Abstract
Cell cycle progression is monitored by highly coordinated checkpoint machinery, which is activated to induce cell cycle arrest until defects like DNA damage are corrected. We have isolated an anti-proliferative cell cycle regulator named G2A (for G2 accumulation), which is predominantly expressed in immature T and B lymphocyte progenitors and is a member of the seven membrane-spanning G protein-coupled receptor family. G2A overexpression attenuates the transformation potential of BCR-ABL and other oncogenes, and leads to accumulation of cells at G2/M independently of p53 and c-Abl. G2A can be induced in lymphocytes and to a lesser extent in nonlymphocyte cell lines or tissues by multiple stimuli including different classes of DNA-damaging agents and serves as a response to damage and cellular stimulation which functions to slow cell cycle progression.
The G1/S transition, the S-phase delay, and the G2/M transition are well defined checkpoints in the cell cycle (1–7). Mitotic entry is monitored by a spindle checkpoint that inhibits anaphase progression when chromosomes are not attached to the mitotic spindle (8). The G2/M transition is regulated by Cdc2 and cyclin B1 as part of the maturation promoting factor that determines entry into mitosis (9). Dephosphorylation of Cdc2 by Cdc25 and association with cyclin B1 results in rapid entry into mitosis (10) whereas cyclin B1 degradation or export to the cytoplasm and phosphorylation of Cdc2 on negative regulatory sites by Wee1 block entry into mitosis (9, 11).
Response to DNA damage is detected by the ataxia-telangiectasia mutated (ATM) (reviewed in refs. 12 and 13), which activates Chk1 and phosphorylates Cdc25C leading to sequestration of Cdc25 by 14-3-3 σ (14–16) and accumulation of the phosphorylated (inactive) form of Cdc2 and G2/M arrest. Cds1 has been demonstrated to function redundantly to Chk-1 by phosphorylating both Wee1 and Cdc25C inactivating both gene products (14, 16, 17). Ataxia-telangiectasia mutated (ATM) also can activate transcription of secondary molecules including p53, which regulate genes including p21Cip/Waf-1 and 14-3-3 σ (reviewed in refs. 18 and 19). The p21 gene product is an inhibitor of cyclin-dependent kinases and arrests the cell cycle in G1. The 14-3-3 σ gene product likely mediates G2/M arrest by binding to Cdc25C to sequester it in the cytoplasm. The ataxia-telangiectasia mutated (ATM) protein physically interacts with the tyrosine kinase c-Abl (20, 21) and is required for its activation.
The general eukaryotic cell cycle control machinery is highly conserved among all cell types, but tissue-specific regulators like GATA-1 play a critical role as an anti-proliferative-signaling molecule with restricted expression in erythroid cells (22, 23). Lymphocytes provide an interesting model system to study cell cycle regulators because their development is marked by the unique property of entering, exiting, and rejoining the cell cycle depending on their internal developmental stages as well as the surrounding environment (24, 25). To generate antigen receptors, lymphocytes also undergo V(D)J recombination by specific DNA double-strand cleavage, which may be coupled to cell cycle checkpoint pathways to minimize genomic instability (reviewed in refs. 26 and 27).
We have isolated a G protein-coupled receptor (GPCR) predominantly expressed in lymphocytes, which we named G2A for G2 accumulation, while searching for genes which were induced by the BCR-ABL tyrosine kinase oncogene (28). G2A appears to function at the G2/M checkpoint to delay mitosis and may serve as a mechanism for T and B cells and other cell types to slow their proliferation and repair damaged DNA to ensure proper replication.
MATERIALS AND METHODS
Cloning of the G2A Gene.
Representational difference analysis (RDA) was carried out as described (29, 30). In this study, the cDNA from bone marrow cells transformed by the wild-type (wt) p185 BCR-ABL (31) was used as tester and by the Src homology 2 (SH2) mutant as a driver to isolate genes that were up-regulated by the wt BCR-ABL. Three rounds of subtractive hybridization were performed with increasing ratios of driver to tester from 1:100 to 1:800 to 1:8,000. The partial G2A sequence from RDA was labeled by using the Primer-It II random primer labeling kit (Stratagene) to screen a mouse spleen cDNA library (CLONTECH). One clone containing the C-terminal portion of G2A was identified after screening 1 × 106 plaques. The 5′-RACE system (GIBCO/BRL) was then used to obtain the N-terminal portion of G2A by using the G2AGSP1 primer (GGTGACAGCAGTCCTCTGGT) and the nested G2AGSP2 primer (TAGCGGTCGCAGGAAATGCAG). The full length G2A was then obtained by reverse transcription–PCR (RT-PCR).
Murine G2A was used to probe a human spleen cDNA library (CLONTECH) to isolate a human homologue. A human genomic G2A clone was isolated from a λ-DASH II lymphocyte library. Human metaphase cells were prepared from phytohemagglutinin-stimulated peripheral blood lymphocytes as described (32) for FISH analysis. A biotin-labeled, 14-kb probe containing the human genomic G2A sequence was prepared by nick-translation by using Bio-16-UTP (Enzo Diagnostics). Hybridization was detected with fluorescein-conjugated avidin (Vector Laboratories), and chromosomes were identified by staining with 4,6-diamidino-2-phenylindole-dihydrochloride (DAPI). To generate G2A-GFP fusion protein, the murine G2A gene was fused to green fluorescence protein (GFP) in pEGFP-N1 vector (CLONTECH). G2A-GFP or GFP also was cloned into the MSCV vector upstream of the internal ribosome entry site and p185 BCR-ABL for bone marrow transformation assays (33).
First-strand cDNA was synthesized by using 5 μg of total RNA and Superscript preamplification system (GIBCO/BRL). Ten percent of the first strand cDNA synthesis product was then used for PCR with G2A-specific primers. Primers HuG2A-C1 (TTAGCGGCCGCTCAGCAGGACTCCTCAATCAG) and HuG2A+6 (CGGTGGTTGTCATCTTCCTA) were used to amplify the human G2A whereas the mouse G2A was amplified with primers MuN2A3′RACE-1 (CAGGACTGGCTTGGGTCATT) and N2AGSP2 (TAGCGGTCGCAGGAAATGCAG). Glyceraldehyde 3-phosphate dehydrogenase control amplimer set (CLONTECH) was included as a control to ensure that equal amounts of templates were used.
