Abstract
Haemophilus influenzae is a major cause of otitis media and other respiratory tract disease in children. The pathogenesis of disease begins with colonization of the upper respiratory mucosa, a process that involves evasion of local immune mechanisms and adherence to epithelial cells. Several studies have demonstrated that human milk is protective against H. influenzae colonization and disease. In the present study, we examined the effect of human milk on the H. influenzae IgA1 protease and Hap adhesin, two autotransported proteins that are presumed to facilitate colonization. Our results demonstrated that human milk lactoferrin efficiently extracted the IgA1 protease preprotein from the bacterial outer membrane. In addition, lactoferrin specifically degraded the Hap adhesin and abolished Hap-mediated adherence. Extraction of IgA1 protease and degradation of Hap were localized to the N-lobe of the bilobed lactoferrin molecule and were inhibited by serine protease inhibitors, suggesting that the lactoferrin N-lobe may contain serine protease activity. Additional experiments revealed no effect of lactoferrin on the H. influenzae P2, P5, and P6 outer-membrane proteins, which are distinguished from IgA1 protease and Hap by the lack of an N-terminal passenger domain or an extracellular linker region. These results suggest that human milk lactoferrin may attenuate the pathogenic potential of H. influenzae by selectively inactivating IgA1 protease and Hap, thereby interfering with colonization. Future studies should examine the therapeutic potential of lactoferrin, perhaps as a supplement in infant formulas.
Acute otitis media is a suppurative infection of the middle ear and is especially common during early childhood (1). By 3 years of age, 80% of children have suffered from acute otitis media and 40–50% have experienced at least three episodes (2). Otitis media accounts for over one-third of pediatric office visits in the United States and is the most common reason for prescription of oral antibiotics (3). After each episode, fluid persists in the middle ear for weeks to months, causing hearing impairment (4) that sometimes results in deficiencies in language acquisition, speech development, and cognitive achievement (5).
Most cases of otitis media are caused by infection with Streptococcus pneumoniae, Haemophilus influenzae, or Moraxella catarrhalis (6, 7). Infection begins with colonization of the nasopharynx, followed by contiguous spread through the eustachian tube to the middle ear. Colonization is a complex process and involves the interplay of bacterial and host factors. Successful colonization requires that an organism evade local immune mechanisms and overcome the mucociliary escalator. Both S. pneumoniae and H. influenzae secrete an IgA1 protease, which specifically cleaves and inactivates human IgA1, the predominant secretory antibody in the upper respiratory tract (8, 9). All three of these respiratory pathogens also elaborate adhesins, which promote attachment to host epithelium and are critical in preventing physical removal from the mucosal surface.
IgA1 proteases are the prototypes of a family of Gram-negative bacterial proteins that undergo a process known as autosecretion (10). The H. influenzae strain Rd protease is synthesized as a 185-kDa protein with four domains, including an N-terminal signal sequence, an internal serine protease domain (Igap), a highly basic and helical alpha domain (Igaα), and a C-terminal beta or helper domain (Igaβ) (11, 12). The signal sequence directs export across the bacterial inner membrane and is then cleaved. Subsequently, the remainder of the protein (hereafter called the preprotein) inserts into the outer membrane by way of the beta domain, which is predicted to form a β-barrel structure with a hydrophilic channel, thus allowing for translocation of the protease and the alpha domain to the extracellular space. Ultimately, the protease domain gains catalytic activity and cleaves within the alpha domain to release itself from the surface of the organism.
The H. influenzae Hap protein is a nonpilus protein that promotes intimate interaction with human epithelium (13). It was originally identified on the basis of its ability to confer a capacity for in vitro attachment and invasion when expressed in a nonadherent, noninvasive laboratory strain of H. influenzae (13). Hap shares significant sequence homology (30–35% identity and 51–55% similarity) with the H. influenzae and Neisseria gonorrhoeae IgA1 proteases and undergoes autosecretion by means of an analogous pathway. Hap is produced as a 155-kDa protein with three functional domains, including an N-terminal signal sequence, a surface-localized serine protease domain (Haps), and a C-terminal outer membrane domain (Hapβ) (14). To date, there is no evidence for an alpha domain. Ultimately, the Haps domain mediates an autoproteolytic cleavage event, releasing itself from Hapβ and from the surface of the organism. Recent evidence indicates that attachment to host cells is a function of the preprotein (Haps linked to Hapβ), before autoproteolytic cleavage (D.R.H. and J.W.St.G., unpublished work).
A number of studies have demonstrated that breast-fed infants have fewer episodes of otitis media and a lower incidence of nasopharyngeal colonization with S. pneumoniae, H. influenzae, and M. catarrhalis than do formula-fed infants (2, 16–20). Human breast milk contains secretory IgA and several nonantibody proteins with antibacterial properties, including lactoferrin, lysozyme, and peroxidase. The precise mechanism by which these factors influence bacterial colonization of the nasopharynx and spread to the middle ear has not been established.
