Abstract
Newly synthesized histones are acetylated prior to their deposition into nucleosomes. Following nucleosome formation and positioning, they are rapidly deacetylated, an event that coincides with further maturation of the chromatin fiber. The histone deacetylases (HDACs) used for histone deposition and de novo chromatin formation are poorly understood. In the ciliate Tetrahymena thermophila, transcription-related deacetylation in the macronucleus is physically separated from deposition-related deacetylation in the micronucleus. This feature was utilized to identify an HDAC named Thd2, a class II HDAC that acts on newly synthesized histones to remove deposition-related acetyl moieties. The THD2 transcript is alternatively spliced, and the major form contains a putative inositol polyphosphate kinase (IPK) domain similar to Ipk2, an enzyme that promotes chromatin remodeling by SWI/SNF remodeling complexes. Cells lacking Thd2, which retain deposition-related acetyl moieties on new histones, exhibit chromatin and cytological phenotypes indicative of a role for Thd2 in chromatin maturation, including the proteolytic processing of histone H3.
In eukaryotes, DNA complexes with an octamer of histone proteins (two each of histones H2A, H2B, H3, and H4) to form a repeating unit of chromatin called a nucleosome (23, 29, 50). The precise positioning of nucleosomes can regulate DNA-templated processes, such as replication, transcription, and recombination. Nucleosome assembly and initial positioning occur during the S phase of the cell cycle and are closely coupled to passage of replication forks (28). During replication, the existing nucleosomes are randomly segregated onto the newly synthesized DNA (47), while new nucleosomes are formed de novo through the assembly of free histones in a specific order. First, two H3/H4 heterodimers are deposited onto the DNA, followed by two H2A/H2B dimers (14). Prior to their deposition, free histones H3 and H4 are acetylated on lysine (Lys) residues by cytoplasmic histone acetyltransferases (HATs) in patterns distinct from transcription acetylation patterns (2, 21, 42, 53). The deposition acetylation promotes histone assembly into nucleosomes through interactions with histone chaperones, such as chromatin assembly factor 1 (CAF-1), Hif1, and Asf1, that incorporate new H3 and H4 specifically onto newly replicated DNA (1, 14, 45, 52). The deposition-related pattern on histone H4 is highly conserved from yeasts to humans and consists of diacetylated Lys5 and Lys12, which correspond to Lys4 and Lys11 in the ciliate Tetrahymena thermophila (2, 8, 46). Recently, acetylation of Lys91 in the globular domain of H4 was identified as another modification important for nucleosome assembly (60). The acetylation pattern on newly synthesized histone H3 is more variable between organisms. In Tetrahymena, Lys9 and Lys14 are acetylated, whereas in Drosophila, Lys14 and Lys23 are the preferred sites (46), and in budding yeast, most new H3 molecules are monoacetylated on Lys9, Lys14, Lys23, Lys27, or Lys56 (25, 32). In contrast to H3 and H4, nascent H2A and H2B histones do not display any acetylation patterns that are distinct from parental forms.
Once assembled into nucleosomes, new histones H3 and H4 are rapidly deacetylated (2, 21), an event important for chromatin maturation. When chromatin is replicated in vivo in the presence of sodium butyrate to inhibit deacetylation, the resulting nucleosome structure and distribution on the chromatin fiber appear normal, but the “mature” fiber is abnormally hypersensitive to DNase I (7, 44). Chromatin assembled in vitro with hyperacetylated histones is also more nuclease sensitive (24). Aside from a recent demonstration that deacetylation of Lys91 on histone H4 facilitates the formation of a salt bridge to histone H2B, little is known about the role of histone deacetylation in chromatin maturation. Further studies in this area have in part awaited identification of the histone deacetylases (HDACs) involved.
Most HDACs identified to date fall into three phylogenetic classes, depending on their homology to the yeast deacetylases Rpd3 (class I), Hda1 (class II), or NAD-dependent Sir2 (class III). Enzymes in these classes can differ in localization and tissue-specific expression (11). Generally, class I HDACs reside in the nucleus, while many class II enzymes shuttle between the nucleus and cytoplasm in response to cellular signals. Some class I yeast HDACs that regulate transcription deacetylate Lys5/Lys12, in addition to other residues on H4, and thus may also play a role in chromatin maturation (41). However, their transcription function complicates investigations into possible maturation functions, since the two processes occur simultaneously in the same organelle. Most systems share the difficulty of separating chromatin maturation-related from transcription-related deacetylation events. To circumvent this barrier to identifying a maturation-related HDAC, we utilized the ciliated protozoan T. thermophila for its two distinct nuclei, called the macronucleus and the micronucleus. Global chromatin in the macronucleus is highly acetylated, while that in the micronucleus is entirely unacetylated. Both nuclei divide and assemble new chromatin, but transcription-related acetylation occurs only in the macronucleus during vegetative growth; the micronucleus is transcriptionally silent throughout the cell cycle (2, 51). The only acetylation observed in micronuclei is highly transient and occurs in deposition-related patterns (8). Thus, HDACs that act on micronuclear chromatin during vegetative growth must function in the removal of deposition acetylation. Compelling evidence for such a micronuclear HDAC activity was observed during a period in the sexual-conjugation pathway when the macronucleus remains in a nonreplicative state (G1) while the micronucleus undergoes successive rounds of mitosis and DNA replication (12). Throughout this period, newly synthesized histones were deacetylated by an unknown micronuclear HDAC (2, 8). Following deacetylation, micronuclear chromatin is thought to mature through evolutionarily conserved pathways, but one unique feature is the proteolytic processing of a fraction of new histone H3 molecules into a truncated form (3, 5, 6). Production of this form is related to mitotic chromosome condensation and segregation in the micronucleus (54).
Here, we report the identification of a class II HDAC called Thd2 (Tetrahymena histone deacetylase 2) that localizes to the micronucleus, where it removes deposition-related acetylation from histones H3 and H4. Cells lacking Thd2 display micronuclear chromatin phenotypes, including reduced H3 proteolysis. THD2 is an alternatively spliced gene, the first reported for a coding sequence in Tetrahymena. Of the two resulting splice variants, the predominant form encodes an HDAC enzyme containing a putative inositol polyphosphate kinase (IPK) domain that in other systems has been implicated in the regulation of chromatin remodeling. From this study, possible roles for HDACs in the maturation of new chromatin are revealed.
MATERIALS AND METHODS
Bioinformatics.
Eighteen HDACs (Thds) were identified as containing putative HDAC domains from The Institute for Genomic Research-annotated genes in the Tetrahymena macronuclear genome database (http://www.ciliate.org). Their respective gene sequence identification numbers are listed in Table 1. Since experimental confirmation of the coding sequences was lacking, only the HDAC domains were used for phylogenetic analysis. The sequences of the 18 Tetrahymena HDAC domains were aligned with the S. cerevisiae Rpd3, Hda1, Sir2, Hst2, and Hst4 HDAC domains using Multiple Sequence Alignment-CLUSTALW (Kyoto University Bioinformatics Center [http://align.genome.jp/]). The CLUSTAL protein alignment was performed using a gap open penalty of 10, a gap extension penalty of 0.05, a hydrophobic gap, no weight transition, and a BLOSUM weight matrix. Distances were computed using the Poisson Correction Distance method in Molecular Evolutionary Genetic Analysis (MEGA) software version 4.0 (MEGA4) (49). The unweighted-pair group method using average linkages tree was constructed from the matrix of distances according to the model using MEGA4, and the robustness of the tree topology was tested with 1,000 bootstrap replicates.
TABLE 1.
Putative Tetrahymena HDACs with similarity to human and S. cerevisiae proteins
| TGD gene ID (TTHERM_ no.) | Name | Human homolog (E value) | S. cerevisiae homolog (E value) |
|---|---|---|---|
| 00647330 | THD1 | HDAC1 (1.0e−126) | RPD3 (1.0e−116) |
| 00621610 | THD2 | HDAC6 (1.0e−61) | HDA1 (4.0e−61) |
| 00663840 | THD3 | HDAC1 (1.0e−101) | RPD3 (4.0e−96) |
| 01014700 | THD4 | HDAC6 (1.0e−39) | HDA1 (6.0e−43) |
| 00395710 | THD5 | HDAC11 (2.0e−63) | HDA1 (1.0e−09) |
| 00502610 | THD6 | MRP1 (1.0e−261) | RPD3 (1.0e−109) |
| 00031680 | THD7 | HDAC6 (1.0e−14) | UBP14 (2.0e−03) |
| 01018420 | THD8 | SIRT6 (9.0e−44) | HST2 (6.0e−08) |
| 00242580 | THD9 | SIRT6 (2.0e−32) | SIR2 (6.0e−12) |
| 00859330 | THD10 | SIRT5 (5.0e−09) | SIR2 (2.0e−03) |
| 00313730 | THD11 | SIRT6 (1.0e−53) | HST2 (4.0e−09) |
| 01018450 | THD12 | SIRT6 (3.0e−49) | HST2 (1.0e−07) |
| 00672190 | THD13 | SIRT3 (3.0e−44) | HST2 (3.0e−29) |
| 00526990 | THD14 | SIRT3 (6.0e−54) | HST2 (2.0e−38) |
| 00112480 | THD15 | SIRT3 (1.0e−39) | HST2 (2.0e−31) |
| 00194150 | THD16 | SIRT2 (2.0e−42) | HST2 (1.0e−28) |
| 00833750 | THD17 | SIRT5 (3.0e−05) | HST4 (9.0e−03) |
| 00833740 | THD18 | SIRT5 (6.0e−06) | HST4 (1.0e−03) |
Strains and cell culture conditions.
