Abstract
Chlorophyll b is an ubiquitous accessory pigment in land plants, green algae, and prochlorophytes. Its biosynthesis plays a key role in the adaptation to various light environments. We isolated six chlorophyll b-less mutants by insertional mutagenesis by using the nitrate reductase or argininosuccinate lyase genes as tags and examined the rearrangement of mutant genomes. We found that an overlapping region of a nuclear genome was deleted in all mutants and that this encodes a protein whose sequence is similar to those of methyl monooxygenases. This coding sequence also contains putative binding domains for a [2Fe-2S] Rieske center and for a mononuclear iron. The results demonstrate that a chlorophyll a oxygenase is involved in chlorophyll b formation. The reaction mechanism of chlorophyll b formation is discussed.
Chlorophyll’s central role in photosynthesis is harvesting light energy and its conversion to chemical energy. All land plants, green algae, and prochlorophytes possess chlorophyll a and chlorophyll b. Chlorophyll a alone occurs in the core complexes of photosystems whose composition and organization are highly conserved. On the other hand, chlorophyll a and chlorophyll b are components of peripheral antenna complexes. Antenna complexes show controlled changes in adapting to various growth conditions, enabling optimal utilization of available light (1). It is known that the chlorophyll a to b ratio is higher in high-light growth conditions than in low-light growth conditions, which is accompanied by larger sizes of antenna complexes in low-light conditions (2). Thus the regulation of chlorophyll b synthesis is important for understanding the mechanisms of adaptation of plants to various light intensity.
Prochlorophytes are prokaryotic photosynthetic organisms that have chlorophyll b as a light-harvesting pigment similar to chlorophytes. However, molecular sequence data of ribosomal RNAs and DNA-dependent RNA polymerase indicate a paraphyletic origin of prochlorophytes and that the prochlorophyte lineage does not include the direct ancestor of the chloroplast (3, 4). Chlorophyll a/b and chlorophyll a/c binding proteins of plants and chlorophyte algae form a superfamily of genetically related proteins; however, chlorophyll a/b binding proteins of prochlorophytes are closely related to IsiA and to CP43, the chlorophyll a antenna protein of photosystem II (5). These observations might suggest that use of chlorophyll b as a light-harvesting pigment has developed independently several times in the evolution of prochlorophytes and chlorophytes.
Recently, many genes encoding proteins involved in chlorophyll synthesis have been isolated through the use of sequences of purified proteins (6) or related bacterial genes for heme and/or bacteriochlorophyll synthesis enzymes (7). However, these strategies are not appropriate for isolation of genes encoding components for chlorophyll b synthesis because there is no in vitro assay system for chlorophyll b formation, and the organisms in which tetrapyrrole biosynthesis is most thoroughly understood do not produce chlorophyll b.
Chlorophyll b appears to be synthesized by oxidation of the methyl group of chlorophyll a to a formyl group (8). Although many studies have been done on the biosynthesis of chlorophyll b, success in elucidating the biosynthetic pathway for chlorophyll b formation or the proteins involved have not been achieved.
Recently, reliable transformation procedures have been developed for the nuclear genome of Chlamydomonas (9, 10), and these provide a powerful experimental tool for studying the photosynthetic apparatus. In this report, we used molecular genetics of Chlamydomonas to isolate a gene for chlorophyll b synthesis. Chlorophyll b-less mutants of Chlamydomonas were isolated by transformation with plasmid DNA carrying either nitrate reductase or argininosuccinate lyase genes as selectable markers, and a gene involved in chlorophyll b synthesis was isolated. Its deduced amino acid sequence contains binding domains for a [2Fe-2S] Rieske center and for a mononuclear iron and shows similarity to methyl monooxygenases.
MATERIALS AND METHODS
Strains and Growth Conditions.
Strain 5D (nit1–305, cw15) and cc1931 (arg7) of Chlamydomonas reinhardtii were provided by the Chlamydomonas Genetic Center (Duke University, Durham, NC). Both strains were grown in constant light at 20°C on TAP medium (1 ml/liter acetate, 2.42 g/liter Tris, 0.75 g/liter KNO3, 0.1 g/liter MgSO4⋅7H2O, 0.05 g/liter CaCl2⋅2H2O, and micronutrients) supplemented when appropriate with 200 μM arginine.
Transformation and Screening of the Mutant.
