Abstract
Biotin protein ligases (BPLs) are enzymes of extraordinary specificity. BirA, the BPL of Escherichia coli biotinylates only a single cellular protein. We report a mutant BirA that attaches biotin to a large number of cellular proteins in vivo and to bovine serum albumin, chloramphenicol acetyltransferase, immunoglobin heavy and light chains, and RNAse A in vitro. The mutant BirA also self biotinylates in vivo and in vitro. The wild type BirA protein is much less active in these reactions. The biotinylation reaction is proximity-dependent in that a greater extent of biotinylation was seen when the mutant ligase was coupled to the acceptor proteins than when the acceptors were free in solution. This approach may permit facile detection and recovery of interacting proteins by existing avidin/streptavidin technology.
Keywords: biotin protein ligase, protein modification, biotinylation, acyl adenylate, BirA
Biotinylation of proteins has routinely been done by chemical means, usually by modification of protein amino groups with biotin-N-hydroxysuccinimide or similar acylating agents. In contrast, enzymatic biotinylation has been limited to the few proteins that normally carry this modification, which are largely biotin-dependent carboxylases and decarboxylases of central metabolism (Chapman-Smith and Cronan Jr. 1999). We supposed that if enzymatic biotinylation could be made less specific, it might provide a means to detect weak (i.e., having dissociation constants >10−7 M) protein–protein interactions. In this scenario the biotinylating enzyme physically coupled to one of the interacting proteins (the target protein) would be used to biotinylate and thereby tag proteins that interact with the target protein. The biotinylation reaction should tag only those protein molecules that are close in space to the target protein. That is, unlike chemical acylation, enzymatic biotinylation should be proximity-dependent. The specific and extremely tight binding of biotin (KD 10−13 to 10−15 M) to streptavidin and avidin would then allow very sensitive detection of biotinylated proteins by a wide variety of robust protocols and low affinity forms of streptavidin and avidin allow efficient purification of biotinylated proteins under mild conditions. However, the biotin protein ligases (BPLs) catalyzing protein biotinylation have exceptional specificities for their protein substrates and thus are not general protein modification enzymes. For example, in vivo, BirA, the BPL of Escherichia coli, biotinylates only a single protein, the BCCP (AccB) subunit of acetyl-CoA carboxylase and other organisms contain less than five biotinylated protein species, indicating these BPLs are similarly specific (McAllister and Coon 1966; Samols et al. 1988; Chapman-Smith and Cronan Jr. 1999). Therefore, in order to use a BPL as a general biotinylating enzyme, this extraordinary specificity must somehow be overcome. A possible route to this end is based upon the mechanism of the BPL reaction, which proceeds in two steps:
Biotin + ATP ⇌ Bio-5′-AMP + PPi
Bio-5′-AMP + apo-Protein →Biotinoyl-Protein + AMP
In the first partial reaction, BPLs catalyze the synthesis of biotinoyl-AMP (bio-5′-AMP, which is also called biotinyl-adenylate) from ATP and biotin (McAllister and Coon 1966; Chapman-Smith and Cronan Jr. 1999). The enzyme then sequesters bio-5′-AMP in the active site until the second partial reaction proceeds. In the second partial reaction, the nucleophilic ɛ-amino group of the target lysine residue of a biotin-accepting domain attacks the mixed anhydride of the bio-5′-AMP bound within the BPL active site to form an amide bond between biotin and the lysine side chain that remains intact for the life of the protein (McAllister and Coon 1966; Chapman-Smith and Cronan Jr. 1999).
A possible means to convert a BPL to a promiscuous protein biotinylation enzyme would be to alter the protein such that the mutant enzyme releases bio-5′-AMP from the active site. Bio-5′-AMP is a mixed anhydride and therefore should act as a nonspecific chemical protein biotinylation reagent. A precedent for this scenario is the nonenzymatic acylation with serine of noncogate acceptor proteins by seryl-AMP released from an adenylation domain derived from EntF (Ehmann et al. 2000). Moreover, the instability of acyl-adenylates such as bio-5′-AMP should result in proximity-dependent biotinylation. This is because any acyl-adenylate molecules that diffuse far from the enzyme should be inefficient protein acylation reagents due to their low concentration and the high rate of acyl-adenylate hydrolysis.
The best studied BPL is E. coli BirA, a monomeric protein of 35.3 kDa. BirA is a multifunctional protein that catalyzes biotinylation of apo-BCCP and also acts as the transcriptional repressor that regulates biotin biosynthesis (Cronan Jr. 1989; Beckett and Matthews 1997). The X-ray crystallographic structure of BirA has been determined (Wilson et al. 1992; Weaver et al. 2001b) and mutations that affect biotinligase activity are located in a disordered loop that becomes more ordered when biotin occupies the active site (Weaver et al. 2001b). Two mutant proteins, G115S and R118G, having alterations within the loop, are defective in binding of both biotin and bio-5′AMP whereas ATP is bound normally. The dissociation constants of the G115S and R118G proteins for bio-5′-AMP binding are, respectively, 3000- and 400-fold greater than that of wild type BirA whereas the reported changes in the biotin binding constants are less dramatic (250- and 100-fold greater than wild type, respectively; Kwon and Beckett 2000). BirA Δ1–34, a protein lacking the N-terminal DNA-binding domain, also binds biotin and bio-5′-AMP weakly (dissociation constants 100- and 1000-fold greater than those of the wild type protein, respectively; Xu and Beckett 1996). These data suggested that these mutant proteins might leak bio-5′-AMP into the solvent where it would act as a promiscuous chemical biotinylation reagent in a proximity-dependent manner.
We report that a mutant BirA protein, R118G, acts as a promiscuous biotinylation reagent that shows proximity-dependence in a model system. Indeed, the R118G protein was found to efficiently biotinylate itself as well as a variety of proteins that normally lack this modification.
