Abstract
Programmed cell death regulates a number of biological phenomena, and the apoptotic signal must itself be tightly controlled to avoid inappropriate cell death. We established a genetic screen to search for molecules that inhibit the apoptotic signal from the Fas receptor. Here we report the isolation of a gene, LFG, that protects cells uniquely from Fas but not from the mechanistically related tumor necrosis factor α death signal. LFG is widely distributed, but remarkably is highly expressed in the hippocampus. LFG can bind to the Fas receptor, but does not regulate Fas expression or interfere with binding of an agonist antibody. Furthermore LFG does not inhibit binding of FADD to Fas.
Apoptosis, or programmed cell death, is used in a number of biological processes from modeling the embryo, regulating the immune system, to tumor regression (1–4). Conversely, dysfunction of this system has been implicated in oncogenesis, development of autoimmunity and degenerative diseases. Intense study of this process in the last few years has identified membrane-bound receptors and their cognate ligands that together begin the program that ultimately leads to cell death (5). One class of receptors falls into the tumor necrosis factor (TNF) receptor superfamily of which the Fas receptor (also called CD95 or Apo-1) is a member (6, 7). This receptor has three cysteine-rich extracellular domains and an intracellular death domain required for signaling (8). Ligation of the receptor by its cognate ligand FasL (9) or an agonistic antibody (10) leads to the recruitment of an adapter molecule FADD/MORT-1 (11, 12). This interaction is mediated by a death domain in FADD/MORT1 with the death domain of Fas. Additionally FADD/MORT1 contains a death effector domain that recruits the protease caspase-8 (also called FLICE, MACH, and Mch5) to this signaling complex (13–15). This zymogen, through proximity with other caspase-8 molecules, is cleaved, rendering it fully active, thus beginning a protease cascade that leads to cell death (16). A counterpoint to this death activation is inhibition of cell death. A number of molecules have been identified that act on various components in the pathway. These include a soluble form of Fas (17) and a secreted decoy receptor that binds and competes for the ligand (18). At the level of the receptor, adenoviral proteins E3 10.4K/14.5K force endocytosis of Fas and other receptors, thus protecting infected cells and aiding the virus to complete its life cycle (19, 20). A group of proteins called FLIPs, which possess a death effector domain, compete for the recruitment of FADD/MORT1 and caspase-8 at the ligated receptor (21). An extension to this control is the molecule toso (22) that induces c-FLIP expression uniquely in hematopoietic lineages. Finally, a number of molecules have been identified called inhibitors of apoptosis proteins (23), a protein from the cowpox virus (crmA) (24), and adenovirus (E3–14.7K) (25), all of which inhibit caspase activity. The majority of these mechanisms inhibit the signal pathway downstream of the receptor and hence will inhibit the death signal from several receptors, i.e., Fas and TNF-α receptor.
In this study we report the cloning of a molecule that uniquely inhibits death mediated by Fas, but not TNF-α receptor. The cloning relies on a technology for stable gene transfer of representative cDNA libraries to diverse cell types. The high transfer efficiencies enable genetic screens to identify cDNAs either by complementation of mutant cell lines or by virtue of ectopic expression.
Materials and Methods
cDNA and Library Construction.
Total RNA was isolated from the human lung fibroblast cell line MRC-5 (ATCC CCL-171) (26), and poly(A)+ RNA was prepared and converted to double-stranded DNA by using the Superscript Choice system (GIBCO/BRL) according to the manufacturer’s instructions. Size-fractionated cDNA species (>500 bp) was ligated into the retroviral vector pCLMFG.MCS (N.S., M.J.S., and I.M.V., unpublished work). The ligation was transformed into supercompetent DH10Bα Escherichia coli (GIBCO/BRL), which generated a cDNA library of 2 × 106 individual clones.
Retroviral Vector Generation.
Fifteen micrograms of plasmid DNA prepared from the pooled cDNA clones was transfected with 5 μg of an expression vector for vesicular stomatitis virus G protein into a cell line 293 gp/bsr (N.S. and I.M.V., unpublished work). Forty-eight hours later, supernatant containing the cDNA viral vectors was recovered. The supernatant was used to infect 5 × 106 HeLa cells and left for 24 hr, after which the media were replaced with a second collection from the transfection for 24 hr.
Selection for Resistant Cells.
A mouse anti-human Fas mAb, clone CH-11 (10) (Kamiya Biomedical, Thousand Oaks, CA), was used at a final concentration of 500 μg/ml. The media were changed every 3–4 days. After 1 week, the surviving clones were pooled into one plate, and selection was maintained for an additional 15 days, with a change of media every 3–4 days. After this time the surviving pool was expanded in the presence of selection, and genomic DNA was isolated.
Rescue of Complementing cDNA.
PCR was performed by using the TaqPlus Precision system (Stratagene). A cycle of 95°C for 30 sec, 58°C for 30 sec, and 72°C for 5 min was repeated 35 times. The amplimers used were: forward, 5′-TGCCGACCCCGGGGGTGGACCATCCTCTAGACTGCCATGG-3′, and back, 5′-CTGATATCCTGTCTTTAACAAATTGGACTAATCCGGATCC. The most prominent PCR product was purified, cloned, and sequenced. The cDNA was subcloned from this vector back into pCLMFG.MCS in the sense orientation. Retrovirus was generated and used for infection as described.
Northern Blot Analysis.
Total RNA, 20 μg for HeLa cells and 10 μg for MRC-5 cells, was separated on a 1% agarose-formaldehyde gel and blotted onto Hybond-N filter (Amersham Pharmacia) according to standard protocols. A 560-bp 5′ fragment of the LFG cDNA (up to methione 179) was labeled with 32P-dCTP by using MegaPrime (Amersham Pharmacia) and hybridized to the filter in Rapid Hybridization buffer (Amersham Pharmacia) according to the manufacturer’s protocols. Filters of human mRNA from different tissues and total RNA from human brain were purchased from Invitrogen and probed as outlined.
