Abstract
Staphylococcus aureus is widely appreciated as an opportunistic pathogen, primarily in hospital-related infections. However, recent reports indicate that S. aureus infections can now occur in other wise healthy individuals in the community setting. The success of this organism can be attributed to the large array of regulatory proteins, including the SarA protein family, used to respond to changing microenvironments. Sequence alignment and structural data reveal that the SarA protein family can be divided into three subfamilies: 1) single domain proteins; 2) double domain proteins and 3) MarR homologs. Structural studies have also demonstrated that SarA, SarR, SarS, MgrA, and thus possibly all members of this protein family are winged helix proteins with minor variations. Mutagenesis studies of SarA disclose that the winged helix motifs are important for DNA binding and function. Recent progress concerning the functions and plausible mechanisms of regulation of SarA and its homologs are discussed.
Keywords: Stapylococcus, SarA, SarA protein family, gene regulation
Introduction
Staphylococcus aureus is a major pathogen in community acquired skin infections as well as in nosocomial bacteremia (Wenzel et al., 2007). In most cases, the bacteremia is a result of a local infection that has gained access to the bloodstream. Once the bacteria are in the bloodstream, patients are at risk for developing endocarditis and other metastatic complications. Of concern is the development of resistance to methicillin and more recently to vancomycin, which is the drug of last resort for many methicillin resistant S. aureus strains. As a result, there is a great need to understand the molecular pathogenesis of S. aureus in order to develop novel prevention and treatment strategies.
2. Overview of Pathogenesis
S. aureus pathogenesis is a complex process involving a diverse array of extracellular and cell wall components that are coordinately expressed during different stages of infection. For instance, the expression of cell wall adhesins, including protein A, fibrinogen and fibronectin binding proteins, collagen binding proteins etc. [reviewed in (Cheung et al., 2004)] occurs during initial colonization while the production of toxins linked to tissue spread (e.g. hemolysins, toxins, lipase, proteases etc.) takes place late in infection (Fig. 1). The coordinated expression of these virulence factors is controlled by global regulatory elements including: 1) two component regulatory systems (TCRS) of which there are 16 in the N315 genome (Cheung et al., 2004; Novick, 2003; Arvidson and Tegmark, 2001); and 2) the SarA protein family, a collection of DNA binding proteins homologous to SarA, a regulator of virulence gene expression in S. aureus (Cheung et al., 2004).
Fig. 1.
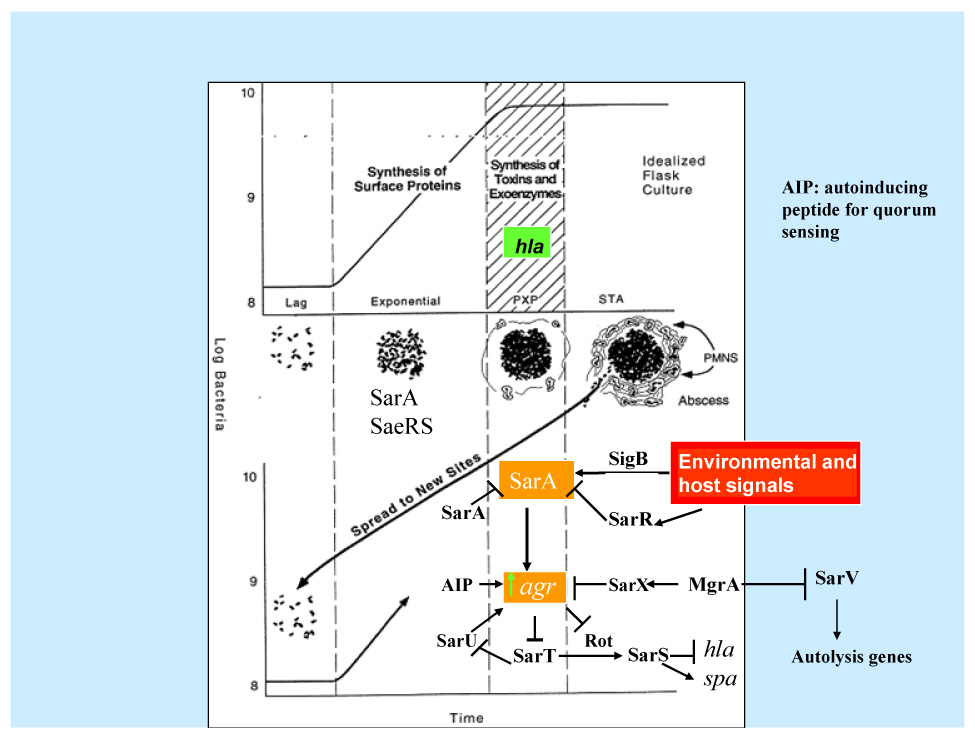