Results
Promiscuous in vivo biotinylation by R118G BirA
We first tested the possibility of obtaining a mutant BirA that gave promiscuous biotinylation by examining the patterns of biotinylation during expression of the mutant proteins in E. coli. We reasoned that any mutant enzyme that had gained promiscuous biotinylation ability would biotinylate E. coli proteins other than BCCP. The wild type and three mutant BirA proteins (R118G, G115S, and Δ 1–34) were expressed from a plasmid-based tac promoter (Kwon et al. 2002). High-level expression of each protein was confirmed by SDS-PAGE (data not shown). To assay for the ability to biotinylate proteins in a promiscuous manner, the extracted proteins of each strain were analyzed for biotin attachment by SDS-PAGE separation followed by Western blotting with a streptavidin-AP conjugate (Fig. 1 ▶). The extracts of all four strains showed the expected biotinylated BCCP band plus a band that corresponded to biotinylated BirA (see below). However, many additional biotinylated protein bands were seen in extracts of the cells in which the R118G mutant protein had been expressed. In contrast, the wild type and N-terminal deletion protein extracts showed weak labeling of noncognate proteins with the mutant protein giving stronger signals. Surprisingly, the G115S protein showed no labeling of noncognate proteins. The labeled bands seen in the R118G extract were strong evidence for promiscuous protein biotinylation and thus we proceeded to purify and study this mutant protein.
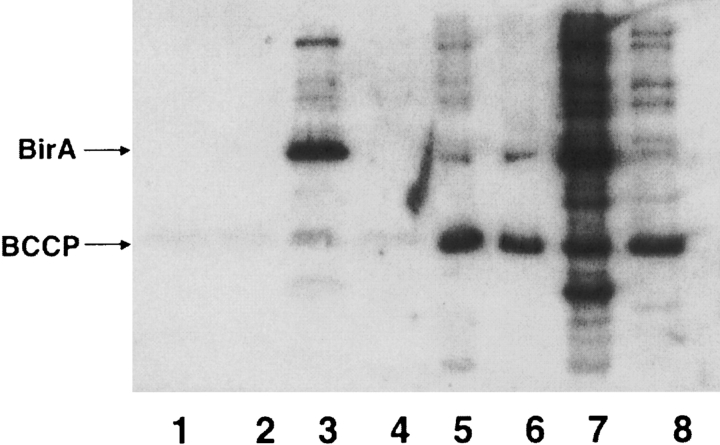
Figure 1.
Streptavidin Western blot analyses of crude cell extracts. Wild type and mutant BirA proteins were expressed in wild type strain JM109 as described in Materials and Methods. The crude extracts were separated by 12% SDS-PAGE, and analyzed by Western blotting with streptavidin-AP. Lanes 5–8 contained 10-fold more extract protein than lanes 1–4. (Lanes 1,5) Wild type BirA; (lanes 2,6) G115S BirA; (lanes 3,7) R118G BirA; and (lanes 4,8) Δ1–34 BirA (the N-terminal deletion mutant). The arrows indicate the bands corresponding to BirA and BCCP.
Promiscuous in vitro biotinylation by R118G
C-terminal hexahistidine-tagged (Histagged) wild type and R118G BirA proteins were purified by nickel-nitrilotriacetic acid (Ni-NTA) agarose chromatography. About 600 μg of each protein was obtained from 100 mL cultures with a purity of >95%. It should be noted that BirA is a robust protein that retains activity even when stored at room temperature. We performed in vitro biotinylation reactions using purified apo BCCP as a specific substrate and commercial BSA and/or RNAse A as promiscuous biotin acceptors. Western blotting with streptavidin-AP showed that both BSA and RNAse A became biotinylated with the R118G protein giving much greater modification than the wild type protein (Fig. 2 ▶). In contrast, both BirA proteins efficiently biotinylated the specific substrate, apo BCCP. The time courses of R118G-dependent biotinylation of apo BCCP and RNAse A were determined (Fig. 3 ▶). Biotinylation of the promiscuous acceptor protein increased throughout the 24 h incubation whereas biotinylation of apo BCCP reached a maximal level within 30 min.
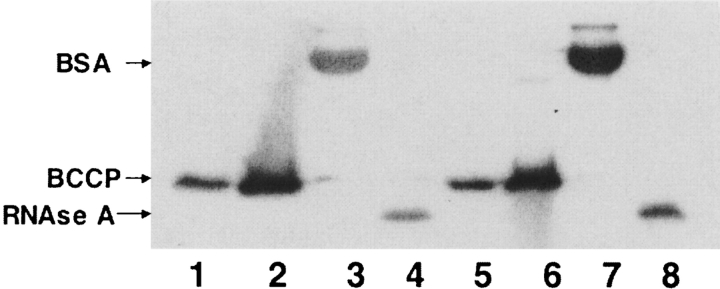
Figure 2.
In vitro biotinylation with purified wild type and R118G proteins. In vitro biotinylation reactions were initiated by addition of purified wild type or R118G mutant BirA proteins (to a final concentration of 20 nM) to a reaction mixture containing a 2 μM final concentration of a specific substrate (apo BCCP) or a promiscuous acceptor (BSA or RNAse A) followed by incubation for 1 h at 37°C as described in Materials and Methods, followed by SDS-PAGE. Biotinylation of each protein was then analyzed by Western blotting with streptavidin-AP. (Lane 1) apo BCCP lacking BirA (note that this apo BCCP preparation was slightly contaminated with holo BCCP accounting for the bands in lanes 1 and 5); (lane 2) apo BCCP with wild type BirA; (lane 3) BSA with wild type BirA; (lane 4) RNAse A with wild type BirA; (lane 5) apo BCCP lacking R118G BirA; (lane 6) apo BCCP with R118G BirA; (lane 7) BSA with R118G BirA; (lane 8) RNAse A with R118G BirA. The arrows show the migration positions of BSA, BCCP, and RNase A.
