Abstract
The 2.0-Å resolution x-ray crystal structure of a novel trimeric antibody fragment, a “triabody,” has been determined. The trimer is made up of polypeptides constructed in a manner identical to that previously described for some “diabodies”: a VL domain directly fused to the C terminus of a VH domain—i.e., without any linker sequence. The trimer has three Fv heads with the polypeptides arranged in a cyclic, head-to-tail fashion. For the particular structure reported here, the polypeptide was constructed with a VH domain from one antibody fused to the VL domain from an unrelated antibody giving rise to “combinatorial” Fvs upon formation of the trimer. The structure shows that the exchange of the VL domain from antibody B1-8, a Vλ domain, with the VL domain from antibody NQ11, a Vκ domain, leads to a dramatic conformational change in the VH CDR3 loop of antibody B1-8. The magnitude of this change is similar to the largest of the conformational changes observed in antibody fragments in response to antigen binding. Combinatorial pairing of VH and VL domains constitutes a major component of antibody diversity. Conformationally flexible antigen-binding sites capable of adapting to the specific CDR3 loop context created upon VH–VL pairing may be employed by the immune system to maximize the structural diversity of the immune response.
The antigen-binding site in antibodies is formed by the hypervariable loop regions (complementarity-determining regions, CDRs) of VH and VL domain pairs to yield a continuous surface mounted on a rigid scaffold provided by the framework regions of these domains. The ultimate diversity of antigen binding is determined by the structural diversity of this surface. From extensive analysis of structures of antibody fragments, it is clear that there is only a small set of “canonical” main-chain loop conformations for five of the six CDRs (1, 2). However, CDR3 of the VH domain, located at the center of the antigen-binding site, has defied attempts at classification due to its large variability in length and sequence, resulting from the junctional diversity generated in somatic VH gene assembly. Furthermore, a range of conformational changes in antigen-binding sites can occur upon antigen binding (3–8). These changes range from minor side-chain adjustments to major rearrangements in the main-chain loop conformations of VH CDR3, as well as to changes in the relative orientation of the VH and VL domains (3). Furthermore, the biphasic kinetics of some antibodies in response to antigen binding suggest that there is conformational heterogeneity even prior to antigen binding (9, 10).
A fundamental component of antibody diversity arises from random VH–VL pairings. “Promiscuous” VH–VL pairings have also been observed in phage-displayed antibody libraries and have been exploited for affinity maturation in vitro (11) or “humanizing” antibodies (12, 13). It has been anticipated that novel VH–VL pairings could influence the conformations of the CDR loops (14). We have determined the structure of an Fv consisting of a VH domain from one antibody paired with a VL domain from an unrelated antibody. This structure shows a large rearrangement of the antigen-binding region of the VH domain and reinforces that the expressed structural diversity is not simply related to the product of the sequence diversity of the VH and VL domains considered independently.
An unrelated but important objective of our work was to show that single chain Fv fragments with very short linkers between the VH and VL domains (zero residues in this instance) can form stable trimers. Previously, we described the structure of a dimeric antibody construct known as a “diabody” (15). In such a construct VH and VL domains are fused to each other with a linker sequence too short to permit intramolecular pairing of the domains, forcing formation of dimers. In several diabody preparations, we have noticed higher-order oligomers. For the B1-8/NQ11 construct in which the VH domain of B1-8 is directly fused to the N terminus of the VL domain of NQ11, the preparation consisted predominantly of oligomers with a gel-filtration estimated size consistent with a trimer. We report here the structure of a trimeric antibody fragment.
MATERIALS AND METHODS
Vector Construction.
An antibody fragment was constructed so that the VH domain of antibody B1-8 was directly fused to the VL domain of antibody NQ11 as illustrated in Fig. 1. Because the VH and VL domains are directly fused to each other, it is not sterically possible for the VH and VL domains in a single polypeptide to pair with each other. Consequently, the polypeptides form oligomers in which the VH–VL pairing is achieved intermolecularly (16–19). This type of multivalent antibody fragment has been referred to as a diabody (15, 17), although dimeric, trimeric, and higher-order multimers can be isolated (16–19).
Figure 1.