Cell Cycle Analysis.
Rat-1 or NIH 3T3 fibroblasts were infected with retroviruses expressing either G2A, G2A-GFP, or GFP. The cells were harvested and analyzed 2 days after infection. For serum starvation, cells were washed, trypsinized, and replated in DMEM with 0.1% FBS. Cell DNA was stained with hypotonic staining buffer (0.1% sodium citrate, 0.3% Triton X-100, 0.01% propidium iodide, and 0.002% ribonuclease A) for 15 min on ice, in the dark. The DNA content was measured by using a Becton Dickinson FACScan machine.
RESULTS
G2A Is an Inducible GPCR.
BCR-ABL is a chimeric tyrosine kinase oncogene generated by a reciprocal chromosomal translocation associated with the pathogenesis of chronic myelogeneous leukemia and acute lymphocytic leukemia (reviewed in refs. 28 and 34). Certain domains of BCR-ABL are critical for its oncogenic activity. (31, 35–37). The R552L substitution within the highly conserved FLVRE motif of the SH2 domain uncouples the SH2 domain from phosphotyrosine-containing proteins without affecting the kinase activity of BCR-ABL (38). This mutation greatly reduces the ability of BCR-ABL to stimulate anchorage-independent growth of fibroblasts in soft agar (31) while retaining the ability to transform primary bone marrow cells in vitro. The SH2 mutant-derived cells are poorly malignant when transferred in vivo (31).
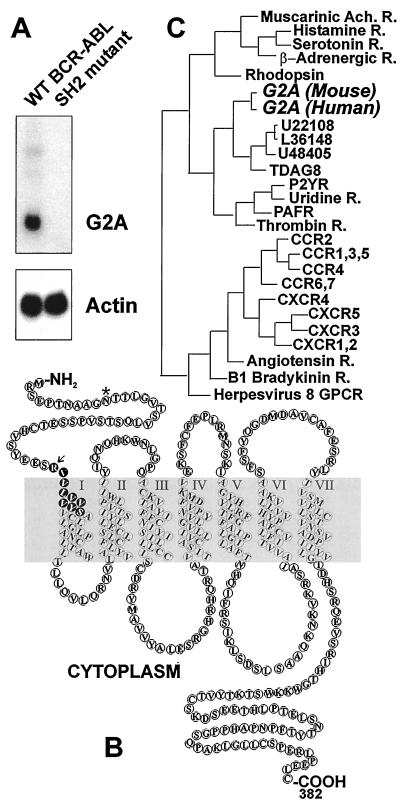
We used RDA (29, 30, 39) to compare genes expressed in murine pre-B cells transformed by wt BCR-ABL or the SH2 mutant variant. One of these clones bore sequence homology to GPCRs (Fig. 1B). Northern blot analysis confirmed that this GPCR is induced by wt BCR-ABL but not the SH2 mutant in pre-B lymphocytes (Fig. 1A).
Figure 1.
G2A is a BCR-ABL-induced GPCR. (A) Induction of G2A by wt BCR-ABL but not by the SH2 mutant of BCR-ABL. Murine bone marrow cells were infected with retrovirus encoding the wt BCR-ABL or the SH2 mutant (R552L) expressed from the long terminal repeat promoter in the pSRαMSV vector (35, 36, 79). Transformed pre-B-lymphoid cells were harvested, and RNA was extracted for Northern blotting. The major G2A transcript is 3 kb in length. Actin was used as a total RNA-loading control. (B) Predicted transmembrane helices of the murine G2A. The potential glycosyslation site (∗) and the putative destruction box (arrow) are indicated. Sequence analysis was performed by using the blast program (Genetics Computer Group, Madison, WI). The transmembrane helices were predicted by using the sosui program available at http://expasy.hcuge.ch/www/tools.html. (C). G2A belongs to a subclass of transcriptionally regulated GPCRs. Phylogenetic analysis of G2A was made with the program pileup (Genetics Computer Group). Phylogenetic relationships were calculated with distances (Jukes–Cantor Distance algorithm) and the tree was made with growtree (UPGMA algorithm).
We used the fragment identified by RDA to obtain full length mouse and human clones. The mouse and human clones had ≈70% overall identity. They were most divergent in the N-terminal extracellular domain (25% identity) and C-terminal cytoplasmic tail (55% identity). G2A contains a putative N-linked glycosylation site (∗) in the N-terminal extracellular domain characteristic of GPCRs and a putative “destruction box” (RVVLVVVYS) that is similar to that of human cyclin A (RAALAVLKS), which serves as a recognition motif for ubiquitin conjugation (40, 41, reviewed in ref. 42).
GPCRs define a large superfamily of >250 members. G2A belongs to a subclass with no defined function (Fig. 1C, reviewed in refs 43 and 44). Other members of this subclass are induced in thymus during T cell receptor engagement-mediated apoptosis (TDAG8, ref. 45) and are transcriptionally up-regulated during T cell activation (P2Y, ref. 46). A human G2A genomic clone isolated from a λ phage library was used to determine the chromosome localization. Hybridization of this probe resulted in specific labeling only of chromosome 14. Specific labeling of 14q32.3 was observed on four (six cells), three (10 cells), two (7) cells, or one (two cells) chromatid(s) of the chromosome 14 homologues in 25 cells examined. Of 99 signals observed, 64 (64%) were located at 14q32.3. Thirty-five background signals (primarily single signals) were observed at other chromosomal sites, but none of these bands were labeled consistently. Thus the G2A gene maps to 14q32.3. This region has been associated with ovarian cancer predisposition and progression (47) but no clear relation to G2A was seen in our preliminary analysis (J. Boyd, Z.W., and O.N.W., unpublished data).
Examination of the expression pattern of murine or human G2A on multiple tissue mRNA blots (CLONTECH) revealed that the highest expression is in hematopoietic tissues rich in lymphocytes like spleen and thymus. Weak G2A expression was seen in heart and lung RNA samples on some blots but not in other tissues. PCR analysis detected variable and weak G2A expression at high cycle numbers in a diverse collection of cell lines and tissue preparations representing epithelial and mesenchymal cell types.
G2A Attenuates the Proliferative Signals of BCR-ABL.
