Abstract
Plants can acclimate rapidly to environmental conditions, including high temperatures. To identify molecular events important for acquired thermotolerance, we compared viability and transcript profiles of Arabidopsis thaliana treated to severe heat stress (45°C) without acclimation or following two different acclimation treatments. Notably, a gradual increase to 45°C (22°C to 45°C over 6 h) led to higher survival and to more and higher-fold transcript changes than a step-wise acclimation (90 min at 38°C plus 120 min at 22°C before 45°C). There were significant differences in the total spectrum of transcript changes in the two treatments, but core components of heat acclimation were apparent in the overlap between treatments, emphasizing the importance of performing transcriptome analysis in the context of physiological response. In addition to documenting increases in transcripts of specific genes involved in processes predicted to be required for thermotolerance (i.e. protection of proteins and of translation, limiting oxidative stress), we also found decreases in transcripts (i.e. for programmed cell death, basic metabolism, and biotic stress responses), which are likely equally important for acclimation. Similar protective effects may also be achieved differently, such as prevention of proline accumulation, which is toxic at elevated temperatures and which was reduced by both acclimation treatments but was associated with transcript changes predicted to either reduce proline synthesis or increase degradation in the two acclimation treatments. Finally, phenotypic analysis of T-DNA insertion mutants of genes identified in this analysis defined eight new genes involved in heat acclimation, including cytosolic ascorbate peroxidase and the transcription factors HsfA7a (heat shock transcription factor A7a) and NF-X1.
Abiotic factors, such as temperature and water availability, have a major impact on successful plant establishment, growth, and reproduction. Consequently, plants have evolved mechanisms to monitor their environment and to respond with cellular, physiological, and developmental changes to optimize growth and reproductive success. These environmental response mechanisms include the ability of plants to acclimate rapidly, within hours or days, to extremes in the abiotic environment that would be damaging or lethal without such acclimation (Levitt, 1972; Jenks and Hasegawa, 2005).
The ability of plants to acclimate to normally lethal high temperatures (to acquire thermotolerance) has been described for over 40 years (Alexandrov et al., 1961; Vierling, 1991; Larkindale et al., 2005a, 2005b). Acquisition of this type of heat tolerance is a cell autonomous phenomenon and results from prior exposure to a conditioning pretreatment, which can be a short, sublethal high temperature or other moderate stress treatment. Even plants growing in their optimal environmental range can experience high temperature extremes that may require short-term acclimation. Furthermore, because plants can experience major diurnal temperature fluctuations, the acquisition of thermotolerance may reflect a more general mechanism that contributes to homeostasis of metabolism on a daily basis.
It is well established that an essential component of acquired thermotolerance in plants, as well as other organisms, is the induction and synthesis of molecular chaperones or heat shock proteins (HSPs; Vierling, 1991). In particular, the Hsp100/ClpB chaperone, Hsp101, is required for heat acclimation in plants, yeast (Saccharomyces cerevisiae), and bacteria (Hong and Vierling, 2000; Lee et al., 2005). Recent research, however, has demonstrated that pathways beyond the induction of HSPs are involved in the acquisition of thermotolerance in plants. Mutations disrupting abscisic acid, salicylic acid, hydrogen peroxide, ethylene, and calcium signaling all decrease the ability of plants to acquire thermotolerance, but plants with these defects still appear to accumulate Hsp101 and other HSPs to wild-type levels (Larkindale et al., 2005a, 2005b). Exogenous application of these signaling agents to plants can also result in some degree of enhanced thermotolerance without an accompanying accumulation of HSPs (Larkindale and Knight, 2002). Thus, similar to acclimation responses to other abiotic stresses, such as cold and drought (Levitt, 1972; Jenks and Hasegawa, 2005; Yamaguchi-Shinozaki and Shinozaki, 2006), acclimation to high temperature is clearly effected by a complex network of changes in plants, only one of which is the production of HSPs/molecular chaperones.
The response of the Arabidopsis (Arabidopsis thaliana) transcriptome to heat stress has been examined by several groups (Rizhsky et al., 2004; Busch et al., 2005; Lim et al., 2006; Schramm et al., 2006; Kilian et al., 2007). These studies, however, have not been designed to determine how the observed transcriptional changes correlate with damage from the stress treatment or with the acquisition of thermotolerance. All of these studies noted the up-regulation of transcripts for well-characterized HSPs, including Hsp101, Hsp70s, and small HSPs. Each study also identified additional transcripts that increased dramatically with heat treatment, a number of which were observed by more than one group. These include members of the DREB2 family of transcription factors, galactinol synthase (At2g47180) and other enzymes in the raffinose oligosaccharide pathway, and ASCORBATE PEROXIDASE2 (APX2; At3g09640).
We have been interested in defining genes whose expression is causally related to the ability of plants to acclimate to high temperature, i.e. to acquire thermotolerance. In previous work, we used both forward and reverse genetics to identify genes/pathways essential for this process (Larkindale and Knight, 2002; Hong et al., 2003; Larkindale et al., 2005a, 2005b). Here, we describe results of transcriptome studies designed to identify additional components important for acquired thermotolerance. Although previous studies have analyzed transcript profiles of plants subjected to high temperatures, these studies have not included parallel quantitative analysis of plant (or cell) survival, nor did they consider the degree to which treated plants develop resistance to further extreme stress. To identify transcript changes specifically associated with thermotolerance, we compared viability and global transcript profiles of plants acclimated to severe heat stress (45°C) using two different acclimation treatments (gradual acclimation [G acclimation], 22°C to 45°C over 6 h, versus step-wise acclimation [S acclimation], 90 min at 38°C plus 120 min at 22°C before 45°C), with profiles from plants treated at 45°C without acclimation. Importantly, we show that the gradual increase to high temperature experienced by G-acclimated plants, which is more typical of the natural environment, is much more effective in protecting plants from subsequent heat stress than the S-acclimation treatment. Transcriptome analysis of heat acclimation not only predicts genes and gene clusters whose increased expression is likely to be involved in thermotolerance but also indicates that decreases in expression of specific genes may be equally critical. Examining insertional mutants of highly heat-regulated genes has also confirmed the importance of new factors in heat acclimation.
RESULTS
Different Heat Acclimation Treatments Induce Different Degrees of Thermotolerance
To identify changes in gene expression that could be causally related to the acquisition of thermotolerance, we first characterized the effectiveness of two different acclimation treatments in inducing tolerance to an otherwise lethal 45°C stress (Fig. 1). We reasoned that changes in gene expression common to more than one acclimation treatment are more likely to be required for thermotolerance than changes seen in a single type of acclimation treatment. In one acclimation treatment, typical of treatments we and others have used in the laboratory and designated S for step acclimation, plants were acclimated by heating at a constant, nonlethal temperature (38°C), followed by a 22°C period prior to the 45°C stress (Hong and Vierling, 2000; Larkindale et al., 2005a). The other treatment was designed to mimic temperature changes in the field; the temperature was increased linearly from 22°C to 45°C over 6 h (designated G for gradient acclimation). Both acclimation treatments were compared to nonacclimated plants placed at 45°C (D, for direct). Figure 2 shows that G- and S-acclimated 7-d-old plants survive 45°C, while, as expected, nonacclimated (D) plants die within 5 d. Although G- and S-acclimated plants show similar viability after 70 or 90 min at 45°C, G-acclimated plants survive longer 45°C treatments. The same temperature treatments were also applied to 2.5-d-old dark-grown seedlings with similar results (Supplemental Fig. S1). Thus, gradual increases in temperature, which are more likely to occur under natural conditions, allow the plant to acclimate better than short, abrupt exposure to high but nonlethal temperatures.
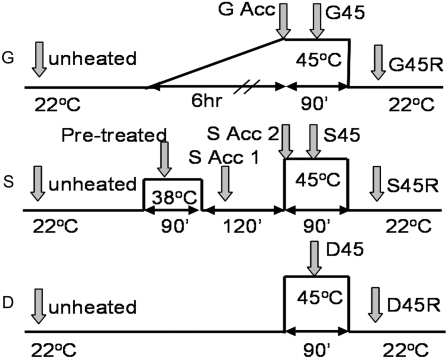
Figure 1.
Heat treatments for comparison of acclimated and nonacclimated plants. Shaded arrows represent sampling times and their designations. All samples except unheated, G Acc, and S Acc2 were taken 30 min after the previous temperature transition. G Acc and S Acc2 were taken at the end of acclimation prior to the shift to 45°C.
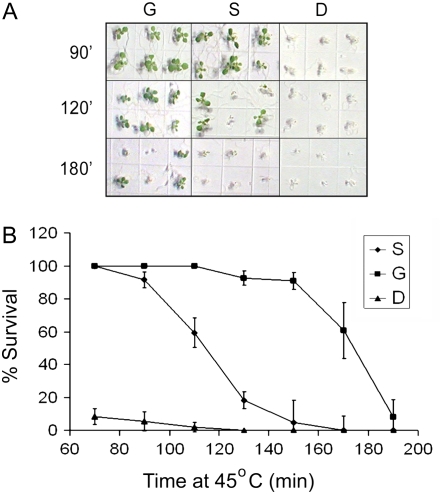
Figure 2.