In the present study, we examined interactions between human milk and H. influenzae and observed that lactoferrin efficiently extracted the IgA1 protease preprotein from the bacterial outer membrane. We also found that lactoferrin specifically degraded and inactivated the Hap adhesin. Both extraction and degradation were inhibited by pretreatment of lactoferrin with broad serine protease inhibitors, suggesting that lactoferrin may have protease activity. Experiments with three additional H. influenzae outer-membrane proteins revealed no effect, suggesting that this newly recognized activity may be specific for certain autotransported proteins.
MATERIALS AND METHODS
Bacterial Strains and Plasmids and Growth Conditions.
H. influenzae strain Rd is a capsule-deficient serotype d strain that secretes IgA1 protease but contains a nonfunctional hap gene because of a spontaneous nonsense mutation at codon 710 (13, 21). H. influenzae strain Rd3–13 is a derivative of Rd with a mutant IgA1 protease that lacks protease activity and is locked in the full-length state; it was constructed by Y. Fishman and A. Wright (Department of Molecular Biology and Microbiology, Tufts University School of Medicine, Boston, MA) and contains a valine residue in place of the active site serine. H. influenzae strain DB117 is a derivative of Rd with a mutation in the rec-1 gene and is deficient in recombination (22). H. influenzae strain 1479 is a clinical isolate that was originally recovered from the sputum of a patient with chronic bronchitis and is serologically nontypable (23).
The plasmid pJS106 contains the wild-type hap gene in the vector pGJB103 and was described previously (13). pHapS243A encodes a Hap derivative that contains an alanine residue in place of the active site serine at position 243 (14); the mutant protein is locked in the full-length form and remains cell-associated (ref. 14; D.R.H. and J.W.St.G., unpublished work). pHapβ encodes a Hap derivative composed of the Hap signal sequence fused directly to the Hap C-terminal domain (Hapβ), thus lacking the central Haps domain (D.R.H. and J.W.St.G., unpublished work).
H. influenzae strains were grown as described previously (24) and were stored at −80°C in brain–heart infusion broth with 20% glycerol.
Preparation of Human Milk Samples.
After obtaining informed consent, human milk was collected between 3 days and 4 months postpartum from healthy mothers taking no antibiotics. All samples were collected in sterile beakers and within 6 hr of collection were centrifuged at 10,000 × g for 20 min at 4°C to remove lipids and cells. The resulting milk whey was stored at −80°C and was prepared for use by thawing slowly, without further modifications.
Milk whey proteins were fractionated by first precipitating with 40 vol of cold acetone, then subjecting to anion exchange (DE-52, Whatman) and molecular sieve chromatography (Bio Gel P-200, Bio-Rad). All steps were carried out in neutral buffers at either room temperature or 4°C.
Human Milk and Lactoferrin Studies.
Recombinant lactoferrin and recombinant lactoferrin N-lobe (residues 1–334) were purified from baby hamster kidney (BHK) cells, as previously described (25, 26). Recombinant lactoferrin purified from Aspergillus awamori was a generous gift from Agennix Corporation (Houston, TX) and was deglycosylated according to the method of Baker et al. (27).
The purity of recombinant protein preparations was confirmed by mass spectroscopy using a linear matrix-assisted laser desorption ionization-time-of-flight instrument (Voyager, PerSeptive Biosystems, Framingham, MA). Lactoferrin iron content was varied as described by Mazurier and Spik (28).
In experiments examining the effect of milk components on bacterial surface proteins, bacteria were grown to mid-logarithmic phase and then pelleted in a microcentrifuge. Subsequently, 2 × 109 organisms were resuspended in 1 ml of human milk whey or in 1 ml of PBS containing lactoferrin or lactoferrin N-lobe, then incubated at 37°C with gentle mixing (29). After incubation, bacteria were collected by centrifugation and resuspended in Laemmli buffer (30) for analysis by Western blot analysis. Proteins that remained in the supernatants after centrifugation were collected by precipitation with 10% trichloroacetic acid, as previously described (13), and then analyzed by Western blotting.
Protein Analysis.
Proteins were resolved by SDS/PAGE, and Western blottings were performed according to standard techniques (30, 31). Antibodies included: rabbit antiserum #331, which recognizes the IgA1 protease preprotein, Igap, and Igaβ (29); rabbit antiserum Rab730, which reacts with the Hap preprotein, Haps, and Hapβ (14); mouse mAb 6G3, which reacts with the P2 outer-membrane protein from H. influenzae strain 1479 (23); mouse mAb 2C7, which reacts with the H. influenzae P5 outer-membrane protein; and mouse mAb 7F3, which reacts with the H. influenzae P6 outer-membrane protein (32).
Adherence Assays.
Adherence assays were performed with Chang epithelial cells [Wong-Kilbourne derivative, clone 1–5c-4 (human conjunctiva), as previously described (33)].
RESULTS
Incubation of Bacteria in Human Milk Whey Results in Extraction and Degradation of the H. influenzae IgA1 Protease.