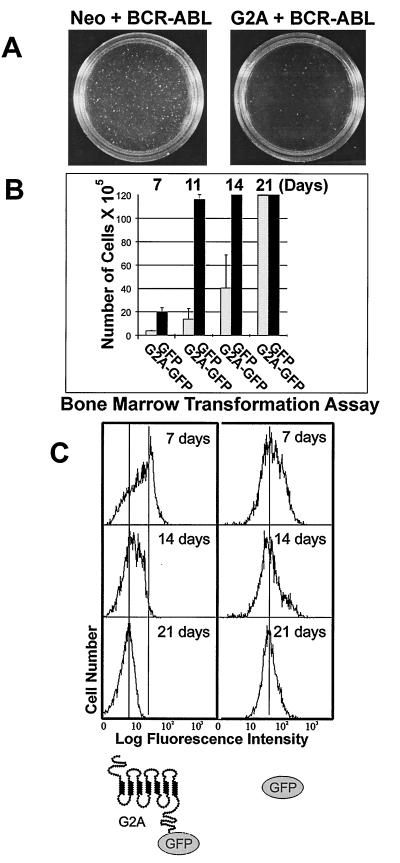
BCR-ABL confers anchorage-independent growth of rodent fibroblasts (Rat-1) in soft agar, and the number of colonies obtained quantitatively reflects its transformation potency (48). G2A strongly antagonized the ability of BCR-ABL to form colonies in soft agar (Fig. 2A). Similar percentages of cells were infected by BCR-ABL-expressing retroviruses as shown by fluorescence-activated cell sorter (FACS) analysis and Western blotting (data not shown). Rat-1/Neo cells exhibited a prominent spindle-like phenotype typical of transformation after infection by BCR-ABL. G2A blocked this gross morphological change by BCR-ABL (data not shown). When these same cell populations were plated in agar, wt BCR-ABL gave rise to >1,000 macroscopic (>1 mm) colonies after 3 weeks. In contrast, BCR-ABL coexpressed with G2A yielded 5-fold fewer colonies suggesting that G2A antagonizes BCR-ABL-mediated transformation but cannot completely block the effect of this oncogene (Fig. 2A). Evaluation of agar colonies expanded in liquid culture showed that the cells that grew in agar had lost expression of G2A but retained expression of BCR-ABL (data not shown). G2A suppressed the transformation potential of other classes of oncogenes including the transcription factor Myc (49) and an activated version of the cytoplasmic tyrosine kinase BTK (50) (data not shown).
Figure 2.
G2A antagonizes the oncogenic potential of the BCR-ABL oncogene. (A) Ectopic expression of G2A attenuates the proliferative signals of BCR-ABL in a fibroblast transformation assay. G2A with the neomycin resistance gene (neo) or Neo only expressing rat fibroblast (Rat-1) cell lines were generated by retroviral infection and G418 selection and were then super-infected with a retroviral stock expressing p185 BCR-ABL. The cells were plated in agar as described (48) and colonies were counted and photographed after 3 weeks. (B) G2A suppresses the induction of pre-B cells by BCR-ABL in bone marrow cells. Bone marrow transformation was performed as described (51) by using bone marrow cells from the tibias and femurs of BALB/c mice. Bone marrow was harvested and infected with retrovirus encoding the wt p185 BCR-ABL along with the G2A-GFP fusion protein (pMSCV G2A-GFP IRES p185 wt) or the GFP control (pMSCV GFP IRES p185 wt). The cells were plated in triplicate at a density of 5 × 106 cells per 6-cm dish in RPMI medium 1640 containing 10% fetal bovine serum and 36-mercapto-ethanol (5 × 10−5 M, Sigma) and monitored for pre-B cell growth at the various time points indicated. Bone marrow cultures reached confluency around 1 × 107 cells/per 3 ml culture. (C) Counter-selection against the G2A protein level in bone marrow cells transformed by BCR-ABL. The protein level of G2A-GFP or GFP was monitored by FACS in BCR-ABL-transformed bone marrow cells at time points indicated.
A bone marrow pre-B cell transformation assay (51) was applied to semi-quantitatively measure the kinetics of the BCR-ABL-mediated hematopoietic transformation in the presence or absence of G2A. To monitor the protein level of G2A, we generated a chimeric fusion protein of G2A and the GFP whose level could be quantitatively measured by cytofluorography. In this assay, mouse bone marrow cells were isolated and infected with a retrovirus doubly encoding wt BCR-ABL and either a G2A-GFP fusion protein or a GFP control. G2A coexpression with BCR-ABL delayed the induction of pre-B cell outgrowth as compared with BCR-ABL plus the GFP (Fig. 2B). Bone marrow cultures transformed by BCR-ABL in the absence of G2A reached confluency (>1 × 107 cells/per 3 ml of culture) within 1.5 weeks, but it took nearly 3 weeks to reach saturation in the presence of G2A (Fig. 2B). G2A-GFP protein levels decreased gradually and after 3 weeks were nearly undetectable (Fig. 2C). BCR-ABL expression was maintained (data not shown). This counter-selection against B cells expressing high levels of G2A strongly suggests an anti-proliferative effect for G2A.
A Wide Variety of Signals Activates G2A Expression.
B lymphocytes are generated from hematopoietic stem cells by successive steps of differentiation during which a diverse repertoire of antigen receptors are generated by Ig gene rearrangement. Mouse bone marrow B cells were fractionated into pro-B (B220+, CD43+), pre-B (B220+, BP1+, CD43−), immature B (B220+, IgM+, IgD-), and mature B (B220+, IgM+, IgD+) cells by FACS (52). A semi-quantitative RT-PCR method (see Materials and Methods) was used to measure the RNA levels of G2A. The G2A transcript was clearly present in pro-B cells, which are undergoing Ig D-J gene recombination (Fig. 3A). Extended PCR cycles revealed a low level of G2A in pre-B and immature B cells (data not shown). Using antibodies directed to the C terminus of G2A and permeabilized cell preparations, a high level of the G2A protein was detected in pro-B cells whereas very weak fluorescent staining was present in the other fractions (data not shown). T lymphocytes mature in the thymus where the T cell receptor genes undergo rearrangement (reviewed in ref. 53). Thymocytes also were isolated and shown by semi-quantitative RT-PCR to express comparable levels of G2A throughout intrathymic maturation defined by CD4 and CD8 surface phenotypes (Fig. 3A).
