Abstract
Pigs are susceptible to both human and avian influenza viruses and have been proposed to be intermediate hosts, or mixing vessels, for the generation of pandemic influenza viruses through reassortment or adaptation to the mammalian host. In this study, we summarize and report for the first time the coexistence of wholly human-like H3N2 viruses, double-reassortant H3N2 viruses, and triple-reassortant H3N2 viruses in pigs in China by analyzing the eight genes of swine influenza A (H3N2) viruses found in China from 1970 to 2006. In 1970, the first wholly human-like H3N2 (Hong Kong/68-like) viruses were isolated from pigs in Taiwan, and then in the next years Victoria/75-like, Sydney/97-like, New York/99-like, and Moscow/99-like swine H3N2 viruses were regularly isolated in China. In the 1980s, two triple-reassortant viruses were isolated from pigs. Recently, the double-reassortant viruses containing genes from the human (HA and NA) and avian (PB2, PB1, PA, NP, M, and NS) lineages and the triple-reassortant viruses containing genes from the human (HA and NA), classical swine (NP), and avian (PB2, PB1, PA, M, and NS) lineages emerged in pigs in China. The coexistence of wholly human-like and reassortant viruses provides further evidence that pigs serve as intermediate hosts, or mixing vessels, and emphasizes the importance of reinforcing swine influenza virus surveillance in China.
Influenza A viruses have been isolated from a large variety of animal species, including humans, pigs, horses, sea mammals, and birds. Aquatic birds are known to serve as the source of all influenza viruses for other species (1). Despite their common origin, influenza A viruses generally are restricted in host range. The host range restriction is a polygenic trait, and the receptor specificity of hemagglutinin (HA) is considered an important determinant (7, 13, 31). The tracheal epithelium in pigs expresses receptors for both human and avian influenza viruses, and this provides a biological basis for the susceptibility of pigs to both avian and human influenza viruses (8, 19). Pigs, therefore, can function as intermediate hosts, or mixing vessels, in establishing new influenza virus lineages by supporting coinfection, replication, and reassortment among human, avian, and swine influenza viruses (1, 13).
Currently, three main subtypes of influenza viruses are circulating in the swine population worldwide: H1N1, H3N2, and H1N2 (1, 15). As one of the main subtypes, swine H3N2 viruses have been reported widely throughout the world. Around 1970, following the human Hong Kong influenza pandemic, the wholly human-like H3N2 virus was transmitted to pigs (28). This human-like swine influenza virus continued to circulate in pigs, particularly in Europe and Asia, but only sporadically caused clinical signs. Since 1984, a new H3N2 influenza virus has started causing clinical disease and replaced the original H3N2 virus in Europe, probably as a result of a reassortment with the avian-like swine H1N1 virus (2). During 1998, severe outbreaks of influenza were observed in the swine herds in the United States, and two antigenically distinct reassortant viruses (H3N2) were isolated: a double-reassortant virus containing genes similar to those of human and swine viruses and a triple-reassortant virus containing genes similar to those of human, swine, and avian influenza viruses (32, 36).
China, especially southern China, is regarded as an epicenter of pandemic influenza viruses throughout history (26). In the past, a number of H3N2 swine influenza viruses were isolated from pigs in China that were similar to human A/Hong Kong/1/68 (A/HK/1/68), A/Victoria/3/75, and A/Sydney/5/97 viruses (25, 27, 28). To gain further epidemiological information and provide necessary data for swine influenza control (and possibly also some useful information for the prediction of and preparedness for future human influenza pandemics), we made full use of sequences available in GenBank from 1970 to 2004 and sequences of our H3N2 swine viruses isolated in China from 2005 to 2006, and we analyzed their genetic evolution. In this study, for the first time, we summarize and report the coexistence of wholly human-like H3N2 viruses, double-reassortant H3N2 viruses, and triple-reassortant H3N2 viruses in pigs in China.
MATERIALS AND METHODS
Viruses.
A total of 500 samples, including nasal swabs, lungs, and trachea, were collected from eight provinces (Heilongjiang, Henan, Shandong, Guangdong, Zhejiang, Anhui, Jiangxi, and Beijing) in China from 2005 to 2006. Four influenza A viruses were isolated by using 10-day-old specific-pathogen-free chicken eggs and were identified as subtype H3N2 (35). Viral gene sequencing was carried out as follows. In brief, viral RNA was directly extracted from infected allantoic fluids using RNeasy mini kits (Qiagen, Chatsworth, CA), and reverse transcriptions were carried out under standard conditions using the Uni12 (AGCAAAAGCAGG) primer. PCR was performed using specific primers for eight genes (primer sequences are available on request). PCR products were purified with the QIAquick PCR purification kit (Qiagen, Inc.), cloned into the pMD18-T vector (TaKaRa; Dalian), and then sequenced using synthetic oligonucleotides (Invitrogen).