Cell Death Assays.
A total of 5 × 104 cells/well of each cell line was plated in triplicate onto 24-well dishes, and after 24 hr challenged with mouse anti-human Fas mAb (CH11 at 500 μg/ml). After 48 or 72 hr the media were aspirated, and the cells were washed twice with PBS. The surviving, adherent cells were stained with 0.05% crystal violet. The stain was eluted from the cells with 200 μl of 100% methanol, and 100 μl was aliquoted to a 96-well microtiter plate and the absorbance was determined at 490 nm. The TNF-α mediated apoptosis was done similarly by using 10 ng/ml TNF-α (Calbiochem) and/or cycloheximide (10 μg/ml) but the cells were fixed 24 hr after treatment.
Immunofluorescence and Biochemical Fractionation.
The LFG coding sequence was cloned in-frame and C terminal to sequences coding for the myc-epitope that is reiterated six times (27). This six myc-LFG was transfected into Hela cells on glass slides, and 48 hr later was fixed in 3% paraformaldehyde and permeabilized in 0.5% Triton X-100. A mouse mAb, 9E10 (28), was used as a primary antibody (asites fluid diluted 1/2,000 incubated at 37°C for 20 min in PBS). Secondary antibody was donkey anti-mouse IgG (Jackson ImmunoResearch) conjugated to Texas red. Nuclei were stained with 4′,6-diamidino-2-phenylindole (0.5 μg/ml). For the biochemical fractionation 293 cells were transfected with 5 μg of expression vectors for myc-LFG, human Fas, or human FADD. The latter two plasmids were cotransfected with 5 μg of an expression plasmid for crmA to prevent cell death. All transfection mixes were brought to 20 μg total DNA with pEGFP (CLONTECH). Forty-eight hours after transfection the cells were lysed for 10 min on ice in 1.5 ml of buffer A (10 mM Hepes-KOH, pH 7.9/1.5 mM MgCl2/10 mM KCl/0.5 mM DTT/0.2 mM PMSF). The nuclei were pelleted at 2,000 rpm for 10 min in a microfuge. The pellet was resuspended in 1.5 ml of buffer C (20 mM Hepes, pH 7.9/25% glycerol/420 mM NaCl/1.5 mM MgCl2/0.2 mM EDTA/0.5 mM DTT/0.2 mM PMSF) and incubated on ice for 20 min, and debris was pelleted at 13,000 rpm in a microfuge. The supernatant (nuclear fraction) was transferred to a new tube. The supernatant from the buffer A lysis was subjected to ultracentrifugation at 35,000 rpm in a SW60 rotor for 30 min, and the supernatant (cytoplasmic fraction) was decanted from the pelleted membrane fraction. The pellet was resuspended in 1.5 ml of RIPA buffer (0.15 mM NaCl/0.05 mM Tris⋅HCl, pH 7.2/1% Triton X-100/1% sodium deoxycholate/0.1% SDS) and briefly sonicated, and debris was pelleted. Fifty microliters of each fraction was separated on a 12% PAGE gel and subjected to immunoblot analysis using the mouse anti-myc antibody, a rabbit anti-Fas antibody (Santa Cruz Biotechnology), a rabbit anti-c-jun antibody (Santa Cruz Biotechnology), a rabbit anti-IκBα antibody (Santa Cruz Biotechnology), a mouse anti-transferrin antibody (a gift from Cathy Charles, Salk Institute), and a rabbit anti-calnexin antibody (a gift from Ari Helenius, Eidgenössiche Technische Hochschule, Zurich).
Immunoprecipitation and Immunoblot Analysis.
Five micrograms of expression plasmids for human Fas receptor, human TNF receptor 1 (TNFR1), or myc-LFG was transfected singly or in combination into 293 cells. Five micrograms of an expression plasmid for crmA was added to all transfections. The total plasmid concentration was brought up to 20 μg with the plasmid pRK5. Forty-eight hours after transfection the cells were lysed (in 20 mM Tris⋅HCL, 1 mM CaCl2, 150 mM NaCl, 1% Triton X-100, and protease inhibitors) on ice for 45 min. Debris was pelleted in a microfuge at 13,000 rpm for 20 min, and the supernatant was precleared with protein-G Sepharose (Amersham Pharmacia) for 1 hr at 4°C. The protein-G beads were pelleted, and rabbit anti-Fas (Santa Cruz Biotechnology), mouse anti-TNFR1 (Biosource International, Camarillo, CA), or rabbit anti-myc (Santa Cruz Biotechnology) was added to 200 μg of the lysate, in a total of 500 μl of lysis buffer. These extracts were incubated on ice for 1 hr, and then protein-G Sepharose was added to bind the antibody complex for 1 hr at 4°C. The protein-G Sepharose was pelleted and washed three times in wash buffer (20 mM Tris⋅HCl, pH 8.0/150 mM NaCl/0.5% NP-40/protease inhibitors). After the final wash the beads were pelleted and resuspended in 50 μl of sample buffer, and the immunoprecipitated proteins were separated on a 12% PAGE gel and subjected to immunoblot analysis, using the mouse anti-myc antibody as a probe. Analysis of FADD binding to Fas was similarly done except the levels of myc-LFG were varied as indicated.
Results
Cloning and Characterization of LFG.