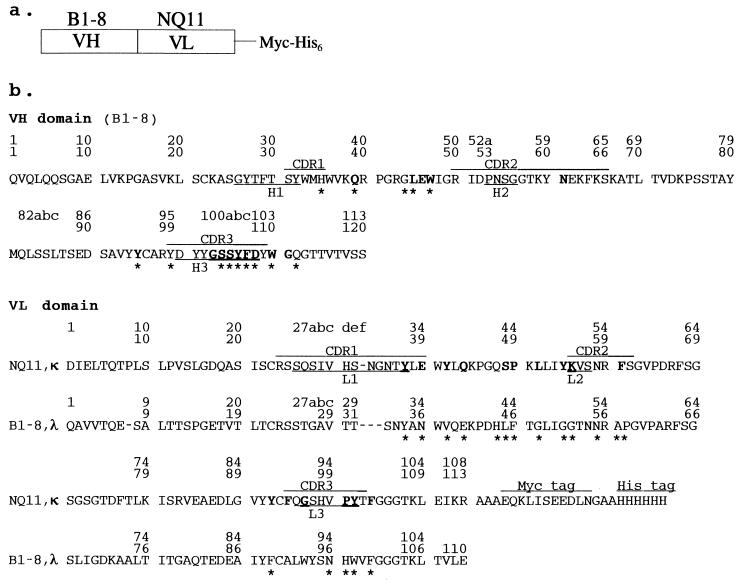
(a) A schematic of the B1-8/NQ11 construct. The polypeptide chain of the homotrimer is composed of an N-terminal VH domain from antibody B1-8 (37) fused directly to a C-terminal VL domain (κ-type) from antibody NQ11 with a myc tag and a (His)6 tag at the C terminus. (b) The sequence of the triabody B1-8/NQ11. The CDRs are indicated by bars above the sequence. The hypervariable loops as defined by Chotia and Lesk (1) are indicated by bars below the sequence. The sequence of the cognate VL domain (λ-type) from B1-8 is indicated below the sequence of the VL domain from NQ11. The sequential sequence numbering used throughout the text and in the Protein Data Bank entry is on the line closest to the sequence. The numbering of Kabat et al. (38) is shown as the uppermost line of each sequence. The residues that are involved in VH–VL interface contacts (closer than 4.0 Å) are shown in bold for B1-8/NQ11 and by asterisks below the sequence for B1-8.
Expression and Purification of Antibody Fragments.
The multivalent antibody fragment B1-8/NQ11 was expressed in Escherichia coli. The protein was purified from culture supernatant using three chromatagraphic steps: Ni2+-NTA agarose (Qiagen, Chatsworth, CA) affinity purification, gel filtration with a HiLoad 16/60 Superdex 75 column (Pharmacia) equilibrated with 20 mM Tris⋅HCl (pH 8.0), 0.1 M NaCl, and finally a salt gradient on a MonoQ HR 5/5 column equilibrated in 20 mM Tris⋅HCl (pH 8.0). Gel filtration analysis indicated that dimers, trimers, and higher-order oligomers are present. The trimeric fraction was pooled and applied to the MonoQ. The protein was concentrated to 18.7 mg/ml and stored at 4°C. Our purification protocol and structure determination focused on a trimeric or “triabody” species.
Crystallization.
Initial small crystals appeared at 4°C in 30% 2-propanol, 0.1 M Na cacodylate (pH 6.5), and 0.2 M Na citrate. Subsequently, pre-equilibrated drops were microseeded. A typical crystal had dimensions of 0.2 mm × 0.2 mm × 0.1 mm. The crystals have R3 symmetry with unit cell dimensions a = 136.32 Å, c = 74.8 Å. The Matthews coefficient (VM) of the crystals is 2.4 Å3/Da.
X-Ray Diffraction Data Collection.
Data from a single crystal were collected at the Daresbury Synchrotron Station 9.6 at a wavelength of 0.882 Å with a MAR-Research (Hamburg, Germany) 300-mm image plate. Data were collected to a resolution of 1.9 Å. The crystal for data collection was flash frozen in a nitrogen cold stream at 95 K. The freezing solution was the same solution as used for growing the triabody crystals with the addition of 20% (wt/vol) 2-methyl-2,4-pentanediol. Diffraction data were processed and refined with the program mosflm (20). Data sorting, scaling and merging were done with the CCP4 package (21). There were a total of 95,579 observations of 33,839 unique reflections giving an average redundancy of 2.8. Data were 96.6% complete to 1.9 Å. The overall Rsym was 0.053 based on intensities with an Rsym of 0.174 for the data from 2.0 Å to 2.05 Å.
