Abstract
The molecular bases of circadian clocks have been studied in animals, fungi, bacteria, and plants, but not in eukaryotic algae. To establish a new model for molecular analysis of the circadian clock, here we identified a large number of components of the circadian system in the eukaryotic unicellular alga Chlamydomonas reinhardtii by a systematic forward genetic approach. We isolated 105 insertional mutants that exhibited defects in period, phase angle, and/or amplitude of circadian rhythms in bioluminescence derived from a luciferase reporter gene in their chloroplast genome. Simultaneous measurement of circadian rhythms in bioluminescence and growth rate revealed that some of these mutants had defects in the circadian clock itself, whereas one mutant had a defect in a specific process for the chloroplast bioluminescence rhythm. We identified 30 genes (or gene loci) that would be responsible for rhythm defects in 37 mutants. Classification of these genes revealed that various biological processes are involved in regulation of the chloroplast rhythmicity. Amino acid sequences of six genes that would have crucial roles in the circadian clock revealed features of the Chlamydomonas clock that have both partially plant-like and original components.
[Keywords: Chlamydomonas reinhardtii, circadian rhythm, bioluminescence, insertional mutagenesis, clock gene]
The molecular bases of circadian clocks have been studied in animals, fungi, bacteria, and plants (Dunlap 1999; Harmer et al. 2001). Despite the striking biochemical features of circadian clocks (e.g., oscillation with long periodicity [∼24 h] and its temperature compensation) (Bünning 1973), their central components are not conserved between these kingdoms (Dunlap 1999; Harmer et al. 2001). To understand the nature of oscillation mechanisms and the evolutionary history of clock components, it is important to understand circadian clock systems of a wide range of organisms.
Circadian rhythms of unicellular algae have been studied extensively (Mittag 2001), but no clock component of eukaryotic algae has yet been identified. Chlamydomonas reinhardtii is one of the best-studied algae in circadian rhythm research. A forward genetic approach to identify circadian clock components of Chlamydomonas was started more than three decades ago (Bruce 1970). Although several clock mutants have been isolated (Bruce 1972, 1974; Mergenhagen 1984), the genes responsible could not be identified because of limitations of tools for genetic analyses. However, since Chlamydomonas is now one of the most attractive model organisms in molecular genetics (Harris 2001), it is possible to re-establish it as a model for studying the molecular mechanism of the circadian clock. For this purpose, we previously developed bioluminescence reporter strains with a codon-optimized luciferase gene in their chloroplast genomes to enable real-time monitoring of circadian rhythms (Breton and Kay 2006; Matsuo et al. 2006).
In this study, we screened ∼16,000 insertional mutants for defects in circadian rhythmicity of the chloroplast bioluminescence reporter, isolated 105 mutants, and identified 30 genes (or gene loci) that would be responsible for rhythm defects in 37 mutants. Classification of these genes revealed that various biological processes, including flagellar function, ubiquitin–proteasome, transcription and transcript metabolism, gene silencing, membrane trafficking and transport, signal transduction, DNA damage response, and apoptosis, are involved in regulation of the chloroplast rhythmicity. Amino acid sequences of six genes that would have crucial roles in the circadian clock revealed features of the Chlamydomonas clock that have both partially plant-like and original components.
Results
Screening of a genetic background suitable for a high-throughput assay of bioluminescence rhythms
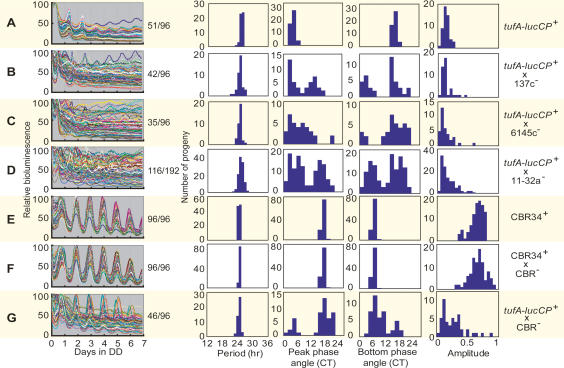
We measured bioluminescence rhythms of a reporter strain (tufA-lucCP+; the plus symbol indicates plus mating type [mt+]) that expresses the luciferase gene under the control of the circadian-regulated chloroplast tufA promoter (Hwang et al. 1996) under high-throughput assay conditions developed in this study (on agar media in constant darkness [DD]) (see Materials and Methods). However, we failed to detect significant circadian rhythms in about half of the preparations because the rhythms were not robust under the conditions (Fig. 1A). To overcome this problem, we screened for a genetic background suitable for the assay conditions. The tufA-lucCP+ strain was crossed with the minus mating type cells of genetically different wild-type strains (137c−, 6145c−, and 11-32a−) (Harris 2001), and the progenies were subjected to the high-throughput bioluminescence assay. Surprisingly, the progenies exhibited various bioluminescence rhythm patterns (Fig. 1B–D). Rhythm amplitudes were quite variable, especially in the offspring from the cross with 11-32a− (Fig. 1B–D). Phase angles of the progenies from crosses with 137c− and 11-32a− were divided into two groups (Fig. 1B,D). On the other hand, period lengths were relatively maintained in the circadian range (Fig. 1B,D). We chose a clone that exhibited a robust rhythm from the cross with 11-32a−. All 96 independent preparations from the clone exhibited stable and robust rhythms (Fig. 1E). We named the clone CBR34+ and used it as a wild-type strain in this study. To generate a mating type minus equivalent of CBR34+, the 11-32a− strain was crossed four times with CBR34+ to obtain a CBR− strain. All progenies from a cross between CBR34+ and CBR− exhibited robust rhythms (Fig. 1F), whereas the progenies from a cross between the original tufA-lucCP+ and CBR− exhibited various bioluminescence rhythm patterns (Fig. 1G). These results indicate that the genetic background of the reporter strain strongly affects the circadian rhythmicity of chloroplast bioluminescence, and that the CBR34 genetic background is suitable for high-throughput screening for circadian rhythm mutants and for their genetic backcross analysis. Moreover, these results imply that multiple genes are involved in maintaining the robustness of chloroplast bioluminescence rhythms.
Figure 1.
Screening of a genetic background suitable for the high-throughput assay of bioluminescence rhythms. All samples were prepared according to the high-throughput assay protocol (see Materials and Methods). Traces of bioluminescence from the 96 samples on agar media are shown on the left, and numbers of samples (per total samples examined) that exhibited a significant circadian rhythmicity (error value [calculated by the RAP] ≤ 0.1) are indicated on the right of the traces. Histograms of period length, peak phase angle, bottom phase angle, and amplitude of the rhythmic samples are shown. Strains or pairs of the crosses are shown on the right.
Screening and genetic analysis of circadian rhythm mutants
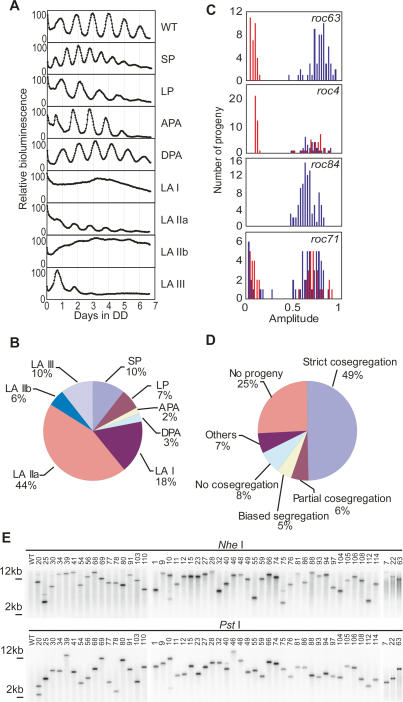
To generate circadian rhythm insertional mutants, the CBR34+ strain was transformed with a DNA fragment containing a hygromycin-resistant gene (Berthold et al. 2002). We screened ∼16,000 transformants for defects in period length, phase angle, and amplitude of bioluminescence rhythm, and isolated 115 candidate rhythm mutants. Then, 91% of the candidates (105 mutants) that reproduced their mutant phenotypes were analyzed further as rhythm of chloroplast (roc) mutants (Supplemental Fig. 1; Supplemental Table 1). The roc mutants were divided into five types (Fig. 2A): short period (SP), long period (LP), advanced phase angle (APA), delayed phase angle (DPA), and low amplitude (LA). The LA mutants could be further divided into four types (Fig. 2A): exhibiting arrhythmic or very weak oscillations for a few cycles (LA I), sustained low-amplitude rhythm with a relatively low bioluminescence level (LA IIa), sustained low-amplitude rhythm with a relatively high bioluminescence level (LA IIb), and damped rapidly (LA III). Interestingly, 78% of isolated mutants were LA (Fig. 2B). This result supports the idea that multiple genes are involved in maintaining the robustness of chloroplast bioluminescence rhythms. Next, all of the mutants were backcrossed to the CBR− strain, and progenies were subjected to bioluminescence rhythm measurements and hygromycin resistance testing (cosegregation analysis) (Supplemental Fig. 2; Supplemental Table 1). If the mutation was caused by a single integration of the marker gene into the genome, the mutant phenotype should have strictly cosegregated with the hygromycin resistance. roc63 is representative of a “strict cosegregation” mutant in which almost all hygromycin-resistant progeny exhibited a mutant phenotype (Fig. 2C). If the integration occurred at multiple loci, only a proportion of hygromycin-resistant progeny (∼50% if integration occurred at two loci) should have exhibited the mutant phenotype. Thus, “partial cosegregation” was observed, as seen in roc4 (Fig. 2C). roc84 is representative of a “biased segregation” mutant in which all progenies were hygromycin-sensitive or -resistant and all exhibited wild-type or mutant phenotypes, respectively (Fig. 2C). If the mutation was independent from the integration of the marker gene, cosegregation should not have been observed. roc71 is representative of “no cosegregation” (Fig. 2C). Strict cosegregation was observed in 52 mutants (49%), whereas no cosegregation was observed in only eight mutants (8%) (Fig. 2D). The mutant phenotypes of most of the strict-cosegregation mutants were segregated into mt+ and mt− progenies (Supplemental Table 1), excluding the possibility that the strict-cosegregation pattern is an artifact of the random progeny analysis caused by surviving parental strains. We could not obtain progenies from crosses in 27 mutants (25%) (Fig. 2D). Most of these mutants exhibited a palmelloid phenotype or precipitated at the bottom in liquid cultures (Supplemental Table 1), suggesting an abnormality in hatching processes or flagellar functions. Although we used 300 ng and 30 ng of marker DNA to generate insertional mutants, there was no apparent difference in the rate of appearance of strict-cosegregation mutants (300 ng, 47.9% [23 of 48]; 30 ng, 50.9% [29 of 57]). Strict cosegregation does not necessarily mean the single integration of a marker gene, because the cosegregation analysis would not reveal additional integrations of the marker gene into silent loci. To determine the number of integration sites, we performed a Southern blot analysis of genomic DNA digested with NheI or PstI, which do not cut the marker gene itself. Except for the PstI-digested roc20 genome, single bands were detected in 50 strict cosegregation mutants (Fig. 2E). These results suggest that the responsible genes of the 50 mutants were tagged by a single integration of the marker gene.