Strains 5D and cc1931 were used as recipients for 2 μg of pMN24, a plasmid containing the cloned wild-type nitrate reductase gene, and p389, a plasmid containing the cloned argininosuccinate lyase gene, respectively (9, 10). Cells were transformed by the glass beads method (9) and spread on TAP agar plates. Pale-green or small-size transformant colonies were selected by visual observation. For the first screening about one chlorophyll b-less mutant was obtained per 1,000 transformants. Their chlorophyll was extracted with a 80% acetone and analyzed by HPLC as described below. Six chlorophyll b-less mutants (cbs1–cbs6) were obtained (see Fig. 1).
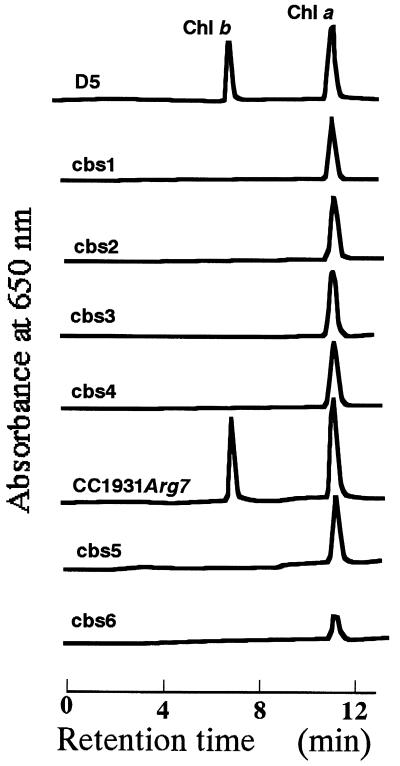
Figure 1.
HPLC analysis of chlorophyll from the transformant mutants. Chlorophylls of transformants colonies were extracted with 80% acetone and were determined by HPLC. Four chlorophyll b-less mutants (cbs1–cbs4) and two mutants (cbs5 and cbs6) were isolated by using pMN24 and p389, respectively.
Preparation of Genomic DNA and Poly(A)+ RNA.
Total DNA was extracted from Chlamydomonas and purified on CsCl gradient (11). Total RNA was extracted by the phenol/SDS method, and poly(A)+ RNA was isolated by chromatography on oligo(dT)-cellulose (12).
Construction of cDNA and Genomic Library.
A cDNA library was constructed from poly(A)+ mRNA by using the Lambda Zap cDNA synthesis kit (Stratagene) according to the manufacturer’s instructions. A genomic library was constructed by using the Lambda DashII/BamHI vector Kit (Stratagene) according to the manufacturer’s instructions.
Southern and Northern Blot Analyses.
Genomic DNA (2 μg) was digested with restriction enzymes, separated on agarose gels, transferred onto Hybond-N+ nylon membrane, and hybridized in 0.25 M sodium phosphate, 7% SDS, and 1 mM EDTA at 65°C. Two micrograms of poly(A)+ RNA were electrophoresed on a 1% agarose/formamide gel and blotted onto Hybond-N nylon membrane. Hybridization with 32P-labeled probe were performed as described (13).
Screening of DNA Fragments Derived from the NIT1-Integrated Regions and Genomic Clone.
Genomic DNA from strain 5D and transformant cbs3 (see Fig. 3) were digested by SacI and subjected to Southern blot analysis by using a NIT1 gene as a probe. DNA from cbs3 has additional two bands compared with that from strain 5D. These additional bands may represent DNA located at the junctions between integrated NIT1 gene and host genomic DNA. By using the entire gene or a portion of NIT1 gene as probes, we cloned the bands from a cbs3 SacI genomic library. By using these clones as hybridization probes, we cloned the DNA fragments from a wild-type genomic library.
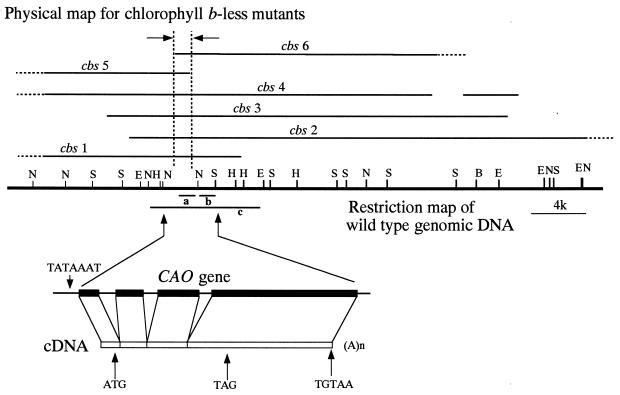
Figure 3.