Initial Phase Determination.
Initial phases were obtained by molecular replacement method using the program amore (22). An initial rotation function was calculated using search models derived from a library of 37 Fv structures. The 10 highest rotation function solutions for each Fv were used for translation function searches. Monoclonal antibody fragment JE142 (Protein Data Bank entry 1jel) yielded the highest correlation coefficient (21.9%) and lowest R factor (51.8%) and was used for the next step of the analysis. This initial rotation/translation solution was refined as a rigid-body and was subsequently fixed for a second round of the translation function searches to locate a second molecule. In the second round of the translation function searches, all of the 37 Fv structures were again used. The second round of translation function searches yielded a correlation coefficient of 29.1% and an R factor of 49.8%. Again, JE142 was the Fv that gave the strongest signal. The two molecules in the asymmetric unit were refined as four rigid-bodies (two VH and two VL domains). This yielded a correlation coefficient of 61.3% and R factor of 36.5%.
Structure Refinement.
The structure was refined using data to 2.0-Å resolution. Unrestrained refinement was carried out with the program arp (23) to generate an electron density map that was used for initial model building. An initial model was built using the program o (24). Noncrystallographic symmetry restrained positional and temperature factor refinement were carried out with the program x-plor. A few cycles of x-plor (25) least-squares refinement (using all data not flagged as “free” from 6.0 Å to 2.0 Å resolution, 31,030 reflections) alternated with model rebuilding into difference electron density maps produced as a result of unrestrained arp refinement, yielded an R factor of 0.235 and a free R factor of 0.262 (using 5% of the data, 1,649 reflections). After switching off the NCS restraints, the R factor was 0.187 with a free R factor of 0.230. Simulated annealing with x-plor did not improve the free R factor. For one molecule in the asymmetric unit, residues 101–108 of the VH domain are well defined (Fig. 2). For the other molecule in the asymmetric unit, this region is disordered. The myc and (His)6 tags at the C terminus of the polypeptide are not visible for either of the molecules in the asymmetric unit. The junction between the N-terminal VH and C-terminal VL domains is well ordered and clearly discernible for both molecules in the asymmetric unit. The electron density is disordered for the main-chain in region ProH41 to ArgH43 of both polypeptides in the asymmetric unit. The model has good stereochemistry as evaluated by procheck (26). Root-mean squared deviations from ideal geometry are 0.008 Å, 1.41°, and 26.9° for bond lengths, bond angles, and dihedral angles, respectively.
Figure 2.
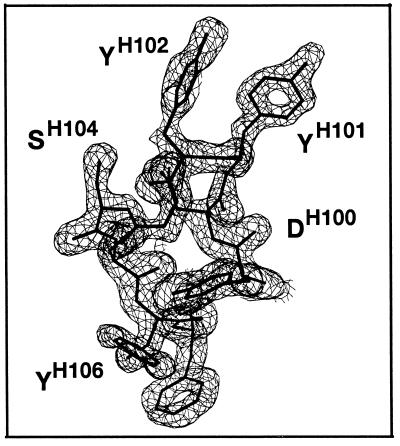
A simulated annealing OMIT difference electron density map contoured at 3σ for VH CDR3 of the B1-8/NQ11 triabody [drawn with bobscript (R. Esnouf, personal communication), a modified version of molscript (39)].
RESULTS AND DISCUSSION
The Structure of a Trimeric, Combinatorial Antibody Fragment.
The overall fold of the antibody fragment is of a type that has been referred to as a domain-swapped trimer (27). The trimer has three Fv heads with the polypeptides in the trimer arranged in a cyclic head-to-tail fashion. The VH domain of each polypeptide pairs with the VL domain of a neighboring polypeptide in the trimer giving rise to three antigen-binding sites (Fig. 3). Monomers of single chain Fvs with short linker sequences between the VH and VL domains are not functional and constitute little if any of the material expressed. However, the structure of NQ11/B1-8 and the structure of a diabody reported previously (15) show that this type of antibody fragment readily forms domain-swapped dimers and trimers.
Figure 3.
