Abstract
Drosophila melanogaster S2 cells were co-transfected with plasmid vectors containing the enhanced green fluorescent protein gene (EGFP), under the control of metallothionein promoter (pMt), and the hygromycin selection gene, in view of establishing parameters for optimized gene expression. A protocol of transfection was worked out, leading after hygromycin selection, to ∼90% of S2MtEGFP fluorescent cells at day 5 after copper sulfate (CuSO4) induction. As analyzed by confocal microscopy, S2MtEGFP cell cultures were shown to be quite heterogeneous regarding the intensity and cell localization of fluorescence among the EGFP expressing cells. Spectrofluorimetry kinetic studies of CuSO4 induced S2MtEGFP cells showed the EGFP expression at 510 nm as soon as 5 h after induction, the fluorescence increasing progressively from this time to attain values of 4.6 × 105 counts/s after 72 h of induction. Induction with 700 μM of CuSO4 performed at the exponential phase of the S2MtEGFP culture (106 cells/mL) led to a better performance in terms of cell growth, percent of fluorescent cells and culture intensity of fluorescence. Sodium butyrate (NaBu) treatment of CuSO4 induced S2MtEGFP cell cultures, although leading to a loss of cell culture viability, increased the percent of EGFP expressing cells and sharply enhanced the cell culture fluorescence intensity. The present study established parameters for improving heterologous protein expression in stably transfected Drosophila S2 cells, as assessed by the EGFP expression.
Keywords: Schneider Drosophila cells, EGFP, Cell transfection, Gene expression, Sodium butyrate
Introduction
Drosophila melanogaster Schneider 2 (S2) cells have become increasingly utilized over the past few years for the expression of heterologous proteins. High levels of protein expression with pharmacological and biotechnological interest can be achieved using the Drosophila expression system (DES) procedure whereby a plasmid encoding a gene of interest is co-transfected with a plasmid encoding resistance to hygromycin B. After three weeks of selection this system, using Drosophila promoters, is able to generate a stable polyclonal cell line processing up to 1,000 gene copies per cell. Proteins of prokaryotic, eukaryotic and viral origin have been expressed in this system showing to be appropriately processed and biologically active (Angelichio et al. 1991; Culp et al. 1991; Han 1996; Li et al. 1996; Delm et al. 1999; Nilsen and Castellino 1999; Lee et al. 2000; Hill et al. 2001; Shin et al. 2003; Perret et al. 2003; Chang et al. 2005). Based on these evidences DES has an important potential as a gene expression system.
Nevertheless, in view of higher expression, some bioprocess parameters have to be optimized. In this context, the transfection procedure, aiming the introduction of foreign DNA into cells, is an important step. The calcium phosphate method has been recommended for gene transfection of S2 cells (Van der Straten et al. 1989) but procedures based on liposome reagent have been reported to be more efficient (Han 1996; Park et al. 1999; Shin and Cha 2002)
Besides the metallothionein promoter induction by copper sulfate (CuSO4) (Huang et al. 2004) it has been studied the enhancement of gene expression by agents, such as the sodium butyrate (NaBu), a histone deacetylase inhibitor that lead to chromatin exposure and allow the transcriptional factors binding and increased gene expression (Dorner et al. 1989; Li and Li 2006; Zhao et al. 2006; Ping et al. 2006; Song et al. 2006).
The heterologous gene encoding the enhanced green fluorescent protein (EGFP), a mutant from jellyfish Aquorea Victoria (Ward 2006; Phillips 2006), has been proved to be very helpful for the estimation of gene expression in several systems, becoming very attractive as a reporter gene expression (Shin and Cha 2002; Shin et al. 2003; Ward and Lippincott-Schwartz 2006; Hazelrigg and Mansfield 2006). This protein is a 238-amino acid polypeptide that undergoes an intramolecular cyclization to give a brightly fluorescent molecule. It remains remarkably stable in the presence of many denaturants and proteases, as well as over a broad range of pH and temperature (Ward 2006).
We report here in this study an optimization of EGFP stable transfection and inducible conditions for expression using the S2 cells. A protocol for EGFP stable transfection was established by using a lipotransfection method taking into account the cell density, the ratio of DNA concentration and the amount of cellfectin reagent. Conditions for inducible EGFP expression were established by using the metallothionein promoter and CuSO4 and NaBu as expression inducers. We have evaluated the kinetics cell growth and EGFP expression in a stable transfected, hygromycin selected, polyclonal S2 cell line by confocal microscopy, flow cytometry, and spectrofluorimetry. Our data point to optimized experimental conditions for heterologous protein expression by Drososphila melanogaster S2 cells.