Restriction map of wild-type genomic DNA and deletion maps for mutants. Different genomic DNA clones were isolated that covered the deleted region of cbs3. Deletions in each of the mutants, indicated by heavy lines, were determined with probes consisting of 13 different genomic DNA fragments clones from wild type. In some cases, the ends of the deletions were not determined (shown by dotted lines). Open and closed boxes represent the cDNA sequence and exons in the gene, respectively. The location of a TATA box (TATAAAT), an initiation codon (ATG), a termination codon (TAG), and a polyadenylation signal sequence (TGTAA) are marked by the vertical arrows. E, EcoRI; H, HindIII; N, NotI; S, SacI; A(n), a poly(A) tail; a, DNA fragment for the isolation of cDNA; b, DNA fragment for the Southern blot analysis in this figure; c, DNA fragment for the complementation of Fig. 4.
Isolation of cDNA.
The cDNA library was screened with the genomic DNA fragment a in Fig. 3. Because full-length cDNA was not obtained by screening the library, we designed oligonucleotide primers according to the genome DNA sequence and amplified the 5′ part of the cDNA. Poly(A)+ mRNA was used to synthesize first-strand cDNA at 70°C with rTth DNA polymerase. PCR amplification was performed with the oligonucleotide primers complementary to the genomic DNA upstream of the sequence corresponding to that part of the cDNA isolated by screening.
Sequencing.
DNA fragments were subcloned into the pBluescript II plasmid vector (Stratagene) and sequenced by using the Dye Terminator DNA Sequence kit (Applied Biosystems).
Complementation.
The DNA fragment containing 4 kb of the protein coding sequence and 1 kb of untranslated sequence on the 5′ end, and 3 kb of untranslated sequence on the 3′ end was subcloned into pSP109, which carries the ble gene as a selectable marker (14). The plasmid (pSP109-E8) was linearized by ScaI and introduced into the genome of strain cbs3. Strain cbs3 was transformed with the 2 μg of the linearized plasmids and grown on plates containing 2 μg/ml phleomycin.
Analysis of Chlorophyll by HPLC.
Chlorophylls were extracted from colonies with 80% acetone and subjected to HPLC on an octadecyl silica column (6 mm, inner diameter × 150 mm) by using methanol as the eluent at a flow rate of 1.5 ml/min at 40°C. Chlorophylls were monitored by their absorbance at 650 nm.
RESULTS
Isolation of Chlorophyll b-Less Mutants.
Because insertion of a DNA fragment is accompanied by a large deletion of genomic DNA upon transformation in Chlamydomonas (15), we obtained many chlorophyll b-less mutants and analyzed their genomic DNA rearrangements. To obtain tagged mutants that have a defect in chlorophyll b biosynthesis, we transformed ammonium or arginine requiring Chlamydomonas cells with plasmid DNA carrying either nitrate reductase (NIT1) or argininosuccinate lyase (ARG7) gene as a selectable marker, respectively. We isolated four chlorophyll b-less mutants (cbs1, cbs2, cbs3, cbs4) by using NIT1 and two (cbs5, cbs6) by using ARG7. In these mutants, no chlorophyll b was detected by HPLC (Fig. 1). Chlorophyll b-less mutants of cbs1, cbs2, cbs3, and cbs4 accumulate chlorophyll a to the wild-type level; however, cbs5 and cbs6 were pale green on a plate.
DNA Rearrangement in the Chlorophyll b-Less Transformants.