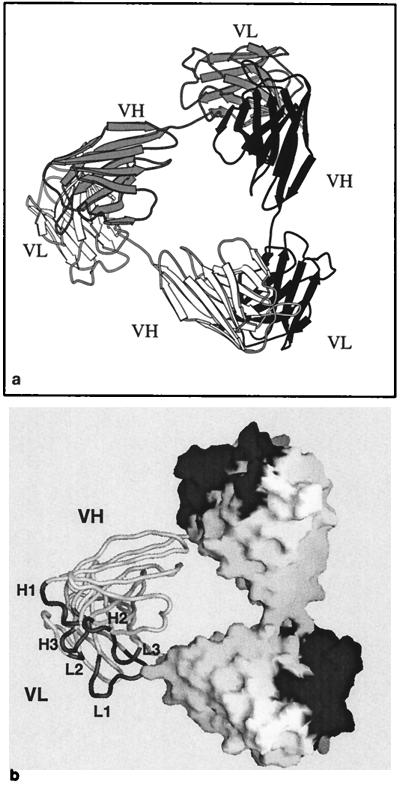
The overall structure of the triabody. (a) Three polypeptides in one triabody are related to each other by crystallographic symmetry and are rendered in different shades of gray. The triabody has three Fv heads, each consisting of a VH domain from one polypeptide paired with the VL domain from a neighboring polypeptide. (b) A view of a triabody with the molecular surface superimposed on two of the Fv heads in the triabody and the third Fv head shown in worm representation. The CDRs of each Fv head are shaded dark gray.
The present structure may be used as a blueprint for the design and construction of trivalent or even trispecific antibody fragments. Triabodies could bind three different or identical epitopes on the same molecule leading to higher specificities and higher functional affinities. Furthermore, by cross-linking three different types of antigens trispecific triabodies might, in principle, be useful immunotherapeutic agents by retargeting resting cytotoxic lymphocytes in a manner that has been demonstrated for other trispecific antibody fragments (28, 29). However, because of random pairing in a triabody system expressing three different polypeptides, only a small fraction of correctly paired, trispecific triabodies would be produced. The fraction of trispecific triabody might be improved by engineering specificity into the VH–VL domain interface as has been described for diabodies (30).
The polypeptide forming the trimer used in this study has a Vκ domain from the NQ11 anti-hapten phOx (2-phenyl-5-oxazolone) antibody fused to the VH domain from an unrelated antibody, a B1-8 anti-hapten NIP (4-hydroxy-5-iodo-3-nitrophenyl acetyl) antibody. Although the “combinatorial” Fvs in the triabody we have crystallized are not capable of binding either of the haptens for which the parental antibodies were specific, they have allowed us to examine the influence of the VL domain on the conformations of the CDRs of the VH domain.
The Effect of Exchanging One VL Domain for Another on the VH Conformation.
The structure of Fv B1-8 fragment had been solved previously both in the presence and absence of antigen (T. Simon and K. Henrick, personal communication). The structure of a very closely related NIP-binding Fab, N1G9, which differs by only one conservative replacement in the VH domain (V116L), has also been determined both in the presence and absence of antigen (31) (Protein Data Bank entries 1 ngp and 1 ngq) and these four structures are nearly identical to each other. We have compared the structure of the B1-8 VH domain paired to the B1-8 Vλ chain with the structure of it paired to the Vκ domain from NQ11. Inspection of the combinatorial B1-8/NQ11 Fv structure shows that substitution of one VL domain for another had little influence on the conformations of the first two hypervariable loops of the VH domain but had a profound influence on VH CDR3 with deviations of 0.17 Å, 0.15 Å, and 1.8 Å for H1, H2, and H3, respectively.
The triabody context per se in which the combinatorial Fvs were expressed does not appear to have an influence on the conformation of the CDRs. As was observed for a diabody (15), both the VH and VL domains in each Fv head have a conformation similar to those of other antibody fragments having the same canonical classes of hypervariable regions. The main-chain rms deviation between the VH domains of the triabody B1-8/NQ11 and the Fv B1-8 is only 0.44 Å. The profound difference in the conformation of CDR3 is largely due to conformational changes in the triabody caused by differences in VH–VL packing arising as a consequence of exchanging one VL domain for another unrelated one.
Disposition of the VH Domain Relative to the VL Domain.
Associated with the exchange of one VL domain for another unrelated one is a pronounced change in the relative orientation of the VH and VL domains. The relative shift of the VL domain in B1-8/NQ11 in comparison with Fv B1-8 is a 9.1° rotation around an axis perpendicular to the VH–VL interface and 2.4-Å translation (Fig. 4). The change in orientation that we have observed upon exchanging one VL domain for an unrelated one is similar to the largest of the antigen induced interdomain movements that have been reported: 16° for the HIV-1 neutralizing Fab 50.1 and 7.5° for the anti-DNA Fab BV04 (32).