Materials and methods
Cell line, plasmids and enzymes
Drosophila melanogaster Schneider 2 cells (Invitrogen) were grown at 28 °C under normal atmosphere in T-25 flasks in SF900-SFMII medium (Invitrogen). For the EGFP expression, we used the pMT/BiP/V5-His (3.6 KB) expression vector (Invitrogen) which has a metallothionein promoter (pMt) for heavy metals inducible expression. It contains also a sequence of C-terminal peptide containing V5 epitope and a polyhistidine (6× His) tag, besides the elements (pUC ori and ampicillin) for replication in bacteria. For selection of co-transfected cells, we used the pCoHygro (Invitrogen) selection vector containing the hygromycin resistance gene. This gene is under control of the constitutive Drosophila copia 5′LTR promoter. E. coli DH5α was used as a primary host for constructing and propagating plasmids. E. coli were grown and maintained in LB medium containing 50 μg/mL of ampicillin. DNA restriction enzymes and ligases were used according to manufacturer’s instructions (Promega and Fermentas).
Recombinant plasmid construction
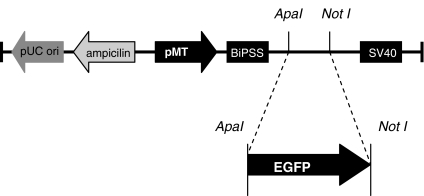
cDNA encoding the EGFP was originally isolated from pGie1EGFP plasmid (kindly provided by Dr. Bergmann Morais from the Brasilia University, Brasília, Brazil). The fragment was isolated from electrophoresis gel of pGeie EGFP digested with Apa I and Not I restrictions enzymes (Fig. 1). The fragment was extracted from the gel and purified using a commercial kit (Promega). Cloning was done according to standard techniques (Sambrook and Russel 2001) and the construct (pMtEGFP) was sequenced to check for in-frame insertion.
Fig. 1.
Schematic construction of pMtEGFP vector
Cell culture and transfection
Drosophila S2 cells were grown at 28 °C in SF900-SFMII medium and co-transfected with pMtEGFP and pCoHygro using cellfectin reagent (Invitrogen) for stable expression. Both plasmids (pMtEGFP and pCoHygro) at a ratio of 20:1 and cellfectin reagent (20 μL) were diluted separately in 500 μL of SF900-SFMII medium and then mixed for 15 min at room temperature, constituting the transfection solution (Table 1).
Table 1.
Optimized protocol for transfection of Drosophila S2 cells
| Parameters | Established transfection protocol |
|---|---|
| S2 cell concentration | 5 × 106 cells/ T-25 flasks |
| Ratio expression/selection vectors | 20:1 |
| Total DNA | 2 μg |
| Cellfectin® reagent | 20 μg |
| SF900 SFMII medium (dilution) | 500 μl |
| Incubation time (complex formation) | 15 min RT |
| Incubation time (transfection) | 5 h |
5 × 106 cells were seeded in T-25 flasks and 24 h after the medium was discarded and 1 mL of transfection solution added. After 5 h of incubation with slowly agitation at room temperature, 4 mL of SF900-SFMII medium was added. The cells were incubated for 2 days and the medium replaced with 600 μg/mL of hygromycin B for selection. Hygromycin B resistant cells were then selected as stable polyclonal populations.
Kinetics experiments of EGFP expression by S2MTEGFP cells were performed upon CuSO4 induction (500 or 700 μM) and/or NaBu (5 mM) treatment of cell cultures performed in 100 mL shake flasks with 20 mL of SF900-SFMII medium incubated at 100 rpm and 28 °C. The cell seeding was the same for all experiments (5 × 105 cells per mL). Cell concentration was determined by viable cells (trypan blue exclusion) counting in hematocytometer.