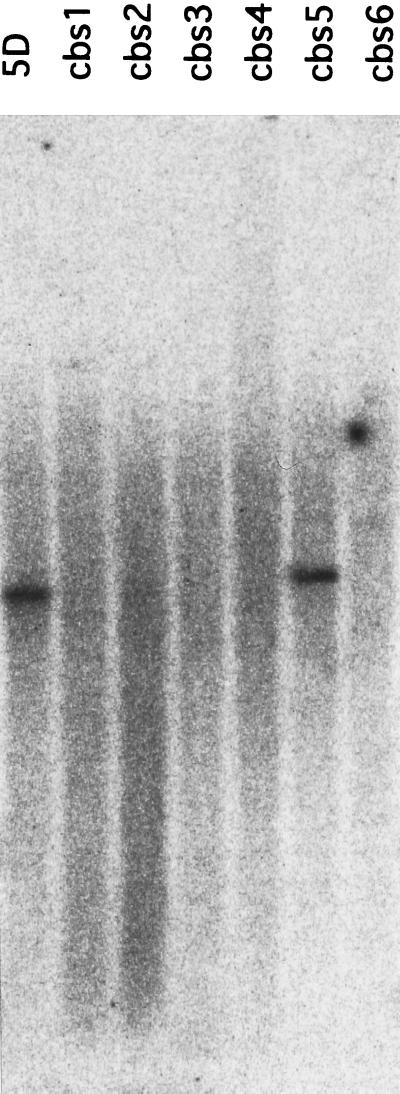
By using the NIT1 gene as a probe, we isolated two DNA clones from a cbs3 SacI genomic library. We determined the nucleotide sequences of these clones and found that they contained sequences other than that of NIT1 gene, and that these were presumed to be a part of the genomic DNA where NIT1 gene had inserted. By using these parts of the cloned fragments as probes, we isolated different overlapping clones from a wild-type library that covered the deleted region of strain cbs3. We examined the rearrangement of the other mutant genomes by Southern blot hybridization with 13 different fragments from these overlapping clones. Some probes did not hybridize to the genomic DNAs from the mutants (Fig. 2), indicating that the DNA insertion was accompanied by the deletion of genomic DNA. As shown in the physical map in Fig. 3, all of the mutants had a large deletion in a common region of the genomic DNA. The genomic DNA region indicated by arrows was deleted in all of the mutants (cbs1–cbs6). These results strongly suggest that the deleted genomic DNA includes a gene for chlorophyll b synthesis.
Figure 2.
Southern blot analysis of genomic DNA from the chlorophyll b-less mutants. DNA from the mutants were digested by SacI and hybridized with radioactive probe for DNA fragment b as indicated in Fig. 3.
Complementation of Chlorophyll b-Less Phenotype.
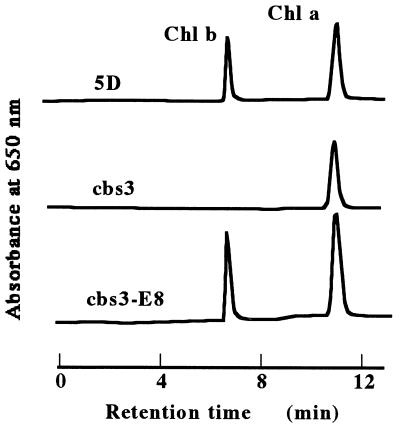
Next, we attempted to rescue the chlorophyll b-less phenotype of cbs3 (Fig. 4) by transformation with an 8-kb genomic DNA fragment (fragment c in Fig. 3) subcloned into pSP109, which has the ble gene as a selectable marker (14). The DNA fragment contained the region that was deleted in all the mutants (Fig. 3). The plasmid (pSP109-E8) was linearized by ScaI and introduced into the genome of cbs3. Transformants that accumulated chlorophyll b had integrated both ble and the genomic DNA fragments as determined by PCR (data not shown).
Figure 4.
Rescue of the chlorophyll b-less phenotype by transformation of wild-type CAO gene. An 8-kb genomic DNA fragment was subcloned into plasmid pSP109 and the resultant plasmid (pSP109-E8) was linearized with ScaI. Strain cbs3 was transformed with the 2 μg of linearized plasmids and selected on plates containing 2 μg/ml phleomycin. Pigments were extracted from the transformant (b3-8E), cbs3, and 5D with 80% acetone and determined by HPLC.
cDNA Cloning and Sequence Analysis.