Geographical distribution of the Chinese isolates examined.
Besides the four isolated viruses described above, we also utilized the sequences of eight gene segments of 40 H3N2 swine influenza viruses that were isolated in China and deposited in GenBank (Table 1). Among all of the 44 swine influenza viruses, the Hong Kong isolates were mainly from provinces in southern China, including Guangdong, Hunan, Jiangxi, Hubei, Henan, Fujian, Zhejiang, Guangxi, and Hong Kong (19), and the rest of the isolates were from Guangdong, Taiwan, Fujian, Inner Mongolia, and Heilongjiang provinces.
TABLE 1.
GenBank accession numbers and origins of gene segments of swine H3N2 influenza viruses isolated in China from 1970 to 2006a
Partial sequences of each gene of our isolates, together with sequences from GenBank, were analyzed by the distance-based neighbor-joining method using phylogenetic analysis software (MEGA 3.1). Nucleotide sequences of HA (positions 98 to 1059), NA (positions 746 to 1386), PB2 (positions 1037 to 2219), PB1 (positions 950 to 2177), PA (positions 1434 to 1580), NP (positions 747 to 1504), M (positions 149 to 888), and NS (positions 27 to 864) genes were examined in this study. GenBank accession numbers in boldface indicate that the gene segments are of avian origin; underlined accession numbers denote that the gene segments are of swine origin; the rest of the genes are of human origin.
The origin of these genes was reported previously (29).
Sequence analysis indicates that this gene is equally closely related to the HA gene of human and duck H3 viruses and may come from either origin.
A dash indicates that the sequence was not available from GenBank.
Sequence analysis.
Sequence data were compiled and edited using the Lasergene sequence analysis software package (Dnastar Inc., Madison, WI). Multiple-sequence alignment was carried out by using CLUSTAL W, and the unrooted phylogenetic trees were generated by the distance-based neighbor-joining method using MEGA 3.1. Bootstrap values were calculated based on 1,000 replicates of the alignment.
RESULTS
Phylogenetic analysis of swine H3N2 influenza viruses isolated in China from 1970 to 2006.
To characterize more precisely the genetic relationships of the swine H3N2 influenza viruses isolated in China from 1970 to 2006, we used the nucleotide sequences of eight segments of the 44 isolates to construct phylogenetic trees.
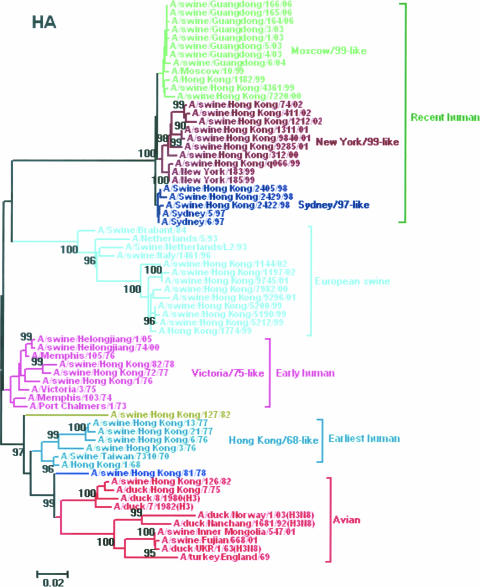
A phylogenetic analysis of the HA gene of the H3N2 influenza viruses showed that the swine H3N2 HA genes could be segregated into five lineages, including avian strains, earliest human strains, early human strains, European swine strains, and recent human strains (Fig. 1). The avian lineage included three swine viruses (A/swine/Hong Kong/126/82, A/swine/Inner Mongolia/547/01, and A/swine/Fujian/668/01). The swine viruses of the earliest human lineage seemed to be derived from the human strain A/Hong Kong/1/68. The early-human-derived swine viruses were closely related to A/Victoria/3/75-like viruses, including one of our isolates (A/swine/Heilongjiang/1/05). The swine viruses of European swine lineage probably were derived from European H3N2 swine influenza viruses for which the HA genes originated from A/Port Chalmers/1/73. The recent human lineage included three sublineages (Sydney/97-like viruses, New York/99-like viruses, and Moscow/99-like viruses), and our three isolates from Guangdong province belonged to the Moscow/99-like viruses sublineage. In addition, A/swine/Hong Kong/81/78 was equally closely related to the avian and human H3 strains and so was an intermediate swine virus between the avian lineage and the earliest human lineage; A/swine/Hong Kong/127/82 also was an intermediate swine virus between the earliest human lineage and the early human lineage.