Regulation of virulence determinants in S. aureus by the SarA protein family adapted from Crossley and Archer (Crossley and Archer, 1997). Normally, the synthesis of cell surface adhesins such as fibronectin binding protein A (FnbA) during the exponential phase coincides with the expression of SarA and SaeRS. In transition from exponential to postexponential phase, the synthesis of cell wall proteins is disrupted and the production of extracellular toxins such as a-toxin would begin. This transition corresponds to the maximal expression of SarA and the ensuing activation of agr. SarA expression is repressed by SarA and SarR. SigB, a stress-induced transcription factor, also activates one of the sarA promoter (the P3 promoter). On the other hand, agr is controlled by SarA, a quorum sensing autoinducing peptide, other TCRS (see Table 1), MgrA, SarX and SarU. Activation of agr would lead to up-regulation of another TCRS system called SaeRS and down-regulation of a SarA protein homolog called Rot. This will eventually lead to repression of two gene products called SarT and subsequently SarS. SarT is an activator of SarS, which is a repressor of alpha toxin production and an activator of protein A synthesis, thus explaining the elevated production of α-toxin and repression of protein A upon agr activation. Activation of agr would also result in the amplification of the original signal by activating SarU, which is a positive regulator of agr.
In this review, we provide an overview of S. aureus pathogenesis, focusing in particular on the newest findings on the regulation cascades and the structure and function of the SarA protein family members (Cheung et al., 2004). The prototypic member of the SarA protein family is SarA, a 124-residue DNA binding protein that enhances postexponential expression of α, β and δ toxins both directly and indirectly via up-regulation of agr (Cheung et al., 2004). The agr locus is a TCRS that acts as molecular switch to turn on transcription of toxin genes while repressing synthesis of cell wall protein postexponentially in S. aureus. For additional analysis of TCRS and other regulatory factors, readers are referred to three recent reviews (Novick, 2003; Arvidson and Tegmark, 2001; Cheung et al., 2004).
3. Structure of the SarA protein family
SarA is a 124-residue DNA binding protein encoded by the sarA locus, which consists of three overlapping transcripts, driven by three distinct romoters, P1, P3 and P2 (Bayer et al., 1996). Phenotypically, the sarA locus promotes synthesis of fibronectin and fibrinogen binding proteins (for adhesion) and also toxins (for tissue spread) while repressing expression of protein A and proteases (Cheung et al., 2004). DNA binding and profiling studies suggest that the SarA protein may regulate target genes by directly binding to target gene promoters or indirectly via downstream effects on regulons (e.g. binding to the agr promoter) or by stabilizing mRNA during log phase (Cheung et al., 2004; Roberts et al., 2006).
