8 × 10−4 |
| 878K3f | 4.1 × 10−4 | 0.2 × 10−4 | 0.3 × 10−4 |
| 759K2e | 25.6 × 10−4 | 3.8 × 10−4 | 1.7 × 10−4 |
| 759K1d | 14.3 × 10−4 | 3.9 × 10−4 | 3.8 × 10−4 |
| 759K1f | 1.4 × 10−4 | 3.5 × 10−4 | 2.2 × 10−4 |
The first three numbers represent the tag numbers of the 4 mice used. Each mouse was heterozygous for both Aprt and Tk.
Cells selected in 80 μg/ml 2,6-diamopurine (DAP).
Cells selected in 4 μg/ml 2-fluoroadenine (FA).
Cells selected in 15 μg/ml trifluorothymidine (TFT).
Tk mutant cells were selected with trifluorothymidine (TFT); the mutant frequencies are presented in Table 1. A molecular analysis showed retention of both the wild type and knockout Tk alleles (i.e. intragenic events) in 20 of 88 mutant cells and LOH for the wild type allele in 56 mutants. The remaining 12 mutant cells exhibited loss of the knockout allele and retention of the wild type allele. This unexpected mutational pattern suggested that two events had occurred, an intragenic event causing loss of expression from the wild type Tk allele and a large mutational event leading to physical loss of the knockout allele. To demonstrate that an intragenic event affecting Tk expression had occurred, two of these cell lines were tested in HAT medium, which requires Tk expression for survival [17], and the cells did not survive. Next, to determine if loss of the knockout Tk allele required TFT selection, or alternatively if it was a frequent and random event unlinked to TFT selection, we ascertained the Tk genotype in 85 Aprt mutants. All Aprt mutants examined were heterozygous for Tk. Together these results demonstrate that the novel combination of retention of the wild type Tk allele and loss of the knockout Tk allele required both TFT selection and loss of Tk activity.
Finally, FA and TFT resistance frequencies were compared to determine if Aprt and Tk mutant frequencies correlated within the individual clones. Although on average the TFT mutant frequencies were 1.3-fold higher than the FA mutant frequencies, this difference was not significant (p = 0.38).
Molecular analysis for chromosomes 8 and 11 in Aprt and Tk mutant cells, respectively
A chromosome-wide molecular analysis was performed for mutant cells exhibiting LOH for Aprt or Tk to identify the type of large mutational event that had occurred. For this analysis, six additional polymorphic loci on chromosome 8 and five additional polymorphic loci on chromosome 11 were examined for loss or retention of heterozygosity in 54 Aprt mutants and 57 Tk mutants, respectively. These loci extended from the centromere to the telomere for both chromosomes (Table 2). DNA preparations from the remaining 10 Aprt and 11 Tk mutants exhibiting LOH were not examined further, nor were most DNA preparations that did not reveal LOH for Aprt or Tk after a sampling showed retention of heterozygosity for all syntenic polymorphic loci on chromosome 8 and 11, respectively, as expected based on prior studies [3,8]. Figure 1 shows how the different molecular patterns can be used to place the mutational events into one of four categories: intragenic event, mitotic recombination, intragenic deletion, and chromosome loss. The mutational spectrum represents the relative proportion of these mutational events in a series of mutant cells.
Table 2.
Polymorphic loci used for LOH analysis.
| Chromosome 8 | Location(cM)a | Chromosome 11 | Location(cM)a |
|---|---|---|---|
| D8Mit124 | 4.4 | D11Mit226 | 2.2 |
| D8Mit125 | 15.4 | D11Mit30 | 37.2 |
| D8Mit75 | 38.3 | D11Mit360 | 64.5 |
| D8Mit166 | 57.9 | D11Mit256 | 77.6 |
| Aprtb | 67.0 | Tkb | 78.0 |
| D8Mit13 | 68.9 | D11Mit104 | 82.0 |
| D8Mit56 | 75.3 | telomerec | 83.1 |
| telomerec | 75.4 |
Relative genetic distance from centromere in centimorgans.