We generated a cDNA library in a retroviral vector from a human lung fibroblast cell line, MRC-5. This cell line is not sensitive to FasL, but, like many other cell lines, is sensitive to the signal in the presence of the protein synthesis inhibitor cycloheximide (data not shown) (10). This observation suggests the existence of a labile protein and/or the need for new protein synthesis to protect cells from the Fas signal. Retroviral vectors containing the cDNA library were generated and used to transduce 5 × 106 HeLa cells. This cell line is sensitive to FasL or agonist antibody in the absence of cycloheximide. The transduced cells were maintained in the presence of a mouse anti-human Fas agonistic antibody (CH11) to induce apoptosis through a Fas-mediated cell death signal. The rationale for the experiment is that a cell will survive this selection either because of the overexpression or the ectopic expression of a cDNA provided by the retrovirus vector. However, survival can be mediated by mutation or loss of genes involved in the apoptotic pathway or up-regulation of a gene that mediates survival. This latter phenomenon can account for the residual survival of HeLa cells when challenged with the agonist anti-Fas antibody (see HeLa cells in Fig. 2B). Genomic DNA was prepared from the surviving pool of cells (HeLa FasR pool) and used as a template to amplify cDNA inserts by PCR, using primers straddling the cDNA cloning site in the retroviral vector. The most prominent PCR product was cloned and subjected to sequence analysis. Nucleotide sequence analysis of the cDNA clone revealed a long ORF encoding a protein of 316 aa with a calculated molecular mass of 34.6 kDa (Fig. 1). Sequence analysis of genomic clones revealed the existence of an upstream stop codon in-frame with the initiating methionine (data not shown), confirming that this is the full-length protein. Homology search of the existing databases revealed that this protein, which we designate as LFG (for lifeguard) is the human homologue of a recently reported rat protein, neural membrane protein 35 (NMP35) (29). This latter protein was identified by using differential display to find genes that are regulated during development of the rat sciatic nerve. However, we demonstrate that although expression of the human gene is very high in neural tissues it is not limited there. Hence we have designated the name LFG to this gene to reflect its function. LFG has significant homology in its C-terminal half to a rat glutamate binding protein (30), the Drosophila NMDA receptor-associated protein (31), and two Caenorhabditis elegans proteins of unknown function (Fig. 1). Hydropathy plots predict that LFG is a seven-membrane spanning protein. Indeed the pfam program assigned it to an uncharacterized protein family UPF005 whose members are predicted to contain seven-membrane spanning domains and that share a signature in the region beginning with the third spanning domain and ending in the middle of the fourth (32).
Figure 2.
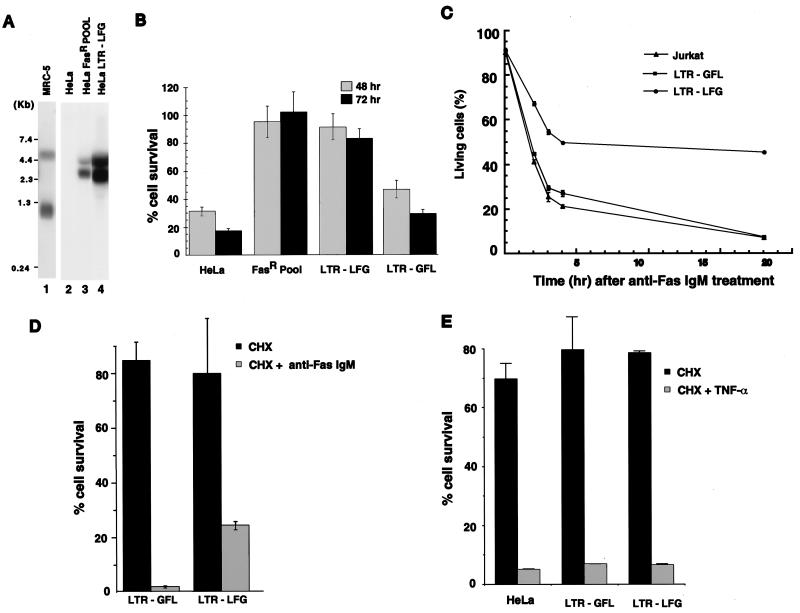
Ectopic expression of LFG in HeLa and Jurkat cells protect cells from Fas but not from TNF-α-mediated apoptosis. (A) Northern blot analysis for expression of LFG after retroviral transduction. Two transcripts are detected because of a splice site in the retroviral vector. Total RNA was derived from the following cell lines: MRC-5, HeLa, HeLa FasR pool (cells surviving the selection), and HeLa LTR-LFG (cells transduced with cDNA for LFG) and probed with the cDNA for LFG. (B) LFG protects cells from Fas-mediated apoptosis. The indicated cell lines were challenged with mouse anti-human Fas antibody (CH11), and surviving cells were monitored 48 hr and 72 hr later. Percent cell survival is expressed relative to cells that were not treated with anti-Fas antibody (mean ± SD; n = 3). (C) Jurkat cells (human T cell line), Jurkats infected with a control virus LTR-GFL, and those transduced with a LFG retroviral vector, LTR-LFG, were challenged with CH11 antibody, and annexin V expression on the cell surface was measured over time. Cell survival is plotted as a percentage of untreated cells. (D) Expression of LFG retards the kinetics of cell death by Fas. HeLa cells expressing LFG or control cells were challenged with cycloheximide (CHX) or the CH11 antibody and CHX. Percentage cell survival is plotted relative to untreated cells (mean ± SD; n = 3). (E) LFG does not protect cells from TNF-α-mediated apoptosis. HeLa cells, HeLa cells transduced with a control virus, or those expressing LFG were treated with cycloheximide (CHX) or TNF-α and CHX, and cell survival was measured 24 hr later. Percentage of cell survival is expressed relative to untreated cells (mean ± SD; n = 3).