Figure 4.
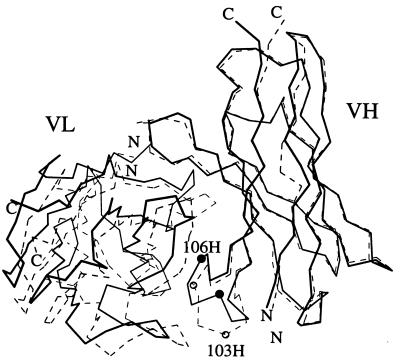
A comparison of the VL–VH orientations of the B1-8 (dashed lines) and B1-8/NQ11 (solid lines) Fvs. The Cα values of the B1-8 VH framework were superimposed on the B1-8/NQ11 VH framework. The Cα positions of residues GlyH103 and TyrH106 are indicated by spheres (white for B1-8 and black for B1-8/NQ11).
The residues involved in the VH–VL interface are quite common in comparison with other antibodies (Table 1). Among these interface residues are most of the residues of the VH CDR3 loop. The extent of the VH–VL interface is similar to the interface found in Fv B1-8 and fairly large (1,720 Å2) relative to other antibodies (solvent accessible surface area buried in the interface calculated using a 1.4-Å probe and the default atomic radii of the program grasp). Most of the VH–VL interactions in B1-8 involve residues in L3 and H3 (31). The residues of the VH domain that are involved in the interface of B1-8/NQ11 are generally the same residues involved in the B1-8 interface (Fig. 1). Those that differ between B1-8/NQ11 and B1-8 are residues packing against CDR3 of the VL domain. These differences in packing are not surprising. All of the mouse Vλ domains whose structures have been determined, including B1-8, have similar L3 conformations, and these differ from canonical “type-1” L3 conformation found in Vκs such as that of B1-8/NQ11 (1).
Table 1.
Framework residues (40) found at the VL–VH interface (sequential B1-8/NQ11 numbering) for several antibodies whose structures have been reported
| Antibody (PDB entry) | VL domain
|
VH domain
|
||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 39 | 41 | 43 | 49 | 51 | 92 | 94 | 101 | 103 | 35 | 37 | 39 | 45 | 47 | 95 | 97 | 107 | 110 | |
| B1-8/NQ11 triabody | E | Y | Q | P | L | Y | F | Y | F | H | V | Q | L | W | Y | A | F | W |
| B1-8 | N | V | E | F | G | F | A | W | F | H | V | Q | L | W | Y | A | F | W |
| JE142 (1jel) | E | Y | Q | P | L | Y | F | Y | F | H | V | Q | L | W | Y | A | F | W |
| ANO2 (1baf) | Y | Y | Q | P | L | Y | Q | I | F | N | I | Q | L | W | Y | T | F | W |
| HIL (8fab) | Y | Y | Q | P | M | Y | Q | S | F | H | V | Q | L | W | Y | A | F | W |
| A/CYC1 (likf) | N | Y | Q | V | L | F | Q | P | F | Y | V | Q | L | W | Y | T | F | W |
| D1.3 (1vfa) | A | Y | Q | P | L | Y | Q | R | F | N | V | Q | L | W | Y | A | L | W |
| MCPC603 (1mcp) | A | Y | Q | P | L | Y | Q | L | F | E | V | Q | L | W | Y | A | F | W |
| NEW (7fab) | K | Y | Q | P | L | Y | Q | R | F | T | V | Q | L | W | Y | A | I | W |
| 4-4-20 (4fab) | R | Y | Q | P | V | F | S | W | F | N | V | Q | L | W | Y | T | M | W |
| C3 (1fpt) | H | Y | Q | P | L | F | S | Y | F | Q | I | Q | L | W | F | A | F | W |
| NC6.8 (1cgs) | H | Y | Q | P | L | F | S | Y | F | E | V | E | L | W | Y | T | M | W |
| L5MK16 diabody (11mk) | H | Y | K | P | L | F | S | F | F | E | V | Q | L | W | Y | A | L | W |
PBD, Protein Data Bank.
Rearrangement of the VH CDR3 Loop.