Thermotolerance of plants given different heat treatments. Percent survival of 7-d-old seedlings acclimated (G and S) or not (D) as shown in Figure 1 and then treated at 45°C for the indicated time. A, Appearance of plants at the time of scoring for viability after the indicated heat treatments. B, Graph showing numerical data. Error bars represent sd from three replicate experiments. [See online article for color version of this figure.]
Thermotolerance Treatments Affect Many Gene Transcripts
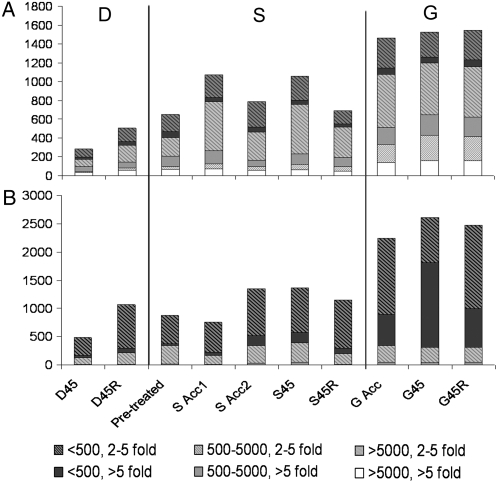
To identify genes that could be involved in the acquisition of thermotolerance, whole-genome microarrays (Affymetrix At-H1) were used to analyze transcript levels. Plant samples were taken over the time course of the G, S, and D heat treatments as shown in Figure 1 in two separate experiments. The 45°C heat stress was limited to 90 min, conditions that result in similar viability of G- and S-acclimated plants (Fig. 2). Recovery samples were taken when plants were still green and not visibly damaged (G, S, and D 45R). Array data were averaged for duplicate samples and filtered as described in “Materials and Methods.” Of the 22,746 genes on the arrays, 4,724 genes passed the filtering criteria and were analyzed further. As compared to unheated samples, large numbers of transcripts increased or decreased during heating, but the G, S, and D treatments affected transcript levels differently. Figure 3A shows the total number of transcripts that increased in abundance either 2- to 5-fold or >5-fold and also the proportions of transcripts that showed different absolute levels of accumulation. At 45°C, many fewer transcripts change in abundance in nonacclimated plants (D45) than in acclimated plants (S45 or G45), consistent with acclimation-protecting processes essential to transcription and RNA stability. Notably, plants subjected to G acclimation showed the largest number of altered transcripts (approximately 1,600 up, approximately 2,500 down), as well as more transcripts with greater fold-changes and higher absolute expression levels than in the S or D treatments. The higher transcript levels in G compared to S samples may contribute to the increased heat tolerance of G-acclimated plants.
Figure 3.
Large numbers of transcripts are increased or decreased by heat treatment. Number of transcripts increased (A) or decreased (B) at the indicated sampling time points as defined in Figure 1. Bars are divided to show classes of transcripts with 2- to 5-fold or >5-fold change in level and to show different absolute expression levels (<500, 500–5,000, >5,000 AU).
The category of transcripts that increased 5-fold or more and accumulated to >5,000 arbitrary units (AU) are likely to represent major effector proteins involved in repair of and recovery from heat damage. There are 185 of these transcripts that are up-regulated by 5-fold or greater in at least one sample (as listed in Supplemental Table S1), approximately one-third of which are HSPs/molecular chaperones. In addition to HSPs and other chaperones, there are five petidyl prolyl isomerases and nine transcripts associated with energy metabolism, in particular, enzymes involved in glycolysis (At4g26270, At2g36460, At5g17310, At5g56630, and At1g79550). There were also 10 non-HSP stress proteins, including two universal stress proteins and several cold- and drought-regulated proteins. Transcripts noted by others as heat regulated are, not surprisingly, part of this list, including HSPs, DREB2A (At5g05410), galactinol synthase (At2g47180), and APX2 (At3g09640).
Significantly more transcripts decreased in abundance under all of the heat treatments than increased in abundance, but again, the greatest change occurred in the most heat tolerant, G-acclimated plants (Fig. 3B). These data suggest RNA turnover and suppression of transcription are also critical for heat tolerance. In contrast to transcripts that increased, 70% to 90% of the decreased transcripts were of low abundance (<500 AU), and there were few transcripts in the >5,000 AU abundance classes. Interestingly, the >5,000 AU expression class included seven pathogen defense-related transcripts (At4g11600, At3g50950, At4g21870, At4g36010, At4g36040, At5g06320, and At5g20630) and four pEARLI-like proteins, which are up-regulated by aluminum and cold stress (Bubier and Schläppi, 2004).
Total Genome Changes in Transcript Levels Vary with Acclimation Treatment
We sought to identify genes common to G and S acclimation that were unaffected in plants heated directly to 45°C (D samples), as these may be critical to thermotolerance. We first used statistical clustering of the complete filtered array data sets to compare overall patterns of transcript abundance. Figure 4A shows the result of Euclidean clustering using average linkage of the different array samples (BRB ArrayTools). For reference, biological replicates had similarities of less than 30 AU. Unexpectedly, the G- and S-acclimation treatments showed significant differences. The overall pattern of transcript changes during the G-acclimation treatment is similar for all three G samples (GAcc, G45, and G45R) but distinct from the S and D samples (Euclidian distance of 95). Thus, although both G- and S-acclimated plants are thermotolerant, more and different changes occurred during G acclimation than in S acclimation. The nonacclimated D samples are also more similar to S-acclimated samples than to the unheated samples, emphasizing that large numbers of transcripts are observed to increase even when plants are treated at temperatures that ultimately lead to plant death.
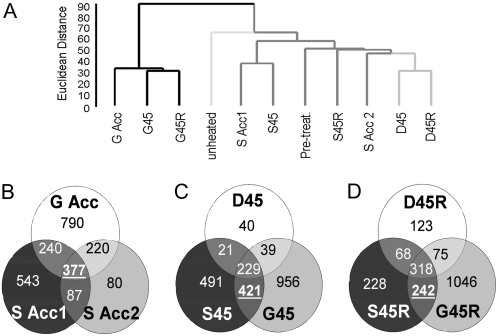
Figure 4.
Comparison of transcripts increased by different heat treatments. A, Dendrogram showing relative similarity between different samples. Total array data were clustered using Euclidean distance and average linkage in BRB array tools. G, S, and D sample branches are each shown with differently shaded lines. B to D, Venn diagrams showing the intersections of total numbers of transcripts increased in different samples. B, Acclimation period samples. C, 45°C samples. D, Recovery samples.
Changes in Specific Sets of Transcripts Are Unique to Thermotolerant Plants
To define a subset of transcripts unique to thermotolerant plants, we first compared sets of transcripts that accumulated at least 2-fold more in the three thermotolerant samples, S Acc1, S Acc2, and G Acc, relative to the unheated sample (Fig. 4B). We suggest that transcripts common to the thermotolerant samples are likely critical to the acclimation process. These include 377 transcripts common to all three samples, 240 transcripts increased in both S Acc1 and G Acc, and 220 common to S Acc2 and G Acc. We next compared transcript sets increased during 45°C stress in acclimated (G45 and S45) and nonacclimated plants (D45; Fig. 4C). There were 229 transcripts increased in all heated samples, which not surprisingly included 33 HSPs/molecular chaperones and six heat shock transcription factors (HSFs). We also found 242 transcripts that increased during recovery in thermotolerant plants (S45R, G45R) but not in nonacclimated plants (Fig. 4D).
Using the comparisons above, we defined transcripts that increased uniquely in thermotolerant plants as those genes found in the intersection of the subsets highlighted in bold and underlined in Figure 4, B to D. There were 57 up-regulated genes that fit these criteria (Supplemental Table S2). Using this stringent definition, there were only two transcription factors in this thermotolerance-specific subset: HsfA3 (At5g03720) and the DREB2B transcription factor, which has been linked to drought stress (At3g11020; Nakashima et al., 2000). The list also includes three known stress proteins: chloroplast-localized Hsp70-7 (At4g24280), cold stress-related kin1/COR6.6 (At5g15960), and a universal stress protein (At3g03270). Analysis with Munich Information Center for Protein Sequences (MIPS) FunCat indicates that compared to the whole genome, this up-regulated subset contains a significantly greater percentage of genes related to protein fate (At2g15790, At4g03320, At5g27660, and At1g01940, two of which are petidyl prolyl isomerases) or involved in energy metabolism (At4g10040, At2g34590, At5g03340, At5g17310). About one-quarter of the genes have no assigned function, as is typical of the entire genome.
When the same analysis was performed to identify down-regulated genes unique to thermotolerant samples (Supplemental Fig. S2, A–C), 69 genes were identified (Supplemental Fig. S3; Supplemental Table S3). The down-regulated genes define diverse functions that are notably distinct from those of up-regulated genes (Supplemental Fig. S2, D and E). Genes found are related to pathogenesis or disease, five cytochrome P450 genes, and an unusually large number of protein kinases (nine). There are no genes in the protein fate or energy metabolism categories, but there are five in the cell defense and virulence category (At5g36910, At4g16860, At4g16880, At3g44970, and At5g55990).