Figure 3.
Multiple stimuli induce G2A. Mouse bone marrow B cells were isolated, stained with the B220, CD43, BP-1, IgM, and IgD antibodies, and FACS sorted into pro-B (B220+, CD43+), pre-B (B220+, BP1+, CD43−), immature B (B220+, IgM+, IgD−), and mature B cells (B220+, IgM+, IgD+). Similarly, mouse thymocytes were isolated from thymus, stained with CD4 and CD8 antibodies, and fractionated into CD4-CD8−, CD4+CD8+, CD4+CD8−, or CD4-CD8+ populations. RNA samples were extracted from 105 of cells of each fraction, and RT-PCR was then performed. A semi-quantitative RT-PCR method was used to measure the RNA levels of G2A (see Materials and Methods). Glyceraldehyde 3-phosphate dehydrogenase was used as a control to ensure that equal amounts of templates were used for RT-PCR. (B). Human Ramos B cells (54) were activated by either phorbol 12-myristate 13-acetate (PMA 2, ug/ml) plus ionomycin (20 ng/ml) or goat anti-IgM (final concentration of 10 μg/ml, Southern Biotechnology Associates). RNA samples were isolated before or after activation at time points indicated. RT-PCR was then performed by using primers specific for the human G2A cDNA sequence. (C–E) Induction of G2A in B cells by different classes of DNA-damaging agents: Ramos cells were irradiated with increasing doses of x-ray, UV, or chemotherapy drugs and then were grown in medium for additional 16 hr before RNA samples were extracted for RT-PCR.
The induction of G2A in pre-B cells by BCR-ABL led us to search for alternative stimuli, which may regulate G2A. We used a human IgM+ B cell line (54) as a model system in which G2A expression is at a low basal level. BCR crosslinking induced G2A transcripts within 6 hr (Fig. 3B). The simultaneous addition of ionomycin to increase the intracellular calcium and PMA to activate protein kinase C also significantly increased G2A mRNA within 2–4 hr. (Fig. 3B). The analogous behavior of G2A to tumor suppressor genes and its transcriptional regulation led us to examine whether G2A is induced by DNA-damaging agents. Chemical agents that either inhibit the de novo synthesis of DNA precursors (hydroxyurea and 5′-fluorouracil), directly block DNA synthesis (cytosine arabinoside, etoposide), alter microtubules (taxol), or induce lesions by intercalating into DNA (doxorubicin) were evaluated (Fig. 3C). The G2A transcript was found to be up-regulated by all these treatments. Varying doses of x-rays or UV were tested. The G2A transcript was induced in a dosage dependent manner (Fig. 3 D and E).
Ectopic Expression of G2A Accumulates Cells at G2 and M and Blocks the Progression of Mitosis.
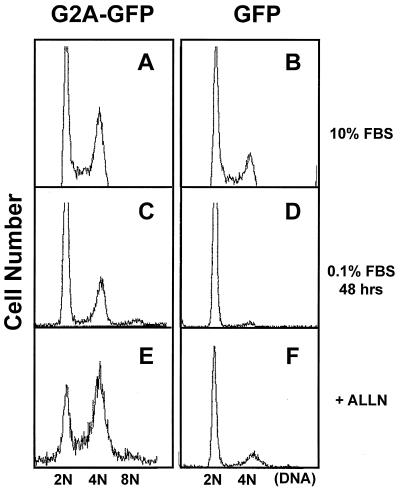
The anti-oncogenic effect of G2A in fibroblasts and lymphoid cells as well as the induction of G2A by cell cycle arrest-associated DNA-damaging agents strongly suggested that G2A may be involved in cell cycle regulation. Retroviruses expressing a G2A-GFP fusion or a GFP control were used to infect >90% of NIH 3T3 cell populations. G2A-GFP expression increased the fraction of cells at G2/M by ≈15% under normal growing conditions (10% FBS) (Fig. 4A). This increase is comparable in magnitude with the 14% elevation in G1 arrest caused by overexpression of p53 (55).
Figure 4.
Ectopic expression of G2A accumulates cells at G2/M and blocks the progression of mitosis. Retroviruses expressing either a G2A-GFP fusion or a GFP control were used to infect NIH 3T3 cells. NIH 3T3 cells expressing G2A-GFP or GFP control were grown in 10% serum (A and B) or starved in media with 0.1% FBS for 48 hr (C and D). Cells were then harvested, and the DNA content was analyzed. Mouse fibroblasts were infected with retroviruses expressing G2A-GFP or GFP and then incubated with 50 μM of ALLN overnight in media with 10% FBS. The cells were harvested and analyzed for DNA content (E and F).
Serum starvation induces fibroblasts to move through the cell cycle until they return to G1. Introduction of a block at another checkpoint will be accentuated in low serum conditions as cells do not have positive growth signals to complete replication. NIH 3T3 cells expressing the G2A-GFP fusion or a GFP control were cultured in the presence of 0.1% FBS for 48 hr. Ninety-five percent of the control cells expressing GFP alone contained a 2 N DNA content (Fig. 4D). However, cells expressing the G2A-GFP fusion protein still exhibited a large percentage of cells with a 4 N DNA content (35%, Fig. 4C), suggesting that G2A is blocking the exit of cells from G2/M during growth factor deprivation. We also observed the formation of bi- and multi-nucleated cells (>8 N DNA content) during growth factor deprivation in cells expressing G2A-GFP (and additional microscopic data not shown). This suggests that there is endoduplication of DNA in the presence of G2A, but failure to undergo cytokinesis, implicating an additional role of G2A in perturbing the mitotic spindle checkpoint.
The G2A sequence contains a destruction box found in cyclins that serves as a recognition motif for the conjugation of ubiquitin and proteolytic degradation (56). There is evidence that GPCRs such as yeast Ste2 are ubiquitinated to direct their internalization for endosome-mediated degradation (57). N-acetyl-l-leucinyl-l-leucinyl-l-norleucinal (ALLN) is a potent inhibitor of the catalytic core of the 26S protease complex (58). Cells were infected with retroviruses expressing G2A-GFP or GFP. After incubation with a low dose of ALLN that alone did not affect cell growth control (50 μM), fibroblasts were harvested and analyzed for GFP expression. ALLN increased the protein level of G2A-GFP by 3- to 4-fold as measured by the fluorescent intensity whereas the inhibitor had no discernible effect on the protein level of GFP (data not shown). The increased protein level of G2A by ALLN correlated with an increase in G2/M cells to 62% (Fig. 4E). In contrast, ALLN did not significantly change the cell cycle profile of GFP-expressing cells (Fig. 4F).