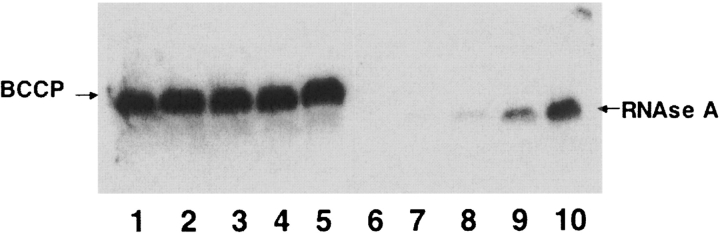
Figure 3.
Time course of in vitro biotinylation of apo BCCP and RNAse A with the wild type and R118G proteins. The experiment was done as described in Figure 2 ▶ and in Materials and Methods. Lanes 1–5 contained the specific substrate apo-BCCP whereas lanes 6–10 contained a promiscuous acceptor, RNAse A. Samples were taken after various time intervals. These were 30 min (lanes 1,6), 1 h (lanes 2,7), 2 h (lanes 3,8), 6 h (lanes 4,9), and 24 h (lanes 5,10). Biotinylation was analyzed by Western blotting with streptavidin-AP. The arrows indicate the bands corresponding to BCCP and RNase A.
BirA self-biotinylation activity
The crude extract of the bacterial strain (JM109, a strain that has a normal biotin synthetic pathway) that expressed high levels of the R118G protein showed many biotinylated protein bands (Fig. 1 ▶). One of the stronger bands had the molecular weight of BirA suggesting that R118G had become biotinylated. A less intense band was found at the same migration position for the other enzymes. We therefore tested the possibility that the purified wild type and R118G proteins had become biotinylated in vivo and found that only the R118G protein was highly modified (Fig. 4A ▶). When the R118G protein was expressed in an E. coli biotin auxotroph in the presence of 14C-labeled biotin, the molar ratio of covalently attached biotin to BirA was 0.4. Therefore, appreciable protein modification occurred during expression under biotin-sufficient conditions.
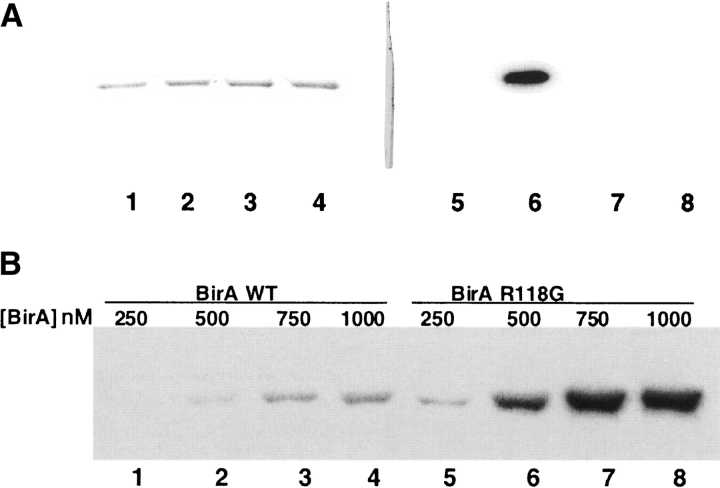
Figure 4.
Preparation and self-biotinylation of unbiotinylated BirA. (A) Wild type and R118G mutant BirA proteins were over expressed in either strain JM109 or in the bio strain BM4092 and purified as described in Materials and Methods. When strain BM4092 was used, the medium was depleted of biotin by addition of avidin to the 2XYT medium prior to induction of protein expression. An SDS-PAGE gel is shown. The left panel (lanes 1–4) was stained with Coomassie blue (1.4 mg protein per lane), whereas the right panel (lanes 5–8) was submitted to streptavidin-AP Western blot analysis (100 ng protein per lane). (Lanes 1,5) Wild type BirA expressed in JM109; (lanes 2,6) R118G BirA expressed in JM109; (lanes 3,7) wild type BirA expressed in BM4092; (lanes 4,8) R118G BirA expressed in BM4092. (B) The unbiotinylated preparations of the wild type (lanes 1–4) and R118G (lanes 5–8) proteins were used for in vitro self-biotinylation reactions (Materials and Methods). The BirA concentrations were varied as given on the figure. (Lanes 1,5) 250 nM; (lanes 2,6) 500 nM; (lanes 3,7) 750 nM; (lanes 4,8) 1 μM. After 1 h at 37°C a volume of each reaction corresponding to 2 pmole of protein was taken from of the various samples, run on SDS-PAGE, and analyzed by Western blotting with streptavidin-AP.
To obtain unbiotinylated R118G for further in vitro characterization of the self-biotinylation reaction, we expressed the protein in a strain (BM4092) defective in biotin biosynthesis and attachment (Barker and Campbell 1981). Avidin was added prior to protein induction to deplete the growth medium of free biotin. The absence of biotin available for the self biotinylation resulted in preparations of unmodified BirA proteins (Fig. 4A ▶). The in vitro biotinylation activity of these preparations toward apo BCCP, BSA, and RNase A was similar to that of the in vivo biotinylated R118G protein (data not shown). Upon incubation of the unmodified proteins with biotin and ATP, both proteins became biotinylated, although the mutant protein showed appreciably higher levels of self modification (Fig. 4B ▶).