In initial experiments, H. influenzae strain Rd was resuspended in human milk whey and incubated for 1 hr. After incubation, whole cells and supernatants were examined by Western immunoblotting using an antiserum that recognizes the IgA1 protease preprotein, Igap, and Igaβ. As shown in Fig. 1A, the entire IgA1 protease preprotein and any Igaβ generated from processing of the preprotein were removed from cells. The extracted preprotein was readily detected in the supernatant, while the extracted Igaβ domain was undetectable, suggesting rapid degradation.
Figure 1.
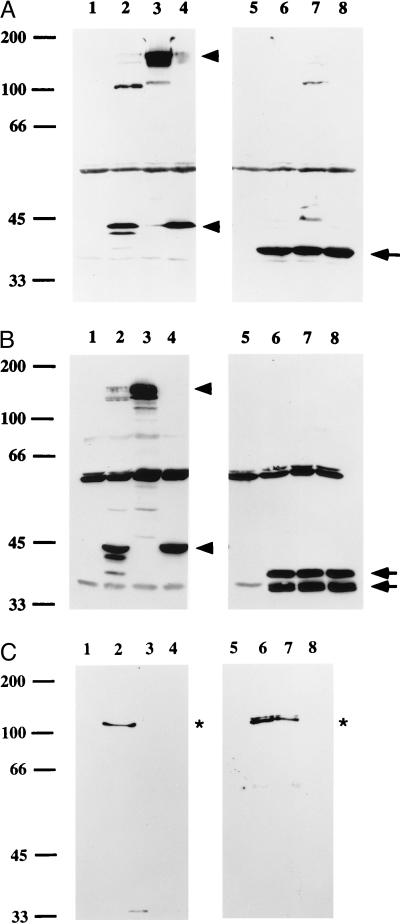
Effect of human milk whey and human lactoferrin on H. influenzae IgA1 protease. (A) Western immunoblot showing that human milk whey removes the native IgA1 protease preprotein and the remnant Igaβ domain from H. influenzae strain Rd. The immunoblot was prepared with rabbit antiserum #331, which reacts with the IgA1 protease preprotein, Igap, and Igaβ. Samples were loaded as follows: lane 1, untreated Rd, whole cells; lane 2, untreated Rd, supernatant; lane 3, milk whey-treated Rd, whole cells; lane 4, milk whey-treated Rd, supernatant. Lane 1 shows the preprotein (P) and the remnant Igaβ domain (β) generated from processing of the preprotein. Lane 2 shows the two active IgA1 protease species released from the surface of the cell. Lanes 3 and 4 demonstrate that the preprotein and Igaβ were removed from whole cells, with the preprotein detectable in the supernatant (∗). The preprotein in the supernatant remained unprocessed, since milk contains antibody that inhibits autoproteolysis. The secondary antibody crossreacts with lactoferrin (Lf), which is indicated by the open arrow. (B) Western immunoblot showing removal of the IgA1 protease preprotein from H. influenzae strain Rd3–13 as a function of time. Bacteria were grown to mid-logarithmic phase and incubated with milk whey, and aliquots were then sampled at the indicated times. Whole cells (Upper) and the corresponding supernatants (Lower) were examined by using rabbit antiserum #331. The preprotein was progressively removed from cells and transferred to the supernatant. In control samples prepared from strain Rd3–13 after incubation in buffer alone (lane C), the preprotein remained associated with cells. Arrowheads point to the preprotein, and the bracket indicates a degradation product of the preprotein. ∗ marks the upward-shifted preprotein. (C) Western immunoblot showing that recombinant lactoferrin is sufficient to remove the IgA1 protease preprotein from H. influenzae strain Rd3–13. The immunoblot was prepared with rabbit antiserum #331. Lanes 2 and 3 contain samples that were treated with human milk lactoferrin, and lanes 6 and 7 contain samples that were treated with recombinant human lactoferrin prepared from BHK cells. Samples were loaded as follows: lanes 1 and 5, untreated Rd3–13, whole cells; lanes 2 and 6, milk lactoferrin or recombinant lactoferrin-treated Rd3–13, whole cells; lanes 3 and 7, milk lactoferrin or recombinant lactoferrin-treated Rd3–13, supernatants; lanes 4 and 8, milk lactoferrin or recombinant lactoferrin, without bacteria. Both milk lactoferrin and recombinant lactoferrin removed the preprotein (∗), which was then partially degraded (brackets). Both sources of lactoferrin resulted in an upward shift of the preprotein. The secondary antibody crossreacts with lactoferrin (Lf), which is indicated by the open arrow.
To examine extraction of the preprotein more closely, we used strain Rd3–13, an Rd derivative that expresses enzymatically inactive IgA1 protease, which accumulates in the bacterial outer membrane as the unprocessed preprotein. As shown in Fig. 1B, when strain Rd3–13 was incubated in milk whey, the preprotein was extracted from cells in a time-dependent manner and was again transferred to the supernatant. Assessment at early time points (2, 5, and 10 min) revealed an increase in apparent molecular mass of the cell-associated preprotein just before extraction, presumably reflecting a change in conformation and resultant anomalous migration. After 10 min of incubation in milk, only a small amount of IgA1 protease preprotein remained cell-associated, and by 60 min, extraction was complete. After transfer to the supernatant, the protein was slowly degraded to lower molecular-weight species. Quantitation of colony-forming units of Rd or Rd3–13 after incubation in milk for 2 hr demonstrated no effect on viability (not shown).