For Aprt, the neo insert in exon 3 and a polymorphic sequence 1 kb upstream of the promoter were used for the LOH analysis; for Tk the neo insert within the promoter was used.
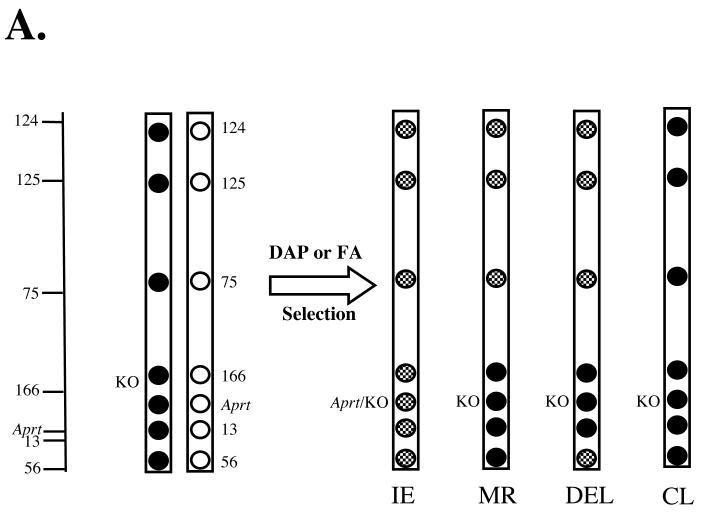
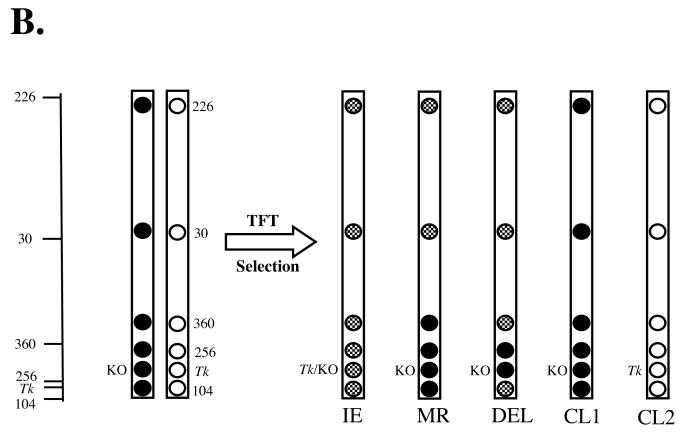
Figure 1. Polymorphic loci examined to determine mutational spectra.
A. Chromosome 8 mutations were analyzed in DAP or FA selected mutants by examining loss or retention of heterozygosity for 7 polymorphic loci, which are represented on the left hand side of the figure by relative cM mappings (see Table 2). Black circles represent loci inherited from the C57BL/6 parent, including the Aprt knockout allele (KO) and white circles represent loci inherited from the DBA/2 parent including the expressed Aprt allele. Heterozygosity for a given locus is represented in a mutational pattern with a mottled black and white (grey) pattern. LOH is represented by a black circle, which represents loss of the DBA/2 allele and retention of the C57BL/6 allele. Representative patterns for intragenic events (IE), mitotic recombination, (MR), intragenic deletion (DEL), and chromosome loss (CL) are shown.
B. Chromosome 11 mutations were analyzed in TFT selected mutants by examining loss or retention of heterozygosity for 6 polymorphic loci on chromosome 11, which are represented on the left hand side of the figure by relative cM mappings (see Table 2). The same analysis as part was followed as part A of this figure. The only difference in the results obtained is that two forms of chromosome loss were observed; CL1, in which all DBA/2 loci were lost including the wild type Tk allele, and CL2, in which all C57BL/6 loci were lost including the knockout Tk allele. See text for more details.