Clusters of Genes Up-Regulated during the Acquisition of Thermotolerance
While genes expressed uniquely in thermotolerant plants may provide critical functions for optimal survival of heat stress, genes whose transcripts increase in nonthermotolerant plants, as well as differences in absolute transcript level, must also be considered. It is already established that chaperones/HSPs contribute to heat tolerance, and as noted above, transcripts of many chaperones/HSPs increase during heating even in nonthermotolerant plants that subsequently die. Therefore, to define additional groups of genes that might be essential for thermotolerance, we used cluster analysis to identify genes showing similar patterns of regulation. Cluster analysis was performed using Euclidean distance and average linkage of transcript levels of all genes under all of the conditions.
We identified 73 clusters, containing from one to 2,124 genes (for complete list, see Supplemental Table S4). Each cluster was analyzed with respect to known promoter elements, responses in other available array data sets, and expression patterns relative to the other clusters (Fig. 5; Supplemental Table S5). All clusters with 10 or more genes showed a statistically significant enrichment of specific sequence motifs within 1,000 bp 5′ of the known or predicted transcription start site when compared to the whole genome. We focused on clusters that contained putative promoter binding sites of known transcription factors and that in many cases contained genes reported to be coregulated under other stress conditions (Vogel et al., 2005; Sakuma et al., 2006).
Figure 5.
Summary of gene cluster data. This summary shows some of the bioinformatics data for the clusters discussed in the text. Clusters in column 1 are grouped based on cis-element analysis and show related expression behavior. The transcript accumulation pattern graphed in column 2 is for the cluster indicated in bold in column 1. These graphs show the average log fold-change (base 2) in transcript level at the time points indicated (as in Fig. 1) versus the unheated control. Dark gray bars, S acclimation; light gray bars, G acclimation (three time points; Acc2 includes the G Acc time point); white bars, D treatment (two time points). Error bars represent sd. The complete summary of gene clusters with expression graphs can be found in Supplemental Table S5.
It is not surprising that six clusters (8, 41–45) included many genes with heat shock elements (HSEs; GAAnnTTC), as this motif binds HSFs (Nover et al., 1996). Genes in these clusters, which are up-regulated under all heating conditions (although not necessarily to the same extent), contain the majority of the highest fold-increased transcripts and include more HSPs (28 genes) than any other group of clusters. Transcripts in these clusters have also generally been identified in other microarray studies of heat-treated Arabidopsis plants or cell cultures (Rizhsky et al., 2004; Busch et al., 2005; Lim et al., 2006; Schramm et al., 2006). Cluster 45, which is equally induced in all three heat treatments here (S, G, and D), includes 18 well-characterized HSPs (Hsp101, 14 small HSPs, and three Hsp70s). Genes in clusters 41 to 44 maintain somewhat higher levels in G-acclimated plants than in S-acclimated plants. In D plants, the transcripts generally appear lower than in either acclimation treatment but are still significantly induced. Clusters 41 to 44 comprise approximately 50% unknown proteins but also include stress-related proteins such as chaperonin 60s, organelle Hsp100/ClpB proteins, two DnaJs, petidyl prolyl isomerases, a glutaredoxin, and the cold-regulated protein COR6.6. While many of these transcripts are likely necessary for thermotolerance, their accumulation is not sufficient, as they also accumulate in nonacclimated plants.
The site II motif (TGGGCC; Welchen and Gonzalez, 2005) was another common promoter motif in clusters with up-regulated genes in acclimated plants. Most genes in these clusters (47, 53, 54, 56, and 57; 821 genes total) are only, or predominantly, up-regulated in G-acclimated plants. Those clusters that were also up-regulated in S acclimation (47 and 56) include the majority of transcripts defined in the Venn diagrams as specifically increased only in thermotolerant plants (Supplemental Table S2). Cluster 57 (484 transcripts), which is uniquely induced in G acclimation, includes 39 ribosomal subunits and 26 transcripts associated with photosynthesis, in addition to eight transcripts associated with transcription/translation and nine associated with protein degradation. Four clusters (510 genes) have a DRE promoter element overrepresented in their promoters (CCGAC; Gilmour et al., 2000). Essentially, none of these transcripts are induced in nonacclimated (D) plants, but notably, only cluster 50 (41 genes) is up-regulated in G-acclimated plants. The fact that all other clusters are significantly up-regulated only in S-acclimated plants further emphasizes the difference between the G- and S-acclimation treatments.
Three clusters (36, 40, and 51) showed an overrepresentation of the abscisic acid response element (ABRE) sequence, ACGTG. Cluster 40 included 25 transcripts, which were up-regulated during both S and G acclimation. Fifteen of these had ABRE sequences, most of which encoded late embryogenesis abundant proteins and unknown proteins, in addition to the DRE-binding transcription factor DREB2A. Cluster 51 included 76 ABRE-containing sequences (of 163). These included five oxidative stress-related transcripts (At1g77510, At4g15660, At4g15700, At5g36270, and At1g30870), three xyloglucan endotransglycolsylases (At4g14130, At4g03210, and At5g65730), an endoglucan transferase (At4g37800), and two subtilisin-like proteinases (At4g21650 and At1g01900). These transcripts showed a range of expression levels, with the trend being for higher expression in G than in S acclimation.
Of the clusters not listed in Figure 5, there were a limited number of up-regulated clusters. One small cluster, cluster 17 (six genes), includes two subunits of NADH dehydrogenase and a cytochrome c oxidase, up-regulated in all heated samples. Genes in clusters 4, 5, 34, and 52 (41 genes total) all increase more in acclimated plants and more in G than in S, but these transcripts are predominantly of unknown function.
Clusters of Genes Down-Regulated during the Acquisition of Thermotolerance
Two groups of clusters that were down-regulated exclusively in acclimated plants were identified. One group (clusters 21, 22, 63B, and 71) contained the W-box motif found in disease resistance genes (Raffaele et al., 2006) and contained 46 disease resistance genes (including classical pathogenesis response proteins PR1 and PR5), 10 cytochrome P450s, and 17 transcripts associated with cell detoxification (mainly glutathione S-transferases).
The other group of down-regulated clusters (46, 62, and 68) contained the sequence TATATA, which may be a form of TATA box (Molina and Grotewold, 2005), and included seven disease resistance genes, six transcripts associated with cell detoxification (mainly glutathione S-transferases), 16 auxin-regulated genes, five expansins, and eight cytochrome P450s. Interestingly, comparison with other available array data showed that these transcripts were also down-regulated in most of the stress and chemical treatment arrays (Genevestigator), suggesting that these may represent short-lived transcripts. Most of the remaining clusters of genes not discussed above (as listed in Supplemental Table S4) were down-regulated and contained primarily disease resistance genes, auxin-induced genes, and enzymes for general metabolism (see Supplemental Table S4).
Transcripts Associated with Cytosolic Protein Synthesis Increase in Acclimated Plants
We utilized MAPMAN array data visualization software (Thimm et al., 2004) as another approach to uncover processes related specifically to acclimation. A very dramatic difference between acclimated and nonacclimated plants was revealed in comparisons of transcripts associated with protein synthesis. Many cytosolic ribosomal proteins are significantly up-regulated throughout G acclimation and are at least transiently up-regulated (though to a lesser extent) during S acclimation but remain notably unchanged in nonacclimated plants (Supplemental Fig. S3). This observation is consistent with results of the cluster analysis, as these genes fall primarily in clusters containing site II promoter elements. At the same time, chloroplast ribosomal protein transcripts are significantly repressed in acclimated plants only. Transcripts associated with translation initiation, elongation, and release are induced by both acclimation treatments, but more so during G acclimation. The fact that none of these transcripts were induced at either the pretreated or the D45 time points explains why they were not identified in previous heat stress transcriptome studies. These data are consistent with the observation that protein synthesis can be acclimated to higher temperatures (Black and Subjeck, 1989) and indicate that early events of acclimation enable enhancement of this transcriptional program.
Differential Responses to Other Abiotic and Biotic Stresses and to Programmed Cell Death with Acclimation
Visualization of changes in stress-related transcripts using MAPMAN also provides insight into potential interactions of stress response networks (Supplemental Fig. S4). Transcripts induced by other abiotic stresses show varied patterns of change. Cold-regulated transcripts accumulate more in S acclimation than in G acclimation, which reinforces the observation that clusters identified as enriched for genes with putative DRE elements are mostly increased in S acclimation only (Fig. 5). Many transcripts associated with oxidative stress are also primarily increased in S acclimation only, although peroxiredoxins and catalase transcripts accumulate under G acclimation as well. These data are consistent with the fact that S-acclimated plants are less thermotolerant than G-acclimated plants and suggest that S-acclimated plants experience greater stress than G-acclimated plants.
Another striking transcriptional change in stress transcripts observed in this MAPMAN analysis concerns biotic stress. There is an obvious decrease in large numbers of transcripts associated with biotic stress in acclimated plants only. The magnitude of the decrease is also greatest for G-acclimated plants. This visualization draws together the down-regulation of disease-related transcripts seen in multiple clusters, including those with W-box motifs (Fig. 5).