FIG. 1.
Phylogenetic tree of the HA (positions 98 to 1059) gene of the H3N2 swine influenza viruses in China from 1970 to 2006. The unrooted phylogenetic tree was generated by the distance-based neighbor-joining method using MEGA 3.1. The reliability of the tree was assessed by bootstrap analysis with 1,000 replications; only bootstrap values of >90% are shown. Different lineages and sublineages are indicated by different colors.
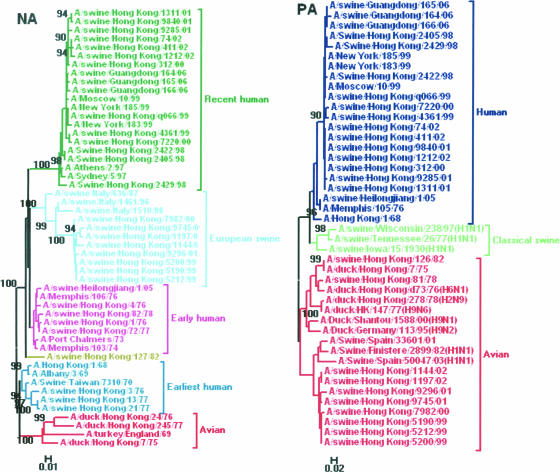
A phylogenetic analysis of the NA gene showed that all of the NA genes from the swine H3N2 viruses tested could be separated into four lineages, including earliest human strains, early human strains, European swine strains, and recent human strains (Fig. 2); this list of lineages is slightly different from that for the HA gene because of the lack of sequence data for the avian lineage in GenBank.
FIG. 2.
Phylogenetic trees of the NA (positions 746 to 1386) and PA (positions 1434 to 1580) genes of the H3N2 swine influenza viruses in China from 1970 to 2006. The method used is described in the legend to Fig. 1. Different lineages are indicated by different colors.
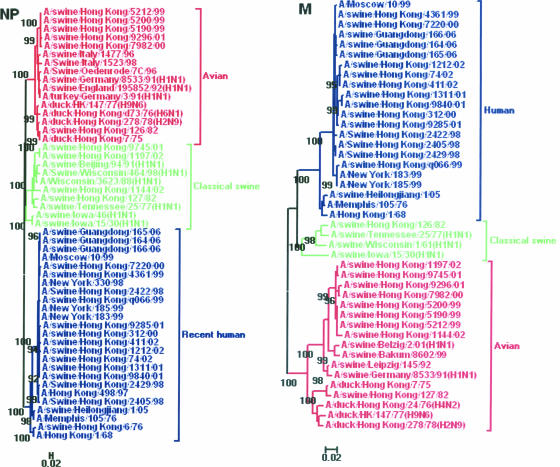
Phylogenetic analyses of PA, NP, and M genes revealed a clear division of swine H3N2 viruses into different lineages (Fig. 2 and 3). In the PA tree, the swine viruses were divided into an avian lineage and a human lineage. The swine viruses, such as A/swine/Hong Kong/126/82, A/swine/Hong Kong/81/78, and A/swine/Hong Kong/1144/02, belonged to the avian lineage, while the remaining swine viruses clustered together as part of the human lineage. In the NP tree, the swine viruses seemed to form three lineages, including avian-like, classical swine, and human-like lineages. The four swine H3N2 viruses (A/Hong Kong/127/82, A/Hong Kong/9745/01, A/swine/Hong Kong/1197/02, and A/swine/Hong Kong/1144/02) belonged to the classical swine lineage, while the remaining swine viruses were incorporated into the avian-like and human-like lineages. In the M tree, just as for the NP gene, the swine viruses were separated into three lineages. Most of the H3N2 swine viruses belonged to avian-like and human-like lineages, while A/swine/Hong Kong/126/82 was located in the classical swine lineage.
FIG. 3.