BirA self biotinylation was expected to be primarily an intramolecular reaction since both the wild type and R118G proteins are monomeric at the concentrations we have tested (Kwon et al. 2002). To test the possible role of dimerization, we added a double-stranded oligonucleotide containing the bio operon regulatory region to the biotinylation reactions because specific DNA binding drives BirA dimerization (Streaker and Beckett 2003) and BirA dimerization in the presence or absence of DNA is thought to use the same protein interface (Weaver et al. 2001a). No increase in self biotinylation was seen (data not shown). Also, the levels of self biotinylation (obtained by quantitation of the data of Fig. 4B ▶; data not shown) were found to show a linear increase with BirA concentration. From these data self biotinylation seems predominantly an intramolecular reaction. The extent of R118G BirA self biotinylation in vitro was measured by assay of attachment of 14C-biotin. When incubated with 14C-biotin and ATP for 24 h, the molar ratio of covalently attached biotin to BirA was 0.2. Note that self biotinylation in vivo was also not caused by dimerization driven by specific DNA binding, as it occurred during expression in a host strain lacking the bio operator (data not shown).
Proximity-dependent biotinylation
As discussed above, biotinylation of promiscuous acceptor proteins by bio-5′-AMP released by BirA R118G would be expected to modify acceptors in a proximity-dependent manner. According to our original hypothesis this could result from the rapid hydrolysis of a mixed anhydride when in free solution at elevated pH, a property that should limit the ability of bio-5′-AMP to react with distant proteins. Another possibility is that the lysine side chains of promiscuous acceptor proteins somehow access the R118G active site. Our first indication of proximity-dependent biotinylation was the finding that a 10-fold molar excess of BSA failed to completely suppress self biotinylation of R118G (data not shown; similar data are given in Fig. 5 ▶) suggesting that R118G was preferentially modified compared to the acceptor protein. For a more general test system we used the His-tag present on the R118G C-terminus. In this case the biotin acceptors were the heavy and light chains of an antibody that recognizes a pentahistidine tag. We compared the relative biotinylation rates of the antibody when it was either bound to the R118G C-terminus or free in solution when bound to a His-tagged derivative of chloramphenicol acetyltransferase (CAT) (the antibody was blocked with a His-tagged CAT protein prior to addition of the His-tagged R118G). Hence, in the first case the acceptor protein would be located close to the BirA active site whereas in the second case the acceptor protein would be randomly distributed in solution. In some experiments we added BSA or RNAse A as promiscuous competitor proteins. The heavy and light chains of the antibody were found to be fair biotinylation acceptors when bound to the R118G C-terminus (Fig. 5 ▶), but were significantly less modified when the antibody was first blocked with His-tagged CAT to prevent binding of the antibody to the BirA His-tag. The blocking step reduced biotinylation of the antibody heavy chain to 22% of that seen in the absence of His-tagged CAT (the intensity of signal for light chain in the presence of His-tagged CAT was too weak for quantitation). The decrease in antibody biotinylation upon blocking cannot be attributed to competition for biotinylation between the His-tagged CAT protein and the antibody since addition of BSA or RNAse A (both of which are much better biotin acceptors than CAT) or CAT lacking a His-tag (data not shown), had little or no effect on biotinylation of the R118G-bound antibody (Fig. 5 ▶).
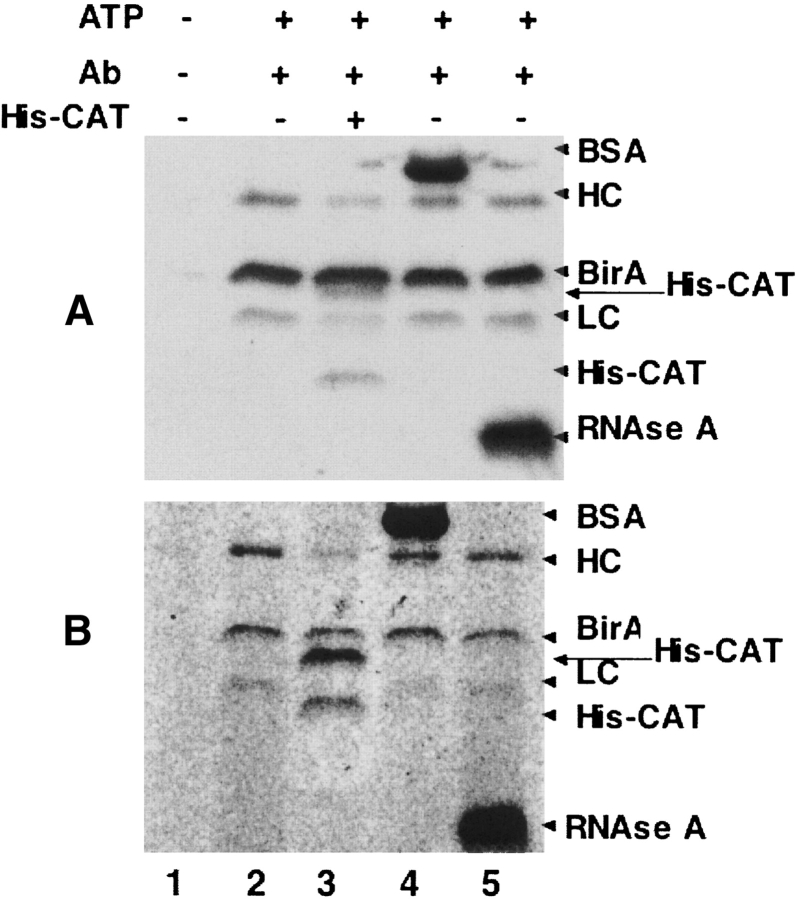
Figure 5.