Figure 2.
Screening and genetic analysis of circadian rhythm mutants. (A) Representative bioluminescence traces of several types of roc mutants. (SP) roc55; (LP) roc66; (APA) roc83; (DPA) roc25; (LA I) roc77; (LA IIa) roc39; (LA IIb) roc109; (LA III) roc76. The maximum values are expressed as 100. (B) Pie chart showing the distribution of all mutants by phenotype. (C) Representative results of the cosegregation analysis. Histograms of hygromycin-resistant (red) and hygromycin-sensitive (blue) progenies are shown. (D) Pie chart showing the distribution of all mutants by segregation pattern. (E) Southern blot analysis. The 3′ region of the aph7″ coding sequence was used as a probe. The positions of size markers (2 and 12 kb) are also shown on the left.
Circadian rhythms in the growth rate of isolated mutants
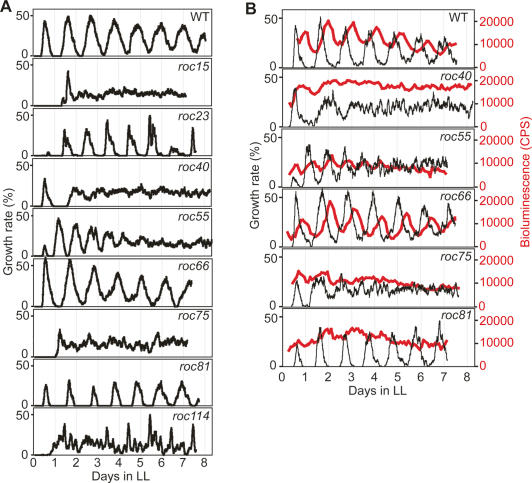
To characterize isolated mutants further, we measured circadian rhythms in cell growth in liquid cultures under continuous light (LL) in several mutants. If these mutants had a defect in the circadian clock itself, the circadian rhythms in growth would also be affected. The wild-type strain exhibited a robust growth rhythm (Fig. 3A). The mutants roc55, roc75, and roc114 showed defects in growth rhythm similar to those in chloroplast bioluminescence rhythm on agar in DD (i.e., short period for roc55 and low amplitudes for roc75 and roc114) (Fig. 3A; Supplemental Fig. 1). In contrast, growth rhythms of the other mutants exhibited different phenotypes to some extent from their bioluminescence rhythms. The short period mutant roc15 and slightly short period mutant roc40 no longer exhibited robust growth rhythmicity (Fig. 3A; Supplemental Fig. 1). Interestingly, the interval between the first and second peaks of growth rhythm for roc40 was extremely long, although roc40 was a short period mutant in terms of bioluminescence rhythm (Fig. 3A; Supplemental Fig. 1). roc66 exhibited a slightly long period (∼26.8 h) in growth rhythm, although the period of its bioluminescence rhythm was very long (29.9 h) (Fig. 3A; Supplemental Fig. 1). The mutants roc23 and roc81 exhibited robust growth rhythms despite their very low-amplitude rhythms in chloroplast bioluminescence (Fig. 3A; Supplemental Fig. 1). To clarify whether these unexpected phenotypes are specific to growth rhythm, we measured chloroplast bioluminescence and growth rhythms simultaneously in identical liquid cultures. The wild-type strain exhibited robust rhythms in both growth and bioluminescence (Fig. 3B). As expected, roc55 and roc75 exhibited short period and low-amplitude phenotypes, respectively, in both rhythms (Fig. 3B). Phenotypes of the bioluminescence rhythms of roc40 and roc66 were similar to those of their growth rhythms (Fig. 3B). This indicates that their unexpected phenotypes are not specific to the growth rhythm, but are thought to be due to a change in the oscillation of the circadian clock in these mutants depending on the culture conditions. The interval between the first and second peaks of roc40 rhythms was extremely long again (Fig. 3B). To determine the period length precisely, we prepared roc40 samples according to the high-throughput assay protocol and measured bioluminescence rhythm in LL. The periodicity of the rhythm was barely detectable under the conditions, and we confirmed that the period length was extremely long in LL (Supplemental Fig. 3). Interestingly, roc81 exhibited different phenotypes of rhythm between growth and bioluminescence. The bioluminescence rhythm was very low amplitude, although the growth rhythm appeared to be normal (Fig. 3B). This indicates that the low-amplitude phenotype of roc81 is specific to chloroplast bioluminescence rhythm, and suggests the possibility that roc81 has a defect in a specific biological process to regulate circadian rhythmicity of chloroplast gene expression. Taken together, these results suggest that our screening isolated not only circadian clock mutants, but also mutants of specific processes for circadian rhythmicity of chloroplast bioluminescence.
Figure 3.
Growth rhythms of isolated mutants. (A) Growth rhythms of mutants. Growth rhythms were monitored by a continuous culture system (see Materials and Methods) in LL. Two-hour moving averages of growth rates (added fresh medium per culture volume per day) are shown. (B) Simultaneous measurement of growth and bioluminescence rhythms. Growth and bioluminescence rhythms were monitored simultaneously from an identical liquid culture by the continuous culture system.
Systematic identification of the responsible genes
To identify integration sites of the marker gene, we determined flanking genomic sequences by the thermal asymmetric interlaced (TAIL)–PCR method (Liu et al. 1995). We analyzed 50 mutants tagged by a single integration of the marker gene (Fig. 2E). We succeeded in identifying flanking genomic sequences on both sides of the marker gene in 18 mutants and on one side in 21 mutants. The opposite flanking sequences in the 21 mutants were analyzed by PCR using specific primer pairs designed for each mutant and were identified in 14 mutants. Overall, we identified both flanking sequences in 32 mutants and a single flanking sequence in seven mutants. The rate of success in identification of both flanking sequences was apparently high in mutants generated with 30 ng of transforming DNA (300 ng, 42.9% [nine of 21]; 30 ng, 79.3% [23 of 29]). A major problem in TAIL–PCR of the mutants generated with 300 ng of transforming DNA was the integration of concatameric marker DNA that led to amplification only of the marker sequence.
Flanking sequences were compared with the Chlamydomonas genome sequence (JGI Chlamydomonas reinhardtii version 3.0: http://genome.jgi-psf.org/Chlre3/Chlre3.home.html; Merchant et al. 2007). All of the flanking sequences were found in the database. In most mutants (27 of 32), the marker genes were integrated without any large changes in their original genome sequences (Supplemental Table 2). On the other hand, the integration site of roc97 was accompanied by a 65.6-kb deletion of genomic DNA, and each flanking sequence of roc9, roc55, roc88, and roc114 was found in different scaffolds of the genome sequence (Supplemental Table 2). These results suggest that a genomic deletion or rearrangement occurred in these mutants accompanied by integration of the marker gene. Two allelic mutants (roc11 and roc55, roc15 and roc74, roc23 and roc112, roc28 and roc59, roc30 and roc54, roc77 and roc105, and roc108 and roc114) were found at seven gene loci (Supplemental Tables 2, 3). This result strongly suggests that the disruptions of these gene loci are responsible for their mutant phenotypes.