T. thermophila strains B2086 (II), CU428 (chx1/chx1 [cy-s] VII), and SB1969 (mpr1/mpr1 [6-mp-s] II) were used as the wild-type strains (Tetrahymena Stock Center, Cornell University). Unless otherwise indicated, cells were grown at 30°C with shaking in either 1% (wt/vol) enriched proteose peptone (17) or 2% proteose peptone, 0.2% yeast extract, and 0.1% sequestrene (61) liquid medium with 1× penicillin, streptomycin, and fungizone (Gibco-BRL) to a density of 1 × 105 to 3 × 105 cells/ml for all experiments. The cells were starved in 10 mM Tris-HCl (pH 7.4) for 14 to 20 h at a density of 3 × 105 cells/ml at 30°C with no shaking.
RT-PCR.
Genomic DNA was isolated as described previously (61). Total RNA was isolated from vegetatively dividing, starved, and conjugating cells using the RNeasy Total RNA kit (Qiagen, Valencia, CA). The cDNA was made as previously described (31) using 2 μg of total RNA for each reaction. PCR was performed on dilutions of the cDNA to determine the linear range of conditions for the sample set (data not shown). Experiments were performed using conditions that were within the linear amplification range to obtain semiquantitative results. PCR and reverse transcriptase (RT)-PCR were performed using 2× GoTaq master mixture (Promega, Madison, WI) following the manufacturer's directions and using the following primer sets: THD2a (+) (5′-GTTTATTTTGATATCTGCTG-3′) and THD2b (+) (5′-TTAGACTTCAAATGAATTTAC-3′); THD2 (−) (5′-CTAAATGCGATCCTTTAATTC-3′) and HHP1 (+) (5′-TTAGCAATGATAAACCTTAGAC-3′); HHP1 (−) (5′-TGTGTAAAGAGATTTTCCATC-3′) and ACT1 (+) (5′-GAACAGAGAAAAGATGACCAAG-3′); ACT1 (−) (5′-GGTAAGTTCGTGGATACCAGG-3′) and CYP1 (+) (5′-AGAGTAACCCTAATAACACC-3′); and CYP1 (−) (5′-CCGTTGAAAATTCCAGACG-3′).
THD2 gene and splice variant analysis.
The 5′ terminus of the THD2 mRNA was deduced by RT-PCR with the THD2 (−) primer listed above and the following primers upstream of the predicted translation start site: THD2-4 (+) (5′-CCAAAACAGACAAACTATGCAAC-3′), THD2-132 (+) (5′-ATTTCATATGATAAGATTGAATTCATTTAC-3′), THD2-219 (+) (5′-CATTTTTTCTATCTATTATGAGTCATAATC-3′), THD2-313 (+) (5′-ATGTTAAATTTTTACGTAATTTTAAATTGCG-3′), and THD2-360 (+) (5′-ATTAACTAATGCTTACATGATATTATTC-3′).
The 3′ poly(A) addition site for the THD2a and THD2b mRNA sequence was determined by RT-PCR using THD2E2 (5′-CATGACGATGGATCATTCTATCC-3′), a plus-sense primer in exon 2 (E2), and a poly(dT)20 primer as the reverse primer. The PCR product was subcloned using the Invitrogen TA cloning kit (Invitrogen, California) and sequenced on an Applied Biosystems 3130xl Genetic Analyzer (Rancho Santa Ana Botanic Garden Molecular Laboratory, Claremont, CA) to determine the correct intron junctions and the poly(A) addition sites for the splice variants.
Cell elutriation and conjugation.
Logarithmically growing cells (1 × 105 to 3 × 105 cells/ml) were synchronized using a centrifugal elutriator rotor as previously described (31), and RNA was collected every 20 min after the elutriated cells were resuspended in fresh 2% proteose peptone, 0.2% yeast extract, and 0.1% sequestrene. For conjugation experiments, starved CU428 (VII) and SB1969 (II) were mixed in equal concentrations (3 × 105 cells/ml), and 10-ml aliquots were placed in 100-mm by 15-mm prewarmed petri dishes and incubated in a humidity chamber at 30°C (no shaking). RNA was collected from 10 ml of conjugating cells every hour initially after the mixing (0 h) through 14 h and again at 24 h after the mixing.
Plasmid construction.
The NEO construct used to replace THD2 was made following a previously described method (56), except that the THD2 3′ and 5′ flanking regions were amplified by PCR using the following primers: 5′FlankBamHI (+) (5′-GAGGGATCCAAAGTGTGATGTGTTTGAAG-3′), 5′FlankXmaI (−) (5′-GTGCCCGGGCTAAAAAGAAATCAAGAAAG-3′), 3′FlankClaI (+) (5′-CGCATCGATAGAAACAACTCCTGTCTGTC-3′), and 3′FlankXhoI (−) (5′-AGACTCGAGTCAAATACCTAGCCCTCTCC-3′).
The PCR products were digested with appropriate restriction enzymes and ligated into p4T2-1 containing the Neor drug resistance cassette to make pTHD2-NEO (56). For transformation, pTHD2-NEO was digested to produce a linear fragment containing the 5′ and 3′ THD2 flanking sequence interrupted by the neomycin drug resistance cassette. Gold beads were coated with this, as previously described (31).
Construction of a strain expressing green fluorescent protein (GFP) fused to the THD2 genomic sequence was done as previously described (59), using primers THD2GFP (+) (5′-CAACAGGATAATATAGAAGGTATAG-3′) and THD2GFP (−) (5′-TCATTTTGCCTTTTTGTTAAGAG-3′), and transformed via electroporation as previously described (15).
Macronuclear THD2 replacement.
Tetrahymena cells were transformed through biolistic bombardment to delete THD2 as previously described (30). Transformants were selected by growth in the presence of 100 μg/ml paromomycin (Sigma Chemicals) after 5 h of incubation at 30°C without paromomycin (no shaking). The thd2Δ allele was assorted to 100% by growing the transformants in media with increasing concentrations of paromomycin to a final concentration of 10 mg/ml. Complete macronuclear replacement of THD2 was confirmed by PCR as previously described (56) using the following primers: THD2Flank (+) (5′-TGGAAGTCAAGCATTTCTTCC-3′), THD2-3 (−) (5′-GTCATCATTCAGGTGAATC-3′), NEOF2 (+) (5′-CTGGGCACAACAFACAATCG-3′), NEOR2 (−) (5′-GTTTCGCTTGGTGGTCGAAT-3′), and NEOS (−) (5′-CCTTAAATTAAAAATTCAATATATATTTTAC-3′).
The resulting strain lacking macronuclear copies of THD2 was named JS01. Lack of THD2 expression in strain JS01 was confirmed by RT-PCR using the THD2a (+) and THD2 (−) primers listed above.
Immunoblot analysis.
Nuclei from B2086 (wild type) and JS01 (thd2Δ) were isolated as previously described (17). The nuclei (1 × 106) were lysed by incubation in 30 μl of sodium dodecyl sulfate (SDS) gel loading buffer (50 mM Tris-HCl [pH 6.8], 100 mM dithiothreitol, 2% [wt/vol] SDS, 0.1% bromophenol blue, 10% glycerol) and boiled for 5 min. Micronuclei (5 × 105) and macronuclei (1.6 × 105) were resolved by SDS-polyacrylamide gel electrophoresis (PAGE) on a 15% (for histone H4) or 22% (for histone H3) polyacrylamide gel, transferred to a nitrocellulose membrane, and probed with the following antibodies: α-histone H4 (1:5,000; a gift from C. D. Allis), α-histone H3 (1:5,000; a gift from C. D. Allis), α-histone H4K16ac (1:2,000; Upstate Biotechnology catalog no. 06-762), and α-histone H3S10ph (1:5,000; a gift from C. D. Allis).
Indirect immunofluorescence and DAPI staining of cells.
B2086 (wild-type) and JS01 (thd2Δ) cells in logarithmic growth (1 × 105 to 3 × 105 cells/ml) were fixed in paraformaldehyde and processed for immunofluorescence as previously described (9). For detection of acetylated histones, cells were incubated with α-acetylated histone H4 antiserum (1:200; Upstate Biotechnology catalog no. 06-598), α-acetylated histone H3 antiserum (1:200; a gift from C. D. Allis), or α-histone H3K9ac antiserum (1:200; Upstate Biotechnology catalog no. 06-942), which also detects H3K14ac in Tetrahymena. For detection of only micronuclei, cells were incubated with anti-micronuclear linker histone 1 (α-Mlh1) (micronuclear-specific linker histone) antiserum (1:200; a gift from C. D. Allis). Primary antibodies were detected with rhodamine-conjugated goat anti-rabbit immunoglobulin G (1:100; Jackson ImmunoResearch catalog no. 111-025-003). The cells were counterstained with 0.1 μg/ml 4′,6 diamino-2-phenylindole dihydrochloride (DAPI; Sigma Chemicals) in 0.1% bovine serum albumin-phosphate-buffered saline for 10 min according to the common protocol (37).