Confocal microscopy
For confocal EGFP detection, CuSO4 induced S2MTEGFP cells were scanned using the 488 nm laser line and light emitted between 500 and 600 nm was collected. Images were collected on a META LSM 510 laser scanning confocal microscope equipped with a 63.0 × 1.2 W objective (Carl Zeiss Ltda). The same settings for image acquisition and processing have been applied for all samples to allow comparison of the fluorescence intensities among different samples
Flow cytometry
After induction with the indicated amounts of CuSO4, 400 μL of S2MTEGFP cells (106 cells) growing in suspension were analyzed by flow cytometry using a FACS-CALIBUR (Becton Dickinson). EGFP fluorescence was excited at 488 nm and emission was measured with a 530/30 nm band pass filter. S2 cells were used as a control for auto fluorescence and to set the gate for EGFP positive cells. 10.000 events/cells were gated from the 400 μL samples. The geometric mean values were used to compare the intensity of fluorescence in different samples as well as the percentage of fluorescent cells.
Spectrofluorimetry
Cell suspensions were analyzed in spectrofluorimeter PTI for fluorescence measurements with a 4 nm band pass. Cell cultures prepared in shake flasks were induced with 700 μM of CuSO4 and samples (107 cells) were harvested at different times post induction and washed with PBS (1×). The cell suspension (1 mL) was evaluated for fluorescence emission at 490–750 nm when excited with blue light (470 nm).
Statistical analysis
Data were analyzed by the variance F test and statistical differences among values were established by the Tukey test with a significance level of 5%.
Results
EGFP expression vector construction and transfection in S2 cells
We have generated an expression vector encoding the EGFP cDNA by cloning the EGFP fragment into a pMt expression vector in Apa I/Not I cloning sites (Fig. 1). The EGFP gene, under control of the inducible Drosophila metallothionein gene promoter, was inserted in a Drosophila expression vector. This vector was co-transfected with a Drosophila selection vector (carrying hygromicin B resistance gene) into Drosophila cells and a population was selected for their resistance to hygromicin B.
For the co-transfection of pMtEGFP and pCOHygro into S2 cells, we have developed a transfection protocol (Table 1) which allowed us to select a stable polyclonal cell population (S2MtEGFP) with hygromycin B after 2 weeks of culture.
EGFP expression upon CuSO4 induction
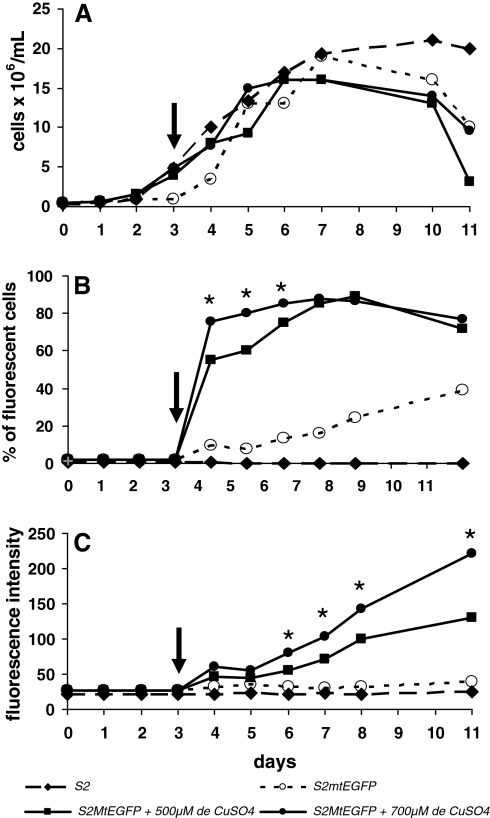
Figure 2 shows the EGFP expression and cell growth of S2MtEGFP cells induced with 500 or 700 μM of CuSO4 at day 3 of culture. We observed that after 2 days of induction, 700 μM of CuSO4 were able to induce the EGFP expression in 80% of the cells, in contrast to 500 μM of CuSO4 that induced the EGFP expression in only 60% of cells. Nevertheless, after 5 days of induction both CuSO4 concentrations led to EGFP expression in about 85% of cells. The fluorescence intensity also increased with the CuSO4 concentration and at day 5 of induction we had an intensity of fluorescence of 100 units for cultures induced with 500 μM of CuSO4 and 142 units for those induced with 700 μM of CuSO4. The cell growth, at the exponential phase, was shown to be comparable for control or CuSO4 induced cultures, indicating that the CuSO4, at these concentrations, had no influence on the cell growth (1.5 to 2 × 107 cells/mL). When the CuSO4 induction was performed together with cell seeding at the beginning of the culture we have observed a significant decrease in the cell multiplication as well as lower expression levels of EGFP as evaluated by the number of fluorescent cell or the intensity of fluorescence of the cell population (data not shown).
Fig. 2.