Caffeine has been used to reverse the DNA damage-induced inhibition of Cdc2 and release G2/M arrest likely by activating cdc25 (59, 60). Mouse fibroblasts infected with retroviruses expressing a G2A-GFP fusion or GFP were incubated in the presence of 5 mM caffeine. The cells were then harvested after overnight incubation for cell cycle analysis. Incubation with caffeine relieved the G2/M arrest, suggesting that the G2A-mediated G2/M block is upstream of Cdc2 (data not shown).
c-Abl and p53 Are Not Required for the G2/M Block by G2A.
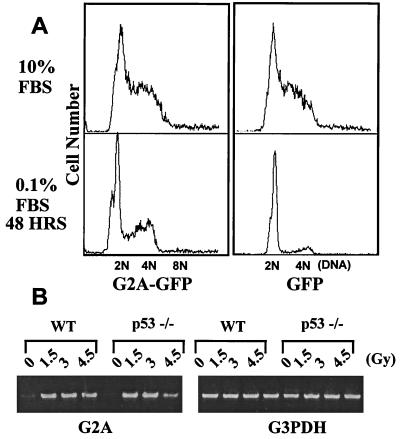
To study the role of p53 and c-Abl in G2A-mediated G2/M arrest, we examined the effect of overexpressing G2A in fibroblasts lacking p53 or c-Abl. DNA content analysis of p53−/− fibroblasts (61) gave an unusually high percentage of cells in S phase supporting a defect in the G1/S transition in these cells (Fig. 5A). Overexpression of G2A in the absence of p53 did not significantly increase the percentage of cells at G2/M under normal growth conditions (10% FBS). Serum starvation for 48 hr revealed a large percentage of G2A-expressing cells still arrested at G2/M (33%) compared with GFP-expressing cells (16%), suggesting that p53 is not required for G2/M arrest mediated by G2A (Fig. 5A).
Figure 5.
p53 is not required for the G2/M block by G2A or the induction of G2A. (A) Retroviruses expressing either G2A-GFP fusion, or a GFP control were used to infect p53−/− fibroblasts (61). Cells were cultured in the presence of 10% or 0.1% FBS for 48 hr and harvested for DNA content analysis by FACS. (B) Bone marrow cells were isolated from the wt and p53-knock-out mice and incubated with interleukin-7 and stem cell factor to stimulate the outgrowth of pre-B cells (63). After 3 days, the pre-B cells were irradiated with varying doses of x-ray followed by overnight incubation. RT-PCR analysis was then performed (see Materials and Methods).
Similar experiments were performed on a c-Abl−/− fibroblast cell line (62). Overexpression of G2A-GFP increased the percentage of cells at G2/M by ≈15% in the presence of 10% FBS when compared with GFP-expressing cells. A high percentage of cells expressing G2A were still arrested at G2/M (39%) after serum starvation (0.1% FBS) compared with GFP-expressing cells (14%), suggesting that c-Abl expression is not functionally required for G2/M arrest by G2A (data not shown).
Because p53 is a sensor of DNA damage and induces the expression of genes such as p21 and 14-3-3 σ in response to DNA damage (reviewed in ref. 18), we tested whether p53 is required for the induction of G2A in response to ionizing irradiation in lymphocytes. Bone marrow cells were isolated from wt and p53 knock-out mice and incubated with interleukin-7 and stem cell factor (63) to stimulate the outgrowth of pre-B cells for 3 days. These pre-B cells were then irradiated with varying doses of x-ray. After overnight incubation, total RNA was isolated and a semi-quantitative RT-PCR was performed. The G2A transcript was low in actively growing nonirradiated pre-B cells (Fig. 5B) but induced in both the wt and the p53−/− mouse lymphocytes.
DISCUSSION
GPCR-mediated growth suppression has been previously described for the platelet-activating factor receptor, which can promote G0 exit and inhibit the G1 to S transition in a ligand-dependent manner. Activation of the platelet-activating factor receptor can block v-Src and v-Ras induced oncogenic morphological changes and anchorage-independent growth (64). Binding of the yeast α-mating factor to its receptor Ste2 induces a G1 arrest via a cyclin-dependent kinase inhibitor FAR1 (65–67). Induction of G2A by oncogenic signals and DNA damage and the ability of G2A to antagonize these signals suggest that G2A functions like a tumor suppressor gene. This is consistent with our data showing that forced overexpression of G2A is counter-selected over time in lymphocyte cell populations, and the loss of G2A expression correlates with enhanced transformation by BCR-ABL in pre-B cells.
The ligands of GPCRs are extremely diverse, ranging from small biogenic amines to large glycoprotein hormones. Signature motifs are present within certain subgroups of GPCRs, which can be used to predict the nature of ligands for an orphan receptor but are not present in G2A (reviewed in ref. 44). Ligand-independent GPCRs have been identified such as the Kaposi virus-associated receptor (68, 69). Further studies are necessary to define a ligand or ligand-independent mode of action for G2A.
G2A resembles p53 in inducible expression by a wide variety of DNA-damaging agents and involvement in the negative regulation of cell cycle progression (reviewed in refs. 18, 70, and 71). However, G2A and p53 exhibit clear differences. The increase in the p53 protein level by DNA-damaging agents is mainly due to a post-translational modification which occurs within 30 min (55, 72). In contrast, G2A is up-regulated at the RNA level (Fig. 3). p53 is responsive to DNA damage in almost all cell types, whereas G2A is most highly expressed in T and B lymphocytes. The p53 gene product induces both G1 and G2/M arrest. We have not excluded that G2A may function at other checkpoints but the clearest effects are seen for the G2/M transition. Although p53 −/− lymphocytes can induce G2A, we have not ruled out the possibility that G2A is under the control of a p53-like response pathway because p73 or other molecules may substitute for p53 (73, 74).