Nucleotide sequence accession numbers.
The cDNA sequences for the splice variants were submitted to GenBank under accession numbers EU254713 (Thd2a) and EU254714 (Thd2b).
RESULTS
Thd2 localizes to the micronucleus.
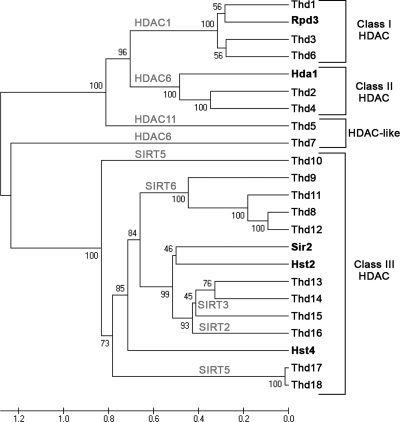
Thds were first identified by searching the Tetrahymena Genome Database to find predicted open reading frames that contained putative HDAC domains. Eighteen putative HDAC proteins were identified (Table 1) and classified by their similarity to the yeast HDACs Rpd3 (class I), Hda1 (class II), and Sir2 (class III) (Fig. 1). The HDAC domains fell into all three classes—3 in class I (Thd1, Thd3, and Thd6), 2 in class II (Thd2 and Thd4), and 11 in class III (Thd8 through Thd18)—and 2 were HDAC-like (Thd5 and Thd7). Most of the putative Tetrahymena class III HDACs (Thd8 through Thd18) were more closely related to human sirtuin homologs than to the yeast sirtuins Sir2, Hst2, and Hst4 (Fig. 1).
FIG. 1.
Phylogenetic tree of the putative Thds (unweighted-pair group method using average linkages). Shown is an alignment of the putative HDAC domains of the 18 Thd protein sequences. S. cerevisiae Rpd3 (class I), Hda1 (class II), Sir2, Hst2, and Hst4 (class III) were used as references to sort the putative Thd proteins into their respective classes (boldface). The positions of human homologs are shown in light-gray capital letters. (Note: HDAC6 and SIRT5 appear twice due to construction of the tree with yeast HDAC domains that are less similar to Thd proteins than those of humans.)
An HDAC that resides in the micronucleus would likely remove deposition-related acetyl marks from newly synthesized histones, since only deposition acetylation occurs on histones in this nucleus. To identify a micronuclear HDAC, several of the putative HDAC genes (Fig. 1) were cloned in frame with GFP coding sequence at their 5′ ends and expressed from the metallothionein promoter. The resulting amino-terminal GFP fusion proteins were induced, and their localization in live cells was assessed by fluorescence microscopy. Of four tested, only GFP-Thd2 localized to the micronucleus in addition to the macronucleus (Fig. 2); the other putative HDACs tested localized exclusively to the macronucleus or to the cytoplasm (data not shown) (57).
FIG. 2.
GFP-Thd2 is localized to the micronucleus. Live-cell images of Tetrahymena transformed with a GFP-Thd2 fusion construct and viewed using the fluorescein isothiocyanate channel to detect GFP signal. The positions of nuclei in each cell were visualized by staining them with DAPI. GFP expressed alone, without fusion to another protein, remained in the cytoplasm; the regions devoid of fluorescence correspond to the macronucleus (M) and the micronucleus (m). Cells expressing GFP-Thd2 showed fluorescence in both the macronucleus and micronucleus, which persisted throughout the cell cycle (see the mitotically dividing cell in the right panel).
THD2 is alternatively spliced.
The THD2 coding sequence was determined by cloning and sequencing its cDNA (as described in Materials and Methods). Amplifying the cDNA with gene-specific primers revealed two cDNAs of different lengths, indicating that the THD2 gene was alternatively spliced. The most abundant splice variant, called THD2a, contained only E1, E2, and E4), while the minor form, called THD2b, contained E1 through E4 (Fig. 3A). Sequence analysis showed that the inclusion of E3 in THD2b caused a frameshift that produced a premature stop codon within E4 and loss of a putative IPK domain (Fig. 3B). The relative transcription level of THD2a was 15- to 20-fold higher than that of THD2b during vegetative growth when normalized to the amplification of genomic DNA for each PCR (Fig. 3C and data not shown). When cells were starved and no longer progressing through the cell cycle, both THD2a and THD2b transcription levels decreased two- to threefold, suggesting that both enzyme forms may have more significant functions during periods of active growth and cell division.
FIG. 3.
Thd2a is a splice variant of THD2 that contains both an HDAC and an IPK domain. (A) Diagram of the THD2 locus with the flanking sequence (thin black), coding sequence (thick gray) with exons labeled below (E1 to E4), and introns (white) labeled below (i1 to i3). The arrows represent primers used to amplify splice variants of THD2. Displayed below the THD2 sequence is a schematic of the mRNA splice variants for Thd2a (only exons E1, E2, and E4) and Thd2b. The locations of putative HDAC (light gray) and putative IPK (dark gray) domains are indicated. (B) Protein sequence alignment of Thd2a and Thd2b. HDAC domain, light gray; IPK domain, dark gray; change in coding sequence due to splicing frameshift, #; and stop codon, *. (C) RT-PCR of wild-type vegetatively growing (V) and starved (S) cells. PCR on a cDNA template using primers Ta(+) and T2(−) was used to detect THD2a transcripts, and primers Tb(+) and T2(−) were used to detect the THD2b variant transcripts. The cDNA was made with (+) and without (−) RT as a control for genomic DNA contamination. Genomic DNA (G) was amplified as a control. RT-PCR yielded two bands for THD2b; the fastest-migrating band corresponds to THD2b (arrow). (Note: THD2b amplification was detected only with an additional seven cycles of PCR amplification over that used for THD2a detection.)
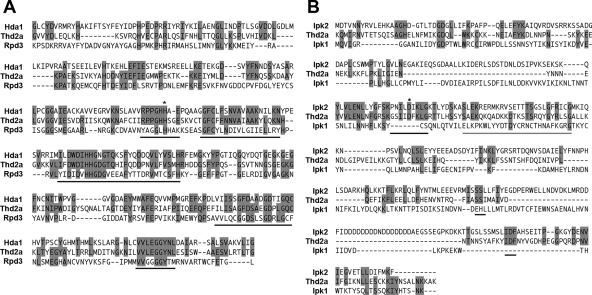
The HDAC domain of Thd2 was aligned with the yeast HDACs Hda1 (class II) and Rpd3 (class I) and found to have greater similarity to Hda1 (37% identical and 56% similar amino acid residues) (Fig. 4A). Thd2 had a high degree of similarity to Hda1 in regions of the HDAC domain that are conserved in most class II enzymes (Fig. 4A) (26, 63). To further classify the putative IPK domain in Thd2, the sequence was compared to two well-characterized yeast IPKs, Ipk1 and Ipk2, the primary enzymes in the pathway for conversion of inositol triphosphate (IP3) to inositol hexaphosphate (IP6) (Fig. 4B). The putative IPK domain of Thd2a was more closely related to Ipk2 (27% identical and 36% similar amino acid residues) than to Ipk1 (10% identical and 22% similar amino acid residues). Notably, the cofactor binding regions and inositol polyphosphate binding region for Ipk2 were similar in Thd2a. Thd2a also contains a conserved aspartate residue that is essential for Ipk2 kinase activity (Fig. 4B) (20).
FIG. 4.
Thd2 is a class II HDAC with a putative IPK domain similar to Ipk2. (A) Alignment of the Thd2a HDAC domain with the HDAC domains of S. cerevisiae Hda1 (class II) and Rpd3 (class I). Residues conserved between Thd2 and either Hda1, Rpd3, or both are shaded in gray. Regions that are highly conserved between most class II HDACs are marked with a black line below the sequence. A conserved histidine residue critical for HDAC activity is marked by an asterisk. (B) Alignment of the Thd2a putative IPK domain with S. cerevisiae Ipk1 and Ipk2. Residues conserved between Thd2 and either Ipk2, Ipk1, or both are shaded in gray. The inositol polyphosphate binding domain is marked with a gray line below the sequence, and regions involved in cofactor binding are marked by a black line below the sequence. A conserved aspartate residue critical for Ipk2 activity in yeast is designated by an asterisk.
Expression of THD2 coincides with DNA replication.