Proximity-dependent biotinylation with R118G BirA. (A) His-tagged R118G BirA (final concentration 400 nM) was incubated in the reaction mixtures containing acceptor (Penta-HIS antibody) at a final concentration of 200 nM with or without the specific competitor (HIS-CAT, final concentration 2 μM) or with a promiscuous acceptor (either BSA or RNAse A, final concentration 2 μM) for 1 h at 37°C as described in Materials and Methods. The samples were then separated on an SDS-PAGE gel and analyzed by Western blot with streptavidin-AP. (Lane 1) His-tagged R118G BirA with omission of ATP; (lane 2) His-tagged R118G BirA plus Penta-HIS antibody; (lane 3) His-tagged R118G BirA plus Penta-HIS antibody preblocked with His-tagged CAT; (lane 4) His-tagged R118G BirA plus Penta-HIS antibody plus BSA; (lane 5) His-tagged R118G BirA plus Penta-HIS antibody plus RNAse A. (B) The reaction conditions were exactly the same as those of Figure 5A ▶, except that 0.25 μCi d-[carbonyl-14C]biotin was used as the biotin source. After the reaction mixtures were resolved in 12% SDS-PAGE, the gel was dried and exposed on X-ray film for one month at −80°C. The lanes are as in panel A. The arrows indicate BSA, the heavy (HC) and light (LC) chains of anti-Penta-HIS antibody, His-tagged BirA, His-tagged CAT (His-CAT), and RNase A. Note that the His-tagged CAT preparation contains a degradation product of the CAT protein in addition to the full-length protein.
We routinely assayed biotinylation by Western blots with detection by streptavidin-AP binding of biotinylated proteins. Although this is an established and very widely used technique, we have verified our results by a more direct assay, attachment of 14C-biotin. Biotin labeled with 14C was used in place of nonradioactive biotin in the in vitro biotinylation reaction. The reaction mixture was then loaded on an SDS-PAGE gel, which was dried and exposed to X-ray film. The results obtained by this direct method (Fig. 5B ▶) confirmed those obtained by the streptavidin-AP Western blot method, although the 14C-biotin method was much less sensitive (a 1 mo exposure was required) and the relative intensities of some bands were somewhat different from those given by streptavidin-AP staining presumably because of the steric problems of binding more than one streptavidin conjugate to a multiply biotinylated protein.
Discussion
We tested three different mutant BirA proteins for the ability to covalently attach biotin to proteins that normally do not carry this modification by assaying for promiscuous protein biotinylation in cells producing these enzymes. Only the R118G mutant protein showed a high level of promiscuous protein biotinylation. The N-terminal deletion mutant enzyme (Δ1–34) showed a lower level of promiscuous biotinylation whereas the third mutant enzyme, G115S, showed the same or a lower rate of biotinylation of noncognate proteins than did the wild type protein. This was surprising since G115S has the highest reported rate of bio-5′-AMP release of the mutant proteins (Kwon and Beckett 2000; Kwon et al. 2002). However, binding was assayed indirectly by loss of the increase in protein intrinsic fluorescence that occurs upon binding of bio-5′-AMP to BirA (Kwon and Beckett 2000). Therefore, the differences in promiscuous biotinylation levels observed for the G115S and R118G mutants could be explained if bio-5′-AMP does not completely exit the G115S active site but only shifts its position within the protein thereby resulting in loss of fluorescence. That is, bio-5′-AMP could be misaligned in G115S rather than released and might block access to the active site (note that the presence of the G115S mutation was confirmed in our experiments by checking for the unique restriction site introduced by the mutation).
All of the BirA proteins we studied undergo self biotinylation, although the R118G enzyme had much higher levels of self modification than the other proteins. The observed self modification was not unexpected, since two other enzymes that synthesize acyl-adenylate intermediates, the methionyl-tRNA synthetases of E. coli and Bacillus stearothermophilus, have been shown to catalyze self methionylation (Gillet et al. 1997; Hountondji et al. 2000). In these enzymes it was reported that once methionine was activated by ATP to form methionine-5′-AMP, the reactive lysine side chains closest to the acyl-adenylate were those modified (Hountondji et al. 2000). We expect that BirA self biotinylation proceeds by a similar mechanism. The low levels of self biotinylation and promiscuous protein biotinylation by wild type BirA observed in vivo seems likely to be a consequence of over expression of the enzyme resulting in an intracellular BirA concentration that greatly exceeds the supply of apo-BCCP, the normal biotin acceptor. In vitro methionyl-tRNA synthetase self modification is suppressed by addition of the tRNA acceptor (Gillet et al. 1997) and we expect the apo BCCP would have a similar effect on BirA self modification in vivo. Indeed, a wild type E. coli cell contains only about ten or so molecules of BirA and BCCP is normally present in great excess. However, this ratio becomes greatly skewed upon over expression of BirA leaving the enzyme without its normal protein substrate.
We have considered two possible mechanisms for promiscuous protein biotinylation. The first possibility is that biotinylation of such proteins is not a direct enzymatic activity of the R118G protein, but is due to chemical acylation by bio-5′-AMP released from the mutant active site. The second possibility is that lysine side chains of promiscuous acceptor proteins somehow gain access to the BirA active site due to the R118G mutation. We favor the first possibility because it seems unlikely that the BirA active site could be rearranged such to allow access by generic lysine residue side chains without losing the ability to sequester bio-5′-AMP from the solvent. Indeed, the target lysine residue of the naturally biotinylated proteins seems designed to be extraordinarily accessible. In the biotin domains of known structure, those of E. coli BCCP and Propionibacterium shermanii 1.3S proteins, the target lysine is located at the very tip of an exposed tight β-turn that markedly protrudes from the surface of the protein (Chapman-Smith and Cronan Jr. 1999; Reddy et al. 2000). Therefore, neither protein secondary structures nor neighboring side chains sterically hinder the ability of this lysine side chain to enter the BirA active site. Indeed, the lysine residue must be in this precise location to productively interact with the BirA active site as moving the residue one place in either direction abolishes biotinylation (Reche et al. 1998). Moreover, introduction of a second lysine residue adjacent to the target residue markedly inhibits biotinylation (Reche et al. 1998). For these reasons plus the precedent of chemical acylation by acyladenylates released by other enzymes (Gillet et al. 1997; Ehmann et al. 2000; Hountondji et al. 2000) we favor bio-5′-AMP release as the mechanism of promiscuous protein biotinylation. Indeed, Beckett and coworkers have shown that wild type BirA releases radioactive bio-5′-AMP synthesized from biotin and ATP (Xu and Beckett 1997; Kwon and Beckett 2000; Kwon et al. 2002). Further support for the involvement of bio-5′-AMP comes from an experiment (data not shown) in which BSA trapped in a dialysis bag was found to be slowly biotinylated in an ATP-dependent manner when R118G, biotin, and ATP were placed outside the bag (which excluded molecules greater than 10 kDa in size). It would seem very difficult for a lysine residue side chains to reach into the BirA active site across a dialysis membrane whereas bio-5′-AMP would diffuse across the membrane. However, further work will be required to define the mechanism of promiscuous protein biotinylation.