The candidate genes are listed in Supplemental Tables 2 and 3. Based on the annotation of the genome sequence (JGI Chlamydomonas reinhardtii version 3.0), sequence similarity to known proteins, or conserved functional domains, we classified the candidate genes into several biological processes: flagellar function (FAP256, ODA4, PF9, DLC7a, FAP131, and FLA14), ubiquitin–proteasome (SKP1 and ROC114), transcription and transcript metabolism (ROC15, ROC40, ROC56, ROC59, ROC66, ROC75, ROC76, ROC93, and XRN1), gene silencing (MUT-9), membrane trafficking and transport (VPS11, ROC81, DRP2, and MFT10), signal transduction (MAPKKKK), DNA damage response (ATR1), and apoptosis (ALIX). These results suggest that various cellular processes are involved in maintenance of the chloroplast circadian rhythmicity.
Structures of ROC genes that would have crucial roles in the circadian clockwork
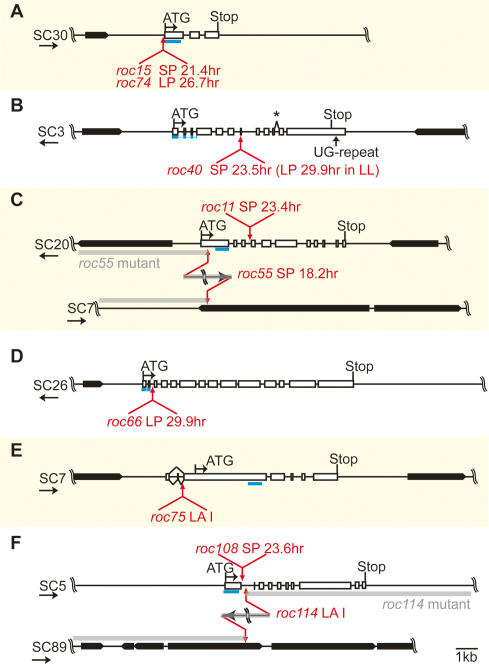
We further focused on six mutants (roc15, roc40, roc55, roc66, roc75, and roc114). Since they exhibited severe defects in circadian rhythmicity, the responsible genes of these mutants would be clock genes closely related to the central oscillatory mechanism of the Chlamydomonas circadian clock. It should be noted, however, that the severe phenotypes of roc40 and roc66 are conditional (Fig. 3). cDNA sequences of the candidate genes were identified by RT–PCR analyses. Comparison of the cDNA sequences to the genome sequence revealed that all of these genes contained introns following the GT–AG rule, except for the splice donor site of an intron of the ROC40 gene (Fig. 4B). Alternative splicing was observed upstream of the predicted initiation codon of the ROC75 gene (Fig. 4E). Since an (UG)≥7 repeat in the 3′-untranslated region that is known to be bound by an RNA-binding protein, CHLAMY1, was found in the ROC40 gene (Fig. 4B; Mittag 1996), it is possible that CHLAMY1 participates in circadian clockwork, as suggested previously (Iliev et al. 2006), via the UG repeat of ROC40. Except for roc15 and roc74, it was evident that the integrations occurred within transcription units (Fig. 4B–F). The integration sites of roc15 and roc74 were upstream from the cDNA sequence identified by RT–PCR (Fig. 4A), and from the most 5′-terminal of available ESTs. Interestingly, roc15 and roc74 exhibited opposite phenotypes, a short (21.3-h) and long (26.6-h) period, respectively, although their integration sites were very close (Fig. 4A; Supplemental Table 2). A comparative analysis of C. reinhardtii and Volvox carteri genome sequences by the Berkeley Genome Pipeline (VISTA track of JGI Chlamydomonas version 3.0) revealed that genomic sequences of the six gene loci are conserved in these two species, suggesting the existence of homologous genes in V. carteri. The gene model for the ROC114 locus has an N-terminal extension compared with our RT–PCR result (Protein ID: 186976, initiation codon locates 14.2 kb upstream); however, we could not amplify the N-terminal extension by an additional RT–PCR.
Figure 4.
Schematic representation of the integration sites of roc15/roc74 (A), roc40 (B), roc11/roc55 (C), roc66 (D), roc75 (E), and roc108/roc114 (F). Boxes indicate exons. Black boxes indicate gene models of JGI Chlamydomonas version 3.0. Predicted (first) initiation codons and stop codons are shown. Red arrows indicate integration sites of the marker in each mutant. The phenotype of the mutants is indicated. Gray bars indicate the predicted genome structures of the roc55 and roc114 mutants. Blue bars indicate the regions used for probes in the Northern blot analysis. The asterisk in B indicates an exon/intron boundary that does not follow the GT–AG rule. Scaffold numbers and directions (arrows) are indicated on the left.
Complementation of the mutant phenotypes by ROC genes
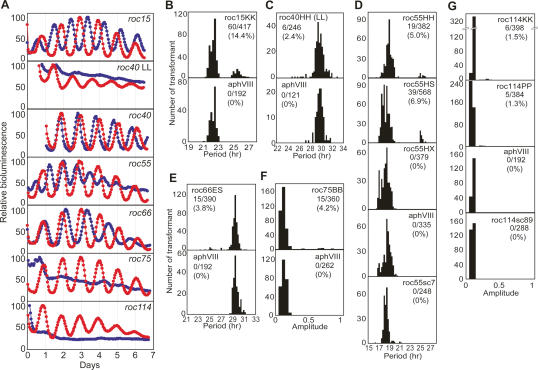
We tested whether these six candidate genes complement their mutant phenotypes or not. The mutants were transformed with genomic DNA fragments (Supplemental Fig. 4) linked to a selectable marker gene, aphVIII (Sizova et al. 2001). All mutants were complemented by the genomic DNA fragments, including their candidate gene, without any other predicted genes (Fig. 5). On the other hand, genomic DNA fragments corresponding to the opposite flanking regions of the integrated marker of roc55 and roc114 (roc55sc7 and roc114sc89, respectively; Supplemental Fig. 4C,F) did not complement their phenotypes (Fig. 5D,G). These results indicate that the ROC15, ROC40, ROC55, ROC66, ROC75, and ROC114 genes are responsible for the circadian rhythm defects of these mutants. Restored rhythmicities of complemented strains of roc114 were, however, at a somewhat low amplitude compared with the wild-type strain, whereas rhythmicities of other mutants were fully restored (Fig. 5A,G).
Figure 5.
Complementation of the mutant phenotypes. Transformants with the genomic DNA fragments were subjected to the high-throughput bioluminescence assay. (A) Representative traces of complemented (red) and control (blue) strains on agar. For roc55 and roc114, traces of complemented strains with roc55HS and roc114KK genomic fragments, respectively, are shown. (B–G) Histograms of the complementation analysis of roc15 (B), roc40 (C), roc55 (D), roc66 (E), roc75 (F), and roc114 (G). For period length mutants (roc15, roc55, and roc66), transformants with period lengths in the range of 24.0–26.0 h were judged to be complemented. For roc40, transformants in the range of 24.0- to 27.0-h periods in LL were judged to be complemented. For low-amplitude mutants (roc75 and roc114), transformants with amplitudes (calculated by the RAP) >0.4 and in the range of 0.2–0.4 were judged to be complemented and partially complemented, respectively. Numbers of complemented (roc15, roc40, roc55, roc66, and roc75) and partially complemented (roc114) transformants among the transformants examined are shown in the graphs.
Confirmation of the disruption of ROC genes by Northern blot analysis
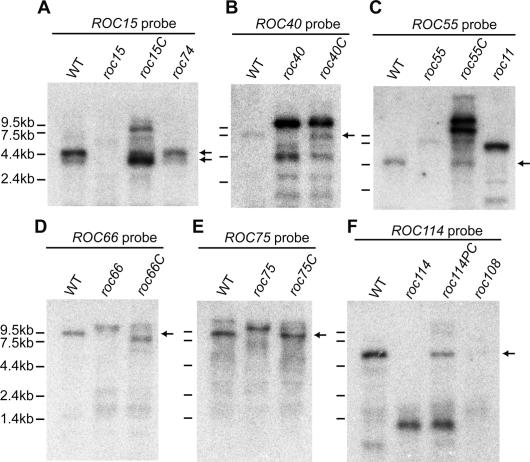
Northern blot analyses of the RNA from these mutants confirmed disruption of these genes. No transcript or transcripts with abnormal sizes were observed in the mutants, whereas transcripts of normal sizes were detected in complemented strains (Fig. 6). One exception was a complemented strain of roc66 (roc66C) in which a slightly short transcript was detected (Fig. 6D). Additional transcripts were detected in the roc15C and roc55C strains (Fig. 6A,C). These transcripts with unexpected sizes might be due to truncation of the transforming DNA fragments. The ROC114 transcript of the partially complemented strain (roc114PC) was comparable in size to that of the wild-type strain (Fig. 6F). This result strongly suggests that the roc114KK genomic DNA fragment contains an entire transcriptional unit of the ROC114 gene (Supplemental Fig. 4F). In addition, the failure of complete restoration in the roc114 mutant implies that roc114 is not a loss-of-function mutant.