A statistical analysis showed that the mutational spectra for large events were similar for DAP (Table 3A) and FA (Table 3B) selected clones (p = 0.61 for large events or 0.64 if the intragenic events were included). As shown in Table 3C, the predominant large mutational event in the Aprt mutants was loss of the chromosome 8 homologue (from the DBA/2 parent) bearing the wild type allele (53.7% of large events). The second most common large event was mitotic recombination between the chromosome 8 homologues (29.6% of large events) and the least common large event was interstitial deletion for a portion of chromosome 8 containing the expressed Aprt allele (16.7% or large events). Two or more independent Aprt mutations were observed in mutant cells isolated from all but three parental clones, which demonstrates that the mutant frequencies result in part from a variety of independent mutational events.
Table 3.
Spectrum of large mutationsa in DAP, FA, and TFT selected cells.
| A. DAP selected mutantsb | ||||
|---|---|---|---|---|
| MR | CL | Del | ||
| N = 25 | 7 | 15 | 3 | |
| % | 28.0% | 60.0% | 12.0% | |
| B. FA selected mutantsc | ||||
| MR | CL | Del | ||
| N = 29 | 9 | 14 | 6 | |
| % | 31.0% | 48.3% | 20.5% | |
| C. Combined DAP and FA selected mutants | ||||
| MR | CL | Del | ||
| N = 54 | 16 | 29 | 9 | |
| % | 29.6% | 53.7% | 16.7% | |
| D. TFT selected mutantsd | ||||
| MR | CL1e | CL2e | Del | |
| N = 57 | 17 | 19 | 12 | 9 |
| % | 29.8% | 33.3 | 21.0% | 15.8% |
Large mutations fell into three classes: mitotic recombination (MR), chromosome loss (CL), and interstitial deletion (Del). Intragenic events were removed for this analysis.
Cells selected in 80 μg/ml 2,6-diamopurine (DAP).
Cells selected in 4 μg/ml 2-fluoroadenine (FA).
Cells selected in 15 μg/ml trifluorothymidine (TFT).
CL1 represents expected form of chromosome loss in which the lost chromosome contained the expressed Tk allele; CL2 represents an unexpected form of chromosome loss in which the lost chromosome contained the knockout Tk allele.
Two distinct LOH patterns consistent with loss of chromosome 11 were observed in the Tk mutants (Table 3D). One pattern (CL1) (see Fig. 1B) was similar to that observed for loss of chromosome 8 in the Aprt mutants, i.e., LOH for all polymorphic loci on the chromosome 11 homologue bearing the wild type Tk allele and retention of the chromosome 11 homologue bearing the knockout allele (33.3% of large events). The second pattern (CL2) was observed in the 12 mutants that exhibited loss of the knockout Tk allele. In these 12 cases (21.0% of large events), LOH was observed for all polymorphic loci on the chromosome 11 homologue containing the knockout Tk allele, which left only the intact Tk allele (Fig. 1B). While this LOH pattern was novel because it involved the “wrong” chromosome, it still represented a clear example of chromosome loss that was linked to TFT selection because, as discussed above, it did not occur in FA or DAP selected clones. Three parental clones gave rise to both CL1 and CL2 mutations, which demonstrated that these mutational events were not exclusive within a given parental clone. When the CL1 and CL2 LOH patterns representing loss of chromosome 11 were combined, chromosome loss was the most common large mutational event in the Tk mutants (54.3% of large events). The second most common large event in the Tk mutants was mitotic recombination between the chromosome 11 homologues (29.8% of large events) and the least common was interstitial deletion (15.8% of large events) (Table 3D). As noted for Aprt, two or more independent Tk mutations were observed in mutant cells isolated from all but two parental clones, which demonstrates that the mutant frequencies result in part from a variety of independent mutational events.