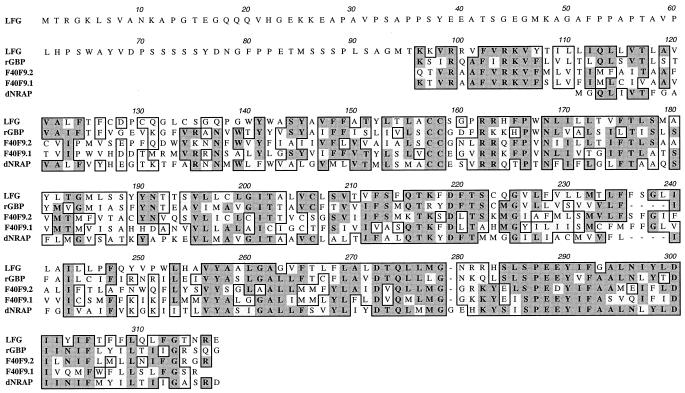
Figure 1.
Sequence comparison and composition of the 316-aa LFG protein. Sequence homology is illustrated for the most significant parts of the 516-aa rat glutamate binding protein (rGBP; GenBank locus S61973; amino acids 129–345 shown), C. elegans ORFs (GenBank locus CEF40F9) F40F9.2 (244 aa total, 25–244 shown) and F40F9.1 (295 aa total, 77–295 shown), and a 203-aa Drosophila NMDA receptor-associated protein (GenBank locus DRONMDA; 1–203 shown). blast homology searches (51) reveal highest homologies in the C terminus of LFG (from amino acid 141). Identity scores are 50% for rGBP, 46% and 42% for F40F9.2, and F40F9.1 respectively, and 42% for dNRAP.
We have determined the chromosomal localization of the LFG gene by using a human/rodent somatic hybrid panel. The result showed that the LFG gene is localized on human chromosome 12 (NIGMS panel 2 from Coriell Cell repositories), which was confirmed by fluorescence in situ hybridization, and this subchromosomal analysis revealed it on 12q13 (Y. Kaneko, N.S., and I.M.V., unpublished data). This finding is also in agreement with the designation of unidentified amplimer WI-15126, which is contained in an expressed sequence tag of LFG by the Whitehead Institute/Massachusetts Institute of Technology Center for Genome Research.
LFG Affords Protection From Fas But Not TNFR1.
To demonstrate the role of LFG in protecting cells from Fas-induced apoptosis, HeLa cells were transduced with a retroviral vector containing LFG cDNA, LTR-LFG, and treated with the CH11 antibody. Northern blot analysis of RNA from transduced cells show the expression of LFG (Fig. 2A, lane 4), whereas expression was not observed in untransduced cells (Fig. 2A, lane 2). LFG expression also was detected in the HeLa FasR pool (Fig. 2A, lane 3), and Fig. 2A (lane 1) shows the expression level of LFG in MRC-5 cells. The relative levels of LFG expression are difficult to quantitate, because MRC-5, like other tissues, reveals three LFG transcripts. Quantitation of the signal, taking into account loading and exposure times, indicated that the HeLa FasR pool expresses equivalent levels of LFG to MRC-5, whereas HeLa LTR-LFG expressed 2- to 3-fold more. Fig. 2B shows cell survival after treatment with the CH11 agonistic antibody. HeLa cells and those infected with a control virus (LFG insert in the reverse orientation, LTR-GFL) have 17% and 29% cell survival, respectively, 72 hr after treatment, whereas the FasR pool and HeLa cells infected with the LFG vector (LTR-LFG) exhibit survival of 102% and 83%, respectively. Furthermore, LFG can protect another cell type, a human Jurkat T cell line, from Fas-mediated apoptosis. This cell line is exquisitely sensitive to the Fas signal. No LFG transcripts were detected in Jurkat cells by Northern blot analysis (data not shown). These cells were infected with LFG retroviral vectors and treated with CH11, and death was monitored by using annexin V as an early index of apoptosis (33). As shown in Fig. 2C, about 50% of the cells infected with LFG retroviral vectors were annexin V negative 20 hr after treatment with CH11, and hence have not initiated apoptosis. Cells transduced with the control retroviral vector (LTR-GFL) show extensive cell death as do untransduced cells. Apoptosis in HeLa cells treated with the agonist antibody is accelerated and more complete on treatment with the antibody and cycloheximide (Fig. 2D). Because HeLa cells are relatively sensitive to Fas (e.g., compared with MRC-5 fibroblasts) this finding suggests that some molecules and/or continued protein synthesis are still able to afford some protection to HeLa cells. The kinetic synergism in killing is retarded by expression of LFG in HeLa cells such that at the 10-hr time point 24% of cells expressing LFG are viable compared with 2% of control cells. That the cells still die reflects either on the stability of LFG or the need for continued synthesis of cofactors needed by LFG to afford protection.
To determine whether LFG also protects cells from TNF-α-mediated apoptosis, HeLa cells, control LTR-GFL cells, and HeLa LTR-LFG cells were treated with TNF-α and cycloheximide. The results showed that LFG did not affect the TNF-α mediated death signal (Fig. 2E). This lack of protection by LFG also is observed for the WEHI-13VAR (ref. 34; ATCC number CRL-2148) cell line that is sensitive to TNF-α alone (data not shown). Furthermore, protection was not afforded by LFG expression to apoptosis mediated by the TNF-related apoptosis-inducing ligand TRAIL (35) or the broad spectrum protein kinase inhibitor staurosporine (data not shown).
Expression Pattern of LFG.