Kinetics of cell growth (A), % of fluorescent cells (B) and fluorescence intensity (C) of S2 and S2MtEGFP cells induced with 500 or 700 μM of CuSO4 at day 3 of culture. •, S2MtEGFP cells + 700 μM of CuSO4; ■, S2MtEGFP cells + 500 μM of CuSO4; ♦, S2 cells; ○, S2MtEGFP cells. Arrows indicate the time of induction at day 3. Results are the average of at least three independent experiments. *: statistically different
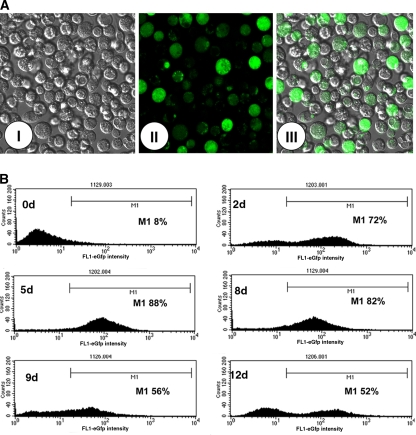
Confocal analysis of CuSO4 induced S2MtEGFP cells showed an heterogeneous EGFP expression among the cell population (Fig. 3A). Kinetics of EGFP expression as assessed by flow cytometry analysis of S2MtEGFP cells (Fig. 3B) showed in a non-induced S2MtEGFP cell population a low percent of fluorescent cells (8%) which sharply increased upon CuSO4 induction attaining high levels (88%) at day 5 of induction. By spectrofluorimetry we could better examine the time response of S2MtEGFP cells upon CuSO4 induction, by analyzing the fluorescence intensity expressed by the cell population in early times after induction. Figure 4 shows that specific fluorescence at 510 mm could be already detected as early as 5 h after induction, showed significant levels at 24 h and increased progressively to attain high levels at 72 h.
Fig. 3.
S2MtEGFP cell culture as evaluated by confocal microscopy (A) and flow cytometry (B). S2MtEGFP cultures were induced with 700 μM of CuSO4. At day 5 of induction they were observed in a confocal microscopy and at different times after induction they were evaluated by flow cytometry. For confocal microscopy: (I), optical microscopy; (II), fluorescence microscopy; (III) merged image. For flow cytometry they were observed at days 0, 2, 5, 8, 9 and 12 of induction and the % of fluorescent cells is indicated as M1. Data are representative of independent experiments
Fig. 4.
Kinetics of EGFP intensity expression by S2MtEGFP cells as evaluated by spectrofluorimetry. S2MtEGFP cell culture was induced with 700 μM of CuSO4 and at different times the EGFP fluorescence emission was detected at 510 nm (A) and the kinetics curve was recorded (B). Data are representative of independent experiments
Improving heterologous protein expression by using sodium butyrate (NaBu).
The effect of NaBu on the EGFP expression by CuSO4 induced S2MtEGFP cells was analyzed. Figure 5 shows that the NaBu (5 mM) treatment led to a decreased cell growth. At day 4 of induction, cell cultures treated with NaBu attained 0.8 × 107 cells/mL in contrast to control cells that attained 1.6 × 107 cells/mL (Fig. 5A). Following NaBu treatment an increased number of cells expressing the EGFP was observed at 24 h of induction (90%) in comparison to control cell cultures (75.6%) (Fig. 5B). Moreover, the intensity of fluorescence showed by the treated culture sharply increased from 54 units in control cultures to 257 units in the treated ones (Fig. 5C), indicating a remarkable effect of NaBu in promoting the EGFP expression in induced cells. When a higher NaBu concentration (7 mM) was used, the enhancement of EGFP expression was of lower magnitude, possibly due to a higher detrimental effect on the cell metabolism (data not shown).
Fig. 5.
Kinetics of cell growth (A), % of fluorescent cells (B) and fluorescence intensity (C) of S2MtEGFP cells induced with 700 μM of CuSO4 and treated with 5 mM of NaBu. At day 3, cell cultures were induced with CuSO4 and/or treated with NaBu. Cell samples were periodically taken for evaluation of viable cell concentration by hematocytometer or fluorescence by flow cytometry. ○, S2MtEGFP cells; •, S2MtEGFP cells + 700 μM of CuSO4; Δ, S2MtEGFP cells + 5 mM NaBu; ▴, S2MtEGFP cells + 700 μM of CuSO4 + 5 mM NaBu. Arrows indicate the time of induction at day 3. Results are the average of at least three independent experiments. *: statistically different
Discussion
The control and improvement of transfected gene expression and recombinant protein synthesis is a central point of study for several purposes in biology, engineering and medicine when one consider fundamental research or immunobiological production.