The mutational spectra for large events affecting chromosome 11 that led to loss of Tk expression (TFT selection) were compared with large events affecting chromosome 8 that led to loss of Aprt expression (DAP and FA selection). This analysis, which combined the CL1 and CL2 events for chromosome 11 because both require TFT selection, showed that the mutational spectra for large events affecting these chromosomes were essentially the same (p = 0.99). If the CL2 chromosome loss events were eliminated from the comparison, chromosome loss remained the most common large mutational event in the TFT selected cells, followed by mitotic recombination and intragenic deletion. Moreover, the chromosome 8 and chromosome 11 mutational spectra would still appear similar at a level that would not be rejected as different by any significance test (p = 0.53).
DISCUSSION
The focus of this study was to determine whether two different mouse autosomes would exhibit similar or different mutational spectra for large events leading to loss of expression of the functional allele in heterozygous cells. In one sense, the structure of different mouse chromosomes is similar. For example, they each have a centromere and two telomeres and they all condense at the same time in preparation for mitosis. In another sense, substantial differences exist between chromosomes, most notably in the different sets of expressed and inactive genes and the distribution of methylation patterns and euchromatic and heterochromatic regions [9-11]. The results of our study demonstrate that with regards to susceptibility to undergo the three main types of large mutational events – chromosome loss, mitotic recombination, and interstitial deletion – two different mouse chromosomes exhibited similar mutational spectra for large mutational events.
Many interrelated factors influence the spectrum of second step autosomal mutations in mammalian cells. For example, we have shown that cell type plays an important role because the spectrum of large autosomal mutations that occurs in kidney epithelial cells is distinct from that occurring in ear mesenchymal cells [8]. Others have shown that significant sequence divergence between homologues will inhibit mitotic recombination [18]. The location of the target gene on a chromosome is also likely to influence the spectrum of second step mutations because closeness to the centromere for a heterozygous gene should reduce the number of mitotic recombinational breakpoints that could lead to a change in genotype. Other factors that influence autosomal mutagenesis are genotoxin exposure [3-5] and DNA repair/DNA damage recognition status [6,7,19]. The latter factors can become impaired as cell lines are established and maintained in culture [20]. The time during the cell cycle when DNA damage occurs can also influence the type of autosomal mutations that can occur [2]. Although Tk is cell cycle regulated [21] whereas Aprt is not [22], which could effect distribution of some base pair substitutions due to transcription coupled repair [23], the large mutational events examined here are not influenced by transcription status of a single gene.
Interestingly, and perhaps somewhat surprisingly, we showed that significant differences in Aprt mutant frequencies resulting from different selective agents (DAP vs. FA) was not accompanied by a difference in the mutational spectra, despite the marked difference in affinities of these two agents for the Aprt enzyme [16]. A potential explanation for the paradoxical observations of similar mutational spectra despite different FA and DAP mutant frequencies is that FA selection causes non-specific loss of some Aprt mutants early during the selection process due to release of toxic privilege metabolites from dying cells [24]. This explanation would not cover, however, the one example in which the FA and DAP resistance frequencies were similar (clone 740K1d) or the one example in which the FA resistance frequency was higher (clone 759K1f). Other potential explanations are random fluctuation from sampling a total of 17 kidney clones or differences in the renal cell types that gave rise to the kidney clones.
Considerable variation was noted for both the Aprt and Tk mutant frequencies. This result is not surprising because prior work from three groups (including ours) has shown such variation for Aprt when comparing mutant frequencies between individual mice for three different cell types (T cells, ear fibroblasts, kidney epithelial) [3,8,25,26]. Moreover, we have shown significant variation for Aprt mutant frequencies when comparing subclones of a mouse kidney cell line [27] and a mouse embryonal carcinoma cell line [28]. In a prior review we have attributed variation in autosomal mutant frequencies to a mixture of the broad spectrum of autosomal mutations and clonal expansion [2] and recently provided evidence for this speculation in an aging study examining a large number of mutant cells from a large number of mice [8]. We did not examine a sufficient number of mutant cells to demonstrate conclusively that clonal expansion was or was not occurring. However, we did show that two or more independent Aprt and Tk mutational events were isolated from the overwhelming majority of parental clones tested, which demonstrates that much of the observed mutant frequencies were due to independent mutational events.