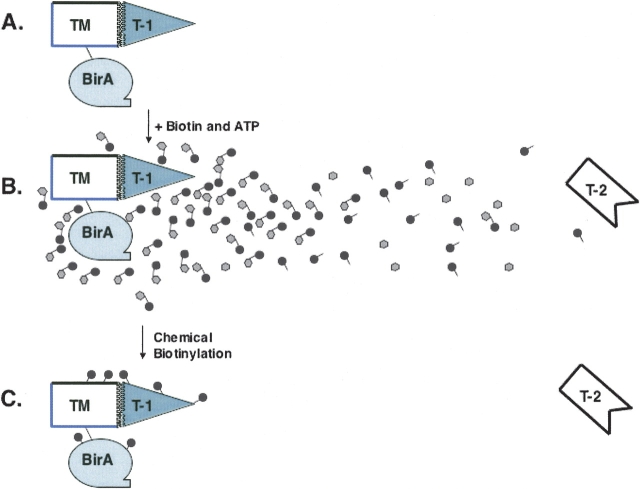
We envision the proximity-dependence of biotin attachment to result from a cloud of bio-5′-AMP emanating from the R118G BirA active site that acylates nearby protein molecules (Fig. 6 ▶). We have not attempted to optimize the proximity-dependence of protein biotinylation, because for unknown reasons acceptor proteins vary markedly in their efficiency of biotinylation (e.g., the antibody light and heavy chains). However, the thickness and range of the bio-5′-AMP cloud should be readily manipulated by choice of the experimental conditions. For example increases in buffer pH will decrease the half-life of bio-5′-AMP (BirA has a broad pH optimum) and small molecule nucleophiles (e.g., amines, thiols) added to the reaction buffer will react with bio-5′-AMP and thereby block labeling of distant proteins. A counter argument to our claim of proximity-dependent biotinylation is that all we have done is to increase the local concentration of the acceptor proteins. However, proximity mandates localization and localization necessarily results in increased concentration.
Figure 6.
A depiction of proximity-dependent biotinylation. (A) A conjugate (or a fusion protein) composed of a mutant BirA and a target module (TM) of interest is exposed to a mixture of target (T) molecules. Some target molecules (T-1) bind the TM whereas others (T-2) fail to bind. (B) Upon addition of biotin and ATP the mutant BirA synthesizes bio-5′-AMP, which is depicted as a solid circle (the biotin moiety) linked to AMP (the shaded hexagon). (C) The localized cloud of bio-5′-AMP chemically bio-tinylates all of the proteins in the vicinity of BirA, T-1, TM, and BirA. The T-2 molecules fail to become biotinylated because most bio-5′-AMP molecules are hydrolyzed (shown as unlinked circles and hexagons) before they can reach the distant T-2 molecules. For simplicity, the process is shown in stages, although in practice, the interaction of TM and T-1, bio-5-AMP synthesis, and chemical biotinylation would proceed simultaneously.
It remains to be seen if promiscuous enzymatic protein biotinylation will be useful. The rates and extents of the BirA R118G-dependent reactions are modest at best. Logical design of more active proteins is hampered by the lack of a crystal structure of BirA complexed with bio-5′-AMP. The only co-crystal structures presently available are BirA complexed to biotinoyl-lysine (Wilson et al. 1992) and to biotin (Weaver et al. 2001b). Neither structure aids study of the protein biotinylation reaction in that the former structure is an enzyme–product complex and the latter structure gives no information on the major conformational changes known to accompany bio-5′-AMP synthesis and binding (Xu and Beckett 1997; Kwon et al. 2000; Weaver et al. 2001b). However, two nonhydrolysable bio-5′-AMP analogs have recently been found to bind BirA and one of these, biotinol-5′-AMP, bound almost as well as bio-5′-AMP (Brown et al. 2004). The stability of these analogs should allow deduction of the structure of the BirA-bio-5′-AMP complex and perhaps design of enzymes more useful than R118G. Another approach to isolate more useful enzymes, that of random screening for mutant enzymes giving higher rates and extents of promiscuous biotinylation should be facilitated by the fact that BirA can be expressed in an active form on the surface of phage M13 (Heinis et al. 2001). Finally, thermophilic and psychrophilic bacteria have BPLs very similar to that of E. coli and the use of mutant forms of these enzymes should allow access to a very wide range of biotinylation conditions.
Materials and methods
Chemicals and reagents
Ni-NTA agarose from Qiagen was used for column chromatography and an anti-pentahistidine antibody (Penta-His Antibody, BSA-free from Qiagen) was used as an acceptor for assay of proximity-dependent biotinylation. A streptavidin-alkaline phosphatase conjugate (Streptavidin-AP) and CDP-Star (Roche) were used for Western blot analysis. BSA, avidin, and RNAse A (Sigma) were used as either promiscuous acceptors or promiscuous competitors. Apo BCCP was prepared as described previously (Choi-Rhee and Cronan 2003). Amersham provided d-[carbonyl-14C]biotin that was dissolved in 100 μL of 0.1 M sodium phosphate buffer (pH 7.0) containing 0.15 M sodium chloride and stored at −20°C.