The current crystal structure of SarA supports the notion of a winged helix protein (Liu et al., 2006) and differs drastically from the previously described SarA structure (Schumacher et al., 2001). SarA, like its homolog SarR, is a dimeric winged helix structure with each monomer consisting of 5 α-helices, 3 β-strands and several loops (α1α2-β1α3α4-β2β3-α5). The SarA dimer possesses a central helical core and two winged helix motifs (Fig. 2). Within each winged helix motif is a helix-turn-helix motif (α3α4) and a β-hairpin turn wing (β2β3), both of which are putative DNA binding domains (Liu et al., 2006).
Fig. 2.
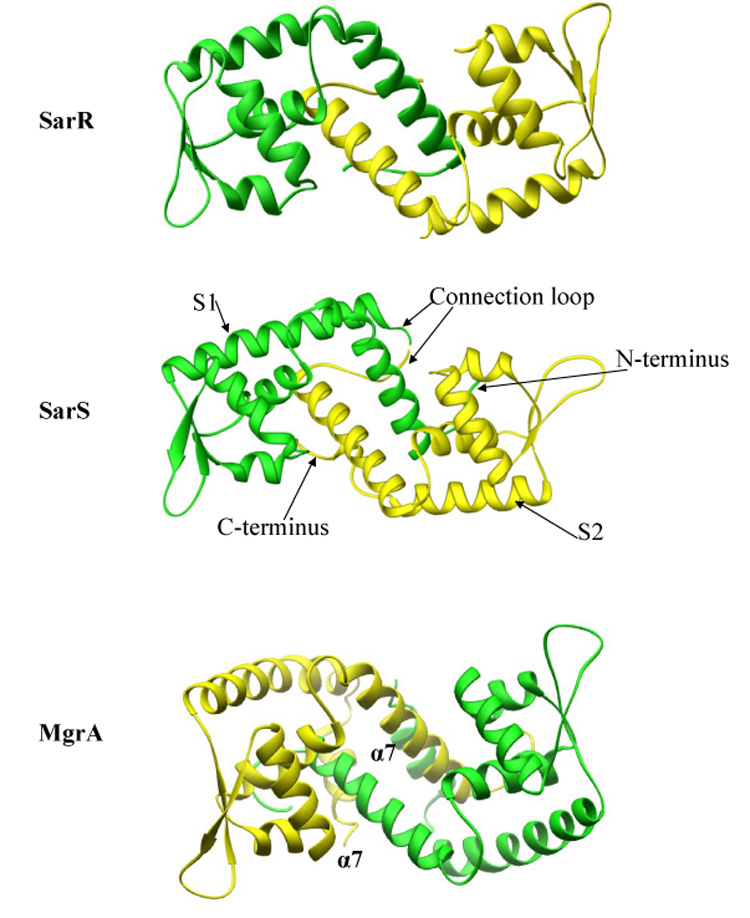
Comparison of the SarR, SarS and MgrA winged helix structures. The SarR structure is composed of two identical monomers, one in green and the other in yellow. Each monomer is composed of five α-helices and three β-strands. In contrast, SarS is composed of two homologous but non-identical halves connected by a linker region. MgrA has 7 α-helices and three β-strands. There are coiled coil interactions between two α7 helices from the monomers.
SarS is a 250-residue DNA binding protein that activates protein A gene (spa) transcription (Table 1) (Tegmark et al., 2000; Cheung et al., 2004). Unlike the dimeric structures of SarA and SarR, SarS forms a monomeric structure with two homologous but non-identical halves (Fig. 3A) connected by a well ordered loop, forming an overall winged helix structure similar to SarA and SarR (Fig. 2).
Table 1.