Figure 6.
Northern blot analysis of ROC15 (A), ROC40 (B), ROC55 (C), ROC66 (D), ROC75 (E), and ROC114 (F). Total RNA (10 μg; A,B,D–F) and polyadenylated RNA (from 100 μg of total RNA; C) from asynchronous cultures of the wild-type, mutant (roc15, roc74, roc40, roc55, roc11, roc66, roc75, roc114, and roc108), complemented (roc15C, roc40C, roc55C, roc66C, and roc75C), and partially complemented (roc114PC) strains were analyzed. The roc55C and roc114PC strains are complemented and partially complemented strains obtained by gene transfer of the genomic DNA fragments roc55HS and roc114KK, respectively. Arrows indicate the band sizes of wild-type mRNA. Probes are indicated in Figure 4.
Predicted amino acid sequences of ROC proteins
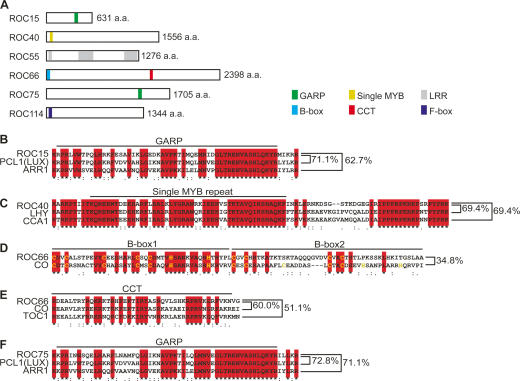
The predicted proteins encoded by ROC15, ROC40, ROC55, ROC66, ROC75, and ROC114 consist of 631, 1556, 1276, 2398, 1705, and 1344 amino acids, respectively (Fig. 7A). Although database searches, except for the V. carteri genome database, did not reveal any obvious homologs showing strong identity to these proteins over their entire length, several conserved domains were detected. ROC15 and ROC75 proteins contain GARP motifs, a subclass of the MYB DNA-binding domain, similar to those of a clock protein PHYTOCLOCK1 (PCL1) (LUX ARRHYTHMO [LUX]) and B-type Arabidopsis response regulators (ARRs) of higher plants (Fig. 7A,B,F; Hwang et al. 2002; Hazen et al. 2005; Onai and Ishiura 2005). ROC40 contains a single MYB repeat that is similar to those of other clock proteins: CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) and LATE ELONGATED HYPOCOTYL (LHY) of higher plants (Fig. 7A,C; Wang et al. 1997; Schaffer et al. 1998). ROC40 also shows similarity to CCA1 and LHY in the region after the MYB domain (Fig. 7C). ROC66 contains a CCT (CONSTANS [CO], CONSTANS-like [COL], TIMING OF CAB EXPRESSION 1 [TOC1]) domain, and a single repeat of the B-box zinc-finger domain that are similar to CO and COL proteins (Fig. 7A,D,E; Robson et al. 2001). An incomplete second B-box that is a feature of some COL proteins (COL9, COL10, COL11, and COL12) was also found in ROC66 (Fig. 7D; Robson et al. 2001). The arrangements of these domains in ROC15/ROC75, ROC40, and ROC66 are similar to those in PCL1(LUX), CCA1/LHY, and CO/COL, respectively (Fig. 7A). On the other hand, it appeared that ROC55 and ROC114 have no significant similarity to plant proteins. ROC114 contains a F-box domain (Fig. 7A), but we could not detect any significant similarity to clock-related F-box proteins (i.e., FWD, FBXL3, β-TRCP, Slimb, JETLAG, ZEITLUPE, and FKF1) (Somers et al. 2000; Grima et al. 2002; Ko et al. 2002; He et al. 2003; Imaizumi et al. 2003; Eide et al. 2005; Shirogane et al. 2005; Koh et al. 2006; Busino et al. 2007; Godinho et al. 2007; Siepka et al. 2007). In the ROC55 sequence, we detected leucine-rich repeat (LRR) regions (Fig. 7A), but could not find any significant similarity to known clock-related proteins. Taken together, these sequence analyses revealed that the circadian clock of Chlamydomonas has both partially plant-like and original components.
Figure 7.
Predicted amino acid sequences of ROC proteins. (A) Schematic view of primary structures of ROC proteins. (B–F) Alignment of amino acid sequences. The amino acid sequences of ROC15, ROC40, ROC66, and ROC75 were aligned to the Arabidopsis proteins using CLUSTAL W (http://clustalw.ddbj.nig.ac.jp). (B) GARP motif of ROC15, PCL1(LUX), and ARR1. (C) Single MYB repeat and following homologous region of ROC40, CCA1, and LHY. (D) B-box zinc-finger domain of ROC66 and CO. The consensus residues of CO-type B-box are denoted in yellow. B-box2 region of ROC66 does not match the consensus of B-box. (E) CCT domain of ROC66, CO, and TOC1. (F) GARP motif of ROC75, PCL1(LUX), and ARR1. Identities of amino acid sequences are indicated.
Circadian expression patterns of ROC mRNAs
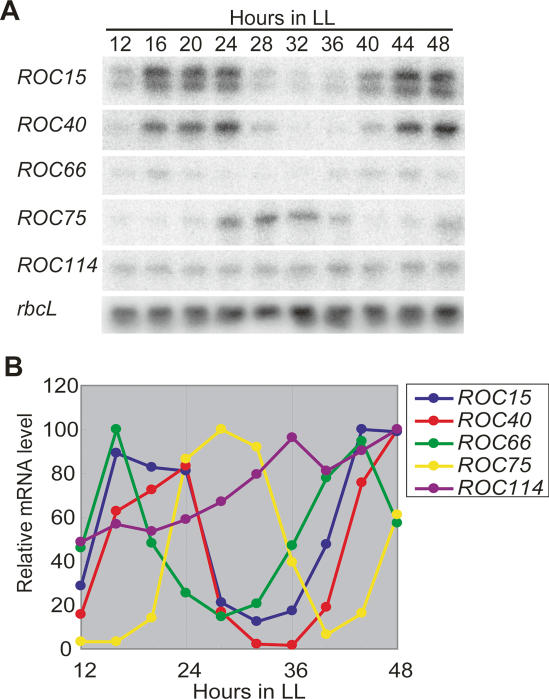
Circadian patterns of ROC gene expression were examined by Northern blot analysis. The mRNA levels of ROC15, ROC40, ROC66, and ROC75 fluctuated in a circadian manner, whereas ROC114 mRNA was expressed constitutively during the circadian cycle (Fig. 8). The ROC66 mRNA level peaked at the mid-subjective night. The ROC15 and ROC40 mRNA levels oscillated with identical or very similar phases peaking during the mid- to late subjective night. The ROC75 mRNA level peaked during the early to mid-subjective day. These results indicate that the expression of the ROC genes encoding putative transcription factors is itself regulated circadianly with different phases.
Figure 8.
Circadian expression patterns of ROC mRNAs. (A) Northern blot analysis of ROC15, ROC40, ROC66, ROC75, and ROC114 mRNAs. Total RNA (5 μg) from a synchronous culture of the wild-type strain in LL (10 μmol m−2 s−1) at 17°C was analyzed. The ROC55 mRNA was undetectable under the experimental conditions. Probes are the same as those used in Figure 6. Equal RNA loading was confirmed by reprobing the blot for rbcL mRNA. (B) Temporal expression profiles of the ROC mRNAs. The maximum values are expressed as 100.
Discussion
In this study, we identified a large number of genes essential for normal circadian rhythmicity of a chloroplast bioluminescence reporter by a systematic forward genetic strategy. Six genes, which would have crucial roles in circadian clockwork, revealed features of the Chlamydomonas circadian clock that have partially plant-like and original components. In addition, close relationships between the chloroplast circadian rhythm and various biological processes, including flagellar function, ubiquitin–proteasome, transcription and transcript metabolism, gene silencing, membrane trafficking and transport, signal transduction, DNA damage response, and apoptosis, were revealed. However, most of the candidate genes, except for the six clock-related genes (ROC15, ROC40, ROC55, ROC66, ROC75, and ROC114), were identified based only on the cosegregation analysis and physical mapping of the integration site by TAIL–PCR. We should take into account that these analyses cannot exclude the possibility that each mutant phenotype was caused by a closely linked mutation in a different gene.