Lactoferrin Is Responsible for Milk-Derived Extraction and Degradation of the H. influenzae IgA1 Protease.
To determine the constituent(s) of human milk whey responsible for extraction of the H. influenzae IgA1 protease preprotein, milk proteins were separated by size-exclusion chromatography, and the resulting fractions were tested for activity. In experiments with both Rd and Rd3–13, only fractions containing lactoferrin reproduced the findings with milk whey (Fig. 1C). Active fractions contained no IgA, indicating that extraction was independent of milk antibody.
To confirm that lactoferrin alone was capable of extraction, assays were performed with preparations of recombinant human lactoferrin produced in either BHK cells or A. awamori. In addition, recombinant lactoferrin N-lobe was examined, again after purification from BHK cells. Recombinant protein was used in a concentration of 1 mg/ml (13 μM), approximating the value reported for lactoferrin in human milk (34). As shown in Fig. 1C, lactoferrin purified from BHK cells removed the IgA protease preprotein from strain Rd3–13 and then degraded the extracted protein. Deglycosylated recombinant lactoferrin from A. awamori and recombinant lactoferrin N-lobe from BHK cells had an identical effect (not shown). To ensure that no other proteins were present in the preparations of recombinant protein, molecular mass measurements were performed by using mass spectroscopy. Intact glycosylated lactoferrin from BHK cells was 79,338 Da, and glycosylated lactoferrin N-lobe was 36,890 Da, in both cases very close to predicted values. Additional analysis demonstrated that lactoferrin iron content had no influence on either extraction or degradation (not shown).
Human Lactoferrin Degrades and Inactivates the H. influenzae Hap Adhesin.
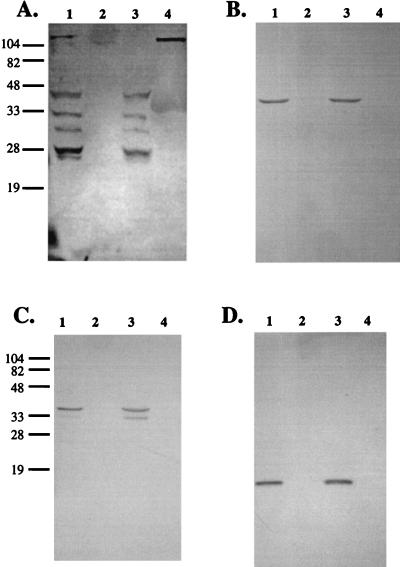
To address whether lactoferrin also extracts other autotransported proteins from H. influenzae, we examined the effect of 13 μM human milk lactoferrin on the Hap adhesin, which bears structural similarity to IgA1 protease. Interestingly, lactoferrin treatment of strain DB117 expressing wild-type Hap resulted in specific proteolysis of Hap. As shown in Fig. 2A, Western blot analysis of whole cells revealed loss of the preprotein and appearance of a C-terminal fragment slightly smaller than Hapβ (39 kDa vs. 45 kDa). To confirm these results and to assess whether proteolysis depends on Hap serine protease activity, we examined the effect of lactoferrin on DB117 expressing Hap with a mutated active site serine (HapS243A). This protein lacks autoproteolytic activity and remains in the outer membrane in the preprotein form (14, 15). Western blot analysis of whole cells again revealed loss of the Hap preprotein and generation of a Hap C-terminal fragment (Fig. 2A, lanes 3 and 7). Treatment of DB117 expressing a Hap derivative containing the Hap signal sequence fused to Hapβ also resulted in generation of the cell-associated 39-kDa C-terminal fragment (Fig. 2A, lanes 4 and 8).
Figure 2.
Western immunoblots demonstrating that treatment of H. influenzae strain DB117 with human milk lactoferrin or deglycosylated A. awamori recombinant human lactoferrin results in degradation of the Hap preprotein and Hapβ, with release of Haps into the culture supernatant. (A) Western immunoblot of whole-cell lysates of H. influenzae strain DB117 derivatives preincubated with either PBS alone (lanes 1–4) or PBS with 13 μM human milk lactoferrin (lanes 5–8). (B) Western immunoblot of whole-cell lysates of H. influenzae strain DB117 derivatives preincubated with either PBS alone (lanes 1–4) or PBS with 13 μM deglycosylated A. awamori recombinant human lactoferrin (lanes 5–8). (C) Western analysis of culture supernatants of H. influenzae strain DB117 derivatives preincubated with either PBS alone (lanes 1–4) or PBS with 13 μM deglycosylated A. awamori recombinant human lactoferrin (lanes 5–8). Western analysis was performed with antiserum Rab730, which reacts with the Hap preprotein, Haps, and Hapβ. The gels in all panels were loaded as follows: lane 1, PBS-treated DB117/vector; lane 2, PBS-treated DB117/wild-type Hap; lane 3, PBS-treated DB117/HapS243A; lane 4, PBS-treated DB117/Hapβ; lane 5, lactoferrin-treated DB117/vector; lane 6, lactoferrin-treated DB117/wild-type Hap; lane 7, lactoferrin-treated DB117/HapS243A; lane 8, lactoferrin-treated DB117/Hapβ. Arrowheads point to the Hap preprotein and Hapβ, arrows point to Hap degradation products, and asterisks indicate Haps.