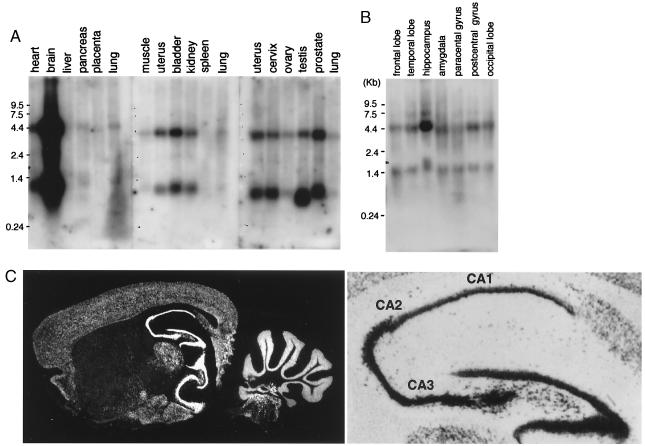
The distribution of the LFG transcripts in various human tissues was examined by Northern blot analysis. LFG is expressed in most tissues except spleen and placenta (Fig. 3A). Two major LFG transcripts of 1.3 kb and 4.4 kb were detected. Further analysis has revealed that the 1.3-kb signal is generated by two transcripts, close in size, that are generated by using an alternative polyadenlyation signal but which code for the same protein product. Similarly, sequence analysis of overlapping cDNA clones that constitute the 4.4-kb transcripts revealed an identical coding sequence to the 1.3-kb transcripts, with a large 5′ untranslated region (confirmed by expressed sequence tag KIAA0950, GenBank accession no. AB023167). The expression levels of LFG were extremely high in the brain. Dissection of the regions of the brain show that the expression is distributed in all of the areas examined (Fig. 3B). In situ hybridization analysis with a mouse cDNA of LFG (N.S. & I.M.V., unpublished work) of mouse brain sections (36) revealed LFG is expressed mostly in neurons, and the highest expression was observed in the hippocampus (Fig. 3C). Within the hippocampus, LFG expression is highest in CA3, compared with CA2 and CA1 (on average 36 silver grains over CA3 pyramidal cells versus 26 grains over CA1 pyramidal cells). No cells exhibiting glial cell morphology (cell body profile under 6 μm when nucleus was visible with large amounts of condensed chromatin at nuclear membrane) expressed LFG as assayed by significant silver grain deposition over these nuclei.
Figure 3.
Tissue distribution of LFG. (A) mRNA from various human organs was analyzed for expression of LFG by Northern blot analysis. (B) Total RNA from various parts of a dissected human brain was analyzed for LFG expression by Northern blot analysis. (C) In situ hybridization for the LFG transcripts in mouse brain (Left) shown in dark field (×7). (Right) A higher magnification (×35) of the hippocampus and dentate gyrus (no counterstain). Greatest expression occurs in the CA3 region.
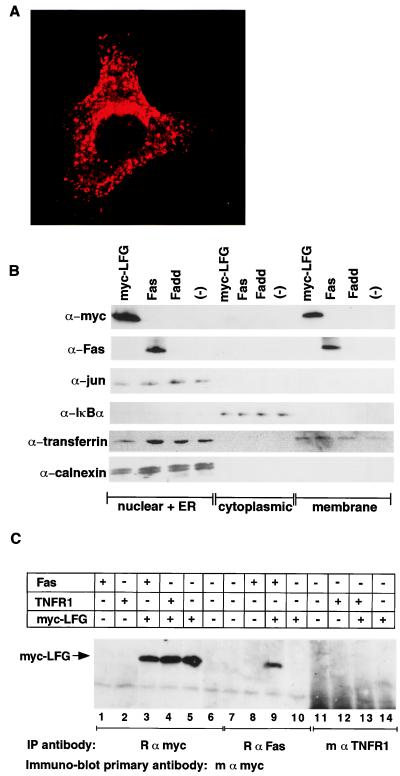
To determine the cellular localization of LFG, HeLa cells were transiently transfected with an amino-terminal myc epitope-tagged LFG (myc-LFG) expression vector. Confocal microscopy reveals LFG to be cytoplasmic (Fig. 4A) and predominantly membrane associated, with peri-nuclear staining reminiscent of endoplasmic reticulum and Golgi structures. Furthermore LFG distributed in vesicles and at the cell membrane. Biochemical fractionation of 293 cells transfected with expression vectors for myc-LFG or human Fas confirms that LFG is membrane associated (Fig. 4B).
Figure 4.
Myc-epitope-tagged LFG is membrane associated and interacts with Fas receptor. (A) Confocal microcopy (×63) of a HeLa cell transfected with an expression plasmid for six myc epitope tagged-LFG. (B) Subcellular fractionation of LFG. 293 cells transfected with myc-LFG, human Fas, human FADD, or mock transfected (−) were separated into nuclear + endoplasmic reticulum (ER) and cytoplasmic and membrane fractions. The fractions then were probed by immunoblot analysis with antibodies to myc (to follow tagged LFG), human Fas receptor, c-jun (a nuclear marker), IκBα (a cytoplasmic marker), transferrin receptor (ER and cell membrane marker), and calnexin (ER marker). (C) Coimmunoprecipitation of LFG with Fas. Expression plasmids for human Fas, human TNFR1, or myc-LFG were transfected into 293 cells singly or in combination. Cell extracts from transfected cells were immunoprecipitated with rabbit anti-myc, mouse anti-human Fas, or mouse anti-TNFR1 antibodies. The precipitates were subjected to immunoblot analysis with mouse anti-myc antibody.
LFG Interacts with Fas But Not TNF-α.
The mechanism of Fas-mediated apoptosis has been elucidated as recruitment of FADD to ligated Fas receptor and subsequent activation of caspase-8. In contrast apoptosis mediated by ligated TNF-α receptor involves the ordered recruitment of TRADD, FADD, and caspase-8 (5). The difference between the two, the adapter molecule TRADD, and the inability of LFG to protect cells from TNF-α mediated death, therefore places the action of LFG upstream of FADD, at Fas or at the level of the Fas/FADD complex. Hence we next investigated whether LFG interacts with components of the Fas signaling pathway. Fas and LFG can be coimmunoprecipitated after transient transfection. Immunoprecipitation and immunoblot analysis revealed that anti-human Fas antibodies can coimmunoprecipitate myc-LFG (Fig. 4C, lane 9), whereas no myc-LFG precipitated with anti-human TNFR1 antibodies (Fig. 4C, lane 13), although these are competent to immunoprecipitate the TNFR1-associated molecule TRADD (data not shown). Furthermore, LFG does not immunoprecipitate FADD.