Expression and purification of the BirA proteins
Plasmids (Xu and Beckett 1996; Kwon and Beckett 2000) carrying wild type and two mutant (G115S and R118G) birA genes each having an extension encoding a C-terminal hexa-histidine sequence (His-tag) plus an untagged ΔN1–34 gene (the kind gifts of Dr. Dorothy Beckett) were introduced into E. coli strain JM109. E. coli strain BM4092 (Barker and Campbell 1981) was used as a host strain for isolation of unbiotinylated BirA protein. Cultures were grown in 2XYT medium containing 100 mg/L sodium ampicillin, and 5 μM biotin. Protein expression was induced at an optical density of 0.5 at 600 nm by addition of isopropyl-β-galactoside (IPTG) to 1 mM and allowed to proceed for 4 h. For biotin depletion of 2XYT medium, cells were harvested at an optical density of 0.5 at 600 nm and resuspended in 2XYT medium lacking added biotin and containing avidin at a final concentration of 2U/mL. IPTG (1 mM) was added to induce protein expression, and induction was allowed to proceed for 4 h. The cells were then collected by centrifugation and either resuspended in SDS sample buffer and analyzed by streptavidin-AP blotting or resuspended in lysis buffer (50 mM sodium phosphate at pH 8.0, 300 mM NaCl, 0.1 mM DTT) for protein purification. Cells were lysed by sonication and extracts were incubated with Ni-NTA agarose for 1 h at 4°C with gentle agitation. After washing the resin with 10 volumes of 25 mM imidazole in lysis buffer, the protein was eluted with 250 mM imidazole in lysis buffer. The eluate was dialyzed against storage buffer (40 mM Tris-HCl at pH 8.0, 100 mM KCl, 10% glycerol) overnight. The concentrations of the purified proteins were adjusted to 2 μM and kept at −80°C until use.
Expression and purification of the His-tagged form of chloramphenicol acetyltransferase (His-CAT)
Plasmid pER50 was constructed by cutting pCY572 (Cronan 2003) with SalI plus XhoI to obtain a 1.2 Kb fragment containing the CAT gene, and ligating this fragment into pET28b (Novagen) that has been cut with XhoI, and screening for an insert having the proper orientation by restriction mapping. Plasmid pER50 was introduced into E. coli strain BL21 (DE3) and cultures were grown in Luria Bertani (LB) medium containing 50 mg/L of kanamycin sulfate and 30 mg/L of chloramphenicol to an optical density of 0.5 at 600 nm. CAT expression was induced by addition of ITPG to 1 mM and allowed to proceed for 30 min at 30°C. The cells were harvested and the protein was purified by Ni-NTA agarose chromatography as described above.
In vitro biotinylation
In vitro biotinylation was done as described previously (Chapman-Smith et al. 1999) with some modifications. Unless otherwise stated, the assays contained 40 mM Tris-HCl (pH 8.0), 3 mM ATP, 5.5 mM MgCl2, 5 μM biotin, 100 mM KCl, 1.4 mM β-mercap-toethanol, plus the indicated concentrations of substrate or acceptor in a final volume of 20 μL. The reaction was initiated by addition of purified BirA protein to a final concentration of 20 nM followed by incubation for up to 24 h at 37°C. All of the BirA proteins were saturated with substrates at these concentrations. For the self-biotinylation reaction, purified BirA proteins were used as both enzyme and substrate. For assay of proximity-dependent bio-tinylation, Penta-HIS antibody (final concentration 200 nM) was incubated with 2 μM (final concentration) of competitor protein (His-tagged-CAT, BSA, or RNAse A) for 30 min at room temperature. The reaction was initiated by addition of purified BirA protein to a final concentration of 400 nM and incubated for 1 h at 37°C. For biotinylation reactions with 14C-biotin, the biotin was replaced with 0.25 μCi of 14C-biotin in a final volume of 20 μL.
Western blot analysis with streptavidin
Crude extracts or purified proteins were loaded on 12% SDS-PAGE gels (Choi-Rhee and Cronan 2003). The separated proteins were transferred to a membrane (Immobilon-P from Millipore) and the membrane was briefly rinsed with maleic buffer (100 mM maleic acid plus 150 mM NaCl adjusted to pH 7.5 with NaOH). The membrane was incubated first with blocking buffer consisting of 1% Western Blocking Reagent (Roche) in maleic acid buffer for 30 min and then with streptavidin-AP-conjugate in maleic acid buffer for another 30 min at room temperature. After rinsing twice with washing buffer (0.3% Tween 20 in maleic acid buffer) for 15 min, the membrane was incubated with CDP-Star in detection buffer (100 mM Tris-HCl at pH 9.5, 100 mM NaCl) for 5 min. The membrane was then exposed to X-ray film.
Acknowledgments
This work was supported by SurroMed, Inc., and NIH grant AI15650.
Article published online ahead of print. Article and publication date are at http://www.proteinscience.org/cgi/doi/10.1110/ps.04911804.