Six genes involved in flagellar function were included in the candidate genes (FAP256, ODA4, PF9, DLC7a, FAP131, and FLA14) (Supplemental Table 2). Phenotypes of all of the corresponding mutants were LA IIa (Supplemental Table 1), suggesting the requirement of normal flagellar function to maintain the robustness of chloroplast rhythmicity. Candidate genes of roc80 and roc108/roc114 were the SKP1 homolog and an unknown F-box protein, respectively (Supplemental Table 2), suggesting the involvement of a Skp1–cullin–F-box (SCF) E3 ubiquitin ligase complex in the circadian system. Since roc114 exhibited circadian rhythm defects in at least two biological processes (Fig. 3; Supplemental Fig. 1), the SCF complex affects the central oscillation mechanism in Chlamydomonas, as well as in other model organisms (Somers et al. 2000; Grima et al. 2002; Ko et al. 2002; He et al. 2003; Eide et al. 2005; Shirogane et al. 2005; Busino et al. 2007; Godinho et al. 2007; Siepka et al. 2007). Several putative transcription factor genes encoding MYB (ROC15, ROC40, ROC59, and ROC75), zinc-finger (ROC56 and ROC66), and basic leucine zipper (ROC76) DNA-binding domain proteins were included (Supplemental Tables 2, 3). These genes may affect each other and form transcriptional feedback loops in the Chlamydomonas clock. Protein kinases are involved in circadian clock systems in all eukaryotic models (Mizoguchi et al. 2006; Gallego and Virshup 2007). Also, in Chlamydomonas it has been demonstrated that casein kinase 1 modulates the circadian period of phototactic rhythms (Schmidt et al. 2006). Our screening additionally revealed the involvement of two putative protein kinase genes MUT-9 and ATR1 (Supplemental Tables 2, 3). MUT-9 has been known to be involved in transcriptional gene silencing and sensitivity to DNA-damaging agents in Chlamydomonas (Jeong Br et al. 2002), and ATR has a central role in the DNA damage response pathway (Zhou and Elledge 2000). Since a close relationship between the circadian clock and DNA damage responses has already been established in mammalian cells and Neurospora (Ünsal-Kaçmaz et al. 2005; Gery et al. 2006; Pregueiro et al. 2006), our findings suggest that the close relationship is a conserved feature among evolutionarily divergent organisms. Several genes identified in this study are conserved in other clock models, including not only Arabidopsis, but also Neurospora, Drosophila, and the mouse (Supplemental Tables 2, 3). It is worth investigating the roles of these genes in the circadian system in these model organisms.
It is noteworthy that our screening detected a mutant of a specific process of the chloroplast bioluminescence rhythm (Fig. 3, roc81). To our knowledge, this is the first study to isolate mutants of a specific output pathway from the circadian clock by forward genetic screens. The candidate gene of the mutant, ROC81, encodes a protein similar to the N-terminal region of Vtc4p, a member of the vacuolar transporter chaperone complex (Cohen et al. 1999; Müller et al. 2003). Because the ROC81 locus contains extended regions of undetermined nucleotides, the sequence of the C-terminal transmembrane domain could not be predicted. Since Vtc4p is known to localize to the membrane of vacuolar and other cellular compartments to control the distribution of membrane proteins in yeast (Cohen et al. 1999; Müller et al. 2003), our results suggest that regulation of membrane proteins is involved in the circadian output pathway for the chloroplast rhythmicity. An interesting question is whether Vtc4p localizes to the outer membrane of chloroplasts, but to date there are no studies assessing its localization in plant cells. Analysis of ROC81 localization in Chlamydomonas will shed light on the molecular mechanisms for synchronization of the circadian rhythmicity between the nucleus and chloroplast.
ROC15/ROC75 and ROC40 have MYB-related DNA-binding domains similar to those of the Arabidopsis clock proteins PCL1(LUX) and CCA1/LHY, respectively (Fig. 7A–C,F). In Arabidopsis, they are critical to the generation of a normal circadian rhythmicity (Schaffer et al. 1998; Wang and Tobin 1998; Hazen et al. 2005; Onai and Ishiura 2005). CCA1/LHY homologs in Oryza sativa and Lemna gibba also play critical roles in the circadian clocks of these species (Miwa et al. 2006; Murakami et al. 2007). The identification of ROC15/ROC75 and ROC40 as components of the circadian clock in Chlamydomonas suggests that the two types of MYB-related DNA-binding domains of ROC15/ROC75/PCL1(LUX) and ROC40/CCA1/LHY are common features of circadian clock proteins over a wide range of species in the green lineage. Are the roles of the Chlamydomonas MYB proteins in the circadian clock the same as those of the Arabidopsis MYB proteins? In contrast to the Arabidopsis cca1/lhy mutants (Green and Tobin 1999; Alabadí et al. 2002; Mizoguchi et al. 2002), the severe phenotype of the roc40 mutant in LL was a long period (Supplemental Fig. 3), and the phenotype was conditional (Supplemental Figs. 1, 3). Furthermore, the roc15 mutant exhibited a short period phenotype, but its rhythm remained robust in DD (Supplemental Fig. 1), whereas the Arabidopsis pcl1(lux) mutant is completely arrhythmic (Hazen et al. 2005; Onai and Ishiura 2005). Concerning the circadian expression profile, Arabidopsis CCA1/LHY and PCL1(LUX) interact genetically with each other, resulting in an antiphase circadian expression peaking at subjective dawn and subjective dusk, respectively (Schaffer et al. 1998; Wang and Tobin 1998; Hazen et al. 2005; Onai and Ishiura 2005). These expression profiles are conserved in the CCA1 and PCL1(LUX) homologs in Oryza sativa (Murakami et al. 2007), and in the LHY homologs in Lemna gibba and Lemna paucicoatata (Miwa et al. 2006). In Chlamydomonas, the expression pattern of ROC40 was comparable with those of CCA1/LHY (Fig. 8). However, in contrast to PCL1(LUX), ROC15 and ROC75 expression did not peak at subjective dusk. The mRNA level of ROC15 oscillated in the same phase with that of ROC40, and the ROC75 mRNA level oscillated in a slightly delayed phase compared with the ROC40 mRNA level and peaked at the early- to mid-subjective day (Fig. 8). Taken together, these results suggest the possibility that the interaction and roles of the MYB proteins in the circadian clock of Chlamydomonas differ, at least to some extent, from those of Arabidopsis.
In the past decade, the transcriptional/translational negative feedback loop model had been believed to be the common mechanism for generation of the circadian oscillation (Dunlap 1999; Harmer et al. 2001). However, recent findings in cyanobacteria (Nakajima et al. 2005; Tomita et al. 2005) indicate that the circadian oscillation is generated by a set of clock proteins without transcription and translation. One of the most interesting questions is whether the new concept found in a bacterium is applicable to eukaryotes or not. In the eukaryotic green alga Acetabularia, the chloroplast photosynthetic rhythm persists even in enucleated cells (Sweeney and Haxo 1961). This fact eliminates the necessity of nuclear transcription for generation of the circadian oscillation, at least in Acetabularia. Although there is no evidence for a transcription/translation-less circadian oscillator in Chlamydomonas, its simple cellular composition will make Chlamydomonas a good system to assess the new concept of circadian oscillation in the eukaryotic cell.
Materials and methods
Strains
Wild-type strains 137c− (CC-124) and 6145c− (CC-1691) were obtained from the Chlamydomonas Center (Duke University, NC), and 11-32a− was kindly provided by Dr. Tatsuaki Saito (Okayama University of Science). Strains were kept as described previously (Matsuo et al. 2006).
Genetic cross
Mt+ and mt− cells were mixed in nitrogen-free Tris-acetate-phosphate medium (TAP-N) in 96-well microtiter plates to allow mating for 2 d. Unmated cells were washed out with fresh TAP-N several times, and zygospores adhering to the walls and bottoms of the wells were allowed to mature for 5 d. Then, the plates were frozen to eliminate residual unmated cells completely, and zygotes were germinated by adding complete TAP medium. After 2–3 d, hatched progenies were spread on TAP plates, and colonies formed on the plates were subjected to random progeny analyses. For cosegregation analysis, the colonies were cultured in TAP in 96-well microtiter plates for 3 d, and 5-μL aliquots of the cultures were spotted on high-salt medium (HSM) plates for the bioluminescence rhythm assay and on TAP plates containing 10 μg/mL hygromycin B (Sigma) for the hygromycin resistance test. In all genetic cross experiments, pairs with the same mating type strain served as negative controls to confirm complete elimination of unmated cells.
Transformation
Nuclear transformation was performed as described (Shimogawara et al. 1998), except that we used a relatively small amount of transforming DNAs and did not use carrier DNAs. To generate insertional mutants, 300 ng and 30 ng of the 1.7-kb HindIII fragment of pHyg3 (Berthold et al. 2002) carrying the hygromycin-resistant gene aph7″ were used per sample (2.5 × 107 cells). Transformants were selected on TAP plates containing 10 μg/mL hygromycin B. For complementation experiments of mutants, 200 ng of PacI fragments carrying a genomic DNA fragment and a paromomycin-resistant aphVIII gene (Sizova et al. 2001) were used. Transformants were selected on TAP plates containing 5 μg/mL paromomycin (Sigma).