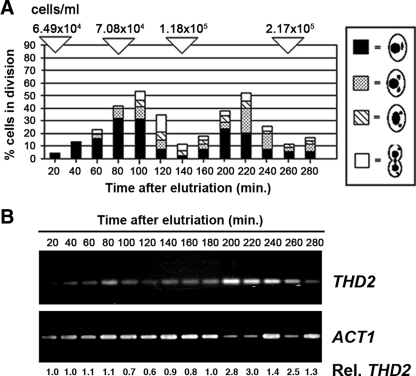
To begin characterizing THD2 function, we first examined its transcription over a normal cell cycle during vegetative growth, since cells produced two- to threefold more THD2 mRNA when growing than in the noncycling, starved state. In Tetrahymena cells, the micronucleus progresses through the cell cycle without resting in G1 phase. Following mitotic division, a cell proceeds directly into S phase and then rests in an extended G2 phase while the macronucleus undergoes amitotic division followed by DNA synthesis (22, 58). Logarithmically growing cells were synchronized by isolating new daughter cells through centrifugal elutriation and then allowing them to proceed through the cell cycle. The synchrony of cells in the culture was monitored at frequent intervals throughout several cell cycles by counting the cells at different stages (Fig. 5A). RT-PCR analysis of THD2 expression at regular intervals throughout two cell cycles (from 20 min to 280 min after elutriation) revealed that THD2 was expressed in a cyclical pattern (Fig. 5B). Maximum expression of THD2 occurred at 80 and 200 min, which were the times when the greatest numbers of cells were in micronuclear mitosis. The expression of ACT1, which does not change throughout the cell cycle, was monitored as a control (34, 62). Since micronuclear DNA synthesis occurs immediately following anaphase, the observed peak of THD2 expression coincident with anaphase was consistent with the possibility that Thd2a might deacetylate newly deposited histones.
FIG. 5.
Thd2 is expressed during DNA replication and cell division in growing cells. (A) Graph of the cell cycle stages represented in samples of a culture synchronized by centrifugal elutriation. Samples were taken every 20 min, and the percentages of cells in each stage of cell division were scored. The concentration of the culture is provided above the graph and a key describing the four different stages of division scored is to the right of the graph. (B) Samples from the synchronized culture were subjected to RT-PCR to detect THD2 transcripts at regular intervals throughout the cell cycle. Primers Ta(+) and T2(−) were used in RT-PCR to detect the predominant form, THD2a. ACT1 cDNA, which remains constant throughout the cell cycle, was used as a control. Relative levels of THD2 (Rel. THD2) were determined by quantifying band intensities of THD2 and ACT1 at each time point, dividing the values for THD2 by the values for ACT1 and then normalizing the resulting value to that at the 20-min time point.
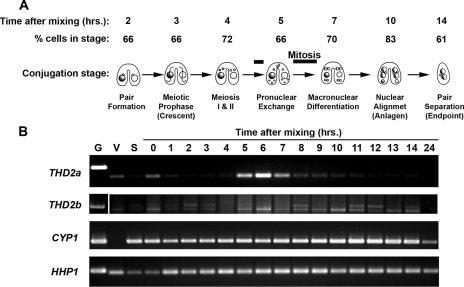
To further examine whether THD2a expression coincides with DNA replication and division, transcript levels were monitored throughout the sexual-conjugation pathway. Early in this process, the transcriptionally silent micronucleus undergoes three successive rounds of DNA replication and two mitoses while the transcriptionally active macronucleus remains in G1 (Fig. 6A). Exploiting this feature, we tested whether THD2a expression coincided with DNA synthesis and histone deposition in the micronucleus at the time (5 to 7 h) when a micronuclear HDAC activity that removes deposition acetylation was previously detected (2). Cells of two different mating types were starved, mixed together, and allowed to conjugate over a 24-hour period. To monitor population synchrony, the percentage of cells in each stage was determined at regular intervals throughout the conjugation time course. By 2 hours after they were mixed, 66% of the cells were paired (Fig. 6A). By 3 hours after they were mixed, 66% were in meiotic prophase and 85% were paired. Throughout the entire conjugation time course, 60 to 70% of the cells were tightly synchronized and an additional 20 to 25% of the culture was within 60 min of this primary synchronized population. RT-PCR on cDNA made from conjugating cells harvested at 1-hour intervals revealed that THD2a was most highly expressed between 5 and 7 h after the mixing (Fig. 6B). At the peak of expression (6 h postmixing), 70% of the cells were in one of the mitotic divisions (30% in prezygotic mitosis and 40% in postzygotic mitosis) (Fig. 6A). This result demonstrated a strong correlation between THD2a expression and periods of DNA replication, evidence that THD2a encodes an HDAC that acts during this time.
FIG. 6.
Thd2 is expressed during conjugation coincident with micronuclear DNA synthesis and mitosis. (A) Diagram of Tetrahymena conjugation stages. Two different mating types were mixed to initiate conjugation. Samples were taken every hour and stained with DAPI, and the percentage of cells in each stage of conjugation was determined by fluorescence microscopy, as indicated above the diagram. The black bars indicate periods of DNA synthesis and mitosis (the short bar is the prezygotic mitosis just prior to the pronuclear exchange; the long bar is the postzygotic mitoses I and II following zygotic fusion). (B) Total RNA was harvested from vegetatively growing (V) cells, starved (S) cells, and cells during conjugation (0 through 14 and 24 h after mixing) and used as a template in RT-PCRs with primers specific for THD2a or THD2b variants. Primers for CYP1 and HHP1, two genes showing consistent expression throughout conjugation, were used as controls in this analysis. Genomic DNA (G) was used to control for genomic-DNA contamination in the RNA samples. (Note: THD2b RT-PCR contained two bands; the faster-migrating band corresponds to the spliced form of Thd2b.)
Similar analysis revealed that Thd2b expression remained low during conjugation compared to that of Thd2a. This minor variant also appeared to be more ubiquitously expressed throughout conjugation, with only a slight peak in expression at 6 h (Fig. 6B). The band corresponding to genomic DNA contamination was amplified to a greater extent at some conjugation points when THD2a levels were relatively low. Curiously, when reactions lacking RT were performed on these mRNA samples and they were used as templates for PCR, no amplification of Thd2 cDNA was observed (Fig. 3C and data not shown). This result raises the possibility that there may be yet another variant, one that retains the intron between E3 and E4. Such a variant would produce an HDAC similar to Thd2b that lacks the putative IPK domain but that has an earlier stop codon than Thd2b.
Thd2 deacetylates micronuclear histones.
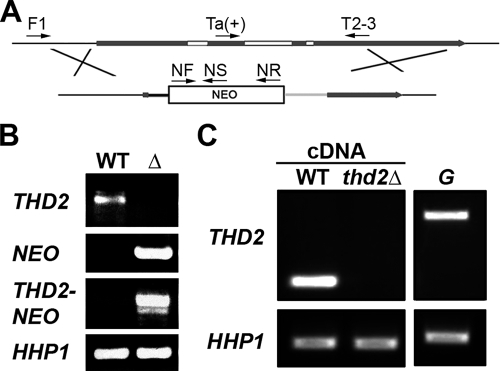
To test whether Thd2 deacetylates micronuclear chromatin, a mutant cell line lacking THD2 was first engineered. A genetic construct for the somatic replacement of THD2 with a paromomycin resistance gene (NEO) was transformed into wild-type cells (Fig. 7A). Complete replacement of the THD2 allele with the NEO construct was confirmed by PCR (Fig. 7B, THD2 and NEO), as was the correct integration of the construct into the THD2 locus (Fig. 7B, THD2-NEO). The resulting cell line lacking all macronuclear copies of THD2 was called thd2Δ. RT-PCR was conducted on the thd2Δ cells to confirm the absence of THD2 expression (Fig. 7C). Obtaining a complete knockout strain indicated that THD2 was a nonessential gene in vegetatively growing cells.
FIG. 7.
THD2 is a nonessential gene. (A) Diagram of the THD2 deletion construct used to replace THD2 with NEO in the somatic macronucleus. Depicted in the diagram are the flanking regions (thin black lines), the coding sequence (thick dark gray lines), introns (white boxes), the histone H4 promoter (thick black lines), the neomycin resistance gene (NEO), and the BTU2 polyadenylation region (light gray line). The arrows represent the primers used to confirm correct integration of the replacement allele. (B) PCR amplification of genomic DNA from wild-type (WT) and thd2Δ (Δ) cells confirmed that all THD2 alleles were replaced with the NEO cassette. THD2 PCR was performed using Ta(+) and T2-3 primers (WT allele), NEO PCR with NF and NR primers (NEO cassette), and THD2-NEO PCR with F1 and NS primers (incorporation of the NEO cassette in the THD2 locus). HHP1 PCR was performed as a positive control for the genomic DNA. (C) Total cDNA derived from WT and thd2Δ cells was used in PCRs to test for the presence of THD2 mRNA in these cells. HHP1 was used as a control for cDNA synthesis and PCR amplification. Genomic DNA (G) was used as a template to control for genomic-DNA contamination in cDNA.