To establish that these results were because of lactoferrin and not some other component of milk whey, we examined the effect of recombinant human lactoferrin. As shown in Fig. 2B, 13 μM deglycosylated recombinant human lactoferrin prepared from A. awamori generated two products, including the same 39-kDa C-terminal fragment observed with milk-derived lactoferrin and a slightly smaller C-terminal fragment. Additional analysis revealed that Haps or a related fragment of the Hap preprotein was liberated into the supernatant (Fig. 2C). Experiments comparing the proteolysis of Hap by 87 nM, 217 nM, 433 nM, and 13 μM recombinant lactoferrin established a dose-response relationship, with proteolysis detectable but incomplete after treatment of cells for 1 hr with the lowest concentration (not shown). Results with BHK-cell recombinant human lactoferrin paralleled those obtained with A. awamori recombinant protein (not shown). Similar to the situation with IgA1 protease, recombinant lactoferrin N-lobe behaved exactly as full-length lactoferrin (not shown).
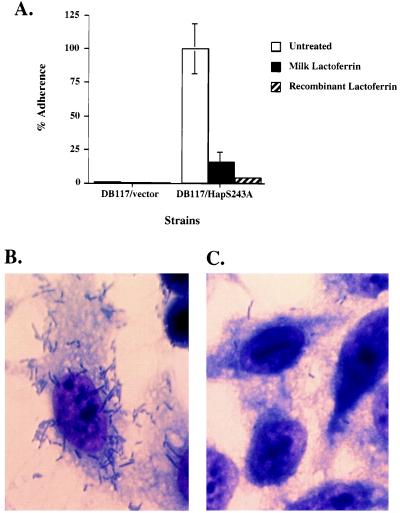
Given that Hap promotes adherence to human epithelial cells, we wondered whether lactoferrin treatment would inhibit Hap-mediated attachment. To address this question, strain DB117 expressing HapS243A was incubated for 1 hr in either PBS alone or PBS with 13 μM lactoferrin, then washed twice and inoculated onto a monolayer of Chang epithelial cells. After incubation for 30 min, adherence was quantitated. [DB117 expressing HapS243A demonstrates augmented in vitro adherence compared with DB117 expressing wild-type Hap, reflecting the fact that attachment is mediated by the preprotein form of Hap (D.R.H. and J.W.St.G., unpublished work), which remains intact and cell-associated when the active site serine is mutated]. As shown in Fig. 3, treatment of DB117/HapS243A with either milk-derived or recombinant lactoferrin resulted in an 85–97% decrease in Hap-mediated adherence. DB117/vector served as a negative control and was nonadherent regardless of lactoferrin treatment. Viability studies demonstrated that lactoferrin treatment had no effect on the survival of either DB117/HapS243A or DB117/pGJB103.
Figure 3.
Effect of human lactoferrin on Hap-mediated H. influenzae adherence to human epithelial cells. (A) Adherence to Chang conjunctival cells by DB117/vector and DB117/HapS243A after incubation in PBS, PBS with 13 μM human milk whey lactoferrin, or PBS with 13 μM deglycosylated A. awamori recombinant lactoferrin. Adherence was measured in a 30-min assay as previously described (33) and is reported relative to DB117/HapS243A after incubation in PBS, which was normalized to 100%. (B and C) Light micrographs of DB117/HapS243A associated with Chang conjunctival cells samples after staining with Giemsa (×2,250). The sample in B was incubated in PBS, and the sample in C was incubated with 13 μM deglycosylated A. awamori recombinant lactoferrin.
Lactoferrin-Associated Degradation of Hap Is Inhibited by Phenylmethylsulfonyl Fluoride (PMSF).
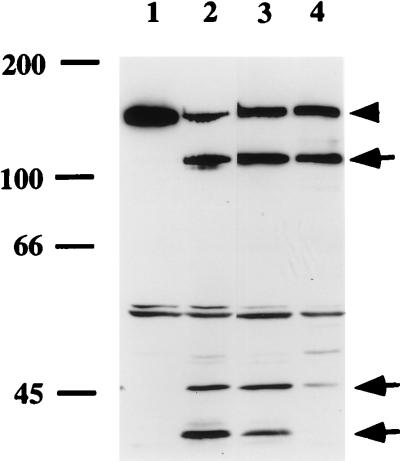
Our findings with DB117 expressing Hap, HapS243A, or Hapβ suggested that lactoferrin has protease activity. With this idea in mind, we examined the lactoferrin crystallographic structure (35) and primary amino acid sequence for features of protease active sites (36). Among the many serine residues in the N-lobe, several are surface-exposed and would be candidate catalytic sites, although none is surrounded by the sequence typical of serine proteases. Lactoferrin lacks nonferrous metal binding sites or features common to other known protease classes, arguing against another catalytic mechanism. To test the possibility that lactoferrin functions as a serine protease, we examined whether PMSF, a broad inhibitor of serine proteases, can inhibit degradation of Hap. As shown in Fig. 4, the partial proteolysis of Hap produced by 433 nM recombinant lactoferrin was significantly inhibited by 7.5 mM PMSF. Similarly, lactoferrin extraction of the IgA protease preprotein was markedly reduced in the presence of either 10 mM PMSF or 10 mM diisopropyl fluorophosphate, another serine protease inhibitor (not shown).