The SarA protein family (N315 genome)
| Locus | Gene symbols | Protein subfamily | Putative function |
|---|---|---|---|
| SA0573 | SarA | single domain (124 residues) | Activates genes via agr and agr independent pathways |
| SA2089 | SarR | single domain (115 residues) | A negative regulator of sarA and positive activator of agr (Cheung et al., 2004) |
| SA0108 | SarS | two domain (250 residues) | An activator of protein A synthesis (Tegmark et al., 2000) |
| SA2286 | SarT | single domain (118 residues) | An activator of sarS and a repressor of alpha hemolysin synthesis (Cheung et al., 2004) |
| SA2287 | SarU | two domain (247 residues) | A positive regulator of agr (Cheung et al., 2004) |
| SA1583 | Rot | single domain (133 residues) | A repressor of toxin synthesis, opposite to agr (Said-Salim et al., 2003) |
| SA0623 | SarX | single domain (141 residues) | A negative regulator of agr and repressed by MgrA (Manna and Cheung, 2006a) |
| SA0641 | MgrA | MarR homolog (147 residues) | A regulator of autolysis and agr (also called Rat or NorR) |
| (Truong-Bolduc et al., 2003) | |||
| SA2174 | SarZ | MarR homolog (148 residues) | A positive regulator of hla (Kaito et al., 2006) |
| SA2062 | SarV | single domain (116 residues) | A regulator of autolysis repressed SarA and MgrA (Manna et al., 2004) |
| SA2091 | SarY | two domain (247 residues) | Function unknown |
Fig. 3.
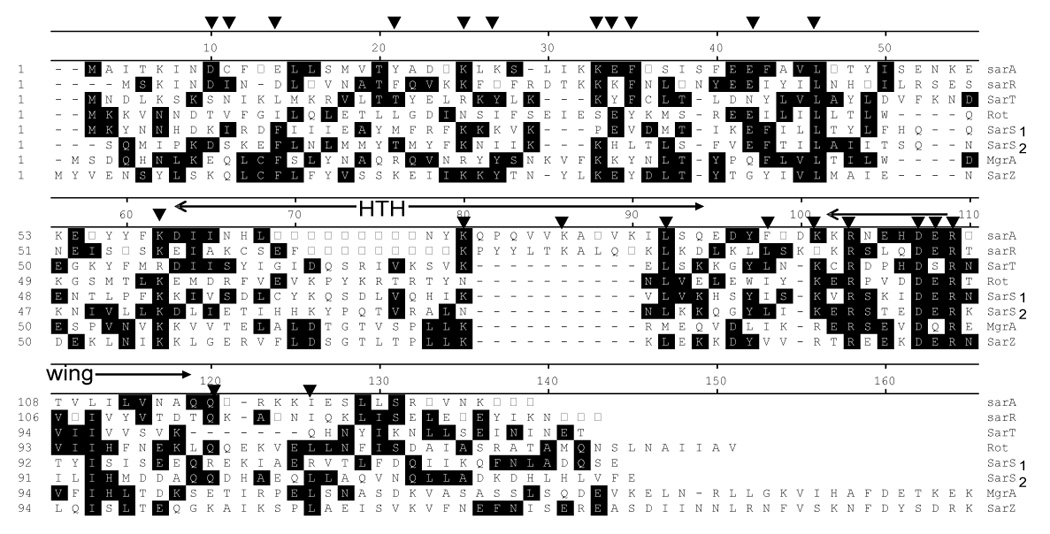
Sequence alignment of members of the SarA protein family. SarA, SarR, SarT and Rot belong to the single domain proteins. Rot is unique for its larger size and an acidic pI. SarS represents a two domain protein, with two homologous halves connected by a linker region. SarS1 and SarS2 represent the N-terminal and C-terminal halves, respectively. MgrA and SarZ are more homologous to the MarR protein family of Gram negative bacteria than to the SarA protein family and thus represent the third subfamily within the SarA protein family. The HTH and the wing regions of the winged helix structures are indicated. The residues (▼) altered by mutagenesis in SarA have been highlighted (Liu et al., 2006).