To reduce potential confounders from our study, as many external variables as possible were eliminated. Firstly, both selectable loci are relatively telomeric; Aprt is 8 cM from the chromosome 8 telomere and Tk is 5 cM from the chromosome 11 telomere. Secondly, spontaneous mutagenesis for these two autosomal loci was studied in parallel in short-term kidney cultures obtained from mice heterozygous for both Aprt and Tk. Thirdly, the mice used were F1 hybrids (B6D2F1) and the amount of sequence divergence between the chromosome 8 homologues was presumed to be roughly equivalent to that between the chromosome 11 homologues. Use of a F1 hybrid made its unlikely that differences in mitotic recombination frequencies, had they been observed, would be due to different levels of inter-homologue sequence divergence. Finally, pooling of data from 17 independent short-term kidney cultures (from 4 mice) essentially eliminated concern that deficiencies in DNA repair/damage recognition or aberrant ploidy levels that can arise in permanent cell lines would distort the results.
Despite the marked similarities for large mutations affecting chromosomes 8 and 11 in mouse kidney cells, one issue remains. This issue is the observation that the “wrong” chromosome was lost in a subset of Tk mutants (i.e., the CL2 bearing mutants). In these mutants, all examined loci were lost for the chromosome 11 homologue that contained the knockout Tk allele, including the knockout allele itself; the wild type allele remained. Nonetheless, the cells were Tk deficient because they could not grow in HAT medium. Combined, these observations demonstrated two distinct events had occurred, i.e., an intragenic event and chromosome loss. Moreover, loss of the “wrong” chromosome was linked to TFT selection, and therefore loss of Tk expression, thereby providing a basis for including this event in the study. The p53 tumor suppressor gene is also located on mouse chromosome 11 [29], which raises the possibility that a mutant p53 allele was linked to the formation of the CL2 clones. However, this possibility is not consistent with the observation that both CL1 (wild type allele bearing chromosome is lost) and CL2 mutant clones were derived from three of the parental clones. Whether the unexpected chromosome loss event observed in the CL2 mutants is also linked to a specific type(s) of intragenic events will require further work.
In summary, we have performed the first direct test of the relation between general chromosome structure and large autosomal mutational events. The results demonstrate similar mutational spectra for large events affecting two mouse autosomes despite differences in primary DNA sequences, DNA methylation patterns, and distribution of heterochromatic and euchromatic regions. Although we only examined two mouse autosomes, these results suggest strongly that the internal and external cellular environments provide the driving force for large mutational events, and that generic chromosome structure per se is the substrate substrate upon which these forces act.
ACKNOWLEDGMENTS
This study was supported by NIH grants CA76528 and DK074742 (MST). We thank Drs. Matt Thayer, Maura Pieretti, and Harvey Mohrenweiser for critical readings of this manuscript, Drs. Robert Heflich and Vasily Dobrovolsky for sharing the Tk knockout mouse strain, and Antoinette Olivas for help out of the bullpen.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
REFERENCES