Figure 4.
Effect of serine protease inhibitor PMSF on lactoferrin-associated proteolysis of H. influenzae Hap. DB117/HapS243A was incubated in PBS (lane 1), PBS with 433 nM deglycosylated A. awamori recombinant lactoferrin (lane 2), PBS with 433 nM recombinant lactoferrin and 7.5% 2-propanol (lane 3), or PBS with 433 nM recombinant lactoferrin and 7.5 mM PMSF in 2-propanol (lane 4). Whole-cell lysates were prepared and examined by Western analysis with antiserum Rab 730, which reacts with the Hap preprotein, Haps, and Hapβ. The arrowhead points to the Hap preprotein, and arrows point to Hap degradation products.
To address the specificity of the interaction between lactoferrin and H. influenzae surface proteins, we examined whether lactoferrin treatment results in extraction or degradation of the H. influenzae major outer-membrane proteins P2, P5, and P6. Like IgAβ and Hapβ, P2, P5, and P6 are predicted to form β-barrel structures comprised of a series of transmembrane antiparallel amphipathic β-sheets (37–40). However, these outer-membrane proteins lack the large extracellular domains that link IgAβ and Hapβ to their N-terminal passenger domains and typify the autotransported proteins. As shown in Fig. 5, treatment with milk-derived lactoferrin under conditions leading to extraction of the IgA1 protease preprotein and proteolysis of Hap had no appreciable effect on P2, P5, or P6. All three of these proteins remained cell-associated and intact.
Figure 5.
The H. influenzae P2, P5, and P6 outer-membrane proteins are not removed or degraded by exposure to human milk whey. Bacteria were grown to mid-logarithmic phase and then incubated with either buffer (lanes 1 and 2 in all panels) or human milk whey (lanes 3 and 4 in all panels). Whole-cell lysates (lanes 1 and 3 in all panels) and corresponding supernatants (lanes 2 and 4 in all panels) were examined by Western immunoblot analysis. A was probed with antiserum #331 against IgA1 protease, B was probed with mAb 6G3 against P2, C was probed with mAb 2C7 against P5, and D was probed with mAb 7F3 against P6. Strain Rd3–13 was used for the analysis of IgA1 protease, P5, and P6. Strain 1479 was used for the analysis of P2, as antibody 6G3 is specific for the strain 1479 P2 protein. The IgA1 protease preprotein was transferred to the supernatant, while P2, P5, and P6 were unaffected.
DISCUSSION
Earlier studies showed that lactoferrin, a bilobed protein found in milk, mucosal secretions, and neutrophils, plays an important role in host defense (15, 41). Lactoferrin efficiently binds iron, starving bacteria of this essential nutrient. In addition, lactoferrin is bactericidal for many Gram-negative species, with lytic activity resulting from a membrane-damaging, N-terminal cationic peptide called lactoferricin.
In the present study, we examined lactoferrin in experiments with H. influenzae and demonstrated a newly recognized antibacterial activity. In particular, we found that lactoferrin removes the H. influenzae IgA1 protease from the bacterial cell wall (making it accessible for inhibition by milk anti-IgA1 protease antibodies) and proteolytically degrades the Hap adhesin. These results suggest that lactoferrin may attenuate the pathogenicity of H. influenzae by inactivating two factors presumed to be essential for colonization. Furthermore, our observations provide at least a partial explanation for earlier reports that breast-feeding protects against otitis media. Whether lactoferrin has similar effects on colonization factors of S. pneumonae and M. catarrhalis remains to be established.
It is interesting that lactoferrin is able to inactivate only certain H. influenzae outer-membrane proteins. Both IgA1 protease and Hap, which are sensitive to lactoferrin, belong to the same family of Gram-negative bacterial autotransporter proteins. These proteins contain a C-terminal domain that is embedded in the membrane and an N-terminal protease domain that resides on the surface of the organism until autoproteolytic cleavage and extracellular release. In contrast, P2, P5, and P6, which are unaffected by lactoferrin, are H. influenzae outer-membrane proteins that are predicted to form β-barrel structures like IgAβ and Hapβ, but lack N-terminal passenger domains or extracellular linker regions. Whether lactoferrin inactivates other IgA1 protease- and Hap-like autotransported proteins is not yet known.