Phylogenetic trees of the NP (positions 747 to 1504) and M (positions 149 to 888) genes of the H3N2 swine influenza viruses in China from 1970 to 2006. The method used is described in the legend to Fig. 1. Different lineages are indicated by different colors.
Phylogenetic analyses of PB2, PB1, and NS genes showed that PB2 and PB1 genes of these swine influenza virus could be segregated into two lineages and the NS gene could be separated into three lineages, which were similar to those for the PA and M genes (data not shown).
Taken together, the phylogenetic analyses of all eight gene segments of H3N2 influenza viruses isolated from pigs in China from 1970 to 2006 revealed the presence of wholly human-like swine H3N2 viruses, double-reassortant swine H3N2 influenza viruses, and even triple-reassortant swine H3N2 influenza viruses in China (Table 1). In 1970, the wholly human-like H3N2 (Hong Kong/68-like) viruses were first isolated from pigs in Taiwan, and then in the next years Victoria/75-like, Sydney/97-like, New York/99-like, and Moscow/99-like swine H3N2 viruses were regularly isolated. In the 1980s, two triple-reassortant viruses were isolated from pigs in China. A/swine/Hong Kong/126/82 contained gene segments similar to those of the avian (HA, PB1, PA, and NP), human (NA), and classical swine (PB2, M, and NS) lineages, whereas A/swine/Hong Kong/127/82 contained genes from the human (HA and NA), classical swine (PB2, PB1, PA, NP, and NS), and avian (M) lineages. Recently, the double-reassortant viruses containing genes from the human (HA and NA) and avian (PB2, PB1, PA, NP, M, and NS) lineages and the triple-reassortant viruses containing genes from the human (HA and NA), classical swine (NP), and avian (PB2, PB1, PA, M, and NS) lineages emerged in pigs in China. Each gene segment of recent double-reassortant viruses was closely related to European swine viruses, indicating that these viruses might come from European pigs. The recent triple-reassortant viruses might derive from recent double-reassortant viruses, with the acquisition of the NP gene through reassortment with the classical swine H1N1 viruses.
Molecular analysis of the HA1 gene of the swine H3N2 influenza viruses from 1970 to 2006.
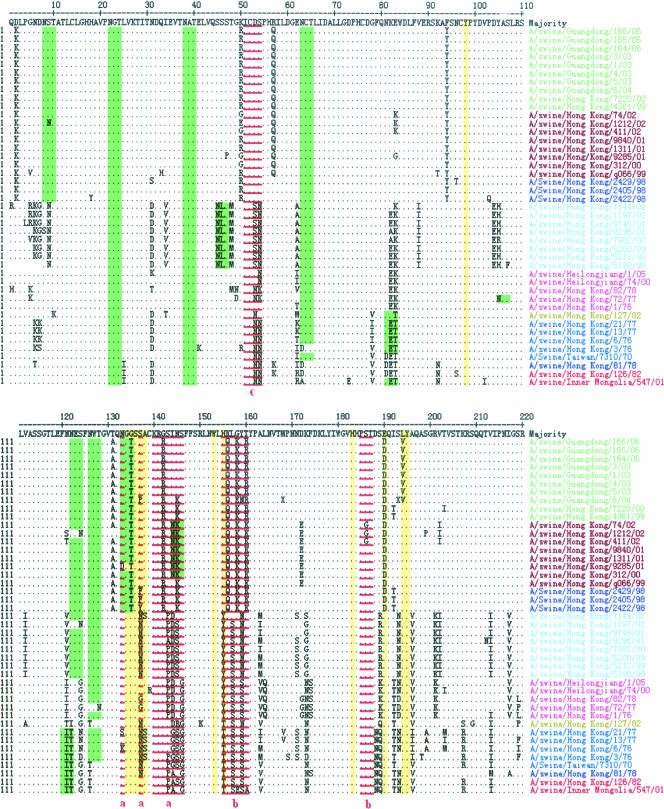
To investigate the differences among the swine H3N2 influenza viruses of the five lineages (avian, earliest human, early human, European swine, and recent human), the amino acid sequences of the HA1 region were aligned. The proposed antigenic sites (3, 14, 34), receptor-binding sites (18), and potential glycosylation sites were analyzed (Fig. 4).