The micronucleus is transcriptionally inactive and lacks acetylated chromatin in growing cells. Acetylated histone H4 in micronuclei was previously detected only when cells were treated with general HDAC inhibitors, such as sodium butyrate and trichostatin A (2, 8, 13). To test the acetylation state of micronuclear chromatin in thd2Δ cells, immunofluorescence was performed using antiserum specific for only acetylated species of histones H3 and H4 (α-H3ac and α-H4ac). Consistent with previous studies of wild-type cells (51), acetylated histones H3 and H4 were detected only within the transcriptionally active macronucleus and not in the micronucleus (Fig. 8, WT). In thd2Δ cells, however, acetylated histones H3 and H4 were detected in the micronucleus, as well (Fig. 8, thd2Δ). Micronuclear histones were acetylated in every cell and in each stage of the cell cycle observed, in contrast to wild-type cells, in which micronuclear acetylation was never detected (Fig. 8). This analysis demonstrated that Thd2 is an HDAC that normally deacetylates histones in the micronucleus. Whether it also acts on macronuclear histones was not determined.
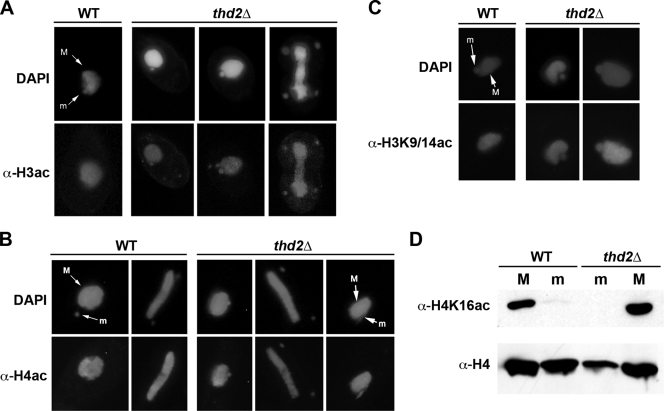
FIG. 8.
Thd2 removes deposition-related acetylation from micronuclear histones. (A) Immunofluorescence using antiserum against acetylated histone H3 (α-H3ac). Cells were counterstained with DAPI to visualize both the macronucleus (M) and the micronucleus (m). Acetylated histone H3 was detected exclusively in the macronuclei of wild-type (WT) cells and additionally in the micronuclei of thd2Δ cells. (B) Immunofluorescence using antiserum against acetylated histone H4 (α-H4ac). DAPI stain was used to detect both the macronucleus and the micronucleus. Acetylated histone H4 was detected exclusively in the macronucleus of WT cells and additionally in the micronuclei of thd2Δ cells throughout every stage of the cell cycle. A high proportion of cells contained elongated micronuclei in close proximity to the macronucleus (the phenotype is depicted in the last panel). (C) Immunofluorescence using antiserum against acetylated Lys9 on histone H3 that additionally detects acetylated Lys14 in Tetrahymena (α-H3K9/14ac). Cells were counterstained with DAPI to visualize both the macronucleus and the micronucleus. Histone H3 Lys9/Lys14 acetylation was detected only in the macronuclei of wild-type cells, but also in the micronuclei of thd2Δ cells. (D) Total nuclear proteins from purified macronuclei and micronuclei were resolved by SDS-PAGE, transferred to a nitrocellulose membrane, and subjected to immunoblot analysis using antiserum against acetylated Lys16 on histone H4 (α-H4K16ac) or against general histone H4 (α-H4). In both wild-type and thd2Δ cells, H4 Lys16 was acetylated only in macronuclei.
It was deduced that the observed micronuclear acetylation in thd2Δ cells resulted from the retention of deposition-related acetyl moieties. To help confirm this, we performed an immunofluorescence assay using an antiserum against acetylated lysine 9 and/or 14 on histone H3 (α-H3K9/14ac), the acetyl marks found on newly synthesized Tetrahymena histones (2, 25). Whereas this antiserum normally does not hybridize with micronuclear chromatin in wild-type cells, it did hybridize with chromatin in thd2Δ micronuclei, indicating that H3Lys9 and/or -14 was acetylated in the mutant cells. Other deposition site-specific acetylation antisera (anti-acetylated Lys5 and anti-acetylated Lys12) were not included in this analysis due to their lack of specificity; Tetrahymena histone H4 is shorter by 1 amino acid (it is missing Arg3) at the N terminus. To help rule out possible aberrant transcription-related acetylation in the micronuclei of thd2Δ cells, an immunoblot analysis was performed on proteins from purified micronuclei or macronuclei with antiserum detecting acetylated Lys16 (α-H4K16ac), a known transcription-specific mark found exclusively in the macronuclei of wild-type cells (8, 25). This antiserum was shown to have correct specificity for a Tetrahymena H4 peptide acetylated at this position (E. Wiley, unpublished data). As shown in Fig. 8D, neither wild-type nor mutant (thd2Δ) micronuclear histone H4 molecules were acetylated on Lys16. Consistent with previous studies, Lys16 acetylation was detected only in macronuclei. Due to high background, results from immunofluorescence experiments with this antiserum are not shown. Combined, these results suggest that Thd2 is normally required for maintenance of the deacetylated state of micronuclear histones H3 and H4 through removal of deposition-related acetyl modifications.
Chromatin defects in thd2Δ cells.
Immunofluorescence studies revealed a number of elongated micronuclei in thd2Δ cells. Although this is the normal appearance of micronuclei in anaphase, many of these elongated micronuclei in mutant cells were closely associated with the macronucleus, an aberrant position for an anaphase nucleus (Fig. 8C, bottom right). In some cases it was difficult to define the micronuclei that stretched across portions of the macronucleus by DAPI staining. To improve the visualization of micronuclei, immunofluorescence was performed with antiserum specific for Mlh1, which is present only in the micronucleus (Fig. 9A). As expected, Mlh1 localized specifically to the micronucleus for both wild-type and thd2Δ cells. Micronuclei were scored for elongated shape when in close proximity to the macronucleus (in the “pocket”) for both wild-type and thd2Δ cells (Fig. 9A). The percentage of cells with abnormal micronuclei in the pocket was approximately 10-fold higher in thd2Δ cells than that in wild-type cells (50.5% ± 3.3% and 4.7% ± 3.2%, respectively). Micronuclei were more variable in size, as well. Although cell division appeared to progress normally, the doubling time for thd2Δ cells was three times that of wild-type cells (9 h and 3 h, respectively) (data not shown). Together, these aberrant morphologies and weak DAPI staining are consistent with chromatin condensation abnormalities in thd2Δ micronuclei.
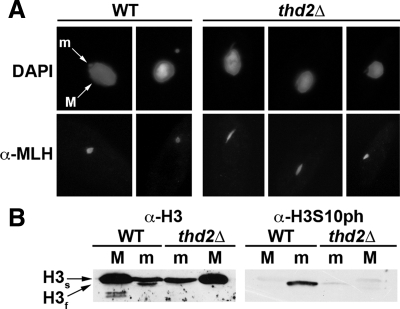
FIG. 9.
Cells lacking Thd2 exhibit chromatin phenotypes. (A) Immunofluorescence using antiserum against micronuclear linker histone H1 (α-Mlh1) was performed on wild-type (WT) and thd2Δ cells. DAPI staining was used to visualize both the macronucleus (M) and the micronucleus (m). A higher incidence of elongated micronuclei in close association with macronuclei was observed in the mutant cells. (B) Total proteins from purified macronuclei and micronuclei were resolved by SDS-PAGE, transferred to a nitrocellulose membrane, and hybridized with antiserum against general histone H3 (α-H3) or with antiserum against phosphorylated serine 10 on histone H3 (α-H3S10ph). The full-length form of histone H3 (H3s) was detected in both macronuclei and micronuclei of all cells, but only wild-type micronuclei contained the faster-migrating proteolytically cleaved form (H3f). Likewise, phosphorylation of Ser10, which is specific for H3f, occurred only in wild-type micronuclei. (Note: the blot was first hybridized with α-H3S10ph and then stripped and hybridized with α-H3.)
In micronuclei, histone H3 (called H3s) is proteolytically cleaved to form a faster-migrating form, called H3f, which is missing 6 amino acids from the extreme amino terminus. H3f is the only form of histone H3 in Tetrahymena that is phosphorylated on Ser10, a mitotic modification necessary for chromosome condensation and segregation (4, 6, 54). In thd2Δ cells, the elongated micronuclear phenotype, reminiscent of either decondensed chromatin or mitotic micronuclei in anaphase, prompted us to examine H3 processing and related phosphorylation on Ser10. Immunoblot analysis revealed that while the abundance of micronuclear H3s was similar to that in wild-type cells, cleavage of histone H3s to H3f was greatly reduced in thd2Δ cells (Fig. 9B). As expected, the reduced amount of H3f correlated with reduced Ser10 phosphorylation, likely the result of the failed H3 processing in these cells (Fig. 9B). These results suggest that Thd2 acts upstream of the proteolytic cleavage and subsequent phosphorylation of Ser10 on histone H3.
DISCUSSION
HDACs in Tetrahymena.