The decrease in transcripts associated with biotic stress led us to more carefully examine behavior of transcripts associated with programmed cell death (PCD). We found that MYB30 (At3g28910, cluster 46; Raffaele et al., 2006) and other pro-PCD transcripts (dnd1, At5g15410, lol1, At1g32540, γ-vacuolar processing enzyme, At4g32940; Clough et al., 2000; Epple et al., 2003; Rojo et al., 2004; Watanabe and Lam, 2006) are down-regulated during acclimation. In contrast, putatively anti-PCD transcripts such as Bax inhibitor 1 (At5g47120) and lsd1 and lsd1-like proteins (At4g20380, At4g21610; Epple et al., 2003; Beere, 2004) are up-regulated, particularly in G acclimation.
Different Transcriptional Programs May Achieve Similar Physiological Changes
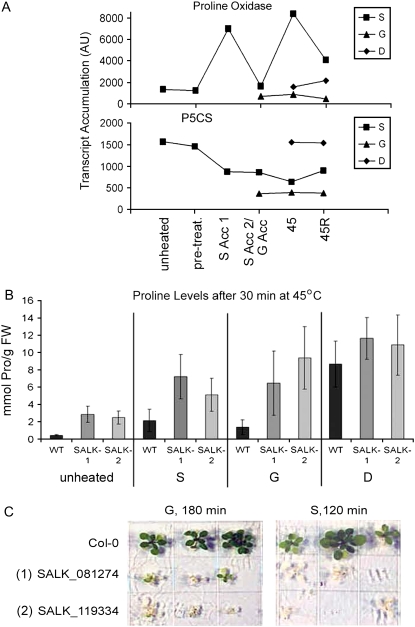
Transcripts increased in only G or S acclimation cannot themselves be essential for thermotolerance. However, they may control similar protective physiological processes by different mechanisms. Levels of transcripts involved in Pro metabolism illustrate this point. It was recently shown that Pro accumulation is toxic to plants at high temperatures (Rizhsky et al., 2004). Free Pro levels are controlled primarily by the balance of synthesis and degradation catalyzed by the key enzymes Δ-1-pyrroline-5-carboxylate synthetase (P5CS; At3g55610) and Pro oxidase/dehydrogenase (ProOx; At3g30775; Hayashi et al., 2000). Notably, during S acclimation, the transcript for ProOx (cluster 63A) accumulates over 5-fold, consistent with an increased ability to remove Pro. In contrast, the ProOx transcript remains low during G acclimation (Fig. 6A). However, in G-acclimated plants, P5CS transcript level is >4-fold reduced compared to unheated plants, while P5CS remains much higher and declines only later in S acclimation. Thus, Pro levels may be controlled in G acclimation by reduced synthesis and in S acclimation by a combination of degradation and reduced synthesis.
Figure 6.
Regulation of Pro metabolism is important for heat tolerance. A, Pro oxidase (At3g30775) and P5CS (At3g55610) transcript levels for the different heat treatments. B, Levels of Pro in plants given the following treatments: unheated, S45, G45, and D45 (as in Fig. 1). Data are averages of five biological replicates, error bars represent sd. C, Survival of 7-d-old S- or G-acclimated plants stressed at 45°C for 120 or 180 min, respectively. Wild type (Col-0) and two Pro oxidase T-DNA insertion lines (Supplemental Fig. S4) are compared. [See online article for color version of this figure.]
To determine if Pro levels were modulated in thermotolerant plants compared to nonacclimated plants, we measured Pro levels at the unheated, S45, D45, and G45 time points (Fig. 6B). Results show that during direct heat stress, leaf Pro content increased on the order of 10-fold or more in the first 30 min of direct heating at 45°C. In S- and G-acclimated plants, the increase in Pro was at least 5 times less than in nonacclimated plants. Thus, control of Pro accumulation appears to be a component of acquired thermotolerance.
To further test the importance of Pro regulation for thermotolerance, we studied two T-DNA insertion alleles of the ProOx gene (Supplemental Fig. S5) and tested the ability of the mutants to acquire thermotolerance. Both alleles were unable to acquire thermotolerance normally under either S- or G-acclimation conditions (Fig. 6, B and C), and both alleles accumulated even higher levels of Pro than wild type, both under control and heated conditions (Fig. 6B). These results further confirm the importance of controlling Pro levels during heat stress. We suggest that a different balance in the regulation of Pro synthesis versus degradation prevents Pro accumulation at high temperature in S- and G-acclimated plants.
Identification of Genes Contributing to Thermotolerance
To test the significance of some of the observed transcript changes, early on in this study we selected 30 genes for mutant analysis (Supplemental Table S6) from the list of 185 genes up-regulated more than 5-fold by heat treatment and showing >5,000 AU of expression (Supplemental Table S1). The criteria for selection included the availability of T-DNA insertion mutant lines and excluded, in general, members of gene families with predicted redundancy (although there are exceptions). This list included the ProOx gene described above (At3g30775), and we also added a putative choline kinase gene (for a total of 31 genes), which showed only >4,000 AU of expression at maximum induction but was over 10-fold induced and known to be regulated by other stresses (Summers and Weretilnyk, 1993; Tasseva et al., 2004). T-DNA insertion lines for 16 of these genes were screened by PCR to identify homozygous mutant lines. Of the 16 tested to identify homozygotes, we were not successful in recovering homozygous lines for two mutants (At5g15450 and At3g24500). Failure to obtain homozygotes of At5g15450, ClpB-p1, was clearly due to lethality of the homozygous mutant as determined from subsequent studies (Lee et al., 2007), whereas insufficient material may have been screened in the case of At3g24500, MBF1c, as others have successfully obtained homozygotes from the same lines (Suzuki et al., 2005). Of the 14 homozygous lines obtained, four were found to have a heat acclimation phenotype (At3g09640, At1g79929, At3g51910, and At3g30775; Table I).
Table I.
Mutants defective in acquired thermotolerance
Homozygous T-DNA insertion mutants, backcrossed once to wild-type Col-0, were tested for acquired thermotolerance as 7-d-old light-grown seedlings after either S acclimation and 120 min at 45°C (7 d [step] column) or gradual acclimation and 150 min at 45°C (7 d [grad] column). Assays were repeated a minimum of three times with at least 12 seedlings and values are expressed as seedling viability as a percent of wild type. For genes for which two insertion mutants were obtained, values for the two lines are indicated. The maximum sd was ±20%, and all values are significantly different from wild type, with a P value ≥ 0.5. Documentation of the homozygosity of the mutants and absence of full-length RNA is provided in Supplemental Figure S5. Ten other mutants shown to be homozygous but to not have a heat stress phenotype were HSFB1 At4g36990; expressed protein At5g67600; LTI78 At5g52310; DREB2A At5g05410; Fer1 At5g01600; immunophilin At4g25340; stress-induced protein At4g12400; ROF1 At3g25230; galactinol synthase At2g47180; and HSFA2 At2g26150. These mutants were not backcrossed to wild type because of the absence of phenotype.
| cis-Elements | Cluster | Description | Accession | 7 d (Step) | 7 d (Grad) | Max Fold Induction |
|---|---|---|---|---|---|---|
| % | ||||||
| HSE | 45 | Hsp101 (hot1-3) | At1g74310 | 0 | 0 | 106 |
| HSE | 7 | APX2 (two insertions) | At3g09640 | 30/20 | 20/20 | 744 |
| 58 | HSFA7a | At3g51910 | 20 | 30 | 25 | |
| 25 | NF-X1 | At1g10170 | 20 | 30 | 12 | |
| 63A | Pro oxidase (two insertions) | At3g30775 | 20/20 | 40/40 | 7 | |
| HSE | 42 | SGT1a | At4g23570 | 40 | 30 | 8 |
| HSE | 41 | Hsp110 (HSP70-15) | At1g79920 | 40 | 60 | 13 |
| 42 | Choline kinase | At1g74320 | 60 | 40 | 10 | |
| DRE | 25 | Thaumatin (two insertions) | At4g36010 | 60/60 | 60/60 | 9 |
For the other 15 of the 31 lines, at least 12 progeny from each line were tested for phenotype directly, using segregating material obtained from the stock center. Of these 15 mutants, four showed a distinct heat acclimation phenotype (At4g23570, At1g10170, At4g36010, and At1g74320) and were subsequently found to be homozygous insertion lines. The other 11 mutants showed no evidence of heat sensitivity among the seedlings tested. It remains possible that some of these genes may prove important for acclimation, but they have not been further analyzed. In total, those lines analyzed as homozygotes but not having a phenotype were HSFB1 At4g36990, expressed protein At5g67600, LTI78 At5g52310, DREB2A At5g05410, Fer1 At5g01600, immunophilin At4g25340, stress-induced protein At4g12400, ROF1 At3g25230, galactinol synthase At2g47180, and HSFA2 At2g26150 (Table I; Supplemental Table S1). We have not rigorously demonstrated that these homozygous mutants without phenotype are indeed null mutants, although the position of the T-DNA insertions early within the gene structure for all but At5g52310, At4g25340, and At4g12400 is consistent with this interpretation (data not shown).