High-throughput assay of bioluminescence rhythms
Colonies on agar plates were transferred into 100 μL of TAP medium in each well of 96-well microtiter plates and cultured for 3 d under LL conditions (30 μmol m−2 s−1) at 24°C. Then, 5-μL aliquots of the cultures were spotted on HSM agar plates and cultured under the same conditions for 5 d. The spots were punched out with the agar medium by using a glass tube (6-mm inner diameter) and transferred into each well of the 96-well microtiter plates. Luciferin was added at a final concentration of 200 μM. To synchronize the circadian clock, the spots were subjected to a single cycle of 12-h darkness/12-h light (30 μmol m−2 s−1) at 17°C before bioluminescence monitoring. Bioluminescence rhythms were monitored by our bioluminescence-monitoring apparatus (Okamoto et al. 2005a) in DD at 17°C, and data were analyzed by a cosiner method of the rhythm analyzing program (RAP) (Okamoto et al. 2005b). We added new functions to the RAP to calculate phase angles and amplitudes of the rhythms. Phase angles were represented by circadian time (CT), which was calculated by dividing the phase value by the period value and multiplying by 24. Amplitudes were represented by the amplitude of a fitted cosine curve normalized by dividing by the average bioluminescence level.
Growth rhythm monitoring
Cells were grown in 0.3 HSM (Jacobshagen and Johnson 1994) with aeration at 17°C under LL (200 μmol m−2 s−1) until the optical density at 750 nm reached 0.35, and then exposed to 12 h of darkness to synchronize the circadian clock. After the darkness, cultures were kept in LL (30 μmol m−2 s−1), and the growth rate and bioluminescence were monitored by a continuous culture and bioluminescence monitoring system (K. Okamoto and M. Ishiura, unpubl.). This system continuously monitored the optical density of cultures with an optical sensor and automatically diluted the cultures with fresh medium to the preset optical density. Data on the volume of fresh medium added to the culture were collected every 5 min, and growth rates (added fresh medium per culture volume per day) were calculated. To monitor bioluminescence, aliquots of the culture were automatically sampled every 2 h, mixed with luciferin, and subjected to bioluminescence measurements by using a photomultiplier tube detector (H7360-01, Hamamatsu Photonics).
Analysis of nucleic acids
Genomic DNA was extracted from 1-mL liquid cultures according to the method of Newman et al. (1990) and dissolved in 30 μL of Tris-EDTA buffer (∼50 ng of DNA per microliter). RNA extraction and Southern and Northern blot analyses were performed as described previously (Matsuo et al. 2006).
For the TAIL–PCR analysis of upstream flanking regions, we used specific primers UP3 (5′-GACTCACCTCCCAGAATTC CTGG-3′), UP2 (5′-TCGTTCCGCAGGCTCGCGTAGG-3′), and UP1 (5′-TCGAGAAGTAACAGGGATTCTTGTGTCATG-3′) for primary, secondary, and tertiary reactions, respectively. Similarly, for downstream flanking regions, we used DP4 (5′-CTTCGAGGTGTTCGAGGAGACCC-3′), DP3 (5′-CGCTG GATCTCTCCGGCTTCACC-3′), and DP2 (5′-GCCCCCGGC GCCTGATAAGG-3′) for primary, secondary, and tertiary reactions, respectively. We used the degenerate primer RMD227 (Dent et al. 2005). The genomic DNA was diluted 10-fold, and 1-μL aliquots were used. Primary reactions (20 μL) containing 1× GC buffer (TaKaRa), 0.2 mM dNTPs, 5 pmol of specific primer, 60 pmol of RMD227, and 1 U of LA Taq (TaKaRa) were performed under the following conditions: 2 min, 95°C; 5× (1 min, 94°C; 1 min, 62°C; 2.5 min, 68°C); 1 min, 94°C; 3 min, 25°C; ramping over 3 min to 68°C; 2.5 min, 68°C; 15× (30 sec, 94°C; 3.5 min, 68°C; 30 sec, 94°C; 3.5 min, 68°C; 30 sec, 94°C; 1 min, 44°C; 2.5 min, 68°C); 5 min, 68°C. The primary reaction products were diluted 50-fold, and 1-μL aliquots were subjected to secondary reactions with identical components to the primary reaction under the following conditions: 12× (30 sec, 94°C; 3.5 min, 68°C; 30 sec, 94°C; 3.5 min, 68°C; 30 sec, 94°C; 1 min, 44°C; 2.5 min, 68°C); 5 min, 68°C. The secondary reaction products were diluted 50-fold, and 1-μL aliquots were subjected to tertiary reactions with identical components to the primary reaction under identical conditions to the secondary reaction. The tertiary reaction products were separated by agarose gel electrophoresis, fragments were extracted from the gel, and their sequences were analyzed. We used UP-S (5′-TCATGTTTGCGGGTTGTGACTG-3′) and DP-S (5′-CCCCGCTCCGTGTAAATGGAGG-3′) primers for sequencing of the upstream and downstream products, respectively. To confirm the TAIL–PCR results, we performed a PCR analysis using specific primers located in each flanking genomic region on both sides of the integrated marker. In the case of mutants in which the sequence flanking only one side of the marker gene was determined by TAIL–PCR, the sequence on the opposite side of the marker was predicted from the genome sequence (JGI Chlamydomonas version 3.0) to design specific primers.
For analyses of cDNA sequences, specific primers were designed to amplify the entire ORF based on a gene prediction program, GeneMark.hmm (http://opal.biology.gatech.edu/GeneMark). Reverse transcription was performed by using total or polyadenylated RNA of CBR34+, SuperScript II (Invitrogen), and an oligo-dT primer. PCR amplification was performed by using LA Taq with GC buffer (TaKaRa) and specific primer pairs for ROC15 (Forward, 5′-AAACTCATTCCAAGCGTCCCAAAC CTTCG-3′; Reverse, 5′-CACAGAAGACGTAGGAGCTCAAC GG-3′), ROC40 (Forward, 5′-CCCAGACTCGGCACAGTTAC TTGCCAACG-3′; Reverse, 5′-CCCAGCACGCAAGAGCAAA CCCCTC-3′), ROC55 (Forward, 5′-CCTTCACCTCGTTGCCC GCCTTCAC-3′; Reverse, 5′-ACACACTTTCACACCGCAGC CCCCC-3′), ROC66 (Forward, 5′-GTGCGCCCTGGCGGTGC TACCAACC-3′; Reverse, 5′-AGGCTTGCCCAAGGCTTGCC GCCAC-3′), ROC75 (Forward, 5′-GTTGTTTGGCGGAGGA CACTCGTTGCTTG-3′; Reverse, 5′-TAATTTGGGCGACG ATTGCTGAGGCATGG-3′), ROC114 (Forward, 5′-GAGAGT GCGAGGAGCGCCGTTGAGC-3′; Reverse, 5′-GCATACCCT GGAAGTTGCAAGGCAC-3′), and ROC114 N-terminal extension (Forward 1, 5′-GTTTACAGATAGCTCTGTATCGTCTT GAC-3′; Forward 2, 5′-GCCCTGAGACAAAGTGTGCGAAA GGAATC-3′; Reverse 1, 5′-GCATACCCTGGAAGTTGCAAG GCAC-3′; Reverse 2, 5′-GCGCGCCACTCCGCAAACCAGC-3′ [all four combinations of the forward and reverse primers]).
Plasmid construction
To generate plasmids for genetic complementation experiments, we modified pSI103ΔNotI (Sizova et al. 2001) as follows: The HindIII site was destroyed by blunting/religation, a PacI linker (5′-TTAATTAAGTAC-3′) was inserted into the KpnI site, and a polylinker (top strand, 5′-GGCCTTAA TTAACTAGTGGTACCGATATCAAGCTTGAATTCGC-3′; bottom strand, 5′-GGCCGCGAATTCAAGCTTGATATCGG TACCACTAGTTAATTAA-3′) was inserted into the NotI site in the same direction as the aphVIII gene, yielding pSI103PL. BAC clones containing a gene of interest were searched for on the Genome browser of JGI Chlamydomonas version 3.0 and obtained from the Clemson University Genomics Institute. BAC clones were digested by restriction enzymes, and a genomic DNA fragment containing the gene was cloned into the polylinker region of pSI103PL. The plasmids were digested by PacI, and a DNA fragment containing the genomic DNA with the aphVIII marker was used for the complementation experiments. Names and sizes of the restriction fragments of genomic DNA segments (Supplemental Fig. 4) and clone numbers of BACs carrying the DNA segments used in this study were as follows: roc15KK, 7.4-kb KpnI fragment, 38G22; roc40HH, 10.6-kb HindIII fragment, 3O10; roc55HH, 11.9-kb HindIII fragment, 16E8; roc55HS, 8.9-kb HindIII–SnaBI fragment, 16E8; roc55HX, 4.8-kb HindIII–XhoI fragment, 16E8; roc55sc7, 11.4-kb HindIII–DraI fragment, 25L12; roc66ES, 13.3-kb EcoRI–SpeI fragment, 36M13; roc75BB, 11.8-kb BamHI fragment, 26N16; roc114KK, 14.3-kb KpnI fragment, 21B13; roc114PP, 9.4-kb PvuI fragment, 21B13; roc114sc89, 12.9-kb EcoRI fragment, 17D2.