LFG Does Not Down-Regulate Fas Expression or FADD Binding.
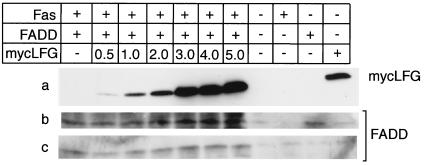
The interaction of LFG with Fas suggests mechanisms such as down-regulation of the Fas receptor, inhibition of ligand binding, or inhibition of FADD binding to Fas. To investigate the first two of these possibilities, control HeLa LTR-GFL cells or LFG expressing HeLa cells were analyzed for Fas expression. In this study the agonist antibody CH11 was bound to the cells and its binding was assayed by fluorescence-activated cell sorting. The mean fluorescence is not altered between control HeLa cells and HeLa cells expressing LFG (data not shown). This result eliminates the possible mechanism of receptor down-regulation or interference to antibody binding, because the same agonist antibody that is used to initiate cell death was used to determine receptor expression. The third possibility is that of interference with Fas/FADD association. To examine this possibility we coimmunoprecipitated FADD with anti-Fas antibodies in the presence of increasing amounts of LFG. Increasing the expression levels of LFG (Fig. 5a) did not affect expression of FADD (Fig. 5b) or affect the extent of coimmunoprecipitation of FADD with anti-Fas antibodies (Fig. 5c). These results eliminate some models that explain the specificity of LFG protection for Fas-mediated apoptosis and imply a novel mechanism.
Figure 5.
LFG does not affect FADD binding to Fas. Five micrograms of expression plasmids for human Fas and human FADD and varying concentrations of myc-LFG (indicated in μg) were transfected singly or in combination into 293 cells. Twenty micrograms of cell extract was subjected to immunoblot analysis with either mouse anti-myc (a) or mouse anti-human FADD (b). Two hundred micrograms of extracts was immunoprecipitated with rabbit anti-human Fas antibody and analyzed by immunoblot analysis with mouse anti-FADD antibodies (c).
Discussion
We report here the cloning of a cDNA that uniquely protects cells from Fas-mediated death. The mechanism of Fas-mediated cell death has been elucidated as recruitment of FADD to ligated receptor and subsequently caspase-8. In contrast apoptosis mediated by ligated TNF-α receptor involves the ordered recruitment of TRADD, FADD, and caspase-8 (5). The difference between the two, the adapter molecule TRADD, and the inability of LFG to protect cells from TNF-α-mediated death, places the action of LFG upstream of FADD, at Fas, or at the level of the Fas/FADD complex.
Fas has been mainly characterized in the immune system and liver and primarily is involved in regulating the immune response (37) and the homeostasis of liver cell number (38). However, Fas and FasL are detectable in other tissues, for example in the central nervous system in normal (39) and brains suffering neurodegeneration (40), raising the possibility that Fas and its ligand have a role in neurological diseases. Consistent with this observation is the high expression of LFG in the brain. Interestingly expression of Fas and FasL is observed in the hippocampus after a single injection of NMDA (41). If transient exposure to NMDA results in transient expression of Fas and FasL, it is tempting to speculate that LFG protects cells from this situation where the cell death program would be initiated and lead to subsequent neurodegeneration. In this regard the most proximal inhibitor molecules in the pathway, cFLIPS are not expressed in the brain (42–45).
The expression of LFG is seen in other tissues and is not restricted to the brain. Interestingly, studies have identified situations where both Fas and FasL are expressed on the same cells or in the same tissue without ensuing “suicide” or “fratricide” (46). It has been shown that a number of tumor cells are resistant to Fas-mediated cell death and may even express both Fas and FasL (47). In these situations expression of LFG may in some part explain the survival of these cells.
Pathologies implicated to involve chromosome 12q13 that may have apoptosis as a component include leukemia (48), lipomas (49), and Alzheimer’s disease (50). Finally, the expression and modulation of LFG may be of practical therapeutic use, when protection from Fas is required, while keeping the biological function of other death receptors intact.
Acknowledgments
We thank Hiroyuki Miyoshi and Heinz Ruffner for critical reading of the manuscript, Gerald Pao for help with confocal microscopy, and members of the Verma lab for helpful discussions. This work was supported by National Institutes of Health Grants CA44360 and HL53670 (I.M.V.) and AG10435 (S.F.H.), the March of Dimes Foundation (I.M.V.), the Wayne and Gladys Valley Foundation (I.M.V.), and the H.N. and Frances C. Berger Foundation (I.M.V.). I.M.V. is a Franklin D. Roosevelt Fellow of the March of Dimes Foundation and an American Cancer Society Professor of Molecular Biology.
Abbreviations
- TNF
tumor necrosis factor
- TNFR1
TNF receptor 1
Footnotes
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AF190461).