In the present study we have investigated the influence of various parameters for the optimization of high gene expression in stably transfected Drosophila melanogaster S2 cells. The enhanced green fluorescent protein (EGFP) reporter gene, under the control of the inducible metallothionein (Mt) promoter and co-transfected with the hygromycin selection gene was employed in the study. Protocols for vector transfection and gene induction in cell cultures were established.
Several parameters have to be considered for improving gene expression optimization. First of all, together with the gene vector construction, an efficient and reliable transfection procedure is required to provide the recombinant cell population with a high percent of properly transfected cells. For this purpose standardizations in transfection or co-transfection protocols must take into account parameters such as the ratio of vectors (expression and selection), the cell concentration, the metabolic state of the cell population, the concentration of transfection and selection reagents and time and temperature of incubation.
In the present study we have established an optimized protocol for gene co-transfection (Table 1) of S2 cells using 20 μg of cellfectin: 5 × 106 cells seeded and cultured for 24 h in T-25 flasks (5 mL), ration of expression: selection vectors of 20:1 (2 μg DNA) and a transfection incubation period of 5 h at 28 °C in a volume of 500 μL. Hygromycin selection was performed with 600 μg/mL of hygromycin for 2 weeks. The above described transfection protocol led to an enriched S2MtEGFP polyclonal cell population (88% fluorescent cells).
Following the transfection step, it was of importance to establish parameters of optimized gene expression based on the promoter induction. Experiments were carried out to optimize the concentration of CuSO4 to induce the Mt promoter. Kinetic parameters of cell growth (cell/mL), cell expressing the gene (% of fluorescent cells) and the degree of gene expression intensity (fluorescence intensity) were analyzed by confocal microscopy, flow cytometry and spectrofluorimetry, in suspension cell cultures, mimicking large scale conditions. S2MtEGFP cells were shown to be not influenced, at their exponential growth phase, by the CuSO4 induction, even though some detrimental effects could be observed after the stationary phase, leading to a higher loss of viability when compared to non-induced cultures. S2MtEGFP cells could, in these conditions, attain a cell concentration of 1.5 to 2 × 107 cells/mL and expressed EGFP as soon as 5 h after CuSO4 induction. 700 μM of CuSO4 were able to induce EGFP expression in about 85% of cells with heterogeneous fluorescence intensity, which reached an average of 142 units, after 5 days of induction (Fig. 2–4). Nevertheless, the overall data of cell growth and EGFP expression were substantially affected when CuSO4 induction was performed at the beginning of cell culture together with cell seeding, indicating a detrimental effect of CuSO4 on the cell metabolism during the lag phase (data not shown) and the importance of CuSO4 induction of the EGFP gene during the exponential cell growth phase.
We found a high heterogeneity of EGFP expression among the cells in the population. This could be due to cells harboring plasmids with rearrangements, since plasmid DNA transfection into a cell culture leads to a high frequency of genetic instability (Calos et al. 1983; Razzaque et al. 1983). Besides, since the expression of an heterologous transfected gene, in the cell context, depends on its properly insertion into the cell genome, it is conceivable that a variable number of gene copies, upon cell transfection and gene integration, became inserted in not transcribed regions of the genome. So, even efficient procedures of gene transfection may lead to gene integrated copies that are not expressed.
Based on published studies (Dorner et al. 1989; Chen et al. 2002; Li and Li 2006; Zhao et al. 2006; Ping et al. 2006; Song et al. 2006), sodium butyrate (NaBu), is a histone deacetylase inhibitor, affecting the chromatin structure and leading to the destabilization of nucleosomal structure. As a consequence, the NaBU influence on chromatin exposure may facilitate binding of transcription factors to DNA resulting in an increased and deregulated cell protein synthesis.