- 1.Knudson AG. Two genetic hits (more or less) to cancer. Nature Rev Cancer. 2001;1:157–162. doi: 10.1038/35101031. [DOI] [PubMed] [Google Scholar]
- 2.Turker MS. Autosomal mutation in somatic cells of the mouse. Mutagenesis. 2003;18:1–6. doi: 10.1093/mutage/18.1.1. [DOI] [PubMed] [Google Scholar]
- 3.Ponomareva ON, Rose JA, Lasarev M, Rasey J, Turker MS. Tissue-specific Deletion and Discontinuous Loss of Heterozygosity Are Signatures for the Mutagenic Effects of Ionizing Radiation in Solid Tissues. Cancer Res. 2002;62:1518–1523. [PubMed] [Google Scholar]
- 4.Von Tungeln LS, Williams LD, Doerge DR, Shaddock JG, McGarrity LJ, Morris SM, Mittelstaedt RA, Heflich RH, Beland FA. Transplacental drug transfer and frequency of Tk and Hprt lymphocyte mutants and peripheral blood micronuclei in mice treated transplacentally with zidovudine and lamivudine. Environ Mol Mutagen. 2006 doi: 10.1002/em.20237. [DOI] [PubMed] [Google Scholar]
- 5.Wijnhoven SW, Sonneveld E, Kool HJ, Van Teijlingen CM, Vrieling H. Chemical carcinogens induce varying patterns of LOH in mouse T- lymphocytes. Carcinogenesis. 2003;24:139–144. doi: 10.1093/carcin/24.1.139. [DOI] [PubMed] [Google Scholar]
- 6.Shao C, Deng L, Chen Y, Kucherlapati R, Stambrook PJ, Tischfield JA. Mlh1 mediates tissue-specific regulation of mitotic recombination. Oncogene. 2004;23:9017–9024. doi: 10.1038/sj.onc.1208148. [DOI] [PubMed] [Google Scholar]
- 7.Liang L, Shao C, Deng L, Mendonca MS, Stambrook PJ, Tischfield JA. Radiation-induced genetic instability in vivo depends on p53 status. Mutat Res. 2002;502:69–80. doi: 10.1016/s0027-5107(02)00029-5. [DOI] [PubMed] [Google Scholar]
- 8.Turker MS, Lasarev M, Connolly L, Kasameyer E, Roessler D. Age-related accumulation of autosomal mutations in solid tissues of the mouse is gender and cell type specific. Aging Cell. 2007;6:73–86. doi: 10.1111/j.1474-9726.2006.00264.x. [DOI] [PubMed] [Google Scholar]
- 9.Stewart A. The functional organization of chromosomes and the nucleus--a special issue. Trends Genet. 1990;6:377–379. doi: 10.1016/0168-9525(90)90282-b. [DOI] [PubMed] [Google Scholar]
- 10.Langer S, Kraus J, Jentsch I, Speicher MR. Multicolor chromosome painting in diagnostic and research applications. Chromosome Res. 2004;12:15–23. doi: 10.1023/b:chro.0000009326.21752.88. [DOI] [PubMed] [Google Scholar]
- 11.Das R, Dimitrova N, Xuan Z, Rollins RA, Haghighi F, Edwards JR, Ju J, Bestor TH, Zhang MQ. Computational prediction of methylation status in human genomic sequences. Proc Natl Acad Sci U S A. 2006;103:10713–10716. doi: 10.1073/pnas.0602949103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Stambrook PJ, Shao C, Stockelman M, Boivin G, Engle SJ, Tischfield JA. APRT: A versatile in vivo resident reporter of local mutation and loss of heterozygosity. Environ Mol Mutagen. 1996;28:471–482. doi: 10.1002/(SICI)1098-2280(1996)28:4<471::AID-EM25>3.0.CO;2-B. [DOI] [PubMed] [Google Scholar]
- 13.Dobrovolsky VN, Casciano DA, Heflich RH. Tk+/− mouse model for detecting in vivo mutation in an endogenous, autosomal gene. Mutat Res. 1999;423:125–136. doi: 10.1016/s0027-5107(98)00234-6. [DOI] [PubMed] [Google Scholar]
- 14.MuCullagh P, Nelder JA. Generalized Linear Methods. 2d ed. Chapman and Hall; London, Great Britian: 1989. [Google Scholar]
- 15.Cox DR. Analysis of Binary Data. Chapman and Hall; New York, New York: 1977. [Google Scholar]