Besides SarA, SarR and SarS, there are eight other SarA homologs in the N315 genome, collectively called the SarA protein family (Table 1). Based on size, the SarA protein family members can be classified into two major groups. The larger proteins such as SarS, SarY and SarU represent two-domain proteins, with each domain homologous to the smaller SarA homologs (e.g. SarA) (Fig. 3). Two of the smaller SarA homologs, MgrA and SarZ, have more sequence similarity to the MarR protein of Gram-negative bacteria than to other smaller SarA proteins (Ingavale et al., 2005). Thus, the SarA protein family can be classified into three subfamilies: 1) single domain proteins (SarA, -R, -T, -V and –X and Rot); 2) double domain proteins (SarS, -U and -Y); and 3) MarR homologs (MgrA and SarZ) (Table 1). Indeed, recent structural data disclosed that MgrA is also a dimeric winged helix protein but contains unique structural features similar to those of MarR than to SarA homologs (Table 1) (Cheung et al., 2004).
4a. Functions and mutational analysis of the SarA protein
Superimposing the secondary structure of SarA with sequence alignment of several SarA homologs (Fig. 3A) reveals the following conserved residues: 1) basic residues within the α3-α4 helix turn helix; 2) DER residues (residues 88-90 in SarA) in the β2-L-β3 wing region; 3) unique cysteine residue in many SarA homologs (SarA, SarR, SarS, SarU, SarX, SarZ, MgrA and Rot); and 4) L40, L74, Q100 and I104 in SarA outside the winged helix motif. Studies with mutated forms of SarA revealed the following: 1) mutations of Y18, E29, E36, K54, L74 and F80 to alanine somewhat reduced binding to the target spa promoter, but did complement the sarA mutant phenotype; 2) R84A and R90A (within the winged region) mutant proteins failed to bind to the spa promoter and, as expected, did not complement the sarA mutant; 3) mutations of D88A and E89A (part of the conserved DER motif) exhibited normal binding to the spa promoter but did not restore the sarA phenotype to a sarA mutant; 4) C9A and K23A mutations, which are not part of the winged helix motif of SarA, exhibited normal DNA binding but did not reestablish the wild type sarA phenotype in the mutant (Liu et al., 2006). K23 lies within the α1 helix involved in dimerization of two SarA monomers, thus implying a plausible dimerization defect in the K23A mutant may interfere with SarA function but not binding activity. Of interest is the C9A mutation which has a phenotype similar to the K23A mutant. Notably, the homologous unique cysteine residue in MgrA has been implicated as an oxidation sensor, required for DNA promoter binding (Chen et al., 2006).
4b. Plausible mechanism of regulation by SarA
Structural data suggest that members of the SarA protein family likely bind to target DNA in a similar fashion (Liu et al., 2006), with the helix-turn-helix binding to the major groove of DNA while the winged region interacts with the minor groove. In contrast to the binding domains, the activation domains differ among family members, probably reflecting the divergent functions of different SarA homologs. Based on the structures of SarA, SarR, SarS and MgrA, there are five possible mechanisms of regulation: 1) the oxidation sensing mechanism for the cysteine residue as proposed for MgrA (Chen et al., 2006); 2) bending of target DNA (e.g. by SarR) to facilitate contact of DNA with the regulatory protein (Cheung et al., 2004); 3) the formation of three SarA dimers encased by the promoter DNA, resulting in closed configuration not amenable to transcription (i.e. repression state of SarA) (Liu et al., 2006); 3) the formation of a heterodimer, similar to SarS, between compatible family members (e.g. SarA and SarR) that may interfere with the function of the homodimer (Cheung et al., 2004); 4) competitive displacement of one homolog by another as has been seen with displacement of SarR by SarA from the agr promoter (Manna and Cheung, 2006b).
4c. Other SarA homologs
SarR is a 115-residue DNA binding protein that has been shown to bind to the sarA promoter to repress SarA expression (Cheung et al., 2004). Inactivation of sarR also affects regulatory genes including agr (Manna and Cheung, 2006b).