Completion of the Tetrahymena genome-sequencing project facilitated the identification and classification of putative HDACs according to the three classical yeast deacetylases, Rpd3, Hda1, and Sir2 (11). All three classes of HDACs (classes I, II, and III, respectively) are represented in Tetrahymena (Fig. 1). Interestingly, their deacetylase domains are more similar to those in human enzymes than to those in budding yeast. Tetrahymena Thd1, Thd3, and Thd6 enzymes are class I HDACs due to their similarity to yeast Rpd3, and their HDAC domains are most similar to human HDAC1. Thd2 and Thd4 are class II homologs with similarity to yeast Hda1 but that have deacetylase domains most similar to human HDAC6. There are two HDAC-like proteins that did not group well with any of the yeast HDACs. One of these, Thd5, is highly similar to HDAC11, a class IV HDAC thought to be a hybrid of class I and II enzymes present in primates, rodents, Drosophila, and plants (16, 18). Tetrahymena also contains 11 sirtuins (class III HDACs), more than have been reported for any other organism to date. The putative Tetrahymena sirtuins are similar to yeast Sir2 but are more closely related to protein sequences for the human SIRT2 (Thd16), SIRT3 (Thd13, Thd14, and Thd15), SIRT5 (Thd10, Thd17, and Thd18), and SIRT6 (Thd8, Thd9, Thd11, and Thd12) proteins. Tetrahymena will likely prove a useful model system for further studies addressing distinct roles for different HDACs, especially the sirtuins, within a single cell.
Identification of an HDAC that deacetylates new histones.
The nuclear dimorphism of Tetrahymena was exploited to identify an HDAC that removes deposition-related acetylation on newly synthesized histones. Previous work demonstrated that the transcriptionally active macronucleus contains highly acetylated histones H2A, H2B, H3, and H4 (51). In contrast, the micronucleus is transcriptionally inactive in growing cells, and little to no acetylation of the core histones is observed (2). The only histone acetylation ever detected in the micronuclei of actively dividing cells was related to histone deposition, and these acetyl moieties were rapidly removed by a micronuclear HDAC activity (2, 40). Surprisingly, the micronucleus was shown to contain HAT activity. However, this activity is distinct from the transcription-related HAT activities of the macronucleus (40). Whereas macronuclear HATs produce transcription-related acetylation patterns (Lys8 and Lys16 on H4 [25, 36]), micronuclear and cytoplasmic HAT activities (40) produce only deposition-related patterns (Lys4/Lys11 on H4 [8, 27, 46], analogous to Lys5/Lys12 in other organisms). Thus, an HDAC(s) in the micronucleus must maintain the unacetylated state of micronuclear chromatin by removing deposition-related acetylation, the only acetylation observed in micronuclei. Previously, GFP tagging was used to show that Thd1, a class I Tetrahymena HDAC, localized exclusively to the macronucleus and not to the micronucleus (57). The same strategy was used in the present study to identify Thd2 as an enzyme (class II) that localized to the micronucleus, in addition to the macronucleus (Fig. 2). As yet, there is no evidence that Thd2, like class II enzymes in other systems, shuttles between the nucleus and cytoplasm.
Our results indicate that in the micronucleus, Thd2 deacetylates histones H3 and H4. Immunofluorescence experiments on deletion mutants of THD2 (thd2Δ) revealed the presence of acetylated micronuclear histones H3 and H4 at different stages throughout the cell cycle, suggesting that deposition acetylation was retained in the mutant cells (Fig. 8A and B). Antiserum that detected deposition-related acetylation on histone H3 (Lys9 and Lys14 in Tetrahymena) showed that acetylation of these residues was retained in thd2Δ micronuclear chromatin (Fig. 8C). Although it is unlikely, these experiments did not rule out the possibility that micronuclear acetylation in thd2Δ cells resulted from aberrant localization of a transcription-related HAT to micronuclei. However, antiserum against the transcription-related acetyl-Lys16 on H4 (α-H4K16ac) confirmed that Lys16 remained unacetylated in thd2Δ micronuclei (Fig. 8D). Combined with the fact that only deposition acetylation has ever been observed in micronuclei, these results were taken as strong evidence that micronuclear Thd2 deacetylates newly deposited histones. Further support was provided by the finding that THD2a is maximally transcribed during periods of DNA replication (late anaphase) (Fig. 5). The clearest example of this was observed early in the sexual-conjugation cycle (hours ∼5 to 7), when micronuclei undergo two rounds of DNA replication and mitotic division in rapid succession while macronuclei remain in G1. Over this period, there was a pronounced peak of THD2 transcription (Fig. 6B), the same window in which rapid deacetylation of newly synthesized histones occurs (2).
Histone deacetylation and chromatin maturation.
Histone deacetylation is an important event in chromatin maturation (7). When chromatin is replicated and assembled in vivo in the presence of sodium butyrate to inhibit deacetylation, it retains the DNase I sensitivity typical of immature chromatin, a feature that is rapidly lost when chromatin is replicated under normal conditions (7, 44). Similarly, chromatin assembled in vitro with hyperacetylated histones displays increased sensitivity to DNase I (24). The nuclease sensitivity can be attributed in part to reduced H1-mediated internucleosomal interactions that require core histone amino termini (38, 39). Recently, it was found that deacetylation of Lys91 in the globular domain of histone H4 is important for interaction of H3-H4 tetramers with H2A-H2B dimers during nucleosome formation. Lys91 is initially acetylated prior to deposition, and Lys91 deacetylation facilitates the formation of a salt bridge between Lys91 and a glutamic acid residue on histone H2B (10, 60). Outside of these observations, little is known about the role of deacetylation in the chromatin maturation process.
Following nucleosome segregation and de novo formation on replicated DNA strands, some nucleosome remodeling is likely required to position nucleosomes for further chromatin maturation. Interestingly, the predominant splice variant of THD2 (THD2a) contains a putative IPK domain related to Ipk2 in yeast. This putative IPK domain is the only one of its type in the predicted Tetrahymena coding sequences (J. Smith, unpublished data). Recent studies demonstrated a functional link between the IPK signaling pathway and chromatin remodeling; Ipk2 was shown to be involved in the recruitment of chromatin-remodeling complexes for transcription (43, 48). In yeast, mutations in Ipk2, the enzyme that normally converts IP3 to IP4 and IP5, prevented specific gene induction due to failed recruitment of the chromatin-remodeling complexes SWI/SNF and INO80 (48). Thd2 contains most of the residues within the conserved cofactor binding and IP binding domains found in polyphosphate kinases (Fig. 4B). In Ipk2, a D131A mutation (underlined) within the IP kinase motif (PXXXDXKXG … SSLL) impaired chromatin remodeling and gene transcription in yeast (43, 48). Thd2a contains this highly conserved region and the critical aspartate residue within the IPK motif (Fig. 4B). Other than regulating chromatin remodeling, other possible nuclear roles for an Ipk2-like domain include promoting mRNA export (33, 35) and regulating nonhomologous end joining (19). Future studies will address these possibilities, especially in relation to the HDAC activity residing on the same enzyme.
Although this is the first report of an IPK motif on an HDAC enzyme, broad gene database searches revealed other Thd2 homologs with putative IPK domains nearly identical in sequence and structure across the putative polyphosphate binding and kinase motifs. Interestingly, they were found only in other protozoans (Paramecium tetraurelia, Plasmodium falciparum, Plasmodium yoelii, Theileria annulata, and Theileria parva) (data not shown). In each case, like Thd2, the putative IPK domain was at the C terminus of the HDAC enzyme. It is possible that in other organisms, instead of residing on the same polypeptide, IPKs exist as separate polypeptides that complex with HDACs to mediate chromatin maturation.
The alternative splicing of THD2 transcripts yields a variant without the putative IPK domain, a process that could be used to regulate the IPK activity on this HDAC. Splice variant THD2a contains the putative IPK domain, whereas splice variant THD2b does not. Moreover, the two variants are differentially expressed throughout sexual conjugation. Whereas they share a common peak of transcription at 6 hours coincident with micronuclear mitosis, THD2b is expressed to almost the same degree at later times, as well (Fig. 6). In this study, GFP was cloned in frame with the THD2 genomic sequence. It is not clear which form, Thd2a or Thd2b (or both), localizes to the macronucleus; future studies will address whether these forms are differentially localized. Our experiments revealed a possible third splice variant that retains intron 3 (Fig. 3C and 6B). This form, like THD2b, would lack the putative IPK domain. We were unable to rule out the possibility that this variant was simply an unprocessed intermediate of THD2b in which intron 3 had not yet been removed (Fig. 3A). Regardless of whether there are two or three splice variants of THD2, this is the first reported example of an alternative spliced coding sequence from Tetrahymena.
The cytological phenotypes of thd2Δ cells were suggestive of chromatin defects. Although the doubling time of thd2Δ cells was ∼3 times that of wild-type cells (data not shown), nuclear division and cytokinesis appeared normal. However, 50% of elongated micronuclei (a shape typical of anaphase) in thd2Δ cells were in close proximity to or associated with the macronucleus, an aberrant position for an anaphase micronucleus. Determination of whether these micronuclei are arrested in anaphase awaits further studies. A prolonged anaphase would account for the longer population doubling time observed in mutant cells. It is noteworthy that micronuclei also elongate normally when their chromatin decondenses over a short interval early in sexual conjugation. It is thus tempting to speculate that chromatin is less condensed in growing thd2Δ cells, something that could result from defects in chromatin maturation. Although the sizes and shapes of micronuclei in mutant cells were more variable than those in wild-type cells (many were quite small), cells completely lacking a micronucleus were not observed, perhaps due to activation of a mitotic checkpoint upon loss of micronuclear DNA.