The precise mechanism by which lactoferrin extracts the IgA1 protease preprotein without affecting bacterial viability is unknown. However, our experiments with serine protease inhibitors suggest that extraction is related to lactoferrin proteolytic activity, localized to the N-lobe of the protein. Our observations with Hap provide additional evidence that lactoferrin has serine protease activity, with activity residing in the N-lobe. An alternative explanation for our results is that lactoferrin complexes with an unidentified protease in milk or cell-culture media during the process of purification. Arguing against this possibility, native and recombinant lactoferrin purified from multiple sources and prepared by several schemes had the same effect. Furthermore, molecular mass measurements of active recombinant lactoferrin and lactoferrin N-lobe closely approximated the predicted values for these species.
The results of the present study suggest that lactoferrin may have significant therapeutic potential and may be valuable as a supplement in infant formulas. However, since colonization involves complex host–bacterial interactions, it remains unclear whether inactivation of selected factors will attenuate virulence. Thus future studies should compare rates of H. influenzae nasopharyngeal colonization and otitis media in infants fed either standard formula or formula supplemented with native or recombinant lactoferrin.
Acknowledgments
We thank Anne Kane for advice and bacterial cultures, Yolanta Fishman and Andrew Wright for strain Rd3–13, Steven Shewry and John Tweedie for help in preparing recombinant lactoferrin, Adam Smith for technical assistance, Denis Headon for A. awamori lactoferrin, Pieter Stouten for assistance in analyzing lactoferrin for potential serine protease domains, and Virginia Miller for critical reading of the manuscript. This work was supported by funding from the National Institutes of Health [Digestive Disease Center NIDDK DK34928 (A.G.P.), DE 09677 (A.G.P.), HD 20859 (E.N.B.), and AI 19641 (T.F.M.)], the American Heart Association (J.W.S.), the March of Dimes (J.W.S.), and Nutricia Corporation.
ABBREVIATIONS
- BHK
baby hamster kidney
- PMSF
phenylmethylsulfonyl fluoride
Footnotes
This paper was submitted directly (Track II) to the Proceedings Office.
References
- 1. Klein J O. Clin Infect Dis. 1994;19:823–833. doi: 10.1093/clinids/19.5.823. [DOI] [PubMed] [Google Scholar]
- 2.Teele D W, Klein J O, Rosner B the Greater Boston Otitis Media Study Group. J Infect Dis. 1989;160:83–94. doi: 10.1093/infdis/160.1.83. [DOI] [PubMed] [Google Scholar]
- 3.Teele D W, Klein J O, Rosner B the Greater Boston Otitis Media Study Group. J Am Med Assoc. 1983;249:1026–1029. [Google Scholar]
- 4.Fria T J, Cantekin E I, Eichler J A. Arch Otolaryngol Head Neck Surg. 1985;111:10–16. doi: 10.1001/archotol.1985.00800030044003. [DOI] [PubMed] [Google Scholar]
- 5.Teele D W, Klein J O, Chase C, Menyuk P, Rosner B A the Greater Boston Otitis Media Study Group. J Infect Dis. 1990;162:685–694. doi: 10.1093/infdis/162.3.685. [DOI] [PubMed] [Google Scholar]
- 6.Giebink, G. S. (1989) Pediatr. Infect. Dis. J. 8, Suppl., S18–S20. [PubMed]
- 7.Del Beccaro M A, Mendelman P M, Inglis A F, Richardson M A, Duncan N O, Clausen C R, Stull T L. J Pediatr. 1992;120:81–84. doi: 10.1016/s0022-3476(05)80605-5. [DOI] [PubMed] [Google Scholar]
- 8.Kilian M, Reinholdt J, Lomholt H, Poulsen K, Frandsen E V G. APMIS. 1996;104:321–328. doi: 10.1111/j.1699-0463.1996.tb00724.x. [DOI] [PubMed] [Google Scholar]
- 9.Brandtzeag, P. (1992) J. Infect. Dis. 165, Suppl., S167–S176. [DOI] [PubMed]
- 10.Jose J, Jahnig F, Meyer T F. Mol Microbiol. 1995;18:378–380. doi: 10.1111/j.1365-2958.1995.mmi_18020378.x. [DOI] [PubMed] [Google Scholar]
- 11.Poulsen K, Brandt J, Hjorth J P, Thogersen H C, Kilian M. Infect Immun. 1989;57:3097–3105. doi: 10.1128/iai.57.10.3097-3105.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Pohlner J, Halter R, Beyreuther K, Meyer T F. Nature (London) 1987;325:458–462. doi: 10.1038/325458a0. [DOI] [PubMed] [Google Scholar]
- 13.St. Geme J W, III, de la Morena M L, Falkow S. Mol Microbiol. 1994;14:217–233. doi: 10.1111/j.1365-2958.1994.tb01283.x. [DOI] [PubMed] [Google Scholar]