We have investigated the enhancement of EGFP expression by S2MtEGFP cell treatment with NaBu. As expected, the cell treatment with NaBu led to loss of cell multiplication and viability, since the NaBu is a potent inducer of G1 cell cycle arrest and provokes apoptosis by activating an intrinsic apoptotic pathway (Li and Li 2006). On the other hand NaBu treatment may expose cell genome regions bearing an heterologous transfected gene and so foster the synthesis of their products (Dorner et al. 1989; Ping et al. 2006; Song et al. 2006). Indeed, our data clearly show that S2MtEGFP cell treatment with NaBu led to a decreased cell multiplication (from 1.6 × 107 in controls with only addition of CuSO4, to 0.8 × 107 cells/mL) and decreased viability, which was accompanied by not only a clear cut enhancement of fluorescence intensity of the cell population (∼5 fold) but also an early increase in the percent of fluorescent cells among the induced cell population (from 75.6% in controls to 90%) at one day post-induction (Fig. 5).
Drosophila S2 cells is a promising cell system for heterologous gene expression due to its characteristics of easy handle cell culture procedures associated to the possibilities of stable and induced gene expression. EGFP, as a reporter gene, is a useful tool to establish suitable parameters for improving heterologous protein expression. We show here in this study an optimized vector transfection protocol that enabled us to establish a primary stable transfected S2MtEGFP cell population that once induced by CuSO4, expressed the recombinant EGFP in a high percentage of the cells. Its expression, by the CuSO4 induced cell population, could be further sharply enhanced by sodium butyrate (NaBu). The procedure shown here was developed in view of a general application for gene expression in the stable transfected Drosophila S2 cell system.
Acknowledgements
This work was supported in part by grants from the FAPESP (02/09482-3, 05/50565-8), CNPq and Fundação Butantan. We thank Dr. Bergmann Morais (UnB, Brasília, Brazil) for pGie1EGFP plasmid supplying and Dr. Jorge MC Ferreira Jr and Dr. Orlando Ribeiro, for flow cytometry analysis. C.A. Pereira is recipient of CNPq research fellowship. S.A.C. Jorge and M.A.G. Santos had scholarships from FAPESP (01/08914-4, 02/04003-0, 03/08978-8).
Abbreviations
- EGFP
Enhanced green fluorescent protein gene
- pMt
Metallothionein promoter
- S2
Schneider 2 Drosophila melanogaster cells
- S2MtEGFP
S2 cells transfected with Mt inducible plasmid containing the EGFP gene
- NaBu
Sodium butyrate
- CuSO4
Copper sulfate
- DES
Drosophila expression system
- Hygro
Hygromycin
References
- Angelichio ML, Beck JA, Johansen HE, Ivey-Hole M (1991) Comparison of several promoters and polyadenylation signals for use in heterologous gene expression in cultured Drosophila cells. Nucleic Acids Res 19:5037–5043 [DOI] [PMC free article] [PubMed]
- Calos MP, Lebkowski JS, Botchan MR (1983) High mutation frequency in DNA transfected into mammalian cells. Proc Natl Acad Sci USA 80:3015–3019 [DOI] [PMC free article] [PubMed]
- Chang KH, Yang JM, Chun HOK, Chung IS (2005) Enhance activity of recombinant β-secretase from Drosophila melanogaster S2 cells transformed with cDNAs encoding human β1,4-galactosyltransferase and Gal β1,4-GlcNac α2,6-sialyltransferase. J Biotechnol 116:359–367 [DOI] [PubMed]
- Chen T, Sun H, Lu J, Zhao Y, Tao D, Li X, Huang B (2002) Histone acetylation is involved in hsp70 gene transcription regulation in Drosophila melanogaster. Archi. Biochem Biophys 408:171–176 [DOI] [PubMed]
- Culp JS, Johansen H, Hellmig B, Beck J, Matthews TJ, Delers A, Rosemberg M (1991) Regulated expression allows high level production and secretion of HIV-1 gp 120 envelope glycoprotein in Drosophila Schneider cells. Bio/technology 9:173–178 [DOI] [PubMed]
- Delm L, Wolf H, Wagner R (1999) High level expression of hepatitis B virus surface antigen in stably transfected in Drosophila Schneider-2 cells. J Virol Methods 79:191–203 [DOI] [PubMed]