References
- Barker, D.F. and Campbell, A.M. 1981. Genetic and biochemical characterization of the birA gene and its product: Evidence for a direct role of biotin holoenzyme synthetase in repression of the biotin operon in Escherichia coli. J. Mol. Biol. 146 469–492. [DOI] [PubMed] [Google Scholar]
- Beckett, D. and Matthews, B.W. 1997. Escherichia coli repressor of biotin biosynthesis. Methods Enzymol. 279 362–376. [DOI] [PubMed] [Google Scholar]
- Brown, P.H., Cronan, J.E., Grotli, M., and Beckett, D. 2004. The biotin repressor: Modulation of allostery by corepressor analogs. J. Mol. Biol. 337 857–869. [DOI] [PubMed] [Google Scholar]
- Chapman-Smith, A. and Cronan Jr., J.E. 1999. The enzymatic biotinylation of proteins: A post-translational modification of exceptional specificity. Trends Biochem. Sci. 24 359–363. [DOI] [PubMed] [Google Scholar]
- Chapman-Smith, A., Morris, T.W., Wallace, J.C., and Cronan Jr., J.E. 1999. Molecular recognition in a post-translational modification of exceptional specificity. Mutants of the biotinylated domain of acetyl-CoA carboxylase defective in recognition by biotin protein ligase. J. Biol. Chem. 274 1449–1457. [DOI] [PubMed] [Google Scholar]
- Choi-Rhee, E., and Cronan, J.E. 2003. The biotin carboxylase-biotin carboxyl carrier protein complex of Escherichia coli acetyl-CoA carboxylase. J. Biol. Chem. 278 30806–30812. [DOI] [PubMed] [Google Scholar]
- Cronan Jr., J.E. 1989. The E. coli bio operon: Transcriptional repression by an essential protein modification enzyme. Cell 58 427–429. [DOI] [PubMed] [Google Scholar]
- Cronan, J.E. 2003. Cosmid-based system for transient expression and absolute off-to-on transcriptional control of Escherichia coli genes. J. Bacteriol. 185 6522–6529. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ehmann, D.E., Shaw-Reid, C.A., Losey, H.C., and Walsh, C.T. 2000. The EntF and EntE adenylation domains of Escherichia coli enterobactin synthetase: Sequestration and selectivity in acyl-AMP transfers to thiolation domain cosubstrates. Proc. Natl. Acad. Sci. 97 2509–2514. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gillet, S., Hountondji, C., Schmitter, J.M., and Blanquet, S. 1997. Covalent methionylation of Escherichia coli methionyl-tRNA synthethase: Identification of the labeled amino acid residues by matrix-assisted laser desorption-ionization mass spectrometry. Protein Sci. 6 2426–2435. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Heinis, C., Huber, A., Demartis, S., Bertschinger, J., Melkko, S., Lozzi, L., Neri, P., and Neri, D. 2001. Selection of catalytically active biotin ligase and trypsin mutants by phage display. Protein Eng. 14 1043–1052. [DOI] [PubMed] [Google Scholar]
- Hountondji, C., Beauvallet, C., Pernollet, J.C., and Blanquet, S. 2000. Enzyme-induced covalent modification of methionyl-tRNA synthetase from Bacillus stearothermophilus by methionyl-adenylate: Identification of the labeled amino acid residues by matrix-assisted laser desorption-ionization mass spectrometry. J. Protein Chem. 19 563–568. [DOI] [PubMed] [Google Scholar]
- Kwon, K., and Beckett, D. 2000. Function of a conserved sequence motif in biotin holoenzyme synthetases. Protein Sci. 9 1530–1539. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kwon, K., Streaker, E.D., Ruparelia, S., and Beckett, D. 2000. Multiple disordered loops function in corepressor-induced dimerization of the biotin repressor. J. Mol. Biol. 304 821–833. [DOI] [PubMed] [Google Scholar]
- Kwon, K., Streaker, E.D., and Beckett, D. 2002. Binding specificity and the ligand dissociation process in the E. coli biotin holoenzyme synthetase. Protein Sci. 11 558–570. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McAllister, H.C., and Coon, M.J. 1966. Further studies on the properties of liver propionyl coenzyme A holocarboxylase synthetase and the specificity of holocarboxylase formation. J. Biol. Chem. 241 2855–2861. [PubMed] [Google Scholar]
- Reche, P., Li, Y.L., Fuller, C., Eichhorn, K., and Perham, R.N. 1998. Selectivity of post-translational modification in biotinylated proteins: The carboxy carrier protein of the acetyl-CoA carboxylase of Escherichia coli. Biochem. J. 329 (Pt. 3) 589–596. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy, D.V., Shenoy, B.C., Carey, P.R., and Sonnichsen, F.D. 2000. High resolution solution structure of the 1.3S subunit of transcarboxylase from Propionibacterium shermanii. Biochemistry 39 2509–2516. [DOI] [PubMed] [Google Scholar]
- Samols, D., Thornton, C.G., Murtif, V.L., Kumar, G.K., Haase, F.C., and Wood, H.G. 1988. Evolutionary conservation among biotin enzymes. J. Biol. Chem. 263 6461–6464. [PubMed] [Google Scholar]
- Streaker, E.D. and Beckett, D. 2003. Coupling of protein assembly and DNA binding: Biotin repressor dimerization precedes biotin operator binding. J. Mol. Biol. 325 937–948. [DOI] [PubMed] [Google Scholar]
- Weaver, L.H., Kwon, K., Beckett, D., and Matthews, B.W. 2001a. Competing protein:protein interactions are proposed to control the biological switch of the E. coli biotin repressor. Protein Sci. 10 2618–2622. [DOI] [PMC free article] [PubMed] [Google Scholar]
- ———. 2001b. Corepressor-induced organization and assembly of the biotin repressor: A model for allosteric activation of a transcriptional regulator. Proc. Natl. Acad. Sci. 98 6045–6050. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wilson, K.P., Shewchuk, L.M., Brennan, R.G., Otsuka, A.J., and Matthews, B.W. 1992. Escherichia coli biotin holoenzyme synthetase/bio repressor crystal structure delineates the biotin- and DNA-binding domains. Proc. Natl. Acad. Sci. 89 9257–9261. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Xu, Y. and Beckett, D. 1996. Evidence for interdomain interaction in the Escherichia coli repressor of biotin biosynthesis from studies of an N-terminal domain deletion mutant. Biochemistry 35 1783–1792. [DOI] [PubMed] [Google Scholar]
- ———. 1997. Biotinyl-5′-adenylate synthesis catalyzed by Escherichia coli repressor of biotin biosynthesis. Methods Enzymol. 279 405–421. [DOI] [PubMed] [Google Scholar]