Although a more extensive analysis of cytological phenotypes in relation to chromatin maturation awaits future studies, one chromatin abnormality detected in thd2Δ cells was defective proteolytic processing of new histone H3 (H3s). It is still unclear what purpose this processing normally serves; both forms (H3s and H3f) are present in mononucleosomes (3). Specific to histone H3f is the phosphorylation of Ser10, a modification present only in micronuclei during the early stages of mitosis (54). Phosphorylation is not required for formation of the H3f species, as demonstrated with an unphosphorylated H3 mutant (H3S10A), but micronuclei in these cells displayed chromosome segregation defects (55). In contrast, proteolytic processing of H3 was defective in thd2Δ cells; H3f was greatly reduced (Fig. 9B). This result was confirmed by immunoblotting with antiserum against phosphorylated Ser10 on histone H3 (α-H3S10ph). As expected, phosphorylation was greatly reduced compared to that in wild-type cells, consistent with there being less H3f in the thd2Δ cells (Fig. 9B). These findings suggest that Thd2 is required for H3 processing and subsequent phosphorylation. As Ser10 phosphorylation is linked to chromatin condensation and progression of mitosis in many organisms, including Tetrahymena, the reduction in phosphorylation may account for some of the observed growth and cytological phenotypes observed in the mutant cells. Future studies will explore these possible relationships in the context of roles for HDAC enzymes in chromatin maturation and function.
Acknowledgments
We thank Dan Romero and Eric Cole for their assistance with cell elutriation and Meng-Chao Yao for his assistance with biolistic transformation. We are grateful to Doug Chalker for providing the pIGF plasmid and protocols for making the GFP-THD2 construct and to C. David Allis for providing various antisera. We give special acknowledgment to Alison Plumley, Paige Chung, Carey Wickham, and Christine Wong for their supporting experimental and intellectual contributions to this study.
This work was supported by NSF Career Award no. 0545560 to E.A.W.
Footnotes
Published ahead of print on 4 January 2008.
REFERENCES
- 1.Ai, X., and M. R. Parthun. 2004. The nuclear Hat1p/Hat2p complex: a molecular link between type B histone acetyltransferases and chromatin assembly. Mol. Cell 14195-205. [DOI] [PubMed] [Google Scholar]
- 2.Allis, C. D., L. G. Chicoine, R. Richman, and I. G. Schulman. 1985. Deposition-related histone acetylation in micronuclei of conjugating Tetrahymena. Proc. Natl. Acad. Sci. USA 828048-8052. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Allis, C. D., C. V. Glover, and M. A. Gorovsky. 1979. Micronuclei of Tetrahymena contain two types of histone H3. Proc. Natl. Acad. Sci. USA 764857-4861. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Allis, C. D., and M. A. Gorovsky. 1981. Histone phosphorylation in macro- and micronuclei of Tetrahymena thermophila. Biochemistry 203828-3833. [DOI] [PubMed] [Google Scholar]
- 5.Allis, C. D., and J. C. Wiggins. 1984. Histone rearrangements accompany nuclear differentiation and dedifferentiation in Tetrahymena. Dev. Biol. 101282-294. [DOI] [PubMed] [Google Scholar]
- 6.Allis, C. D., and J. C. Wiggins. 1984. Proteolytic processing of micronuclear H3 and histone phosphorylation during conjugation in Tetrahymena thermophila. Exp. Cell Res. 153287-298. [DOI] [PubMed] [Google Scholar]
- 7.Annunziato, A. T., and R. L. Seale. 1983. Histone deacetylation is required for the maturation of newly replicated chromatin. J. Biol. Chem. 25812675-12684. [PubMed] [Google Scholar]
- 8.Chicoine, L. G., I. G. Schulman, R. Richman, R. G. Cook, and C. D. Allis. 1986. Nonrandom utilization of acetylation sites in histones isolated from Tetrahymena. Evidence for functionally distinct H4 acetylation sites. J. Biol. Chem. 2611071-1076. [PubMed] [Google Scholar]
- 9.Cole, E. S., K. R. Stuart, T. C. Marsh, K. Aufderheide, and W. Ringlien. 2002. Confocal fluorescence microscopy for Tetrahymena thermophila. Methods Cell Biol. 70337-359. [DOI] [PubMed] [Google Scholar]
- 10.Cosgrove, M. S., J. D. Boeke, and C. Wolberger. 2004. Regulated nucleosome mobility and the histone code. Nat. Struct. Mol. Biol. 111037-1043. [DOI] [PubMed] [Google Scholar]
- 11.de Ruijter, A. J., A. H. van Gennip, H. N. Caron, S. Kemp, and A. B. van Kuilenburg. 2003. Histone deacetylases (HDACs): characterization of the classical HDAC family. Biochem. J. 370737-749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Doerder, F. P., and L. E. Debault. 1975. Cytofluorimetric analysis of nuclear DNA during meiosis, fertilization and macronuclear development in the ciliate Tetrahymena pyriformis, syngen 1. J. Cell Sci. 17471-493. [DOI] [PubMed] [Google Scholar]
- 13.Duharcourt, S., and M. C. Yao. 2002. Role of histone deacetylation in developmentally programmed DNA rearrangements in Tetrahymena thermophila. Eukaryot. Cell 1293-303. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.English, C. M., M. W. Adkins, J. J. Carson, M. E. Churchill, and J. K. Tyler. 2006. Structural basis for the histone chaperone activity of Asf1. Cell 127495-508. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Gaertig, J., and M. A. Gorovsky. 1992. Efficient mass transformation of Tetrahymena thermophila by electroporation of conjugants. Proc. Natl. Acad. Sci. USA 899196-9200. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Gao, L., M. A. Cueto, F. Asselbergs, and P. Atadja. 2002. Cloning and functional characterization of HDAC11, a novel member of the human histone deacetylase family. J. Biol. Chem. 27725748-25755. [DOI] [PubMed] [Google Scholar]
- 17.Gorovsky, M. A., M. C. Yao, J. B. Keevert, and G. L. Pleger. 1975. Isolation of micro- and macronuclei of Tetrahymena pyriformis. Methods Cell Biol. 9311-327. [DOI] [PubMed] [Google Scholar]
- 18.Gregoretti, I. V., Y. M. Lee, and H. V. Goodson. 2004. Molecular evolution of the histone deacetylase family: functional implications of phylogenetic analysis. J. Mol. Biol. 33817-31. [DOI] [PubMed] [Google Scholar]
- 19.Hanakahi, L. A., M. Bartlet-Jones, C. Chappell, D. Pappin, and S. C. West. 2000. Binding of inositol phosphate to DNA-PK and stimulation of double-strand break repair. Cell 102721-729. [DOI] [PubMed] [Google Scholar]
- 20.Holmes, W., and G. Jogl. 2006. Crystal structure of inositol phosphate multikinase 2 and implications for substrate specificity. J. Biol. Chem. 28138109-38116. [DOI] [PubMed] [Google Scholar]
- 21.Jackson, V., A. Shires, N. Tanphaichitr, and R. Chalkley. 1976. Modifications to histones immediately after synthesis. J. Mol. Biol. 104471-483. [DOI] [PubMed] [Google Scholar]
- 22.Jaeckel-Williams, R. 1978. Nuclear divisions with reduced numbers of microtubules in Tetrahymena. J. Cell Sci. 34303-319. [DOI] [PubMed] [Google Scholar]
- 23.Kornberg, R. D., and Y. Lorch. 1999. Twenty-five years of the nucleosome, fundamental particle of the eukaryote chromosome. Cell 98285-294. [DOI] [PubMed] [Google Scholar]
- 24.Krajewski, W. A., and P. B. Becker. 1998. Reconstitution of hyperacetylated, DNase I-sensitive chromatin characterized by high conformational flexibility of nucleosomal DNA. Proc. Natl. Acad. Sci. USA 951540-1545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kuo, M. H., J. E. Brownell, R. E. Sobel, T. A. Ranalli, R. G. Cook, D. G. Edmondson, S. Y. Roth, and C. D. Allis. 1996. Transcription-linked acetylation by Gcn5p of histones H3 and H4 at specific lysines. Nature 383269-272. [DOI] [PubMed] [Google Scholar]
- 26.Leipe, D. D., and D. Landsman. 1997. Histone deacetylases, acetoin utilization proteins and acetylpolyamine amidohydrolases are members of an ancient protein superfamily. Nucleic Acids Res. 253693-3697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 27.Lin, R., J. W. Leone, R. G. Cook, and C. D. Allis. 1989. Antibodies specific to acetylated histones document the existence of deposition- and transcription-related histone acetylation in Tetrahymena. J. Cell Biol. 1081577-1588. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Lucchini, R., R. E. Wellinger, and J. M. Sogo. 2001. Nucleosome positioning at the replication fork. EMBO J. 207294-7302. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Luger, K., A. W. Mader, R. K. Richmond, D. F. Sargent, and T. J. Richmond. 1997. Crystal structure of the nucleosome core particle at 2.8 Å resolution. Nature 389251-260. [DOI] [PubMed] [Google Scholar]