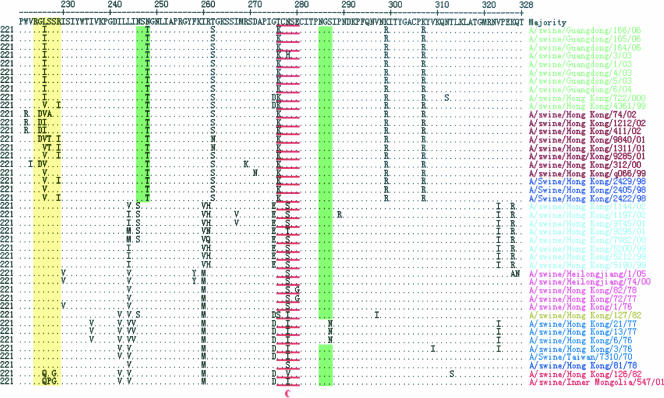
FIG. 4.
Alignment of HA1 amino acid sequences of the swine H3N2 influenza viruses from 1970 to 2006. The underlined residues are antigenic sites (lowercase letters indicate discrete antigenic sites), residues in green are potential glycosylation sites, and residues in yellow shade denote the receptor-binding sites. Swine influenza viruses of different lineages are indicated by different colors, as shown in Fig. 1.
Antigenic sites are regions of molecules involved in antibody binding (34). The majority of difference among these swine viruses occurred in regions other than the proposed antigenic sites. The percentage of mutations in these sites ranged from 21.7 to 36.8%.
The receptor-binding property of the HA protein of influenza virus is an important molecular determinant of host range restrictions (16, 33). All of the swine viruses had the same amino acids at Y98, G134, S136, W153, H183, Y195, and R224 (receptor-binding sites), and the viruses of the earliest human and European swine lineages had the identical amino acid at N137. Three obvious mutations (T135→G135, H155→Y155, and D190→E190) occurred between the recent human lineage and other lineages; these mutations were unique to the recent human lineage. The amino acid sequence of the earliest human, early human, and European swine lineages at residues 226 to 228, a site known to affect receptor-binding properties (21), was L-S-S, while the recent human lineage was I-S-S or V-S-S and the avian lineage was Q-S-G or Q-P-G.
Some glycosylation sites have a significant effect on the antigenic and receptor-binding properties of the influenza virus HA protein, and glycosylation therefore is an important process in the generation of new virus (24). Two potential glycosylation sites (N-X-S/T) were conserved at positions 28 and 38 in the HA1 domains of the swine H3N2 viruses. The majority of these viruses also had conserved glycosylation sites at positions 63, 126, and 285. Three sites at positions 8, 138 (except for A/swine/Hong Kong/9285/01), and 246 and one site at position 45 were unique to the recent human lineage and European swine lineage, respectively. In addition, as shown in Fig. 4, swine viruses of the human (earliest human, early human, European swine, and recent human) lineages had more glycosylation sites than those of the avian lineage.
DISCUSSION
The disease caused by influenza viruses in pigs is similar to that of humans, with low mortality but high morbidity. However, influenza has serious consequences in pigs due to the increased time needed to attain slaughter weight (5). The primary clinical manifestations of influenza virus infection in pigs are fever and acute respiratory distress (nasal discharge, coughing, and dyspnea). Although swine influenza is widespread and is endemic throughout the world, isolating swine influenza viruses is relatively difficult and is dependent on the time of sampling. Sample collection needs to be based on the differentiation of acutely infected pigs producing large amounts of viruses from subacute pigs producing little or no virus. The influenza virus titer peaks at 48 h postinfection, and there is little virus shed 6 to 8 days after infection (12). A total of 500 samples, including nasal swabs, lungs, and trachea, were collected from pigs from eight provinces in China from 2005 to 2006. Four H3N2 swine influenza viruses and one H1N1 swine influenza virus were isolated (35). The reason why only five swine influenza viruses were isolated might be that samples were not collected at the optimal time.
Influenza virus genomes are well known to undergo antigenic drift or antigenic shift that enables them to escape from preexisting immunity and cause new outbreaks of influenza in animals and even humans (4, 20, 30). As the main surface glycoproteins, HA and NA are the targets of the protective immune response and can vary as a result of antigenic drift and antigenic shift (29). Phylogenetic analyses of the HA and NA genes of the H3N2 influenza viruses showed that the HA and NA genes could be segregated into five or four lineages, respectively. This grouping differed somewhat from that reported by Nerome et al. (17) and Zhou et al. (36) because of the emergence of new H3N2 swine influenza viruses. In addition, an amino acid sequence analysis of the HA1 gene revealed multiple changes in the surface proteins of swine influenza viruses. Most of the differences between swine viruses occurred in regions other than the proposed antigenic sites. Swine viruses of the recent human lineage had four unique receptor-binding sites (T135, H155, D190, and I226 or V226). Swine viruses of the human (earliest human, early human, European swine, and recent human) lineages had more glycosylation sites those of avian lineage. These results enhanced our understanding of the genetic evolution of swine influenza A (H3N2) viruses in China from 1970 to 2006.