SarS is a 250-residue two-domain SarA homolog that activates spa and represses hla transcription (Tegmark et al., 2000; Cheung et al., 2004)}. Transcriptional analysis indicated that sarS is repressed by agr and mgrA and activated by ClpXP protease (Frees et al., 2005). A recent study suggested that SarA may repress rather than activate hla by up-regulating sarS (Oscarsson et al., 2006); however, these data were generated primarily from strain SH1000, a derivative of 8325-4 that is not a natural isolate with a repaired rsbU gene and little alpha toxin production. Thus, these data would require additional confirmation in clinical isolates.
SarT is a 118-residue single domain SarA homolog that is normally repressed by sarA and agr. Activation of sarT results in up-regulation of sarS (Cheung et al., 2004), thus leading to hla repression and spa activation.
Adjacent but divergently transcribed from sarT is sarU, which encodes a 247-residue two-domain protein. sarU is repressed by SarT (Cheung et al., 2004), which, in turn, is down-modulated by agr. Since sarU is an activator of agr expression, this will lead to amplification of the original agr signal. This finding is consistent with the finding that agr promoter can be activated in vivo in an agr deletion mutant.
Rot was originally described as a 166-residue SarA homolog that represses toxin synthesis and up-regulates cell wall protein synthesis (e.g. clumping factor, SdrC etc.) (Said-Salim et al., 2003). Recent characterization of the rot transcript by primer extension, coupled with expression studies, reveals that Rot is likely a 133-residue protein (Manna and Ray, 2007). Transcriptional profiling studies indicate that Rot affects the transcription of 168 genes (Said-Salim et al., 2003), many of which reflect an agr minus phenotype. A recent study suggested that the agr RNAIII molecule may interact with rot mRNA to inhibit Rot translation (Geisinger et al., 2006).
SarX is a 141-residue single-domain SarA homolog. MgrA, a MarR homolog (Table 1), is an activator of sarX expression (Manna and Cheung, 2006a). SarX can also act as a repressor of agr, thus implying an additional regulatory loop whereby mgrA can modulate agr expression.
MgrA is a 147-residue SarA homolog within the MarR subfamily. Mutation of mgrA resulted in increased autolysis and decreased in agr, sarS and sarX transcription (Ingavale et al., 2005). Transcriptional profiling studies disclosed that MgrA regulates 355 genes, with up-regulation in 175 and down-regulation in 180 (Luong et al., 2006). In particular, exoproteins are up-regulated and surface proteins are down-regulated by MgrA, consistent with an agr positive phenotype (Ingavale et al., 2005). The effect of MgrA on autolysis may be mediated by sarV which is a positive regulator of several autolytic enzymes (see below).
Another SarA homolog is SarZ, a 148-residue protein within the MarR homolog subfamily (Cheung et al., 2004). Kaito et al identified SarZ (Kaito et al, 2006), based on its ability to restore hemolysis activity in a S. aureus cvfA mutant defective in hla expression. A sarZ mutant of RN4220 had attenuated virulence in both silk worm and mouse infection models. However, these results must be interpreted with caution as RN4220 is a S. aureus strain that has been heavily mutagenized with nitrosoguanidine. When the sarZ mutation was transduced into the 8325-4 strain (rsbU and SigB defective), the decrease in hemolysis was marginal and was only partially complemented with sarZ provided in trans, thus suggesting that gene(s) downstream of sarZ may contribute to the hemolytic phenotype.
SarV is a 116-residue single domain SarA homolog. The expression of sarV is low or undetectable under in vitro growth conditions but is significantly augmented in mgrA and sarA mutants, consistent with repression of sarV by mgrA and sarA (Cheung et al., 2004). Contrary to mgrA and sarA mutants, the sarV mutant was more resistant to detergent- or cell wall antibiotic-mediated lysis compared to the wild type while hyperexpression of sarV rendered the parental strain more susceptible to lysis. Accordingly, sarV may be part of the common pathway by which mgrA and sarA control autolysis in S. aureus.