Our data suggest that G2A is not induced by proliferative signals per se but by excessive stimuli or damage. Activation of a negative feedback loop by a positive signal is a common regulatory mechanism. For example, interleukin-6 induces a protein, SOCS-1, which serves to suppress interleukin-6-induced receptor phosphorylation and cellular differentiation by directly inactivating the Jak kinases (75–77). Smad7, a transforming growth factor β-induced antagonist of signaling, provides another example (78). G2A may function in a similar manner to balance very strong signals and mitigate DNA damage. G2A expression may have evolved to protect cells from genomic instability because it is found in pro-B cells and thymocytes, which are undergoing antigen receptor gene recombination. Hyperexpression or inactivation of G2A in animal models will provide valuable tools to further assess the possible connection of G2A to normal lymphocyte physiology and pathological states.
Acknowledgments
We would like to thank S. Quan, C. Hogg, X. Wang, J. Johnson, and J. McLaughlin for excellent technical assistance; Dr. R. Mohr for editorial assistance and isolation of the human G2A genomic clone; J. C. White and J. Shimaoka for assistance with the manuscript; Drs. C. Sawyers and Y. Whang for providing c-Abl−/− and p53−/− cell lines; Drs. H. Wu, R. Gatti, and B. Braun for helpful discussions; and Drs. A. Berk, D. Black, A. Satterthwaite, S. Smale, K. H. Zavitz, N. Rosenberg, and S. L. Zipursky for review of the manuscript. Z.W. was an Associate of the Howard Hughes Medical Institute and a Fellow of Leukemia Society of America. A.C.F. was supported by a fellowship from the Human Frontier Science Program Organization. M.I.W. is supported by the Cancer Research Fund of the Damon Runyon–Walter Winchell Foundation Fellowship (DRG-086). O.N.W. is an Investigator of the Howard Hughes Medical Institute. This work was supported in part by U.S. Public Health Service grants to O.N.W. (CA76204) and M.M.L. (CA40046).
ABBREVIATIONS
- GFP
green fluorescence protein
- ALLN
N-acetyl-l-leucinyl-l-leucinyl-l-norleucinal
- RDA
representational difference analysis
- GPCR
G protein-coupled receptor
- RT-PCR
reverse transcription–PCR
- wt
wild type
- FACS
fluorescence-activated cell sorter
- SH2
Src homology 2
Footnotes
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AF083442).
References
- 1. Hartwell L H, Weinert T A. Science. 1989;246:629–634. doi: 10.1126/science.2683079. [DOI] [PubMed] [Google Scholar]
- 2.Hartwell L. Cell. 1992;71:543–546. doi: 10.1016/0092-8674(92)90586-2. [DOI] [PubMed] [Google Scholar]
- 3.Murray A W. Nature (London) 1992;359:599–604. doi: 10.1038/359599a0. [DOI] [PubMed] [Google Scholar]
- 4.Hunter T. Cell. 1993;75:839–841. doi: 10.1016/0092-8674(93)90528-x. [DOI] [PubMed] [Google Scholar]
- 5.Hartwell L H, Kastan M B. Science. 1994;266:1821–1828. doi: 10.1126/science.7997877. [DOI] [PubMed] [Google Scholar]
- 6.Nurse P. Cell. 1997;91:865–867. doi: 10.1016/s0092-8674(00)80476-6. [DOI] [PubMed] [Google Scholar]
- 7.Paulovich A G, Toczyski D P, Hartwell L H. Cell. 1997;88:315–321. doi: 10.1016/s0092-8674(00)81870-x. [DOI] [PubMed] [Google Scholar]
- 8.Nicklas R B. Science. 1997;275:632–637. doi: 10.1126/science.275.5300.632. [DOI] [PubMed] [Google Scholar]
- 9.Holloway S L, Glotzer M, King R W, Murray A W. Cell. 1993;73:1393–1402. doi: 10.1016/0092-8674(93)90364-v. [DOI] [PubMed] [Google Scholar]
- 10.Weinert T. Science. 1997;277:1450–1451. doi: 10.1126/science.277.5331.1450. [DOI] [PubMed] [Google Scholar]
- 11.Toyoshima F, Moriguchi T, Wada A, Fukuda M, Nishida E. EMBO J. 1998;17:2728–2735. doi: 10.1093/emboj/17.10.2728. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Westphal C H. Curr Biol. 1997;7:R789–R792. doi: 10.1016/s0960-9822(06)00406-4. [DOI] [PubMed] [Google Scholar]
- 13.Meyn M S. Cancer Res. 1995;55:5991–6001. [PubMed] [Google Scholar]
- 14.Sanchez Y, Wong C, Thoma R S, Richman R, Wu Z, Piwnica-Worms H, Elledge S J. Science. 1997;277:1497–1501. doi: 10.1126/science.277.5331.1497. [DOI] [PubMed] [Google Scholar]
- 15.Peng C Y, Graves P R, Thoma R S, Wu Z, Shaw A S, Piwnica-Worms H. Science. 1997;277:1501–1505. doi: 10.1126/science.277.5331.1501. [DOI] [PubMed] [Google Scholar]
- 16.Furnari B, Rhind N, Russell P. Science. 1997;277:1495–1497. doi: 10.1126/science.277.5331.1495. [DOI] [PubMed] [Google Scholar]
- 17.Boddy M N, Furnari B, Mondesert O, Russell P. Science. 1998;280:909–912. doi: 10.1126/science.280.5365.909. [DOI] [PubMed] [Google Scholar]
- 18.Levine A J. Cell. 1997;88:323–331. doi: 10.1016/s0092-8674(00)81871-1. [DOI] [PubMed] [Google Scholar]