Accession numbers
cDNA sequences of ROC15, ROC40, ROC55, ROC66, longer splice variant of ROC75, shorter splice variant of ROC75, and ROC114 have been deposited in the GenBank/EMBL/DDBJ under accession numbers AB363964, AB363965, AB363966, AB363967, AB363968, AB363969, and AB363970, respectively.
Acknowledgments
We thank W. Mages (Regensburg University) for the pHyg3 plasmid, I. Sizova (St. Petersburg State University) for the pSI103ΔNotI plasmid, T. Saito (Okayama University of Science) for the 11-32a− strain, Y. Nakaoka (Nagoya University) for technical assistance, and D. Mrozek (Medical English Service) for professional editing. This work was supported by grants from the Japanese Ministry of Education, Culture, Sports, Science and Technology (MEXT) to M.I., and grants from Research Fellowships of the Japan Society for the Promotion of Science for Young Scientists to T.M. The Division of Biological Science, Graduate School of Science, Nagoya University, was supported by a 21st century COE grant from MEXT.
Footnotes
Supplemental material is available at http://www.genesdev.org.
Article published online ahead of print. Article and publication date are online at http://www.genesdev.org/cgi/doi/10.1101/gad.1650408.
References
- Alabadí D., Yanovsky M.J., Mas P., Harmer S.L., Kay S.A. Critical role for CCA1 and LHY in maintaining circadian rhythmicity in Arabidopsis. Curr. Biol. 2002;12:757–761. doi: 10.1016/s0960-9822(02)00815-1. [DOI] [PubMed] [Google Scholar]
- Berthold P., Schmitt R., Mages W. An engineered Streptomyces hygroscopicus aph 7″ gene mediates dominant resistance against hygromycin B in Chlamydomonas reinhardtii. Protist. 2002;153:401–412. doi: 10.1078/14344610260450136. [DOI] [PubMed] [Google Scholar]
- Breton G., Kay S.A. Circadian rhythms lit up in Chlamydomonas. Genome Biol. 2006;7:215. doi: 10.1186/gb-2006-7-4-215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruce V.G. The biological clock in Chlamydomonas reinhardi. J. Protozool. 1970;17:328–334. doi: 10.1093/genetics/70.4.537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruce V.G. Mutants of the biological clock in Chlamydomonas reinhardi. Genetics. 1972;70:537–548. doi: 10.1093/genetics/70.4.537. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bruce V.G. Recombinants between clock mutants of Chlamydomonas reinhardi. Genetics. 1974;77:221–230. doi: 10.1093/genetics/77.2.221. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bünning E. The physiologycal clock: Circadian rhythms and biologycal chronometry. Springer-Verlag; New York: 1973. [Google Scholar]
- Busino L., Bassermann F., Maiolica A., Lee C., Nolan P.M., Godinho S.I., Draetta G.F., Pagano M. SCFFbxl3 controls the oscillation of the circadian clock by directing the degradation of cryptochrome proteins. Science. 2007;316:900–904. doi: 10.1126/science.1141194. [DOI] [PubMed] [Google Scholar]
- Cohen A., Perzov N., Nelson H., Nelson N. A novel family of yeast chaperons involved in the distribution of V-ATPase and other membrane proteins. J. Biol. Chem. 1999;274:26885–26893. doi: 10.1074/jbc.274.38.26885. [DOI] [PubMed] [Google Scholar]
- Dent R.M., Haglund C.M., Chin B.L., Kobayashi M.C., Niyogi K.K. Functional genomics of eukaryotic photosynthesis using insertional mutagenesis of Chlamydomonas reinhardtii. Plant Physiol. 2005;137:545–556. doi: 10.1104/pp.104.055244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dunlap J.C. Molecular bases for circadian clocks. Cell. 1999;96:271–290. doi: 10.1016/s0092-8674(00)80566-8. [DOI] [PubMed] [Google Scholar]
- Eide E.J., Woolf M.F., Kang H., Woolf P., Hurst W., Camacho F., Vielhaber E.L., Giovanni A., Virshup D.M. Control of mammalian circadian rhythm by CKIε-regulated proteasome-mediated PER2 degradation. Mol. Cell. Biol. 2005;25:2795–2807. doi: 10.1128/MCB.25.7.2795-2807.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gallego M., Virshup D.M. Post-translational modifications regulate the ticking of the circadian clock. Nat. Rev. Mol. Cell Biol. 2007;8:139–148. doi: 10.1038/nrm2106. [DOI] [PubMed] [Google Scholar]
- Gery S., Komatsu N., Baldjyan L., Yu A., Koo D., Koeffler H.P. The circadian gene per1 plays an important role in cell growth and DNA damage control in human cancer cells. Mol. Cell. 2006;22:375–382. doi: 10.1016/j.molcel.2006.03.038. [DOI] [PubMed] [Google Scholar]
- Godinho S.I., Maywood E.S., Shaw L., Tucci V., Barnard A.R., Busino L., Pagano M., Kendall R., Quwailid M.M., Romero M.R., et al. The after-hours mutant reveals a role for Fbxl3 in determining mammalian circadian period. Science. 2007;316:897–900. doi: 10.1126/science.1141138. [DOI] [PubMed] [Google Scholar]
- Green R.M., Tobin E.M. Loss of the circadian clock-associated protein 1 in Arabidopsis results in altered clock-regulated gene expression. Proc. Natl. Acad. Sci. 1999;96:4176–4179. doi: 10.1073/pnas.96.7.4176. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grima B., Lamouroux A., Chelot E., Papin C., Limbourg-Bouchon B., Rouyer F. The F-box protein slimb controls the levels of clock proteins period and timeless. Nature. 2002;420:178–182. doi: 10.1038/nature01122. [DOI] [PubMed] [Google Scholar]
- Harmer S.L., Panda S., Kay S.A. Molecular bases of circadian rhythms. Annu. Rev. Cell Dev. Biol. 2001;17:215–253. doi: 10.1146/annurev.cellbio.17.1.215. [DOI] [PubMed] [Google Scholar]
- Harris E.H. Chlamydomonas as a model organism. Annu. Rev. Plant Physiol. Plant Mol. Biol. 2001;52:363–406. doi: 10.1146/annurev.arplant.52.1.363. [DOI] [PubMed] [Google Scholar]
- Hazen S.P., Schultz T.F., Pruneda-Paz J.L., Borevitz J.O., Ecker J.R., Kay S.A. LUX ARRHYTHMO encodes a Myb domain protein essential for circadian rhythms. Proc. Natl. Acad. Sci. 2005;102:10387–10392. doi: 10.1073/pnas.0503029102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- He Q., Cheng P., Yang Y., Yu H., Liu Y. FWD1-mediated degradation of FREQUENCY in Neurospora establishes a conserved mechanism for circadian clock regulation. EMBO J. 2003;22:4421–4430. doi: 10.1093/emboj/cdg425. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwang S., Kawazoe R., Herrin D.L. Transcription of tufA and other chloroplast-encoded genes is controlled by a circadian clock in Chlamydomonas. Proc. Natl. Acad. Sci. 1996;93:996–1000. doi: 10.1073/pnas.93.3.996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hwang I., Chen H.C., Sheen J. Two-component signal transduction pathways in Arabidopsis. Plant Physiol. 2002;129:500–515. doi: 10.1104/pp.005504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Iliev D., Voytsekh O., Schmidt E.M., Fiedler M., Nykytenko A., Mittag M. A heteromeric RNA-binding protein is involved in maintaining acrophase and period of the circadian clock. Plant Physiol. 2006;142:797–806. doi: 10.1104/pp.106.085944. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Imaizumi T., Tran H.G., Swartz T.E., Briggs W.R., Kay S.A. FKF1 is essential for photoperiodic-specific light signalling in Arabidopsis. Nature. 2003;426:302–306. doi: 10.1038/nature02090. [DOI] [PubMed] [Google Scholar]
- Jacobshagen S., Johnson C.H. Circadian rhythms of gene expression in Chlamydomonas reinhardtii: Circadian cycling of mRNA abundances of cab II, and possibly of β-tubulin and cytochrome c. Eur. J. Cell Biol. 1994;64:142–152. [PubMed] [Google Scholar]
- Jeong Br B.R., Wu-Scharf D., Zhang C., Cerutti H. Suppressors of transcriptional transgenic silencing in Chlamydomonas are sensitive to DNA-damaging agents and reactivate transposable elements. Proc. Natl. Acad. Sci. 2002;99:1076–1081. doi: 10.1073/pnas.022392999. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ko H.W., Jiang J., Edery I. Role for Slimb in the degradation of Drosophila Period protein phosphorylated by Doubletime. Nature. 2002;420:673–678. doi: 10.1038/nature01272. [DOI] [PubMed] [Google Scholar]