5. Future prospects and potential clinical application
Due to the confluence of growth phase, environmental and host signals, the regulation of virulence determinants in S. aureus is inherently complex. Genomic analysis revealed that many of the SarA homologs are also present in other staphylococcal species including S. epidermidis, S. hemolyticus and S. saprophyticus. We speculate many of these homologs are likely winged helix structures. However, the regulation pathways may differ as many virulence genes present in S. aureus are absent in other staphylococcal species. While most of the known winged helix proteins in prokaryotes are repressive in nature (e.g. MarR of E. coli), the exact mechanism by which SarA and related family members repress and activate target genes remains unknown. While we propose in this review plausible mechanisms for activation and repression, ultimately, co-crystallization studies of SarA with activated and repressed promoters will likely provide insights into the regulatory paradigm.
Finally, we have very incomplete knowledge on the host and environmental signals that trigger the activation or repression of virulence genes. With the availability of GFP and Lux reporter genes in S. aureus, it is now possible to monitor real-time gene expression in vivo during infection. These studies should advance our understanding on the in vivo activation of virulence genes controlled by the SarA protein family in S. aureus. Our long term goal is to disrupt the relevant regulatory pathways as a mean of developing novel antibiotics.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References
- Arvidson S, Tegmark K. Regulation of virulence determinants in Staphylococcus aureus. Int. J. Med. Microbiol. 2001;291:159–170. doi: 10.1078/1438-4221-00112. [DOI] [PubMed] [Google Scholar]
- Bayer MG, Heinrichs JH, Cheung AL. The molecular architecture of the sar locus in Staphylococcus aureus. J. Bacteriol. 1996;178:4563–4570. doi: 10.1128/jb.178.15.4563-4570.1996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chen PR, Bae T, Williams WA, Duguid EM, Rice PA, Schneewind O, He C. An oxidation-sensing mechanism is used by the global regulator MgrA in Staphylococcus aureus. Nat. Chem. Biol. 2006;2:591–595. doi: 10.1038/nchembio820. [DOI] [PubMed] [Google Scholar]
- Cheung AL, Bayer AS, Zhang G, Gresham H, Xiong Y-Q. Regulation of virulence determinants in vitro and in vivo in Staphylococcus aureus. FEMS Microbiol.Lett. 2004;1649:1–9. doi: 10.1016/S0928-8244(03)00309-2. [DOI] [PubMed] [Google Scholar]
- Crossley KB, Archer GL. The staphylococci in human disease. New York: Churchill Livingston; 1997. [Google Scholar]
- Frees D, Sorensen K, Ingmer H. Global virulence regulation in Staphylococcus aureus: pinpointing the roles of ClpP and ClpX in the sar/agr regulatory network. Infect. Immun. 2005;73:8100–8108. doi: 10.1128/IAI.73.12.8100-8108.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Geisinger E, Adhikari RP, Jin R, Ross HF, Novick RP. Inhibition of rot translation by RNAIII, a key feature of agr function. Mol. Microbiol. 2006;61:1038–1048. doi: 10.1111/j.1365-2958.2006.05292.x. [DOI] [PubMed] [Google Scholar]
- Ingavale S, van WW, Luong TT, Lee CY, Cheung AL. Rat/MgrA, a regulator of autolysis, is a regulator of virulence genes in Staphylococcus aureus. Infect.Immun. 2005;73:1423–1431. doi: 10.1128/IAI.73.3.1423-1431.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaito C, Morishita D, Matsumoto Y, Kurokawa K, Sekimizu K. Novel DNA binding protein SarZ contributes to virulence in Staphylococcus aureus. Mol. Microbiol. 2006;62:1601–1617. doi: 10.1111/j.1365-2958.2006.05480.x. [DOI] [PubMed] [Google Scholar]