Infections of pigs with human H3N2 influenza viruses can occur under natural conditions (1), and the wholly human-like H3N2 viruses were isolated frequently from pigs in China. In 1970, the first wholly human-like H3N2 (Hong Kong/68-like) viruses were isolated from pigs in Taiwan, and then in the next years Victoria/75-like, Sydney/97-like, New York/99-like, and Moscow/99-like swine H3N2 viruses were regularly isolated in China. Because of the short average life span of pigs, the human influenza viruses that transferred to pigs appear to be genetically more stable than their counterparts in humans (6). Once the human H3N2 viruses were introduced into pigs, these viruses appeared to have been under less immune selection pressure, and all of the genes evolved more slowly in pigs than in humans (29). Pigs, therefore, have been shown to serve as reservoirs for the older human H3N2 influenza viruses (9, 10). This might be the reason why our four swine H3N2 viruses isolated from 2005 to 2006 were closely related to Moscow/99-like or Victoria/75-like human H3N2 viruses in all gene segments and had not undergone genetic reassortment.
Pandemic strains of influenza A virus might arise by genetic reassortment between viruses from different hosts. The human influenza pandemic strains of 1957 [Asian/57(H2N2)] and 1968 [Hong Kong/68(H3N2)] were the result of reassortment; they were double-reassortant viruses containing genes similar to those of avian and human influenza viruses (11, 23). The reassortment events responsible for the 1957 and 1968 pandemic viruses might have occurred in an animal that served as a mixing vessel. Pigs are susceptible to both human and avian influenza viruses and have been proposed to be intermediate hosts for the generation of pandemic influenza viruses through reassortment or adaptation to the mammalian host (5, 8, 22). Thus, the human influenza pandemic strains of 1957 and 1968 might have been generated in pigs (5). In this study, we report that double-reassortant and triple-reassortant H3N2 swine influenza viruses emerged in pigs in China, and this finding might raise further concerns about whether these viruses will become established in pigs in China and whether they will become a significant pandemic threat.
China, especially southern China, is thought to be the epicenter for the human influenza pandemics throughout history (26). The special environment and lifestyle in southern China provide more chances for wild aquatic birds, domestic poultry, pigs, and humans to come into close contact and create the opportunity for interspecies transmission and the generation of new reassortment influenza viruses. Although it is virtually impossible to prevent new outbreaks of influenza in humans and animals, it is now well recognized that animal influenza virus surveillance can play a key role in the early recognition of outbreak threats. Therefore, carrying out swine influenza virus surveillance is of great significance. The coexistence of reassortant viruses in this study provides further evidence that pigs serve as intermediate hosts, or mixing vessels, and emphasizes the importance of reinforcing swine influenza virus surveillance in China.
Acknowledgments
This work was supported by the National Key Technology Research and Development Program (2004BA519A55) and National Basic Research Program (973) of China (2005CB523200).
Footnotes
Published ahead of print on 16 January 2008.