The VH CDR3 loop, residues 99H–109H, of the triabody has its tip twisted with respect to its conformation in Fv B1-8 (Fig. 5). The twist of the loop is accomplished by a flip of the main-chain dihedral angles for residues half-way between the base and the tip, GlyH103, SerH104, and SerH105 [sequential triabody numbering (Fig. 1)] (Table 2). The conformational change in H3 results in main-chain deviations of up to 5.6 Å (Cα of GlyH103) and a maximal side-chain deviation of 13.9 Å (OH of TyrH106). The magnitude of this rearrangement is similar to the largest of the conformational changes induced by antigen binding (33).
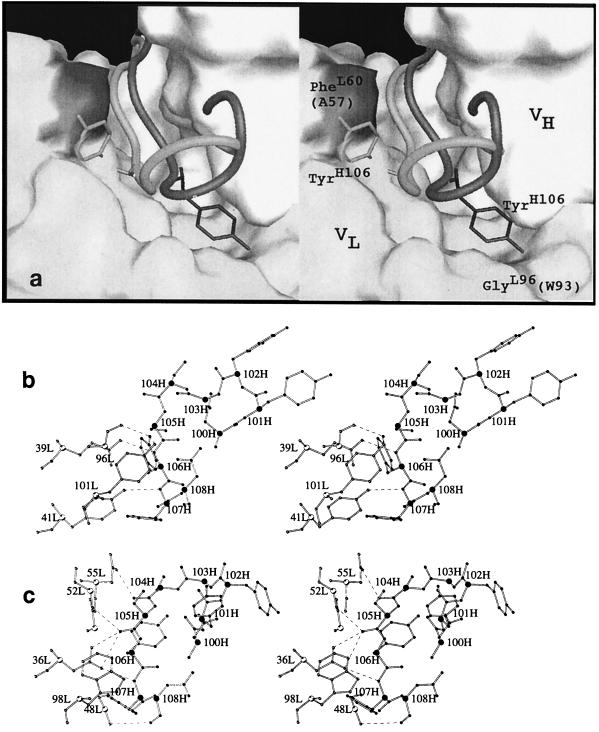
Figure 5.
Stereoview comparisons of the H3 loops in B1-8 and B1-8/NQ11. (a) A worm representation of the H3 loop in B1-8 (shaded lighter) superimposed on the H3 loop of B1-8/NQ11 (shaded darker). The molecular surface shown is that of the B1-8/NQ11 model with the H3 loop omitted. The surface of the VH domain is white and the general surface of the VL domain is light gray. The surface of PheL60 is shaded dark. The analogous residue in B1-8 is AlaL57 that leaves space to accommodate the side chain of TyrH106 (shown in stick representation) in the B1-8 Fv. The presence of GlyL96 (shaded) leaves space to accommodate the side chain of TyrH106 in B1-8/NQ11. The analogous volume in B1-8 is occupied by TrpL93. (b) The H3 loop of B1-8/NQ11 and its interaction with Vκ residues. The Cα positions are indicated with larger spheres. VL atoms are shown as white spheres and VH atoms as black spheres. (c) The B1-8 H3 loop and its interactions with the Vλ domain.
Table 2.
A comparison of the main-chain torsion angles for the H3 loop of B1-8/NQ11 and B1-8 antibodies
| Residue | B1-8/NQ11
|
B1-8
|
Difference
|
|||
|---|---|---|---|---|---|---|
| φ | ψ | φ | ψ | Δφ | Δψ | |
| Tyr-101 | −55 | −35 | −65 | −46.8 | 10 | 12 |
| Tyr-102 | −110 | 11 | −85 | −7.1 | −24 | 18 |
| Gly-103 | 68 | 23 | −111 | −108.7 | 179 | 132 |
| Ser-104 | 23 | −31 | −102 | 12.0 | 125 | −43 |
| Ser-105 | −138 | 160 | 57 | 44.2 | 165 | 116 |
| Tyr-106 | −88 | −59 | −122 | 158.7 | 34 | 143 |
| Phe-107 | −116 | 83 | −91 | 76.3 | −25 | 7 |
| Asp-108 | −75 | −37 | −70 | −55.0 | −5 | 18 |
A notable consequence of the conformational change of H3 is that the side chain of TyrH106 is in a radically different position than in B1-8. The change in the position of TyrH106 appears to be dictated by the space available for it against the surface of the VL domain (Fig. 5a). In the B1-8 structure, both in the presence and absence of antigen, TyrH106 is packed against the VL domain filling the space adjacent to AlaL57 [B1-8 numbering (Fig. 1)]. In the B1-8/NQ11 structure, the analogous space is occupied by PheL60 and by the side chain of TyrL101 from the C terminus of L3 thereby occluding TyrH106 from this site. Conversely, the cavity adjacent to GlyL96 is filled by the side chain of TyrH106 in B1-8/NQ11 whereas in B1-8, the analogous volume is occupied by the side chain of TrpL93. The structures suggest that the H3 loop is able to take on very different conformations in response to the surface provided by the VL domain and that the interactions of TyrH106 may be of central importance in determining which conformation the loop adopts. The importance of this residue in determining the conformation of the H3 loop was proposed from an analysis of the H3 loop conformations observed in known antibody structures (1). The change in conformation that we observe in exchanging a Vκ for a Vλ domain is likely the consequence of general features of the κ and λ light chains. A tryptophane residue analogous to TrpL93 is a feature of all of the mouse Vλ germ-line sequences. It is fairly common for Vκ sequences to have a small residue at the analogous position, and it is a pocket formed by a glycine at this position in the NQ11 Vκ domain that has allowed the side chain of TyrH106 to change its position so dramatically.