- Liu Y, Manna AC, Pan CH, Kriksunov IA, Thiel DJ, Cheung AL, Zhang G. Structural and function analyses of the global regulatory protein SarA from Staphylococcus aureus. Proc. Natl. Acad. Sci. U. S. A. 2006;103:2392–2397. doi: 10.1073/pnas.0510439103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luong TT, Dunman PM, Murphy E, Projan SJ, Lee CY. Transcription Profiling of the mgrA Regulon in Staphylococcus aureus. J. Bacteriol. 2006;188:1899–1910. doi: 10.1128/JB.188.5.1899-1910.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manna AC, Cheung AL. Expression of SarX, a negative regulator of agr and exoprotein synthesis, is activated by MgrA in Staphylococcus aureus. J. Bacteriol. 2006a;188:4288–4299. doi: 10.1128/JB.00297-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Manna AC, Cheung AL. Transcriptional regulation of the agr locus and theidentification of DNA binding residues of the global regulatory protein SarR in Staphylococcus aureus. Mol. Microbiol. 2006b;60:1289–1301. doi: 10.1111/j.1365-2958.2006.05171.x. [DOI] [PubMed] [Google Scholar]
- Manna AC, Ray B. Regulation and characterization of rot transcription in Staphylococcus aureus. Microbiology. 2007;153:1538–1545. doi: 10.1099/mic.0.2006/004309-0. [DOI] [PubMed] [Google Scholar]
- Novick RP. Autoinduction and signal transduction in the regulation of staphylococcal virulence. Mol. Microbiol. 2003;48:1429–1449. doi: 10.1046/j.1365-2958.2003.03526.x. [DOI] [PubMed] [Google Scholar]
- Oscarsson J, Kanth A, Tegmark-Wisell K, Arvidson S. SarA is a repressor of hla (alpha-hemolysin) transcription in Staphylococcus aureus: its apparent role as an activator of hla in the prototype strain NCTC 8325 depends on reduced expression of sarS. J. Bacteriol. 2006;188:8526–8533. doi: 10.1128/JB.00866-06. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Roberts C, Anderson KL, Murphy E, Projan SJ, Mounts W, Hurlburt B, Smeltzer M, Overbeek R, Disz T, Dunman PM. Characterizing the effect of the Staphylococcus aureus virulence factor regulator, SarA, on log-phase mRNA half-lives. J. Bacteriol. 2006;188:2593–2603. doi: 10.1128/JB.188.7.2593-2603.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Said-Salim B, Dunman PM, McAleese FM, Macapagal D, Murphy E, McNamara PJ, Arvidson S, Foster TJ, Projan SJ, Kreiswirth BN. Global regulation of Staphylococcus aureus genes by Rot. J. Bacteriol. 2003;185:610–619. doi: 10.1128/JB.185.2.610-619.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schumacher MA, Hurlburt BK, Brennan RG. Crystal structures of SarA, a pleiotropic regulator of virulence genes in S. aureus. Nature. 2001;409:215–219. doi: 10.1038/35051623. [DOI] [PubMed] [Google Scholar]
- Tegmark K, Karlsson A, Arvidson S. Identification and characterization of SarH1, a new global regulator of virulence gene expression in Staphylococcus aureus. Mol. Microbiol. 2000;37:398–409. doi: 10.1046/j.1365-2958.2000.02003.x. [DOI] [PubMed] [Google Scholar]
- Truong-Bolduc QC, Zhang X, Hooper DC. Characterization of NorR protein, a multifunctional regulator of norA expression in Staphylococcus aureus. J. Bacteriol. 2003;185:3127–3138. doi: 10.1128/JB.185.10.3127-3138.2003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wenzel RP, Bearman G, Edmond MB. Community-acquired methicillin-resistant Staphylococcus aureus (MRSA): new issues for infection control. Int. J. Antimicrob. Agents. 2007;30:210–212. doi: 10.1016/j.ijantimicag.2007.04.003. [DOI] [PubMed] [Google Scholar]