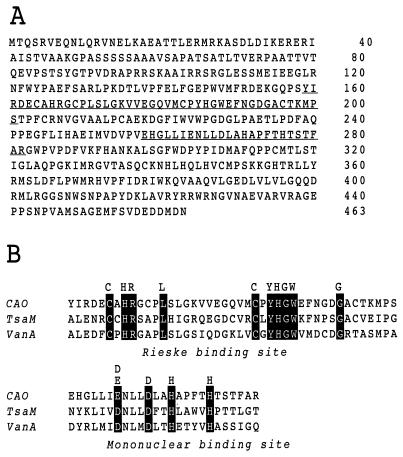
Comparison of cDNA and genomic DNA sequences revealed three introns (data not shown). The splice junctions of these introns agree with the consensus motifs for other Chlamydomonas genes (16). The sequence analysis of the DNA fragment used for complementation indicates that the DNA fragment contained 4 kb of protein-coding sequence, flanked by 1 kb of untranslated sequence on the 5′ end and 3 kb of untranslated sequence on the 3′ end. The cDNA is 2,885 bp in length and has a single ORF. The corresponding cDNA has a coding capacity for 463 amino acids, giving a calculated molecular weight of 51341 (Fig. 5A). Analysis of the deduced amino acid sequence revealed two conserved motifs, a consensus sequence for coordinating a Rieske-type [2Fe-2S] cluster, and a conserved mononuclear nonheme Fe-binding site (Fig. 5B). This finding indicates that the gene encodes an oxygenase (17), in agreement with observations from mass spectrometry that oxygen in the formyl group of chlorophyll b is incorporated from O2 (18, 19). The protein has sequence similarity to vanillate demethylase (20) and ρ-toluenesulfonate monooxygenase (21), which catalyze oxygenation at the methyl side chain of an aromatic ring and convert a methyl group to a hydroxymethyl group. The catalytic pathway is consistent with that proposed for chlorophyll b formation from chlorophyll a by Beale and Weinstein (22), whose hypothesis was based on the chemical properties of the substrate.
Figure 5.
(A) Predicted amino acid sequence of the chlorophyll a oxygenase protein with specific binding sites underlined and expanded in B. (B) Amino acid sequence alignments of the Rieske [2Fe-2S] center and mononuclear iron-binding regions of CAO, toluenesulfonate methyl monooxygenase (TsaM), and vanillate demethylase (VanA). Residues for the binding domain are shaded.
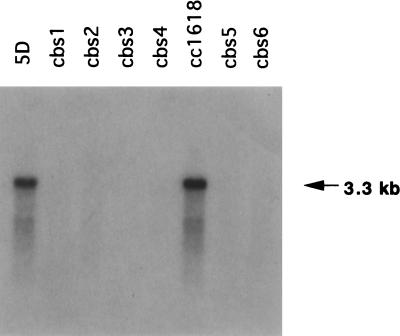
Northern hybridization of wild-type RNA with the cDNA clone detected a single band of 3.3 kb in accordance with the sequence of the cDNA plus addition of a poly(A)+ tail to the mRNA. No hybridization was detected with RNA extracted from the mutants (Fig. 6).
Figure 6.
Northern blot analysis of CAO transcripts. Poly(A)+ RNA isolated from wild-type and mutant cells were examined by RNA gel blot analysis with radioactive CAO probe.
DISCUSSION
From the results presented above, we conclude that we have identified a gene encoding an enzyme for chlorophyll b formation by the oxygenation of chlorophyll a. We designate the gene as CAO (chlorophyll a oxygenase).
For the reaction pathway of the enzyme, it is likely that an electron would be transferred from the Rieske center to the mononuclear iron with subsequent activation of molecular oxygen for oxygenation of the chlorophyll a methyl group. Each of the known oxygenase systems uses an electron transfer system from NADH to ferrous via specific reductases with similarities to ferredoxin-NADPH oxidoreductase (23). Because CAO is chloroplast-localized, we speculate that it could accept electrons from ferredoxin-NADPH oxidoreductase or perhaps ferredoxin by itself. However, we cannot rule out the possibility that there are other, more specific, reducing systems for CAO as in the case for other oxygenase.
For the conversion of ρ-toluenesulfonate to ρ-sulfo-benzaldehyde, methyl monooxygenase, and alcohol dehydrogenase are required (21). It is likely that the conversion of a methyl group to a formyl group in chlorophyll b formation proceeds in successive steps catalyzed by CAO and a dehydrogenase. However, it was reported with Chlamydomonas (24) and barley (25) that chlorophyll b-deficient and chlorophyll b-less mutants belong to a single complementation group. In our mutant collection, a genomic DNA deletion is found at the same locus (Fig. 3). If two enzymes (CAO and dehydrogenase) are needed for chlorophyll b formation (Fig. 7), more than one copy of the gene for 7-hydroxymethyl chlorophyll dehydrogenase must exist to be consistent with these results. Another possibility is that there is a multifunctional dehydrogenase that catalyzes the last step of chlorophyll b synthesis, but mutations affecting this enzyme are lethal because of its role(s) in other pathways. At present, it could not be excluded that CAO converts chlorophyll a to chlorophyll b without dehydrogenase. Enzymatic studies of CAO are needed to clarify the mechanism of chlorophyll b formation.