REFERENCES
- 1.Brown, I. H. 2000. The epidemiology and evolution of influenza viruses in pigs. Vet. Microbiol. 7429-46. [DOI] [PubMed] [Google Scholar]
- 2.Castrucci, M. R., I. Donatelli, L. Sidoli, G. Barigazzi, Y. Kawaoka, and R. G. Webster. 1993. Genetic reassortment between avian and human influenza A viruses in Italian pigs. Virology 193503-506. [DOI] [PubMed] [Google Scholar]
- 3.Caton, A. J., G. G. Brownlee, J. W. Yewdell, and W. Gerhard. 1982. Antigenic structure of the influenza virus A/PR/8/34 hemagglutinin (H1 subtype). Cell 31417-427. [DOI] [PubMed] [Google Scholar]
- 4.Chi, X. S., T. V. Bolar, P. Zhao, J. S. Tam, R. Rappaport, and S. M. Cheng. 2005. Molecular evolution of human influenza A/H3N2 virus in Asia and Europe from 2001 to 2003. J. Clin. Microbiol. 436130-6132. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Fouchier, R. A. M., A. D. M. E. Osterhaus, and I. H. Brown. 2003. Animal influenza virus surveillance. Vaccine 211754-1757. [DOI] [PubMed] [Google Scholar]
- 6.Gorman, O. T., W. J. Bean, Y. Kawaoka, I. Donatelli, Y. Guo, and R. G. Webster. 1991. Evolution of influenza A virus nucleoprotein genes: implications for the origins of H1N1 human and classical swine viruses. J. Virol. 653704-3714. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Ito, T. 2000. Interspecies transmission and receptor recognition of influenza A viruses. Microbiol. Immunol. 44423-430. [DOI] [PubMed] [Google Scholar]
- 8.Ito, T., J. N. Couceiro, S. Kelm, L. G. Baum, S. Krauss, M. R. Castrucci, I. Donatelli, H. Kida, J. C. Paulson, R. G. Webster, and Y. Kawaoka. 1998. Molecular basis for the generation in pigs of influenza A viruses with pandemic potential. J. Virol. 727367-7373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Karasin, A. I., M. M. Schutten, L. A. Cooper, C. B. Smith, K. Subbarao, G. A. Anderson, S. Carman, and C. W. Olsen. 2000. Genetic characterization of H3N2 influenza viruses isolated from pigs in North America, 1977-1999: evidence for wholly human and reassortant virus genotypes. Virus Res. 6871-85. [DOI] [PubMed] [Google Scholar]
- 10.Katsuda, K., T. Shirahata, H. Kida, and H. Goto. 1995. Antigenic and genetic analyses of the hemagglutinin of influenza viruses isolated from pigs in 1993. J. Vet. Med. Sci. 571023-1027. [DOI] [PubMed] [Google Scholar]
- 11.Kawaoka, Y., S. Krauss, and R. G. Webster. 1989. Avian-to-human transmission of the PB1 gene of influenza A viruses in the 1957 and 1968 pandemics. J. Virol. 634603-4608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Kothalawala, H., M. J. M. Toussaint, and E. Gruys. 2006. An overview of swine influenza. Vet. Q. 2845-53. [PubMed] [Google Scholar]
- 13.Landolt, G. A., A. I. Karasin, L. Phillips, and C. W. Olsen. 2003. Comparison of the pathogenesis of two genetically different H3N2 influenza A viruses in pigs. J. Clin. Microbiol. 1411936-1941. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Lubeck, M. D., and W. Gerhard. 1981. Topological mapping antigenic sites on the influenza A/PR/8/34 virus hemagglutinin using monoclonal antibodies. Virology 11364-72. [DOI] [PubMed] [Google Scholar]
- 15.Ma, W., M. Gramer, K. Rossow, and K. J. Yoon. 2006. Isolation and genetic characterization of new reassortant H3N1 swine influenza virus from pigs in the Midwestern United States. J. Virol. 805092-5096. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Matrosovich, M., A. Tuzikov, N. Bovin, A. Gambaryan, A. Klimov, M. R. Castrucci, I. Donatelli, and Y. Kawaoka. 2000. Early alterations of the receptor-binding properties of H1, H2, and H3 avian influenza virus hemagglutinins after their introduction into mammals. J. Virol. 748502-8512. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Nerome, K., Y. Kanegae, K. F. Shortridge, S. Sugita, and M. Ishida. 1995. Genetic analysis of porcine H3N2 viruses originating in southern China. J. Gen. Virol. 76613-624. [DOI] [PubMed] [Google Scholar]
- 18.Nobusawa, E., T. Aoyama, H. Kato, Y. Suzuki, Y. Tateno, and K. Nakajima. 1991. Comparison of complete amino acid sequences and receptor-binding properties among 13 serotypes of hemagglutinins of influenza A viruses. Virology 182475-485. [DOI] [PubMed] [Google Scholar]