Of the remaining eight T-DNA insertion mutants with heat acclimation phenotypes, in addition to ProOx, second insertion alleles were obtained for APX2 (At3g09640) and thaumatin (At4g36010), and all mutants were then backcrossed once to wild type and homozygous mutant lines reselected and used to obtain the data in Table I. Absence of detectable full-length RNA in all mutants was confirmed by reverse transcription-PCR (Supplemental Fig. S5), indicating that the mutants most likely are null alleles. These eight mutants represent multiple important functions and had heat sensitive phenotypes ranging from mild to severe, although none was as severe as a mutation in Hsp101 (At1g74310), which was completely unviable after the treatments used (Table I; Supplemental Fig. S6). The most severely defective mutants were APX2, a cytosolic ascorbate peroxidase, and two transcription factors, HsfA7a (At3g51910) and NF-X1 (At1g10170), all showing 20% to 30% viability compared to wild type under both the S- and G-acclimation treatments. The next most severe mutants were ProOx and SGT1a (At4g23570), a factor implicated in Hsp90 function (Takahashi et al., 2003; Azevedo et al., 2006). The Hsp110 (At1g79920), choline kinase (At1g74320), and thaumatin (At4g36010) mutants showed moderate, but significant phenotypes. These mutants were also tested in the hypocotyl elongation assay, and all behaved as wild type, with the exception of a mild phenotype for the ProOx mutants (Supplemental Fig. S6). This is consistent with previous data showing that there can be significant differences in heat acclimation phenotypes of mutants at different growth stages (Hong and Vierling, 2000; Larkindale et al., 2005a). Notably, all of these mutants grew identically to wild type in the absence of heat stress, indicating that their heat sensitivity is not compounded by poor growth. In total, we conclude that these eight genes, including ProOx, represent genes that are important for acquired thermotolerance.
DISCUSSION
We have defined transcript profiles associated with plant acclimation to heat stress, revealing the complexity of molecular events that contribute to thermotolerance. We examined two different heat acclimation treatments, a gradual increase to 45°C (G acclimation) and a stepped heat pretreatment (S acclimation), and compared viability and gene expression profiles to nonacclimated, 45°C-treated plants (D treatment). As expected, both acclimation treatments allowed much longer plant survival upon exposure to 45°C than no acclimation. Interestingly, G acclimation, designed to mimic the gradual increase in temperature that would be experienced in the natural environment, induced greater heat tolerance than S acclimation, a treatment typical of those used in heat stress and heat acclimation studies in plants, yeast, bacteria, and mammalian cells (Knop et al., 1985; Mackey and Derrick, 1990; Hong and Vierling, 2000). The difference in response to these two treatments indicates the importance of placing transcript profiling results in the context of physiological responses. Our whole-genome transcript profiling defined specific transcripts (Fig. 4; Supplemental Tables S2 and S3), gene clusters (Fig. 5; Supplemental Table S5), and physiological processes (Supplemental Fig. S7; Supplemental Figs. S4 and S5) that are up-regulated as well as down-regulated in thermotolerant plants. In the response to heat and the process of heat acclimation, we suggest that decreases in gene expression may be of more importance than previously recognized, as nonthermotolerant plants fail to make many of these adjustments. Genes whose transcripts specifically increase associated with thermotolerance, as distinct from genes regulated by heat even in plants that fail to acclimate, have also been identified and can now be tested for their role with available mutants. Finally, among the genes with abundant transcripts that are most highly heat induced, we have identified eight new genes required for maximum heat acclimation, including the cytosolic APX2 and the transcription factors HsfA7a and NF-X1.
Several other studies have examined the response of the Arabidopsis transcriptome to heat stress but not as related to heat acclimation. Rizhsky et al. (2004) reported the effects of heat, drought, and a combination of the two stresses on total transcript profiles, after treatment of Arabidopsis plants at 38°C for 6 h. The 38°C treatment alone slightly increased plant respiration rates and stomatal conductance and induced HSP genes but did not significantly affect photosynthetic rate or relative water content. No specific measures of stress damage or ability to survive subsequent stress were reported. Using Arabidopsis cell cultures, Lim et al. (2006) investigated the affects of 37°C for up to 16 h. This treatment allowed cells to survive a subsequent 9-min, 50°C heat shock, and induction of HSPs was observed. Two studies have tested transcriptional responses to heat treatments in wild-type Arabidopsis plants compared to plants carrying mutations of different HSFs (HsfA1a and A1b tested after 1 h at 37°C [Busch et al., 2005] and HsfA2, tested during 3 h at 42°C [Schramm et al., 2006]). An Arabidopsis Functional Genomics Network study (Kilian et al., 2007; http://www.uni-tuebingen.de/plantphys/AFGN/) investigated both heating at 38°C for 3 h, plus changes in transcripts over 21 h of recovery, with root, shoot, and cell cultures examined separately. This latter study did not correlate results with plant survival. Not surprisingly, the transcript profiles from these studies overlap primarily with the transcripts we observe in our pretreated (38°C for 30 min) and direct heating samples (45°C for 30 min without pretreatment), which includes primarily genes associated with HSE-containing clusters (Supplemental Table S5).
Besides the HSE-containing clusters, most of the transcripts in the other clusters showed no simple pattern of change compared to other studies in which the heat stress transcriptome has been examined. As many of these were transiently increased or changed only under specific treatments, it is not surprising that such differences are seen. Interestingly, however, many clusters identified here showed significant correlations with patterns of gene regulation in array studies involving other stresses. Several different clusters of transcripts (HSE clusters; DRE clusters; site II clusters 47, 54, and 56; TATATA clusters; and clusters 4, 59, 65, and 70) showed a similar up- and down-regulation in response to norflurazon and syringolin as they did to heat (Genevestigator; Michel et al., 2006). Other clusters, such as the putatively DRE-regulated clusters, were en masse up-regulated in response to cold and anoxia (Genevestigator; Branco-Price et al., 2005; Gonzali et al., 2005; Vogel et al., 2005; Oono et al., 2006). The putatively W-box-regulated clusters showed induction in response to biotic stress but repression during heat acclimation. These data suggest that there may be common factors coregulating transcripts within a specific cluster under different conditions.
The greater thermotolerance of G-acclimated compared to S-acclimated plants appears to be due to bona fide differences in the molecular events leading to thermotolerance between the two treatments. Differences in the amount of time plants were exposed to an acclimating temperature do not appear to account for the greater viability of G-acclimated plants. In the treatments analyzed here, plants begin to express HSPs during G acclimation when the temperature reaches approximately 30°C at 225 min prior to 45°C, whereas in S acclimation, HSP expression begins immediately at 38°C, 210 min prior to 45°C (data not shown). Shorter G-acclimation treatments in which the temperature was increased from 22°C to 45°C in 4.5 h resulted in similar protection, while extending the S-acclimation treatment to a total 6 h (90, 180, or 240 min at 38°C, followed by 22°C) did not increase survival compared to the S-acclimation treatment reported here (data not shown). In total, better survival of G-acclimated plants appears due to four general factors: greater expression of heat stress-induced transcripts (including many of the classical HSP/molecular chaperone genes), higher expression of transcripts unique to thermotolerant plants, increases in transcripts unique to the specific heat treatment (gradual heating), and more effective repression during stress of many transcripts presumably not needed until plants have recovered and potentially damaging if expressed during stress.
There were a number of transcriptional changes that occurred only during S acclimation. This may be attributed to thermotolerance being induced through different pathways by the two treatments or to differences in the stress being perceived by the plant. Unique S-acclimation transcripts were primarily those in the DRE cluster 63A, which is also up-regulated by cold, drought, hydrogen peroxide, and anoxic stresses (Supplemental Table S5). The significantly greater up-regulation of general stress-related transcripts during S acclimation can be seen clearly in Supplemental Figure S4. This suggests that under S acclimation, the plant is experiencing other stresses secondary to heat stress. S acclimation may be a shock response whereby the plant responds by repairing stress-induced damage. In comparison, the greater numbers and magnitudes of changes in transcript abundance during G acclimation suggest a more adaptive response whereby damage is prevented; the plant reduces its general metabolism and may reduce damage that could occur due to buildup of toxic intermediates. These data further emphasize how the nature of the applied stress significantly affects the outcome of transcriptome studies and supports the need for carefully controlled conditions and documentation of associated physiological responses.
Despite the differences in transcript profiles between the two acclimation treatments, ultimately the same systems must be protected/repaired. We hypothesize that the same physiological outcome might be achieved in different ways. One example of this is the accumulation of Pro, where we suggest that lower levels of this toxic metabolite are achieved by reduction of synthesis in G acclimation and increased degradation in S acclimation. We show that the ability to degrade Pro through Pro oxidase is essential for thermotolerance. Mutants of ProOx showed reduced thermotolerance, had higher Pro levels prior to heat stress, and had similar Pro levels after heat stress in both acclimated and nonacclimated plants. By comparison, wild-type plants subjected to either S or G acclimation did not accumulate high levels of Pro during heat treatment. Previous work has shown that high levels of Pro result in PCD in Arabidopsis at high temperature (Rizhsky et al., 2004). In that study, Pro levels were shown to increase during drought stress but remain low during simultaneous drought and heat stress, or heat stress alone (6 h at 38°C, temperature used in S acclimation). Thus, we show that both acclimation treatments prevent heat-induced Pro induction but that the mechanism employed may be different for the two treatments.