- Koh K., Zheng X., Sehgal A. JETLAG resets the Drosophila circadian clock by promoting light-induced degradation of TIMELESS. Science. 2006;312:1809–1812. doi: 10.1126/science.1124951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu Y.G., Mitsukawa N., Oosumi T., Whittier R.F. Efficient isolation and mapping of Arabidopsis thaliana T-DNA insert junctions by thermal asymmetric interlaced PCR. Plant J. 1995;8:457–463. doi: 10.1046/j.1365-313x.1995.08030457.x. [DOI] [PubMed] [Google Scholar]
- Matsuo T., Onai K., Okamoto K., Minagawa J., Ishiura M. Real-time monitoring of chloroplast gene expression by a luciferase reporter: Evidence for nuclear regulation of chloroplast circadian period. Mol. Cell. Biol. 2006;26:863–870. doi: 10.1128/MCB.26.3.863-870.2006. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merchant S.S., Prochnik S.E., Vallon O., Harris E.H., Karpowicz S.J., Witman G.B., Terry A., Salamov A., Fritz-Laylin L.K., Marechal-Drouard L., et al. The Chlamydomonas genome reveals the evolution of key animal and plant functions. Science. 2007;318:245–250. doi: 10.1126/science.1143609. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mergenhagen D. Circadian clock: Genetic characterization of a short period mutant of Chlamydomonas reinhardii. Eur. J. Cell Biol. 1984;33:13–18. [PubMed] [Google Scholar]
- Mittag M. Conserved circadian elements in phylogenetically diverse algae. Proc. Natl. Acad. Sci. 1996;93:14401–14404. doi: 10.1073/pnas.93.25.14401. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mittag M. Circadian rhythms in microalgae. Int. Rev. Cytol. 2001;206:213–247. doi: 10.1016/s0074-7696(01)06023-5. [DOI] [PubMed] [Google Scholar]
- Miwa K., Serikawa M., Suzuki S., Kondo T., Oyama T. Conserved expression profiles of circadian clock-related genes in two Lemna species showing long-day and short-day photoperiodic flowering responses. Plant Cell Physiol. 2006;47:601–612. doi: 10.1093/pcp/pcj027. [DOI] [PubMed] [Google Scholar]
- Mizoguchi T., Wheatley K., Hanzawa Y., Wright L., Mizoguchi M., Song H.R., Carre I.A., Coupland G. LHY and CCA1 are partially redundant genes required to maintain circadian rhythms in Arabidopsis. Dev. Cell. 2002;2:629–641. doi: 10.1016/s1534-5807(02)00170-3. [DOI] [PubMed] [Google Scholar]
- Mizoguchi T., Putterill J., Ohkoshi Y. Kinase and phosphatase: The cog and spring of the circadian clock. Int. Rev. Cytol. 2006;250:47–72. doi: 10.1016/S0074-7696(06)50002-6. [DOI] [PubMed] [Google Scholar]
- Müller O., Neumann H., Bayer M.J., Mayer A. Role of the Vtc proteins in V-ATPase stability and membrane trafficking. J. Cell Sci. 2003;116:1107–1115. doi: 10.1242/jcs.00328. [DOI] [PubMed] [Google Scholar]
- Murakami M., Tago Y., Yamashino T., Mizuno T. Comparative overviews of clock-associated genes of Arabidopsis thaliana and Oryza sativa. Plant Cell Physiol. 2007;48:110–121. doi: 10.1093/pcp/pcl043. [DOI] [PubMed] [Google Scholar]
- Nakajima M., Imai K., Ito H., Nishiwaki T., Murayama Y., Iwasaki H., Oyama T., Kondo T. Reconstitution of circadian oscillation of cyanobacterial KaiC phosphorylation in vitro. Science. 2005;308:414–415. doi: 10.1126/science.1108451. [DOI] [PubMed] [Google Scholar]
- Newman S.M., Boynton J.E., Gillham N.W., Randolph-Anderson B.L., Johnson A.M., Harris E.H. Transformation of chloroplast ribosomal RNA genes in Chlamydomonas: Molecular and genetic characterization of integration events. Genetics. 1990;126:875–888. doi: 10.1093/genetics/126.4.875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okamoto K., Onai K., Furusawa T., Ishiura M. A portable integrated automatic apparatus for the real-time monitoring of bioluminescence in plants. Plant Cell Environ. 2005a;28:1305–1315. [Google Scholar]
- Okamoto K., Onai K., Ishiura M. RAP, an integrated program for monitoring bioluminescence and analyzing circadian rhythms in real time. Anal. Biochem. 2005b;340:193–200. doi: 10.1016/j.ab.2004.11.007. [DOI] [PubMed] [Google Scholar]
- Onai K., Ishiura M. PHYTOCLOCK 1 encoding a novel GARP protein essential for the Arabidopsis circadian clock. Genes Cells. 2005;10:963–972. doi: 10.1111/j.1365-2443.2005.00892.x. [DOI] [PubMed] [Google Scholar]
- Pregueiro A.M., Liu Q., Baker C.L., Dunlap J.C., Loros J.J. The Neurospora checkpoint kinase 2: A regulatory link between the circadian and cell cycles. Science. 2006;313:644–649. doi: 10.1126/science.1121716. [DOI] [PubMed] [Google Scholar]
- Robson F., Costa M.M., Hepworth S.R., Vizir I., Pineiro M., Reeves P.H., Putterill J., Coupland G. Functional importance of conserved domains in the flowering-time gene CONSTANS demonstrated by analysis of mutant alleles and transgenic plants. Plant J. 2001;28:619–631. doi: 10.1046/j.1365-313x.2001.01163.x. [DOI] [PubMed] [Google Scholar]
- Schaffer R., Ramsay N., Samach A., Corden S., Putterill J., Carre I.A., Coupland G. The late elongated hypocotyl mutation of Arabidopsis disrupts circadian rhythms and the photoperiodic control of flowering. Cell. 1998;93:1219–1229. doi: 10.1016/s0092-8674(00)81465-8. [DOI] [PubMed] [Google Scholar]
- Schmidt M., Gessner G., Luff M., Heiland I., Wagner V., Kaminski M., Geimer S., Eitzinger N., Reissenweber T., Voytsekh O., et al. Proteomic analysis of the eyespot of Chlamydomonas reinhardtii provides novel insights into its components and tactic movements. Plant Cell. 2006;18:1908–1930. doi: 10.1105/tpc.106.041749. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shimogawara K., Fujiwara S., Grossman A., Usuda H. High-efficiency transformation of Chlamydomonas reinhardtii by electroporation. Genetics. 1998;148:1821–1828. doi: 10.1093/genetics/148.4.1821. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shirogane T., Jin J., Ang X.L., Harper J.W. SCFβ-TRCP controls clock-dependent transcription via casein kinase 1-dependent degradation of the mammalian period-1 (Per1) protein. J. Biol. Chem. 2005;280:26863–26872. doi: 10.1074/jbc.M502862200. [DOI] [PubMed] [Google Scholar]
- Siepka S.M., Yoo S.H., Park J., Song W., Kumar V., Hu Y., Lee C., Takahashi J.S. Circadian mutant Overtime reveals F-box protein FBXL3 regulation of cryptochrome and period gene expression. Cell. 2007;129:1011–1023. doi: 10.1016/j.cell.2007.04.030. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sizova I., Fuhrmann M., Hegemann P. A Streptomyces rimosus aphVIII gene coding for a new type phosphotransferase provides stable antibiotic resistance to Chlamydomonas reinhardtii. Gene. 2001;277:221–229. doi: 10.1016/s0378-1119(01)00616-3. [DOI] [PubMed] [Google Scholar]
- Somers D.E., Schultz T.F., Milnamow M., Kay S.A. ZEITLUPE encodes a novel clock-associated PAS protein from Arabidopsis. Cell. 2000;101:319–329. doi: 10.1016/s0092-8674(00)80841-7. [DOI] [PubMed] [Google Scholar]
- Sweeney B.M., Haxo F.T. Persistence of a photosynthetic rhythm in enucleated Acetabularia. Science. 1961;134:1361–1363. doi: 10.1126/science.134.3487.1361. [DOI] [PubMed] [Google Scholar]
- Tomita J., Nakajima M., Kondo T., Iwasaki H. No transcription–translation feedback in circadian rhythm of KaiC phosphorylation. Science. 2005;307:251–254. doi: 10.1126/science.1102540. [DOI] [PubMed] [Google Scholar]
- Ünsal-Kaçmaz K., Mullen T.E., Kaufmann W.K., Sancar A. Coupling of human circadian and cell cycles by the timeless protein. Mol. Cell. Biol. 2005;25:3109–3116. doi: 10.1128/MCB.25.8.3109-3116.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wang Z.Y., Tobin E.M. Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell. 1998;93:1207–1217. doi: 10.1016/s0092-8674(00)81464-6. [DOI] [PubMed] [Google Scholar]
- Wang Z.Y., Kenigsbuch D., Sun L., Harel E., Ong M.S., Tobin E.M. A Myb-related transcription factor is involved in the phytochrome regulation of an Arabidopsis Lhcb gene. Plant Cell. 1997;9:491–507. doi: 10.1105/tpc.9.4.491. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhou B.B., Elledge S.J. The DNA damage response: Putting checkpoints in perspective. Nature. 2000;408:433–439. doi: 10.1038/35044005. [DOI] [PubMed] [Google Scholar]