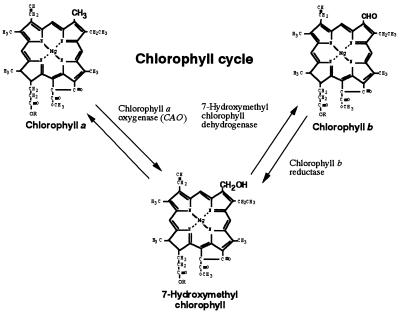
Figure 7.
Proposed chlorophyll cycle, whereby interconversion of chlorophyll a and chlorophyll b levels could be regulated via 7-hydroxymethyl chlorophyll.
When chlorophyll a is oxygenated by CAO, the converted molecule should be 7-hydroxymethyl chlorophyll, which we reported to be the intermediate molecule in chlorophyll b to chlorophyll a conversion (26). Thus chlorophyll a and chlorophyll b could be interconvertable through 7-hydroxymethyl chlorophyll by the chlorophyll cycle (Fig. 7). Regulation of the metabolism of the 7-hydroxymethyl chlorophyll could be important for the adjustment of chlorophyll a/b ratios. Regulating photosynthetic antenna by interconversion of chlorophyll a and chlorophyll b would be more efficient than their degradation and synthesis, and it would be a great advantage to chlorophyll a/b organisms.
In this report, we show that an oxygenase is involved in chlorophyll b formation. To determine the origin of CAO would be important to understand how the photosynthetic organisms obtained chlorophyll b as a photosynthetic pigment. Further investigations on CAO genes from algae to higher plants should give additional information about their control of expression and evolution. Chlorophyll b is present in prochlorophytes as well as chlorophytes. One hypothesis is that chlorophyll b has arisen several times during evolution (3, 4). Alternatively, it has been suggested that a progenitor to both groups possessed chlorophyll a, chlorophyll b, and phycobilins but that the phycobilins were lost during evolution of higher plants (27). Sequence analysis of chlorophyll b binding proteins indicated that these proteins in prochlorophytes and chlorophytes are likely to be the result of convergent evolution (5). Sequence comparison of genes for chlorophyll b synthesis would help to resolve events in the evolution of photosynthetic organisms.
The conversion of a methyl group on ring B to a formyl group is also needed for the formation of chlorophyll c3, chlorophyll ccs-170 and bacteriochlorophyll e (28). Evolutionarily related enzymes to CAO might be responsible for the formation of these chlorophylls. Thus modification of CAO genes might partially account for the diversity of chlorophyll molecules among various photosynthetic organisms.
The investigation of CAO gene would be a great help for the studies on adaptation, evolution, and diversity of chlorophylls.
Acknowledgments
We thank Drs. S. Ishiguro and T. Oyama for their helpful suggestions. We also thank Dr. S. Purton for the pSP109 plasmid. We are grateful to the Chlamydomonas Genetics Center, Duke University, for plasmids and strains used in this work. This work was supported by the Shorai Foundation for Science and Technology.
ABBREVIATIONS
- ble
phleomycin resistant gene
- CAO
chlorophyll a oxygenase
- CP43
43-kDa chlorophyll a binding protein of photosystem II
Footnotes
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AB015139).