It is well accepted that protection and refolding of cellular proteins through the HSP network of molecular chaperones are important for survival of high temperature stress (Vierling, 1991; Larkindale et al., 2005b), and many genes for these components appeared in HSE-containing clusters, which are up-regulated by all heat treatments. Four of the eight genes we found to have a reduced ability to acclimate to high temperature were in these clusters: APX2, Hsp110, SGT1a, and choline kinase (Table I). APX2 has been documented as heat stress induced in several studies (Shi et al., 2001; Larkindale and Huang, 2004; Schramm et al., 2006), and it appears to be regulated by HsfA2 (Schramm et al., 2006). Previous studies also indicate it is involved in survival of high light stress (Rossel et al., 2006; Giacomelli et al., 2007). It remains unclear why this particular cytosolic APX isoform is required for heat tolerance (Arabidopsis has two other cytosolic APX proteins; Panchuk et al., 2005). Determining whether this results from a specific property of the APX2 isoform, or if it is the result of the heat regulation will be of interest to determine. Hsp110, SGT1a, and choline kinase had milder but significant effects on heat acclimation. Hsp110 is a member of the larger Hsp70 family (Lin et al., 2001) but, unlike Hsp70s, does not appear to be essential for normal growth. SGT1a is associated with Hsp90 and RAR1 in the development of R-protein modulate disease resistance (Niikura and Kitagawa, 2003; Takahashi et al., 2003; Azevedo et al., 2006), and this is the first indication it may also be important for abiotic stress. Little is known about choline kinases in plants, but their induction by other stresses, specifically salt stress, is documented (Summers and Weretilnyk, 1993; Shank et al., 2001; Tasseva et al., 2004), and the potential importance of this enzyme in heat tolerance could be due to subtle modulation of membrane structure and/or lipid signaling.
Although it has long been known that Hsfs are major transcription factors involved in gene transcription in response to heat stress, the relative roles of the 21 different Arabidopsis Hsfs as well as other transcription factors in heat acclimation are far from resolved. Of the 21 Hsfs, only Hsfs A1e, A2, A3, A7a, B1, and B2b were shown to be heat induced. Charng et al. (2007) reported that insertion mutants of HsfA2 show a reduced ability to sustain thermotolerance (Charng et al., 2007), although they are not required for tolerance to short-term heat stress as tested here. HsfA1a/A1b double mutants show some reduction in their ability to synthesize Hsps but show little reduced thermotolerance (Busch et al., 2005). Here, we show that HsfA7a is one of the Hsfs that contributes to heat acclimation. Determining whether this Hsf has specific targets or works in conjunction with other Hsfs will be of interest to investigate. Among the other Hsfs, we have recently shown that HsfA3, which is heat induced and found in the DR-containing clusters in our study, is regulated by the DREB transcription factors Dreb2A and Dreb2B. Furthermore, HsfA3 RNAi plants and an HsfA3 T-DNA insertion mutant have reduced heat tolerance (Schramm et al., 2008). Notably, HsfA3 and Dreb2B were the only two transcription factors among the genes whose induction was unique to thermotolerant plants (Supplemental Table S2). Although others have reported heat sensitivity of insertion mutants of DREB2A plants, we have not found a defect in heat acclimation for either DREB2A or DREB2B mutants (J. Larkindale, unpublished data). However, we would expect that double mutants of these two factors would be heat sensitive. Our data also identify another transcription factor involved in heat acclimation, NF-X1. NF-X1 also clustered with the DRE group, although it has no apparent DRE elements. This gene is also up-regulated by a wide range of other abiotic stresses (e.g. drought, hypoxia, oxidative stress, salt stress, and wounding; see Genevestigator), and recent data show NF-X1 mutants have reduced growth and survival under salt stress (Lisso et al., 2006). Thus, we can now begin to link heat acclimation with drought/osmotic stress through the DREB2 and NF-X1 transcription factors.
The potential importance for stress acclimation of genes with the site II motif has not previously been recognized. Site II clusters up-regulated in both S and G acclimation included splicing factors (At1g36730, At2g18510, and At4g03430), a putative elongation factor (At2g38560), ribosomal proteins (At3g27450 and At3g11120), and translation initiation factors (At1g36730 and At2g04520). Other transcripts within these clusters were associated with ubiquitin and proteasomes (10 transcripts) and with protein folding (20 transcripts, including six chaperonins, three petidyl prolyl isomerases, and two Hsp70s). Importantly, site II cluster 57 (484 genes) was up-regulated exclusively in G-acclimated plants. It includes many additional translation initiation and elongation factors and ribosomal proteins, as well as photosystem subunits. The dramatic increase in components associated with cytosolic protein translation was further emphasized by visualization using MAPMAN (Supplemental Fig. S3). Thus, G-acclimated plants appear well positioned to repair and restart translation and photosynthesis quickly after heat stress and to maintain translation and photosynthesis to higher temperatures, no doubt contributing to their higher survival rate. The heat-induced translation initiation components may account in part for decreased ability to acclimate. Indeed, preliminary analysis of T-DNA insertion mutations in eIF1A-2 (At2g04520) and eIF5-2 (At1g36370) indicates that these factors are dispensable for normal growth but contribute to heat tolerance (data not shown).
Little attention has been paid to transcripts decreased during stress treatments, but results here suggest that decreases in specific transcripts play an important role in stress acclimation. Many transcripts associated with growth and general metabolism were down-regulated during S acclimation and even more so in G acclimation. Transcripts containing TATATA in their promoters also remained down-regulated in G45R samples, when other transcripts had returned to normal. Molina and Grotewold (2005) determined that only about 30% of the transcripts in the Arabidopsis genome have A/T-rich sequences in their promoters and that these transcripts tended to be highly expressed under nonstress conditions. MIPS FunCat identified transcripts in these clusters as overrepresented in metabolic genes. Acclimated plants therefore appear to be effectively limiting nonessential cellular processes until the return to temperatures permissive for growth.
Our data further indicate that a decrease in the potential for induction of PCD is important for acclimation to high temperature in plants. High temperatures in nonacclimated plants have been shown to induce a form of PCD (Swidzinski et al., 2002), and in animal systems, PCD is reported to be inhibited by HSPs (Beere, 2004). We saw decreases in cell death promoting transcripts and increases in cell death inhibiting transcripts only in thermotolerant plants. Furthermore, reports on treatments inducing PCD indicate heat-induced genes (especially those in the HSP clusters) are generally repressed, suggesting antagonism between acclimation and PCD pathways (Supplemental Table S5). In contrast, chemical treatments that induce PCD in plants (syringolin, secreted by Pseudomonas syringae pv syringae and isoxaben, a herbicide that inhibits cellulose synthesis), mimicked the effects of heat on both up- and down-regulated clusters of transcripts (Supplemental Table S5; Genevestigator). Thus, there appears to be a complex interaction between heat stress, acclimation to high temperatures, and the induction of PCD in plants.
MATERIALS AND METHODS
Plant Growth and Heat Treatment
Wild-type Arabidopsis (Arabidopsis thaliana) plants, ecotype Columbia-0 (Col-0), were grown on plates as described (Larkindale et al., 2005a). Ten-day-old plants were heat treated as in Figure 1. All samples were taken in the light except the pretreated, S45, and D45 time points. All RNA and protein samples were taken 8.5 h into the 16-h light cycle. For each sample, plants were pooled from three plates (approximately 300 plants). Microarray samples were repeated in duplicate, other assays in at least triplicate. Thermotolerance assays of 2.5-d-dark-grown and 7-d-light-grown seedlings were performed as described (Hong and Vierling, 2000; Larkindale et al., 2005a). Homozygous lines carrying T-DNA insertions were identified, ones with acclimation phenotypes were backcrossed once to wild-type plants, and homozygous F2 lines reselected and tested for expression of full-length mRNA for the mutant gene (Supplemental Fig. S5).
Microarray Analysis
RNA samples were prepared with a Qiagen RNeasy plant mini kit. Quality and quantity of RNA was determined by standard methods. RNA was used for cDNA synthesis, labeling, and hybridization of Affymetrix AtH1 arrays (22,746 genes) according to the Affymetrix GeneChip Expression Analysis technical manual (http://www.affymetrix.com) by the Microarray Core Facility, University of Arizona. Duplicate biological replicates were hybridized to separate arrays. Data file quantification was performed as optimized by Affymetrix. Data quality was confirmed by examining standard parameters for Affymetrix arrays with Microarray Suite (MAS 5.0). Data were normalized by scaling the expression of all probe sets to a median expression level of genes on an array to 500 AU. These data were then analyzed with Data Mining Tool (v 3.0, Affymetrix; http://www.affymetrix.com), Microsoft Excel, and BRB ArrayTools (http://linus.nci.nih.gov/BRB-ArrayTools.html). An average expression value for each gene for each experimental condition was generated from the normalized data from the duplicate arrays. Genes were labeled as present only if the probability of detection was P < 0.05, leaving 17,312 transcripts that were detected on at least two duplicate arrays. Data were validated by comparison with real-time PCR for a representative set of genes (Supplemental Fig. S7). For cluster analysis, data were further filtered such that genes were excluded if there was less than a 2-fold change between any individual array and the median value for that gene on all of the arrays, or if the gene was detected on only one or two of the 11 array samples. A total of 4,724 genes passed this filtering and were analyzed further. Clustering of both total transcript accumulation within a specific treatment and of individual genes was done using Euclidean distance and average linkage in BRB array tools. Genes in each cluster were exported into Data Mining Tool and expression of the selected genes across all arrays was compared.