- 19.Hermeking H, Lengauer C, Polyak K, He T-C, Zhang L, Thiagalingam S, Kinzler K W, Vogelstein B. Mol Cell. 1997;1:3–11. doi: 10.1016/s1097-2765(00)80002-7. [DOI] [PubMed] [Google Scholar]
- 20.Shafman T, Khanna K K, Kedar P, Spring K, Kozlov S, Yen T, Hobson K, Gatei M, Zhang N, Watters D, et al. Nature (London) 1997;387:520–523. doi: 10.1038/387520a0. [DOI] [PubMed] [Google Scholar]
- 21.Baskaran R, Wood L D, Whitaker L L, Canman C E, Morgan S E, Xu Y, Barlow C, Baltimore D, Wynshaw-Boris A, Kastan M B, et al. Nature (London) 1997;387:516–519. doi: 10.1038/387516a0. [DOI] [PubMed] [Google Scholar]
- 22.Briegel K, Lim K C, Plank C, Beug H, Engel J D, Zenke M. Genes Dev. 1993;7:1097–1109. doi: 10.1101/gad.7.6.1097. [DOI] [PubMed] [Google Scholar]
- 23.Briegel K, Bartunek P, Stengl G, Lim K C, Beug H, Engel J D, Zenke M. Development (Cambridge, UK) 1996;122:3839–3850. doi: 10.1242/dev.122.12.3839. [DOI] [PubMed] [Google Scholar]
- 24.Kantor A B, Herzenberg L A. Annu Rev Immunol. 1993;11:501–538. doi: 10.1146/annurev.iy.11.040193.002441. [DOI] [PubMed] [Google Scholar]
- 25.Cumano A, Kee B L, Ramsden D A, Marshall A, Paige C J, Wu G E. Immunol Rev. 1994;137:5–33. doi: 10.1111/j.1600-065x.1994.tb00657.x. [DOI] [PubMed] [Google Scholar]
- 26.Lin W C, Desiderio S. Immunol Today. 1995;16:279–289. doi: 10.1016/0167-5699(95)80182-0. [DOI] [PubMed] [Google Scholar]
- 27.Danska J S, Guidos C J. Semin Immunol. 1997;9:199–206. doi: 10.1006/smim.1997.0072. [DOI] [PubMed] [Google Scholar]
- 28.Kurzrock R, Gutterman J, Talpaz M. N Engl J Med. 1988;319:990–998. doi: 10.1056/NEJM198810133191506. [DOI] [PubMed] [Google Scholar]
- 29.Lisitsyn N, Lisitsyn N, Wigler M. Science. 1993;259:946–951. doi: 10.1126/science.8438152. [DOI] [PubMed] [Google Scholar]
- 30.Hubank M, Schatz D G. Nucleic Acids Res. 1994;22:5640–5648. doi: 10.1093/nar/22.25.5640. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Goga A, McLaughlin J, Afar D E H, Saffran D C, Witte O N. Cell. 1995;82:981–988. doi: 10.1016/0092-8674(95)90277-5. [DOI] [PubMed] [Google Scholar]
- 32.Rowley J D, Diaz M O, Espinoza R, III, Patel Y D, Van Melle E, Ziemin S, Taillon-Miller P, Lichter P, Evans G A, Kersey J H, et al. Proc Natl Acad Sci USA. 1990;87:9358–9362. doi: 10.1073/pnas.87.23.9358. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Hawley R G, Lieu F H L, Fong Z C, Hawley T S. Gene Ther. 1994;1:136–138. [PubMed] [Google Scholar]
- 34.Groffen J, Heisterkamp N. Baillieres Clin Haematol. 1997;10:187–201. doi: 10.1016/s0950-3536(97)80002-9. [DOI] [PubMed] [Google Scholar]
- 35.Pendergast A M, Quilliam L A, Cripe L D, Bassing C H, Dai Z, Li N, Batzer A, Rabun K M, Der C J, Schlessinger J, Gishizky M L. Cell. 1993;75:175–185. [PubMed] [Google Scholar]
- 36.Muller A J, Young J C, Pendergast A-M, Pondel M, Landau N R, Littman D R, Witte O N. Mol Cell Biol. 1991;11:1785–1792. doi: 10.1128/mcb.11.4.1785. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 37.McWhirter J R, Wang J Y J. Mol Cell Biol. 1991;11:1553–1565. doi: 10.1128/mcb.11.3.1553. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Mayer B J, Baltimore D. Mol Cell Biol. 1994;14:2883–2894. doi: 10.1128/mcb.14.5.2883. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Braun B S, Frieden R, Lessnick S L, May W A, Denny C T. Mol Cell Biol. 1995;15:4623–4630. doi: 10.1128/mcb.15.8.4623. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Glotzer M, Murray A W, Kirschner M W. Nature (London) 1991;349:132–138. doi: 10.1038/349132a0. [DOI] [PubMed] [Google Scholar]
- 41.King R W, Peters J M, Tugendreich S, Rolfe M, Hieter P, Kirschner M W. Cell. 1995;81:279–288. doi: 10.1016/0092-8674(95)90338-0. [DOI] [PubMed] [Google Scholar]
- 42.King R W, Deshaies R J, Peters J-M, Kirschner M W. Science. 1996;274:1652–1659. doi: 10.1126/science.274.5293.1652. [DOI] [PubMed] [Google Scholar]
- 43.Strader C D, Fong T M, Graziano M P, Tota M R. FASEB J. 1995;9:745–754. [PubMed] [Google Scholar]
- 44.Strader C D, Fong T M, Tota M R, Underwood D, Dixon R A. Annu Rev Biochem. 1994;63:101–132. doi: 10.1146/annurev.bi.63.070194.000533. [DOI] [PubMed] [Google Scholar]
- 45.Choi J W, Lee S Y, Choi Y. Cell Immunol. 1996;168:78–84. doi: 10.1006/cimm.1996.0051. [DOI] [PubMed] [Google Scholar]
- 46.Koshiba M, Apasov S, Sverdlov V, Chen P, Erb L, Turner J T, Weisman G A, Sitkovsky M V. Proc Natl Acad Sci USA. 1997;94:831–836. doi: 10.1073/pnas.94.3.831. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.Bandera C A, Takahashi H, Behbakht K, Liu P C, LiVolsi V A, Benjamin I, Morgan M A, King S A, Rubin S C, Boyd J. Cancer Res. 1998;57:513–515. [PubMed] [Google Scholar]