- 19.Peiris, J. S. M., Y. Guan, D. Markwell, P. Ghose, R. G. Webster, and K. F. Shortridge. 2001. Cocirculation of avian H9N2 and contemporary “human” H3N2 influenza A viruses in pigs in southeastern China: potential for genetic reassortment? J. Virol. 759679-9686. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Potter, C. W. 2001. A history of influenza. J. Appl. Microbiol. 91572-579. [DOI] [PubMed] [Google Scholar]
- 21.Rogers, G. N., J. C. Paulson, R. S. Daniels, J. J. Skehel, I. A. Wilson, and D. C. Wiley. 1983. Single amino acid substitutions in influenza haemagglutinin change receptor binding specificity. Nature 30476-78. [DOI] [PubMed] [Google Scholar]
- 22.Scholtissek, C., S. Ludwig, and W. M. Fithch. 1993. Analysis of influenza A virus nucleoproteins for the assessment of molecular genetic mechanism leading to new phylogenetic virus lineages. Arch. Virol. 131237-250. [DOI] [PubMed] [Google Scholar]
- 23.Scholtissek, C., W. Rohde, V. V. Hoyningen, and R. Rott. 1978. On the origin of the human influenza virus subtypes H2N2 and H3N2. Virology 8713-20. [DOI] [PubMed] [Google Scholar]
- 24.Schulze, I. T. 1997. Effects of glycosylation on the properties and functions of influenza virus hemagglutinin. J. Infect. Dis. 1(Suppl.)S24-S28. [DOI] [PubMed] [Google Scholar]
- 25.Shortridge, K. F., A. Cherry, and A. P. Kendal. 1979. Further studies on the antigenic properties of H3N2 strains of influenza A viruses isolated from swine in southeast Asia. J. Gen. Virol. 44251-254. [DOI] [PubMed] [Google Scholar]
- 26.Shortridge, K. F., and C. H. Stuart-Harris. 1982. An influenza epicentre? Lancet ii812-813. [DOI] [PubMed] [Google Scholar]
- 27.Shortridge, K. F., and R. G. Webster. 1979. Geographical distribution of swine (Hsw1N1) and Hong Kong (H3N2) influenza virus variants in pigs in southeast Asia. Intervirology 119-15. [DOI] [PubMed] [Google Scholar]
- 28.Shortridge, K. F., R. G. Webster, W. K. Butterfield, and C. H. Campbell. 1977. Persistence of Hong Kong influenza virus variants in pigs. Science 1961454-1455. [DOI] [PubMed] [Google Scholar]
- 29.Shu, L. L., Y. P. Lin, S. M. Wright, K. F. Shortridge, and R. G. Webster. 1994. Evidence for interspecies transmission and reassortment of influenza A viruses in pigs in southern China. Virology 202825-833. [DOI] [PubMed] [Google Scholar]
- 30.Subbaral, K., and T. Joseph. 2007. Scientific barriers to developing vaccines against avian influenza viruses. Nature 7267-278. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Suzuki, Y., I. Toshihiro, T. Suzuki, R. E. Holland, T. M. Chambers, M. Kiso, H. Ishida, and Y. Kawaoka. 2000. Sialic acid species as a determinant of the host range of influenza A viruses. J. Virol. 7411825-11831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Webby, R. J., S. L. Swenson, S. L. Krauss, P. J. Gerrish, S. M. Goyal, and R. G. Webster. 2000. Evolution of swine H3N2 influenza viruses in the United States. J. Virol. 748243-8251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Weis, W., J. H. Brown, S. Cusack, J. C. Paulson, J. J. Skehel, and D. C. Wiley. 1988. Structure of the influenza virus haemagglutinin complexed with its receptor, sialic acid. Nature 333426-431. [DOI] [PubMed] [Google Scholar]
- 34.Wiley, D. C., I. A. Wilson, and J. J. Skehel. 1981. Structural identification of the antibody-binding sites of Hong Kong influenza haemagglutinin and their involvement in antigenic variation. Nature 289373-378. [DOI] [PubMed] [Google Scholar]
- 35.Yu, H., G. H. Zhang, R. H. Hua, Q. Zhang, T. Q. Liu, M. Liao, and G. Z. Tong. 2007. Isolation and genetic analysis of human origin H1N1 and H3N2 influenza viruses from pigs in China. Biochem. Biophys. Res. Commun. 35691-96. [DOI] [PubMed] [Google Scholar]
- 36.Zhou, N. N., D. A. Senne, J. S. Landgraf, S. L. Swenson, G. Erickson, K. Rossow, L. Liu, K. J. Yoon, S. Krauss, and R. G. Webster. 1999. Genetic reassortment of avian, swine, and human influenza A viruses in American pigs. J. Virol. 738851-8856. [DOI] [PMC free article] [PubMed] [Google Scholar]