Promoter analysis was performed using 1,000 bp of upstream gene sequence obtained from The Arabidopsis Information Resource (http://www.arabidopsis.org/). cis-Elements were identified by blind searching for common six-letter words using The Arabidopsis Information Resource's motif analysis selecting only those words with a P value of <10−4 and by direct searching for defined non-six-letter cis-element sequences. These words were then compared to known cis-elements using PlantCare (http://oberon.fvms.ugent.be:8080/PlantCARE/index.html) and Google (www.google.com). When similarities to known cis-elements were identified in clusters of interest, the original sequence was searched for the entire element and its variants. Expression was compared to other conditions using Genevestigator (www.genevestigator.ethz.ch; Zimmermann et al., 2004). For experiments in the database involving heat treatments, the array data were downloaded and compared with array data from this study. Transcript levels relative to physiological pathways were analyzed using MAPMAN (Thimm et al., 2004; http://gabi.rzpd.de/projects/MapMan/). Genes of similar function were identified using gene ontology annotations (http://www.arabidopsis.org/) and FUNCAT (http://mips.gsf.de/).
Pro Measurements
Pro levels were measured using the method described by Bates et al. (1973). Plant tissue was harvested at the time points indicated and immediately frozen in liquid nitrogen. Tissues samples were ground in microfuge tubes in 3% (v/v) sulfosalicyclic acid and 10% celite resin and incubated at room temperature overnight prior to assay. Then 0.5 mL of sample was added to 2 mL ninhydrin (1% w/v in glacial acetic acid:water 60:40, v/v), and the tubes were boiled for 1 h. After cooling, the chromophore was extracted with 5 mL of toluene, and the absorbance was measured at 520 nm against a tissue-free blank. Amounts of Pro were calculated by comparison to a standard curve.
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. Thermotolerance of dark-grown seedlings given different acclimation treatments.
Supplemental Figure S2. Venn diagrams showing down-regulated genes and gene ontology annotations for up- and down-regulated genes.
Supplemental Figure S3. Changes in accumulation of transcripts associated with protein synthesis.
Supplemental Figure S4. Changes in accumulation of transcripts associated with stress responses.
Supplemental Figure S5. Summary of homozygous mutants tested for heat stress phenotype.
Supplemental Figure S6. Quantitative data for the eight insertion mutants with heat stress phenotypes.
Supplemental Figure S7. Real-time PCR validation of array data.
Supplemental Table S1. Transcripts showing a high level of expression (>5,000 AU) and high fold-change in response to high temperature (>5-fold).
Supplemental Table S2. Transcripts up-regulated under all thermotolerance treatments.
Supplemental Table S3. Transcripts down-regulated under all thermotolerance treatments.
Supplemental Table S4. Gene listing for the clusters of genes identified by Euclidean distance analysis.
Supplemental Table S5. Complete summary of gene clusters.
Supplemental Table S6. Summary of mutant data.
Supplementary Material
Acknowledgments
Thanks to Kevin Keisler (Microarray Core Facility, University of Arizona) for help with processing the microarray data and David Mount (University of Arizona) for help with analysis. We also thank many past members of the Vierling lab who helped with initial screening and testing of many of the T-DNA mutants discussed in this study.
This work was supported by the National Science Foundation (grant no. IBN–0213128) and by the U.S. Department of Agriculture (grant no. NRICGP 99–351007618 to E.V.).
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Elizabeth Vierling (vierling@email.arizona.edu).
Some figures in this article are displayed in color online but in black and white in the print edition.
The online version of this article contains Web-only data.
Open Access articles can be viewed online without a subscription.
References
- Alexandrov V, Ouchakov B, Poljansky G (1961) The thermal death of cells in relation to the problem of the adaptation of organisms to the temperature of the environment. Pathol Biol 9 849–854 [PubMed] [Google Scholar]
- Azevedo C, Betsuyaku S, Peart J, Takahashi A, Noel L, Sadanandom A, Casais C, Parker J, Shirasu K (2006) Role of SGT1 in resistance protein accumulation in plant immunity. EMBO J 25 2007–2016 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bates LS, Waldren RP, Teare ID (1973) Rapid determination of free proline for water-stress studies. Plant Soil 39 205–207 [Google Scholar]
- Beere H (2004) ‘The stress of dying’: the role of heat shock proteins in the regulation of apoptosis. J Cell Sci 117 2641–2651 [DOI] [PubMed] [Google Scholar]
- Black AR, Subjeck JR (1989) Involvement of rRNA synthesis in the enhanced survival and recovery of protein synthesis seen in thermotolerance. J Cell Physiol 138 439–449 [DOI] [PubMed] [Google Scholar]
- Branco-Price C, Kawaguchi R, Ferreira RB, Bailey-Serres J (2005) Genome-wide analysis of transcript abundance and translation in Arabidopsis seedlings subjected to oxygen deprivation. Ann Bot (Lond) 96 647–660 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bubier J, Schläppi M (2004) Cold induction of EARLI1, a putative Arabidopsis lipid transfer protein, is light and calcium dependent. Plant Cell Environ 27 929–936 [Google Scholar]
- Busch W, Wunderlich M, Schöffl F (2005) Identification of novel heat shock factor-dependent genes and biochemical pathways in Arabidopsis thaliana. Plant J 41 1–14 [DOI] [PubMed] [Google Scholar]
- Charng YY, Liu HC, Liu NY, Chi WT, Wang CN, Chang SH, Wang TT (2007) A heat-inducible transcription factor, HsfA2, is required for extension of acquired thermotolerance in Arabidopsis. Plant Physiol 143 251–262 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clough SJ, Fengler KA, Yu IC, Lippok B, Smith RK Jr, Bent AF (2000) The Arabidopsis dnd1 “defense, no death” gene encodes a mutated cyclic nucleotide-gated ion channel. Proc Natl Acad Sci USA 97 9323–9328 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Epple P, Mack AA, Morris VR, Dangl JL (2003) Antagonistic control of oxidative stress-induced cell death in Arabidopsis by two related, plant-specific zinc finger proteins. Proc Natl Acad Sci USA 100 6831–6836 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Giacomelli L, Masi A, Ripoll DR, Lee MJ, van Wijk KJ (2007) Arabidopsis thaliana deficient in two chloroplast ascorbate peroxidases shows accelerated light-induced necrosis when levels of cellular ascorbate are low. Plant Mol Biol 65 627–644 [DOI] [PubMed] [Google Scholar]
- Gilmour SJ, Sebolt AM, Salazar MP, Everard JD, Thomashow MF (2000) Overexpression of the Arabidopsis CBF3 transcriptional activator mimics multiple biochemical changes associated with cold acclimation. Plant Physiol 124 1854–1865 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gonzali S, Loreti E, Novi G, Poggi A, Alpi A, Perata P (2005) The use of microarrays to study the anaerobic response in Arabidopsis. Ann Bot (Lond) 96 661–668 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hayashi F, Ichino T, Osanai M, Wada K (2000) Oscillation and regulation of proline content by P5CS and ProDH gene expressions in the light/dark cycles in Arabidopsis thaliana L. Plant Cell Physiol 41 1096–1101 [DOI] [PubMed] [Google Scholar]
- Hong SW, Lee U, Vierling E (2003) Arabidopsis hot mutants define multiple functions required for acclimation to high temperatures. Plant Physiol 132 757–767 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong SW, Vierling E (2000) Mutants of Arabidopsis thaliana defective in the acquisition of tolerance to high temperature stress. Proc Natl Acad Sci USA 97 4392–4397 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jenks M, Hasegawa P (2005) Plant Abiotic Stress. Blackwell Publishing, Oxford
- Kilian J, Whitehead D, Horak J, Wanke D, Weinl S, Batistic O, D'Angelo C, Bornberg-Bauer E, Kudla J, Harter K (2007) The AtGenExpress global stress expression data set: protocols, evaluation and model data analysis of UV-B light, drought and cold stress responses. Plant J 50 347–363 [DOI] [PubMed] [Google Scholar]
- Knop RH, Chen CW, Mitchell JB, Russo A, McPherson S, Cohen JS (1985) Adaptive cellular response to hyperthermia: 31P-NMR studies. Biochim Biophys Acta 845 171–177 [DOI] [PubMed] [Google Scholar]