References
- 1.Jacobson M D, Weil M, Raff M C. Cell. 1997;88:347–354. doi: 10.1016/s0092-8674(00)81873-5. [DOI] [PubMed] [Google Scholar]
- 2.Winoto A. Curr Opin Immunol. 1997;9:365–370. doi: 10.1016/s0952-7915(97)80083-0. [DOI] [PubMed] [Google Scholar]
- 3.Evan G, Littlewood T. Science. 1998;281:1317–1322. doi: 10.1126/science.281.5381.1317. [DOI] [PubMed] [Google Scholar]
- 4.Arai H, Gordon D, Nabel E G, Nabel G J. Proc Natl Acad Sci USA. 1997;94:13862–13867. doi: 10.1073/pnas.94.25.13862. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Ashkenazi A, Dixit V M. Science. 1998;281:1305–1308. doi: 10.1126/science.281.5381.1305. [DOI] [PubMed] [Google Scholar]
- 6.Itoh N, Yonehara S, Ishii A, Yonehara M, Mizushima S, Sameshima M, Hase A, Seto Y, Nagata S. Cell. 1991;66:233–243. doi: 10.1016/0092-8674(91)90614-5. [DOI] [PubMed] [Google Scholar]
- 7.Oehm A, Behrmann I, Falk W, Pawlita M, Maier G, Klas C, Li-Weber M, Richards S, Dhein J, Trauth B C, et al. J Biol Chem. 1992;267:10709–10715. [PubMed] [Google Scholar]
- 8.Itoh N, Nagata S. J Biol Chem. 1993;268:10932–10937. [PubMed] [Google Scholar]
- 9.Suda T, Takahashi T, Golstein P, Nagata S. Cell. 1993;75:1169–1178. doi: 10.1016/0092-8674(93)90326-l. [DOI] [PubMed] [Google Scholar]
- 10.Yonehara S, Ishii A, Yonehara M. J Exp Med. 1989;169:1747–1756. doi: 10.1084/jem.169.5.1747. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Chinnaiyan A M, O’Rourke K, Tewari M, Dixit V M. Cell. 1995;81:505–512. doi: 10.1016/0092-8674(95)90071-3. [DOI] [PubMed] [Google Scholar]
- 12.Boldin M P, Varfolomeev E E, Pancer Z, Mett I L, Camonis J H, Wallach D. J Biol Chem. 1995;270:7795–7798. doi: 10.1074/jbc.270.14.7795. [DOI] [PubMed] [Google Scholar]
- 13.Muzio M, Chinnaiyan A M, Kischkel F C, O’Rourke K, Shevchenko A, Ni J, Scaffidi C, Bretz J D, Zhang M, Gentz R, et al. Cell. 1996;85:817–827. doi: 10.1016/s0092-8674(00)81266-0. [DOI] [PubMed] [Google Scholar]
- 14.Boldin M P, Goncharov T M, Goltsev Y V, Wallach D. Cell. 1996;85:803–815. doi: 10.1016/s0092-8674(00)81265-9. [DOI] [PubMed] [Google Scholar]
- 15.Srinivasula S M, Ahmad M, Fernandes-Alnemri T, Litwack G, Alnemri E S. Proc Natl Acad Sci USA. 1996;93:14486–14491. doi: 10.1073/pnas.93.25.14486. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Muzio M, Stockwell B R, Stennicke H R, Salvesen G S, Dixit V M. J Biol Chem. 1998;273:2926–2930. doi: 10.1074/jbc.273.5.2926. [DOI] [PubMed] [Google Scholar]
- 17.Cheng J, Zhou T, Liu C, Shapiro J P, Brauer M J, Kiefer M C, Barr P J, Mountz J D. Science. 1994;263:1759–1762. doi: 10.1126/science.7510905. [DOI] [PubMed] [Google Scholar]
- 18.Pitti R M, Marsters S A, Lawrence D A, Roy M, Kischkel F C, Dowd P, Huang A, Donahue C J, Sherwood S W, Baldwin D T, et al. Nature (London) 1998;396:699–703. doi: 10.1038/25387. [DOI] [PubMed] [Google Scholar]
- 19.Tollefson A E, Hermiston T W, Lichtenstein D L, Colle C F, Tripp R A, Dimitrov T, Toth K, Wells C E, Doherty P C, Wold W S. Nature (London) 1998;392:726–730. doi: 10.1038/33712. [DOI] [PubMed] [Google Scholar]
- 20.Elsing A, Burgert H G. Proc Natl Acad Sci USA. 1998;95:10072–10077. doi: 10.1073/pnas.95.17.10072. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Tschopp J, Irmler M, Thome M. Curr Opin Immunol. 1998;10:552–558. doi: 10.1016/s0952-7915(98)80223-9. [DOI] [PubMed] [Google Scholar]
- 22.Hitoshi Y, Lorens J, Kitada S I, Fisher J, LaBarge M, Ring H Z, Francke U, Reed J C, Kinoshita S, Nolan G P. Immunity. 1998;8:461–471. doi: 10.1016/s1074-7613(00)80551-8. [DOI] [PubMed] [Google Scholar]
- 23.Deveraux Q L, Roy N, Stennicke H R, Van Arsdale T, Zhou Q, Srinivasula S M, Alnemri E S, Salvesen G S, Reed J C. EMBO J. 1998;17:2215–2223. doi: 10.1093/emboj/17.8.2215. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Enari M, Hug H, Nagata S. Nature (London) 1995;375:78–81. doi: 10.1038/375078a0. [DOI] [PubMed] [Google Scholar]
- 25.Chen P, Tian J, Kovesdi I, Bruder J T. J Biol Chem. 1998;273:5815–5820. doi: 10.1074/jbc.273.10.5815. [DOI] [PubMed] [Google Scholar]
- 26.Chomczynski P, Sacchi N. Anal Biochem. 1987;162:156–159. doi: 10.1006/abio.1987.9999. [DOI] [PubMed] [Google Scholar]