- Dorner AJ, Wasley LC, Kaufman RJ (1989) Increased synthesis of secreted proteins induces expression of glucose-regulated proteins in butyrate-treated Chinese hamster ovary cells. J Biol Chem 264:20602–20607 [PubMed]
- Han K (1996) An efficient DDAB-mediated transfection of Drosophila S2 cells. Nucl Acids Res 24:4362–4363 [DOI] [PMC free article] [PubMed]
- Hazelrigg T, Mansfield JH (2006) Green fluorescent protein applications in Drosophila. Methods Biochem Anal 47:227–257 [PubMed]
- Hill RM, Brennan SO, Birch NP (2001) Expression, purification and functional characterization of the serine protease inhibitor neuroserpin expressed in Drosophila S2 cells. Protein Expr Purif 22:406–413 [DOI] [PubMed]
- Huang EP, Marquis CP, Gray PP (2004) Process development for recombinant chinese hamster ovary (CHO) cell line utilizing a metal induced and amplified metallotionein expression system. Biotechnol Bioeng 88:438–450 [DOI] [PubMed]
- Lee JM, Park JH, Park JO, Chang KH, Chung IS (2000) Expression of recombinant erythropoietin in stably transformed Drosophila melanogaster S2 cells. In Vitro Cell Dev Biol Anim 36:348–350 [DOI] [PubMed]
- Li B, Tising S, Kosaka AH, Nguyen B, Osen EG, Bach C, Chan H, Barnett J (1996) Expression of Human Dopamine β Hydroxylase in Drosophila Schneider 2 Cells. Biochem J 313:57–64 [DOI] [PMC free article] [PubMed]
- Li RW, Li CJ (2006) Butyrate induces profound changes in gene expression related to multiple signal pathways in bovine kidney epithelial cells. BMC genomics 7:234 [DOI] [PMC free article] [PubMed]
- Nilsen SL Castellino FJ (1999) Expression of human plasminogen in Drosophila Schneider S2 cells. Protein Expr Purif 16:136–143 [DOI] [PubMed]
- Park HJ, Kim HY, Han KH, Chung SI (1999) Optimization of transfection conditions for expression of green fluorescent protein in Drosophila melanogaster S2 cells. Enzyme Microb Technol 25:558–563 [DOI]
- Perret BG, Wagner R, Lecat S, Brillet K, Rabut G, Bucher B, Pattus F (2003) Expression of EGFP-amino-tagged human mu opioid receptor in Drosophila Schneider 2 cells: a potential expression system for large-scale production of G-protein coupled receptors. Protein Expr Purif 31:123–132 [DOI] [PubMed]
- Phillips GN Jr (2006) The three-dimensional structure of green fluorescent protein and its implications for function and design. Methods Biochem Anal 47:67–82 [PubMed]
- Ping W, Ge J, Li S, Zhou H, Wang K, Feng Y, Lou Z (2006) Baculovirus-mediated gene expression in chicken primary cells. Avian Dis 50:59–63 [DOI] [PubMed]
- Razzaque A, Mizusawa H, Seidman MM (1983) Rearrangement and mutagenesis of a shuttle vector plasmid after passage in mammalian cells. Proc Natl Acad Sci USA 80:3010–3014 [DOI] [PMC free article] [PubMed]
- Sambrook J. and Russel D.W (2001) Molecular cloning: a laboratory manual. New York Cold Spring Harbor Laboratory Press
- Shin HS, Cha HJ (2002) Facile and statistical optimization of transfection conditions for secretion of foreign proteins from insect Drosophila S2 cells using green fluorescent protein reporter. Biotechnol Prog 18:1187–1194 [DOI] [PubMed]
- Shin HS, Lim HJ, Cha HJ (2003) Quantitative monitoring for secreted production of human interleukin-2 in stable insect Drosophila S2 cells using a green fluorescent protein fusion partner. Biotechnol Prog 19:152–157 [DOI] [PubMed]
- Song J, Liang C, Chen X (2006) Transduction of avian cells with recombinant baculovirus. J Virol Methods 135:157–162 [DOI] [PubMed]
- Van der Straten A, Johansen H, Rosenberg M, Sweet R (1989) Introduction and constitutive expression of gene products in cultured Drosophila cells using hygromicin B selection. Curr Methods Mol Cell Biol 1:1–8
- Ward WW (2006) Biochemical and physical properties of green fluorescent protein. Methods Biochem Anal 47:39–65 [PubMed]
- Ward TH, Lippincott-Schwartz J (2006) The use of green fluorescent protein in mammalian cells. Methods Biochem Anal 47:305–337 [DOI] [PubMed]
- Zhao YM, Chen X, Sun H, Yuan ZG, Ren GL, Li XX, Lu J, Huang BQ (2006) Effects of histone deacetylase inhibitors on transcriptional regulation of the hsp70 gene in Drosophila. Cell Res 16:566–576 [DOI] [PubMed]