- Larkindale J, Hall JD, Knight MR, Vierling E (2005. a) Heat stress phenotypes of Arabidopsis mutants implicate multiple signaling pathways in the acquisition of thermotolerance. Plant Physiol 138 882–897 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larkindale J, Huang B (2004) Thermotolerance and antioxidant systems in Agrostis stolonifera: involvement of salicylic acid, abscisic acid, calcium, hydrogen peroxide, and ethylene. J Plant Physiol 161 405–413 [DOI] [PubMed] [Google Scholar]
- Larkindale J, Knight MR (2002) Protection against heat stress-induced oxidative damage in Arabidopsis involves calcium, abscisic acid, ethylene, and salicylic acid. Plant Physiol 128 682–695 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Larkindale J, Mishkind M, Vierling E (2005. b) Plant responses to high temperature. In M Jenks, P Hasegawa, eds, Plant Abiotic Stress. Blackwell, Oxford
- Lee U, Rioflorido I, Hong SW, Larkindale J, Waters ER, Vierling E (2007) The Arabidopsis ClpB/Hsp100 family of proteins: chaperones for stress and chloroplast development. Plant J 49 115–127 [DOI] [PubMed] [Google Scholar]
- Lee U, Wie C, Escobar M, Williams B, Hong SW, Vierling E (2005) Genetic analysis reveals domain interactions of Arabidopsis Hsp100/ClpB and cooperation with the small heat shock protein chaperone system. Plant Cell 17 559–571 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levitt J (1972) Responses of Plants to Environmental Stresses. Academic Press, New York
- Lim CJ, Yang KA, Hong JK, Choi JS, Yun DJ, Hong JC, Chung WS, Lee SY, Cho MJ, Lim CO (2006) Gene expression profiles during heat acclimation in Arabidopsis thaliana suspension-culture cells. J Plant Res 119 373–383 [DOI] [PubMed] [Google Scholar]
- Lin BL, Wang JS, Liu HC, Chen RW, Meyer Y, Barakat AS, Delseny M (2001) Genomic analysis of the Hsp70 superfamily in Arabidopsis thaliana. Cell Stress Chaperones 6 201–208 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lisso J, Altmann T, Mussig C (2006) The AtNFXL1 gene encodes a NF-X1 type zinc finger protein required for growth under salt stress. FEBS Lett 580 4851–4856 [DOI] [PubMed] [Google Scholar]
- Mackey BM, Derrick C (1990) Heat shock protein synthesis and thermotolerance in Salmonella typhimurium. J Appl Bacteriol 69 373–383 [DOI] [PubMed] [Google Scholar]
- Michel K, Abderhalden O, Bruggmann R, Dudler R (2006) Transcriptional changes in powdery mildew infected wheat and Arabidopsis leaves undergoing syringolin-triggered hypersensitive cell death at infection sites. Plant Mol Biol 62 561–578 [DOI] [PubMed] [Google Scholar]
- Molina C, Grotewold E (2005) Genome wide analysis of Arabidopsis core promoters. BMC Genomics 6 25. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nakashima K, Shinwari ZK, Sakuma Y, Seki M, Miura S, Shinozaki K, Yamaguchi-Shinozaki K (2000) Organization and expression of two Arabidopsis DREB2 genes encoding DRE-binding proteins involved in dehydration- and high-salinity-responsive gene expression. Plant Mol Biol 42 657–665 [DOI] [PubMed] [Google Scholar]
- Niikura Y, Kitagawa K (2003) Identification of a novel splice variant: human SGT1B (SUGT1B). DNA Seq 14 436–441 [DOI] [PubMed] [Google Scholar]
- Nover L, Scharf KD, Gagliardi D, Vergne P, Czarnecka-Verner E, Gurley WB (1996) The Hsf world: classification and properties of plant heat stress transcription factors. Cell Stress Chaperones 1 215–223 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Oono Y, Seki M, Satou M, Iida K, Akiyama K, Sakurai T, Fujita M, Yamaguchi-Shinozaki K, Shinozaki K (2006) Monitoring expression profiles of Arabidopsis genes during cold acclimation and deacclimation using DNA microarrays. Funct Integr Genomics 6 212–234 [DOI] [PubMed] [Google Scholar]
- Panchuk II, Zentgraf U, Volkov RA (2005) Expression of the Apx gene family during leaf senescence of Arabidopsis thaliana. Planta 222 926–932 [DOI] [PubMed] [Google Scholar]
- Raffaele S, Rivas S, Roby D (2006) An essential role for salicylic acid in AtMYB30-mediated control of the hypersensitive cell death program in Arabidopsis. FEBS Lett 580 3498–3504 [DOI] [PubMed] [Google Scholar]
- Rizhsky L, Liang H, Shuman J, Shulaev V, Davletova S, Mittler R (2004) When defense pathways collide: the response of Arabidopsis to a combination of drought and heat stress. Plant Physiol 134 1683–1696 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rojo E, Martin R, Carter C, Zouhar J, Pan S, Plotnikova J, Jin H, Paneque M, Sanchez-Serrano JJ, Baker B, et al (2004) VPEgamma exhibits a caspase-like activity that contributes to defense against pathogens. Curr Biol 14 1897–1906 [DOI] [PubMed] [Google Scholar]
- Rossel JB, Walter PB, Hendrickson L, Chow WS, Poole A, Mullineaux PM, Pogson BJ (2006) A mutation affecting ASCORBATE PEROXIDASE 2 gene expression reveals a link between responses to high light and drought tolerance. Plant Cell Environ 29 269–281 [DOI] [PubMed] [Google Scholar]
- Sakuma Y, Maruyama K, Osakabe Y, Qin F, Seki M, Shinozaki K, Yamaguchi-Shinozaki K (2006) Functional analysis of an Arabidopsis transcription factor, DREB2A, involved in drought-responsive gene expression. Plant Cell 18 1292–1309 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schramm F, Ganguli A, Kiehlmann E, Englich G, Walch D, von Koskull-Doring P (2006) The heat stress transcription factor HsfA2 serves as a regulatory amplifier of a subset of genes in the heat stress response in Arabidopsis. Plant Mol Biol 60 759–772 [DOI] [PubMed] [Google Scholar]
- Schramm F, Larkindale J, Kiehlmann E, Ganguli A, Englich G, Vierling E, von Koskull-Döring P (2008) A novel network of transcription factor DREB2A and heat stress transcription factor HsfA3 regulates the heat stress response of Arabidopsis. Plant Cell (in press) [DOI] [PubMed]
- Shank KJ, Su P, Brglez I, Boss WF, Dewey RE, Boston RS (2001) Induction of lipid metabolic enzymes during the endoplasmic reticulum stress response in plants. Plant Physiol 126 267–277 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shi WM, Muramoto Y, Ueda A, Takabe T (2001) Cloning of peroxisomal ascorbate peroxidase gene from barley and enhanced thermotolerance by overexpressing in Arabidopsis thaliana. Gene 273 23–27 [DOI] [PubMed] [Google Scholar]
- Summers PS, Weretilnyk EA (1993) Choline synthesis in spinach in relation to salt stress. Plant Physiol 103 1269–1276 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Suzuki N, Rizhsky L, Liang H, Shuman J, Shulaev V, Mittler R (2005) Enhanced tolerance to environmental stress in transgenic plants expressing the transcriptional coactivator multiprotein bridging factor 1c. Plant Physiol 139 1313–1322 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swidzinski JA, Sweetlove LJ, Leaver CJ (2002) A custom microarray analysis of gene expression during programmed cell death in Arabidopsis thaliana. Plant J 30 431–446 [DOI] [PubMed] [Google Scholar]
- Takahashi A, Casais C, Ichimura K, Shirasu K (2003) HSP90 interacts with RAR1 and SGT1 and is essential for RPS2-mediated disease resistance in Arabidopsis. Proc Natl Acad Sci USA 100 11777–11782 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tasseva G, Richard L, Zachowski A (2004) Regulation of phosphatidylcholine biosynthesis under salt stress involves choline kinases in Arabidopsis thaliana. FEBS Lett 566 115–120 [DOI] [PubMed] [Google Scholar]
- Thimm O, Blasing O, Gibon Y, Nagel A, Meyer S, Kruger P, Selbig J, Muller LA, Rhee SY, Stitt M (2004) MAPMAN: a user-driven tool to display genomics data sets onto diagrams of metabolic pathways and other biological processes. Plant J 37 914–939 [DOI] [PubMed] [Google Scholar]
- Vierling E (1991) The role of heat shock proteins in plants. Annu Rev Plant Physiol Plant Mol Biol 42 579–620 [Google Scholar]
- Vogel JT, Zarka DG, Van Buskirk HA, Fowler SG, Thomashow MF (2005) Roles of the CBF2 and ZAT12 transcription factors in configuring the low temperature transcriptome of Arabidopsis. Plant J 41 195–211 [DOI] [PubMed] [Google Scholar]
- Watanabe N, Lam E (2006) Arabidopsis Bax inhibitor-1 functions as an attenuator of biotic and abiotic types of cell death. Plant J 45 884–894 [DOI] [PubMed] [Google Scholar]
- Welchen E, Gonzalez DH (2005) Differential expression of the Arabidopsis cytochrome c genes Cytc-1 and Cytc-2: evidence for the involvement of TCP-domain protein-binding elements in anther- and meristem-specific expression of the Cytc-1 gene. Plant Physiol 139 88–100 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamaguchi-Shinozaki K, Shinozaki K (2006) Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses. Annu Rev Plant Biol 57 781–803 [DOI] [PubMed] [Google Scholar]
- Zimmermann P, Hirsch-Hoffmann M, Hennig L, Gruissem W (2004) GENEVESTIGATOR: Arabidopsis microarray database and analysis toolbox. Plant Physiol 136 2621–2632 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.