- 27.Vojtek A B, Cooper J A. J Cell Sci. 1993;105:777–785. doi: 10.1242/jcs.105.3.777. [DOI] [PubMed] [Google Scholar]
- 28.Evan G I, Lewis G K, Ramsay G, Bishop J M. Mol Cell Biol. 1985;5:3610–3616. doi: 10.1128/mcb.5.12.3610. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Schweitzer B, Taylor V, Welcher A A, McClelland M, Suter U. Mol Cell Neurosci. 1998;11:260–273. doi: 10.1006/mcne.1998.0697. [DOI] [PubMed] [Google Scholar]
- 30.Kumar K N, Tilakaratne N, Johnson P S, Allen A E, Michaelis E K. Nature (London) 1991;354:70–73. doi: 10.1038/354070a0. [DOI] [PubMed] [Google Scholar]
- 31.Pellicena-Palle A, Salz H K. Biochim Biophys Acta. 1995;1261:301–303. doi: 10.1016/0167-4781(95)00031-b. [DOI] [PubMed] [Google Scholar]
- 32.Walter L, Marynen P, Szpirer J, Levan G, Gunther E. Genomics. 1995;28:301–304. doi: 10.1006/geno.1995.1145. [DOI] [PubMed] [Google Scholar]
- 33.Martin S J, Reutelingsperger C P, McGahon A J, Rader J A, van Schie R C, LaFace D M, Green D R. J Exp Med. 1995;182:1545–1556. doi: 10.1084/jem.182.5.1545. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Khabar K S, Siddiqui S, Armstrong J A. Immunol Lett. 1995;46:107–110. doi: 10.1016/0165-2478(95)00026-2. [DOI] [PubMed] [Google Scholar]
- 35.Wiley S R, Schooley K, Smolak P J, Din W S, Huang C P, Nicholl J K, Sutherland G R, Smith T D, Rauch C, Smith C A, et al. Immunity. 1995;3:673–682. doi: 10.1016/1074-7613(95)90057-8. [DOI] [PubMed] [Google Scholar]
- 36.Simmons D M, Arriza J L, Swanson L W. J Histotechnol. 1989;12:169–181. [Google Scholar]
- 37.Nagata S. Cell. 1997;88:355–365. doi: 10.1016/s0092-8674(00)81874-7. [DOI] [PubMed] [Google Scholar]
- 38.Adachi M, Suematsu S, Kondo T, Ogasawara J, Tanaka T, Yoshida N, Nagata S. Nat Genet. 1995;11:294–300. doi: 10.1038/ng1195-294. [DOI] [PubMed] [Google Scholar]
- 39.Park C, Sakamaki K, Tachibana O, Yamashima T, Yamashita J, Yonehara S. Biochem Biophys Res Commun. 1998;252:623–628. doi: 10.1006/bbrc.1998.9572. [DOI] [PubMed] [Google Scholar]
- 40.de la Monte S M, Sohn Y K, Wands J R. J Neurol Sci. 1997;152:73–83. doi: 10.1016/s0022-510x(97)00131-7. [DOI] [PubMed] [Google Scholar]
- 41.Shin S W, Park J W, Suh M H, Suh S I, Choe B K. J Neurochem. 1998;71:1773–1776. doi: 10.1046/j.1471-4159.1998.71041773.x. [DOI] [PubMed] [Google Scholar]
- 42.Shu H B, Halpin D R, Goeddel D V. Immunity. 1997;6:751–763. doi: 10.1016/s1074-7613(00)80450-1. [DOI] [PubMed] [Google Scholar]
- 43.Hu S, Vincenz C, Ni J, Gentz R, Dixit V M. J Biol Chem. 1997;272:17255–17257. doi: 10.1074/jbc.272.28.17255. [DOI] [PubMed] [Google Scholar]
- 44.Srinivasula S M, Ahmad M, Ottilie S, Bullrich F, Banks S, Wang Y, Fernandes-Alnemri T, Croce C M, Litwack G, Tomaselli K J, et al. J Biol Chem. 1997;272:18542–18545. doi: 10.1074/jbc.272.30.18542. [DOI] [PubMed] [Google Scholar]
- 45.Irmler M, Thome M, Hahne M, Schneider P, Hofmann K, Steiner V, Bodmer J L, Schroter M, Burns K, Mattmann C, et al. Nature (London) 1997;388:190–195. doi: 10.1038/40657. [DOI] [PubMed] [Google Scholar]
- 46.Bonfoco E, Stuart P M, Brunner T, Lin T, Griffith T S, Gao Y, Nakajima H, Henkart P A, Ferguson T A, Green D R. Immunity. 1998;9:711–720. doi: 10.1016/s1074-7613(00)80668-8. [DOI] [PubMed] [Google Scholar]
- 47.Husain N, Chiocca E A, Rainov N, Louis D N, Zervas N T. Acta Neuropathol. 1998;95:287–290. doi: 10.1007/s004010050799. [DOI] [PubMed] [Google Scholar]
- 48.Dierlamm J, Wlodarska I, Michaux L, Vermeesch J R, Meeus P, Stul M, Criel A, Verhoef G, Thomas J, Delannoy A, et al. Genes Chromosomes Cancer. 1997;20:155–166. [PubMed] [Google Scholar]
- 49.Merscher S, Marondel I, Pedeutour F, Gaudray P, Kucherlapati R, Turc-Carel C. Genomics. 1997;46:70–77. doi: 10.1006/geno.1997.4993. [DOI] [PubMed] [Google Scholar]
- 50.Knuutila S, Bjorkqvist A M, Autio K, Tarkkanen M, Wolf M, Monni O, Szymanska J, Larramendy M L, Tapper J, Pere H, et al. Am J Pathol. 1998;152:1107–1123. [PMC free article] [PubMed] [Google Scholar]
- 51.Altschul S F, Madden T L, Schaffer A A, Zhang J, Zhang Z, Miller W, Lipman D J. Nucleic Acids Res. 1997;25:3389–3402. doi: 10.1093/nar/25.17.3389. [DOI] [PMC free article] [PubMed] [Google Scholar]