- 48.Lugo T, Witte O N. Mol Cell Biol. 1989;9:1263–1270. doi: 10.1128/mcb.9.3.1263. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 49.Stone J, De Lange T, Ramsay G, Jakobovits E, Bishop J M, Varmus H, Lee W. Mol Cell Biol. 1987;7:1697–1709. doi: 10.1128/mcb.7.5.1697. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Li T, Rawlings D J, Park H, Kato R M, Witte O N, Satterthwaite A B. Oncogene. 1997;15:1375–1383. doi: 10.1038/sj.onc.1201308. [DOI] [PubMed] [Google Scholar]
- 51.McLaughlin J, Chianese E, Witte O N. Proc Natl Acad Sci USA. 1987;84:6558–6562. doi: 10.1073/pnas.84.18.6558. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 52.Marcos M A, Godin I, Cumano A, Morales S, Garcia-Porrero J A, Dieterlen-Lievre F, Gaspar M L. Immunol Rev. 1994;137:155–171. doi: 10.1111/j.1600-065x.1994.tb00663.x. [DOI] [PubMed] [Google Scholar]
- 53.von Boehmer H. Annu Rev Immunol. 1993;6:309–326. doi: 10.1146/annurev.iy.06.040188.001521. [DOI] [PubMed] [Google Scholar]
- 54.Klein G, Giovanella B, Westman A, Stehlin J S, Mumford D. Intervirology. 1975;5:319–334. doi: 10.1159/000149930. [DOI] [PubMed] [Google Scholar]
- 55.Zhang Y, Xiong Y, Yarbrough W G. Cell. 1998;92:725–734. doi: 10.1016/s0092-8674(00)81401-4. [DOI] [PubMed] [Google Scholar]
- 56.Hochstrasser M. Cell. 1996;84:813–815. doi: 10.1016/s0092-8674(00)81058-2. [DOI] [PubMed] [Google Scholar]
- 57.Terrell J, Shih S, Dunn R, Hicke L. Mol Cell. 1998;1:193–202. doi: 10.1016/s1097-2765(00)80020-9. [DOI] [PubMed] [Google Scholar]
- 58.Rock K L, Gramm C, Rothstein L, Clark K, Stein R, Dick L, Hwang D, Goldberg A L. Cell. 1994;78:761–771. doi: 10.1016/s0092-8674(94)90462-6. [DOI] [PubMed] [Google Scholar]
- 59.Schlegel R, Pardee A B. Science. 1986;232:1264–1266. doi: 10.1126/science.2422760. [DOI] [PubMed] [Google Scholar]
- 60.Poon R Y, Chau M S, Yamashita K, Hunter T. Cancer Res. 1997;57:5168–5178. [PubMed] [Google Scholar]
- 61.Lowe S W, Schmitt E M, Smith S W, Osborne B A, Jacks T. Nature (London) 1993;362:847–879. doi: 10.1038/362847a0. [DOI] [PubMed] [Google Scholar]
- 62.Tybulewicz V L J, Crawford C E, Jackson P K, Bronson R T, Mulligan R C. Cell. 1991;65:1153–1163. doi: 10.1016/0092-8674(91)90011-m. [DOI] [PubMed] [Google Scholar]
- 63.McNiece I K, Langley K E, Zsebo K M. J Immunol. 1991;146:3785–3790. [PubMed] [Google Scholar]
- 64.Kume K, Shimizu T. J Biol Chem. 1997;272:22898–22904. doi: 10.1074/jbc.272.36.22898. [DOI] [PubMed] [Google Scholar]
- 65.Zanolari B, Riezman H. Mol Cell Biol. 1991;11:5251–5258. doi: 10.1128/mcb.11.10.5251. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 66.Peter M, Herskowitz I. Science. 1994;265:1228–1231. doi: 10.1126/science.8066461. [DOI] [PubMed] [Google Scholar]
- 67.Henchoz S, Chi Y, Catarin B, Herskowitz I, Deshaies R J, Peter M. Genes Dev. 1997;11:3046–3060. doi: 10.1101/gad.11.22.3046. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 68.Arvanitakis L, Geras-Raaka E, Varma A, Gershengorn M C, Cesarman E. Nature (London) 1997;385:347–350. doi: 10.1038/385347a0. [DOI] [PubMed] [Google Scholar]
- 69.Bais C, Santomasso B, Coso O, Arvanitakis L, Raaka E G, Gutkind J S, Asch A S, Cesarman E, Gershengorn M C, Mesri E A. Nature (London) 1998;391:86–89. doi: 10.1038/34193. [DOI] [PubMed] [Google Scholar]
- 70.Gottlieb T M, Oren M. Biochim Biophys Acta. 1996;1287:77–102. doi: 10.1016/0304-419x(95)00019-c. [DOI] [PubMed] [Google Scholar]
- 71.Ko L J, Prives C. Genes Dev. 1996;10:1054–1072. doi: 10.1101/gad.10.9.1054. [DOI] [PubMed] [Google Scholar]
- 72.Shieh S Y, Ikeda M, Taya Y, Prives C. Cell. 1997;91:325–334. doi: 10.1016/s0092-8674(00)80416-x. [DOI] [PubMed] [Google Scholar]
- 73.Jost C A, Marin M C, W. G. Kaelin J. Nature (London) 1997;389:191–194. doi: 10.1038/38298. [DOI] [PubMed] [Google Scholar]
- 74.Zeng X, Levine A J, Lu H. Proc Natl Acad Sci USA. 1998;95:6681–6686. doi: 10.1073/pnas.95.12.6681. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 75.Nicholson S E, Hilton D J. J Leukocyte Biol. 1998;63:665–668. doi: 10.1002/jlb.63.6.665. [DOI] [PubMed] [Google Scholar]
- 76.Starr R, Willson T A, Viney E M, Murray L J L, Rayner J R, Jenkins B J, Gonda T J, Alexander W S, Metcalf D, Nicola N A, et al. Nature (London) 1997;387:917–921. doi: 10.1038/43206. [DOI] [PubMed] [Google Scholar]
- 77.Endo T A, Masuhara M, Yokouchi M, Suzuki R, Sakamoto H, Mitsui K, Matsumoto A, Tanimura S, Ohtsubo M, Misawa H, et al. Nature (London) 1997;387:921–924. doi: 10.1038/43213. [DOI] [PubMed] [Google Scholar]
- 78.Naka T, Narazaki M, Hirata M, Matsumoto T, Minamoto S, Aono A, Nishimoto N, Kajita T, Taga T, Yoshizaki K, et al. Nature (London) 1997;387:924–929. doi: 10.1038/43219. [DOI] [PubMed] [Google Scholar]
- 79.Pear W S, Nolan G P, Scott M L, Baltimore D. Proc Natl Acad Sci USA. 1993;90:8392–8396. doi: 10.1073/pnas.90.18.8392. [DOI] [PMC free article] [PubMed] [Google Scholar]