- 30.Marsh, T. C., E. S. Cole, and D. P. Romero. 2001. The transition from conjugal development to the first vegetative cell division is dependent on RAD51 expression in the ciliate Tetrahymena thermophila. Genetics 1571591-1598. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Marsh, T. C., E. S. Cole, K. R. Stuart, C. Campbell, and D. P. Romero. 2000. RAD51 is required for propagation of the germinal nucleus in Tetrahymena thermophila. Genetics 1541587-1596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Masumoto, H., D. Hawke, R. Kobayashi, and A. Verreault. 2005. A role for cell-cycle-regulated histone H3 lysine 56 acetylation in the DNA damage response. Nature 436294-298. [DOI] [PubMed] [Google Scholar]
- 33.Miller, A. L., M. Suntharalingam, S. L. Johnson, A. Audhya, S. D. Emr, and S. R. Wente. 2004. Cytoplasmic inositol hexakisphosphate production is sufficient for mediating the Gle1-mRNA export pathway. J. Biol. Chem. 27951022-51032. [DOI] [PubMed] [Google Scholar]
- 34.Mitchell, E. J., S. Zimmerman, and A. M. Zimmerman. 1985. Actin and tubulin in Tetrahymena. Can. J. Biochem. Cell Biol. 63389-396. [DOI] [PubMed] [Google Scholar]
- 35.Nalaskowski, M. M., C. Deschermeier, W. Fanick, and G. W. Mayr. 2002. The human homologue of yeast ArgRIII protein is an inositol phosphate multikinase with predominantly nuclear localization. Biochem. J. 366549-556. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Ohba, R., D. J. Steger, J. E. Brownell, C. A. Mizzen, R. G. Cook, J. Cote, J. L. Workman, and C. D. Allis. 1999. A novel H2A/H4 nucleosomal histone acetyltransferase in Tetrahymena thermophila. Mol. Cell. Biol. 192061-2068. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.Parker, K., J. Maxson, A. Mooney, and E. A. Wiley. 2007. Class I histone deacetylase Thd1p promotes global chromatin condensation in Tetrahymena thermophila. Eukaryot. Cell 61913-1924. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Perry, C. A., and A. T. Annunziato. 1989. Influence of histone acetylation on the solubility, H1 content and DNase I sensitivity of newly assembled chromatin. Nucleic Acids Res. 174275-4291. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Perry, C. A., and A. T. Annunziato. 1991. Histone acetylation reduces H1-mediated nucleosome interactions during chromatin assembly. Exp. Cell Res. 196337-345. [DOI] [PubMed] [Google Scholar]
- 40.Richman, R., L. G. Chicoine, M. P. Collini, R. G. Cook, and C. D. Allis. 1988. Micronuclei and the cytoplasm of growing Tetrahymena contain a histone acetylase activity which is highly specific for free histone H4. J. Cell Biol. 1061017-1026. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Robyr, D., Y. Suka, I. Xenarios, S. K. Kurdistani, A. Wang, N. Suka, and M. Grunstein. 2002. Microarray deacetylation maps determine genome-wide functions for yeast histone deacetylases. Cell 109437-446. [DOI] [PubMed] [Google Scholar]
- 42.Ruiz-Carrillo, A., L. J. Wangh, and V. G. Allfrey. 1975. Processing of newly synthesized histone molecules. Science 190117-128. [DOI] [PubMed] [Google Scholar]
- 43.Shen, X., H. Xiao, R. Ranallo, W. H. Wu, and C. Wu. 2003. Modulation of ATP-dependent chromatin-remodeling complexes by inositol polyphosphates. Science 299112-114. [DOI] [PubMed] [Google Scholar]
- 44.Shimamura, A., and A. Worcel. 1989. The assembly of regularly spaced nucleosomes in the Xenopus oocyte S-150 extract is accompanied by deacetylation of histone H4. J. Biol. Chem. 26414524-14530. [PubMed] [Google Scholar]
- 45.Smith, S., and B. Stillman. 1991. Stepwise assembly of chromatin during DNA replication in vitro. EMBO J. 10971-980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 46.Sobel, R. E., R. G. Cook, C. A. Perry, A. T. Annunziato, and C. D. Allis. 1995. Conservation of deposition-related acetylation sites in newly synthesized histones H3 and H4. Proc. Natl. Acad. Sci. USA 921237-1241. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Sogo, J. M., H. Stahl, T. Koller, and R. Knippers. 1986. Structure of replicating simian virus 40 minichromosomes. The replication fork, core histone segregation and terminal structures. J. Mol. Biol. 189189-204. [DOI] [PubMed] [Google Scholar]
- 48.Steger, D. J., E. S. Haswell, A. L. Miller, S. R. Wente, and E. K. O'Shea. 2003. Regulation of chromatin remodeling by inositol polyphosphates. Science 299114-116. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Tamura, K., J. Dudley, M. Nei, and S. Kumar. 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 241596-1599. [DOI] [PubMed] [Google Scholar]
- 50.Van Holde, K. E. 1989. Chromatin. Springer-Verlag, New York, NY.
- 51.Vavra, K. J., C. D. Allis, and M. A. Gorovsky. 1982. Regulation of histone acetylation in Tetrahymena macro- and micronuclei. J. Biol. Chem. 2572591-2598. [PubMed] [Google Scholar]
- 52.Verreault, A., P. D. Kaufman, R. Kobayashi, and B. Stillman. 1996. Nucleosome assembly by a complex of CAF-1 and acetylated histones H3/H4. Cell 8795-104. [DOI] [PubMed] [Google Scholar]
- 53.Waterborg, J. H., S. R. Fried, and H. R. Matthews. 1983. Acetylation and methylation sites in histone H4 from Physarum polycephalum. Eur. J. Biochem. 136245-252. [DOI] [PubMed] [Google Scholar]
- 54.Wei, Y., C. A. Mizzen, R. G. Cook, M. A. Gorovsky, and C. D. Allis. 1998. Phosphorylation of histone H3 at serine 10 is correlated with chromosome condensation during mitosis and meiosis in Tetrahymena. Proc. Natl. Acad. Sci. USA 957480-7484. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 55.Wei, Y., L. Yu, J. Bowen, M. A. Gorovsky, and C. D. Allis. 1999. Phosphorylation of histone H3 is required for proper chromosome condensation and segregation. Cell 9799-109. [DOI] [PubMed] [Google Scholar]
- 56.Wiley, E. A., T. Myers, K. Parker, T. Braun, and M. C. Yao. 2005. Class I histone deacetylase Thd1p affects nuclear integrity in Tetrahymena thermophila. Eukaryot. Cell 4981-990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 57.Wiley, E. A., R. Ohba, M. C. Yao, and C. D. Allis. 2000. Developmentally regulated rpd3p homolog specific to the transcriptionally active macronucleus of vegetative Tetrahymena thermophila. Mol. Cell. Biol. 208319-8328. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 58.Wu, M., C. D. Allis, and M. A. Gorovsky. 1988. Cell-cycle regulation as a mechanism for targeting proteins to specific DNA sequences in Tetrahymena thermophila. Proc. Natl. Acad. Sci. USA 852205-2209. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 59.Yao, M. C., C. H. Yao, L. M. Halasz, P. Fuller, C. H. Rexer, S. H. Wang, R. Jain, R. S. Coyne, and D. L. Chalker. 2007. Identification of novel chromatin-associated proteins involved in programmed genome rearrangements in Tetrahymena. J. Cell Sci. 1201978-1989. [DOI] [PubMed] [Google Scholar]
- 60.Ye, J., X. Ai, E. E. Eugeni, L. Zhang, L. R. Carpenter, M. A. Jelinek, M. A. Freitas, and M. R. Parthun. 2005. Histone H4 lysine 91 acetylation: a core domain modification associated with chromatin assembly. Mol. Cell 18123-130. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 61.Yu, G. L., and E. H. Blackburn. 1990. Amplification of tandemly repeated origin control sequences confers a replication advantage on rDNA replicons in Tetrahymena thermophila. Mol. Cell. Biol. 102070-2080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 62.Zimmerman, A. M., S. Zimmerman, J. Thomas, and I. Ginzburg. 1983. Control of tubulin and actin gene expression in Tetrahymena pyriformis during the cell cycle. FEBS Lett. 164318-321. [DOI] [PubMed] [Google Scholar]
- 63.Zou, H., Y. Wu, M. Navre, and B. C. Sang. 2006. Characterization of the two catalytic domains in histone deacetylase 6. Biochem. Biophys. Res. Commun. 34145-50. [DOI] [PubMed] [Google Scholar]