- 16.Krenitsky TA, Neil SM, Elion GB, Hitchings GH. Adenine phosphoribosyltransferase from monkey liver. Specificity and properties. J Biol Chem. 1969;244:4779–4784. [PubMed] [Google Scholar]
- 17.Capecchi MR. High efficiency transformation by direct microinjection of DNA into cultured mammalian cells. Cell. 1980;22:479–488. doi: 10.1016/0092-8674(80)90358-x. [DOI] [PubMed] [Google Scholar]
- 18.Shao C, Stambrook PJ, Tischfield JA. Mitotic recombination is suppressed by chromosomal divergence in hybrids of distantly related mouse strains. Nat Genet. 2001;28:169–172. doi: 10.1038/88897. [DOI] [PubMed] [Google Scholar]
- 19.Wijnhoven SW, Kool HJ, Mullenders LH, Slater R, van Zeeland AA, Vrieling H. DMBA-induced toxic and mutagenic responses vary dramatically between NER-deficient Xpa, Xpc and Csb mice. Carcinogenesis. 2001;22:1099–1106. doi: 10.1093/carcin/22.7.1099. [DOI] [PubMed] [Google Scholar]
- 20.Romanov SR, Kozakiewicz BK, Holst CR, Stampfer MR, Haupt LM, Tlsty TD. Normal human mammary epithelial cells spontaneously escape senescence and acquire genomic changes. Nature. 2001;409:633–637. doi: 10.1038/35054579. [DOI] [PubMed] [Google Scholar]
- 21.Dou QP, Markell PJ, Pardee AB. Thymidine kinase transcription is regulated at G1/S phase by a complex that contains retinoblastoma-like protein and a cdc2 kinase. Proc Natl Acad Sci U S A. 1992;89:3256–3260. doi: 10.1073/pnas.89.8.3256. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.La Bella F, Brown EH, Basilico C. Changes in the levels of viral and cellular gene-transcripts in the cell cycle of SV40 transformed mouse cells. J Cell Physiol. 1983;117:62–68. doi: 10.1002/jcp.1041170110. [DOI] [PubMed] [Google Scholar]
- 23.Vrieling H, van Zeeland AA, Mullenders LH. Transcription coupled repair and its impact on mutagenesis. Mutat Res. 1998;400:135–142. doi: 10.1016/s0027-5107(98)00064-5. [DOI] [PubMed] [Google Scholar]
- 24.Dickerman LH, Tischfield JA. Comparative effects of adenine analogs upon metabolic cooperation between Chinese hamster cells with different levels of adenine phosphoribosyltransferase activity. Mutat Res. 1978;49:83–94. doi: 10.1016/0027-5107(78)90080-5. [DOI] [PubMed] [Google Scholar]
- 25.Shao C, Deng L, Henegariu O, Liang L, Raikwar N, Sahota A, Stambrook PJ, Tischfield JA. Mitotic recombination produces the majority of recessive fibroblast variants in heterozygous mice. Proc Natl Acad Sci U S A. 1999;96:9230–9235. doi: 10.1073/pnas.96.16.9230. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Van Sloun PP, Wijnhoven SW, Kool HJ, Slater R, Weeda G, van Zeeland AA, Lohman PH, Vrieling H. Determination of spontaneous loss of heterozygosity mutations in Aprt heterozygous mice. Nucleic Acids Res. 1998;26:4888–4894. doi: 10.1093/nar/26.21.4888. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Turker MS, Gage BM, Rose JA, Elroy D, Ponomareva ON, Stambrook PJ, Tischfield JA. A novel signature mutation for oxidative damage resembles a mutational pattern found commonly in human cancers. Cancer Res. 1999;59:1837–1839. [PubMed] [Google Scholar]
- 28.Turker MS, Pieretti M, Kumar S. Molecular evidence for the induction of large interstitial deletions on mouse chromosome 8 by ionizing radiation. Mutat Res. 1997;374:201–208. doi: 10.1016/s0027-5107(96)00230-8. [DOI] [PubMed] [Google Scholar]
- 29.Rotter V, Wolf D, Pravtcheva D, Ruddle FH. Chromosomal assignment of the murine gene encoding the transformation-related protein p53. Mol Cell Biol. 1984;4:383–385. doi: 10.1128/mcb.4.2.383. [DOI] [PMC free article] [PubMed] [Google Scholar]