- 14.Hendrixson D R, de la Morena M L, Stathopoulos C, St. Geme J W., III Mol Microbiol. 1997;26:505–518. doi: 10.1046/j.1365-2958.1997.5921965.x. [DOI] [PubMed] [Google Scholar]
- 15.Levay P F, Viljoen M. Haematologica. 1995;80:252–267. [PubMed] [Google Scholar]
- 16.Aniansson G, Alm B, Andersson B, Hakansson A, Larsson P, Nylen O, Peterson H, Rigner P, Svanborg M, Sabharwal H, et al. Pediatr Infect Dis J. 1994;13:183–188. doi: 10.1097/00006454-199403000-00003. [DOI] [PubMed] [Google Scholar]
- 17.Cunningham A S. J Pediatr. 1977;90:726–729. doi: 10.1016/s0022-3476(77)81236-5. [DOI] [PubMed] [Google Scholar]
- 18.Duncan B, Ey J, Holberg C J, Wright A L, Martinez F D, Taussig L M. Pediatrics. 1993;91:867–872. [PubMed] [Google Scholar]
- 19.Saarinen U M. Acta Pediatr Scand. 1982;71:567–571. doi: 10.1111/j.1651-2227.1982.tb09476.x. [DOI] [PubMed] [Google Scholar]
- 20.Kaleida P H, Nativio D G, Chao H P, Cowden S N. J Clin Microbiol. 1993;31:2674–2678. doi: 10.1128/jcm.31.10.2674-2678.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Fleischmann R D, Adams M D, White O, Clayton R A, Kirkness E F, Kerlavage A R, Bult C J, Tomb J F, Dougherty B A, Merrick J M, et al. Science. 1995;269:496–512. doi: 10.1126/science.7542800. [DOI] [PubMed] [Google Scholar]
- 22.Setlow J K, Brown D C, Boling M E, Mattingly A, Gordon M P. J Bacteriol. 1968;95:546–558. doi: 10.1128/jb.95.2.546-558.1968. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Haase E M, Yi K, Morse G D, Murphy T F. Infect Immun. 1994;62:3712–3722. doi: 10.1128/iai.62.9.3712-3722.1994. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Anderson P, Johnston R B, Jr, Smith D H. J Clin Invest. 1972;51:31–38. doi: 10.1172/JCI106793. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Stowell K M, Rado T A, Funk W D, Tweedie J W. Biochem J. 1991;276:349–355. doi: 10.1042/bj2760349. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Day C L, Stowell K M, Baker E N, Tweedie J W. J Biol Chem. 1992;267:13857–13862. [PubMed] [Google Scholar]
- 27.Baker H M, Day C L, Norris G E, Baker E N. Acta Crystallogr D. 1994;50:380–384. doi: 10.1107/S0907444993013435. [DOI] [PubMed] [Google Scholar]
- 28.Mazurier J, Spik G. Biochim Biophys Acta. 1980;629:399–408. doi: 10.1016/0304-4165(80)90112-9. [DOI] [PubMed] [Google Scholar]
- 29.Plaut A G, Qiu J, Grundy F, Wright A. J Infect Dis. 1992;166:43–52. doi: 10.1093/infdis/166.1.43. [DOI] [PubMed] [Google Scholar]
- 30.Laemmli U K. Nature (London) 1970;227:680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 31.Towbin H, Staehelin T, Gordon J. Proc Natl Acad Sci USA. 1979;76:4350–4354. doi: 10.1073/pnas.76.9.4350. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 32.Murphy T F, Nelson M B, Dudas K C, Mylotte J M, Apicella M A. J Infect Dis. 1985;152:1300–1307. doi: 10.1093/infdis/152.6.1300. [DOI] [PubMed] [Google Scholar]
- 33.St. Geme J W, III, Falkow S, Barenkamp S J. Proc Natl Acad Sci USA. 1993;90:2875–2879. doi: 10.1073/pnas.90.7.2875. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Masson P L, Heremens J F, Dive C H. Clin Chim Acta. 1966;14:735–739. [Google Scholar]
- 35.Anderson B F, Baker H M, Norris G E, Rice D W, Baker E N. J Mol Biol. 1989;209:711–734. doi: 10.1016/0022-2836(89)90602-5. [DOI] [PubMed] [Google Scholar]
- 36.Brenner S. Nature (London) 1988;334:528–530. doi: 10.1038/334528a0. [DOI] [PubMed] [Google Scholar]
- 37.Vachon V, Laprade R, Coulton J W. Biochim Biophys Acta. 1986;861:74–82. doi: 10.1016/0005-2736(86)90373-1. [DOI] [PubMed] [Google Scholar]
- 38.Deich R A, Metcalf B J, Finn C W, Farley J E, Green B A. J Bacteriol. 1988;170:489–498. doi: 10.1128/jb.170.2.489-498.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Munson R S, Jr, Grass S, West R. Infect Immun. 1993;61:4017–4020. doi: 10.1128/iai.61.9.4017-4020.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Nelson M B, Apicella M A, Murphy T F, Vankeulen L D, Spotila L D, Rekosh D. Infect Immun. 1988;56:128–134. doi: 10.1128/iai.56.1.128-134.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Ellison R T., III . In: Lactoferrin: Structure and Function. Hutchens T W, Rumball S V, Lönnerdal B, editors. New York: Plenum; 1994. pp. 71–90. [Google Scholar]