References
- 1. Anderson J M. Annu Rev Plant Physiol. 1986;37:93–136. [Google Scholar]
- 2.Björkman O, Boardman N K, Anderson J M, Thorne S W, Goodchild D J, Pyliotis N A. Carnegie Inst Washington Yearb. 1972;71:115–135. [Google Scholar]
- 3.Palenik B, Haselkorn R. Nature (London) 1992;355:265–267. doi: 10.1038/355265a0. [DOI] [PubMed] [Google Scholar]
- 4.Urbach E, Robertson D L, Chisholm S W. Nature (London) 1992;355:267–270. doi: 10.1038/355267a0. [DOI] [PubMed] [Google Scholar]
- 5.Roche J L, van der Staay G W M, Partensky F, Ducret A, Aebersold R, Li R, Golden S S, Hiller R G, Wrench P M, Larkum A W D, Green B R. Proc Natl Acad Sci USA. 1996;93:15244–15248. doi: 10.1073/pnas.93.26.15244. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Grimm B. Proc Natl Acad Sci USA. 1990;87:4169–4173. doi: 10.1073/pnas.87.11.4169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Bollivar D W, Suzuki J Y, Beatty T, Dobrowolski J M, Bauer C E. J Mol Biol. 1994;237:622–640. doi: 10.1006/jmbi.1994.1260. [DOI] [PubMed] [Google Scholar]
- 8.von Wettstein D, Gough S, Kannangara C G. Plant Cell. 1995;7:1039–1057. doi: 10.1105/tpc.7.7.1039. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Kindle K L. Proc Natl Acad Sci USA. 1990;87:1228–1232. doi: 10.1073/pnas.87.3.1228. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Debuchy R, Purton S, Rochaix J-D. EMBO J. 1989;8:2803–2809. doi: 10.1002/j.1460-2075.1989.tb08426.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Sambrook J, Fritsch E F, Maniatis T. Molecular Cloning: A Laboratory Manual. 2nd Ed. Plainview, NY: Cold Spring Harbor Lab. Press; 1989. [Google Scholar]
- 12.Ausubel F M, Brent R, Kingston R E, Moore D D, Seidmann J G, Smith J A, Struhl K, editors. Current Protocols in Molecular Biology. New York: Wiley; 1987. p. 4.3.1. ., 4.5.1. [Google Scholar]
- 13.Yoshida K, Chen R-M, Tanaka A, Teramoto H, Tanaka R, Timko M P, Tsuji H. Plant Physiol. 1995;109:231–238. doi: 10.1104/pp.109.1.231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Stevens D R, Rochaix J-D, Purton S. Mol Gen Genet. 1996;271:23–30. doi: 10.1007/BF02174340. [DOI] [PubMed] [Google Scholar]
- 15.Tam L-W, Lefebvre P A. Genetics. 1993;135:375–384. doi: 10.1093/genetics/135.2.375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Lee V D, Stanpleton M, Huang B. J Mol Biol. 1991;221:175–191. [PubMed] [Google Scholar]
- 17.Harayama S, Kok M. Annu Rev Microbiol. 1992;46:565–601. doi: 10.1146/annurev.mi.46.100192.003025. [DOI] [PubMed] [Google Scholar]
- 18.Schneegurt M K, Beale S I. Biochemistry. 1992;31:11677–11683. doi: 10.1021/bi00162a002. [DOI] [PubMed] [Google Scholar]
- 19.Porra R J, Schäfer W, Cmiel E, Katheder I, Scheer H. Eur J Biochem. 1994;219:671–679. doi: 10.1111/j.1432-1033.1994.tb19983.x. [DOI] [PubMed] [Google Scholar]
- 20.Brunel F, Davison J. J Bacteriol. 1988;179:4924–4930. doi: 10.1128/jb.170.10.4924-4930.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Junker F, Kiewitz R, Cook A M. J Bacteriol. 1997;179:919–927. doi: 10.1128/jb.179.3.919-927.1997. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Beale S I, Weinstein J D. In: Biosynthesis of Heme and Chlorophylls. Dailey H A, editor. New York: McGraw-Hill; 1990. pp. 287–392. [Google Scholar]
- 23.Gassner G T, Ludwig L M, Gatti L D, Correll C, C, Ballou P D. FASEB J. 1995;9:1411–1418. doi: 10.1096/fasebj.9.14.7589982. [DOI] [PubMed] [Google Scholar]
- 24.Chunayev A S, Mirnaya O N, Maslov V G, Boschetti A. Photosynthetica. 1991;27:291–301. [Google Scholar]
- 25.Simpson D, Machold O, Hoyer-Hansen G, von Wettstein D. Carlsberg Res Commun. 1985;50:223–238. [Google Scholar]
- 26.Ito H, Ohtsuka T, Tanaka A. J Biol Chem. 1997;271:1475–1479. doi: 10.1074/jbc.271.3.1475. [DOI] [PubMed] [Google Scholar]
- 27.Hess W R, Partensky F, van der Staay G W M, Garcia-Fernandez J M, Borner T, Vaulot D. Proc Natl Acad Sci USA. 1996;93:11126–11130. doi: 10.1073/pnas.93.20.11126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Porra R J. Photochem Photobiol. 1997;65:492–516. [Google Scholar]