The conformation of the VH CDR3 in B1–8/NQ11 is stabilized by hydrogen bonds with residues in the framework and L3 of the VL domain: the OH of TyrH106 interacts with the side chain of TyrL101 and the carbonyl oxygen of GlyL96, and the amide N of PheH107 interacts with the OH of TyrL41. These interactions are not present in B1-8. Instead, in the B1-8 structure, VH CDR3 is stabilized by hydrogen bonds with the framework and hypervariable loops L1 and L2 (Fig. 5). The canonical conformation of L1 unique to mouse Vλ domains (34) makes it likely that H3–L1 interactions will be altered upon replacing a Vκ domain with a Vλ domain.
Combinatorial VH–VL pairing constitutes a fundamental source of antibody diversity in the immune response. However, the B1-8/NQ11 structure provides evidence that the expressed diversity is not simply related to the genetic diversity. The exchange of one variable domain for another through the process known as “receptor editing” may be an important feature of development of the Ig repertoire (35, 36). Thus a novel VH–VL pairing may “remodel” the binding site through conformational changes of VH CDR3. This may abolish antigen binding, but it may also allow the immune response to escape from a dead-end antigen-binding site that cannot be improved beyond a certain limit by somatic mutation, providing a fresh start with an entirely reshaped antigen-binding site. Plasticity of the H3 loop may be an important source of diversity of antigen-binding sites and may be an important consideration for antibody design and structure prediction.
Acknowledgments
We thank O. Perisic for help with experiments and for comments on the manuscript; T. Simon and K. Henrick for providing the coordinates of B1-8; C. Chothia, A. Lesk, and G. Winter for helpful discussions; and I. Fearnley for mass spectroscopy. Daresbury synchrotron radiation source and the staff of Station 9.6 are gratefully acknowledged for their assistance. The work was supported by the MRC/DTI/ZENECA/LINK program.
ABBREVIATION
- CDR
complementarity-determining region
Footnotes
Data deposition: The atomic coordinates and structure factors have been deposited in the Protein Data Bank, Biology Department, Brookhaven National Laboratory, Upton, NY 11973 (reference 1nqb).
References
- 1.Chothia C, Lesk A M. J Mol Biol. 1987;196:901–917. doi: 10.1016/0022-2836(87)90412-8. [DOI] [PubMed] [Google Scholar]
- 2.Chothia C, Lesk A M, Tramontano A, Levitt M, Smith-Gill S J, Air G, Sheriff S, Padlan E A, Davies D, Tulip W R, Colman P M, Spinelli S, Alzari P M, Poljak R J. Nature (London) 1989;342:877–883. doi: 10.1038/342877a0. [DOI] [PubMed] [Google Scholar]
- 3.Wilson I A, Stanfield R L. Curr Opin Struct Biol. 1993;3:113–118. [Google Scholar]
- 4.Wilson I A, Stanfield R L. Curr Opin Struct Biol. 1994;4:857–867. doi: 10.1016/0959-440x(94)90267-4. [DOI] [PubMed] [Google Scholar]
- 5.Sheriff S, Chang C Y, Jeffrey P D, Bajorath J. J Mol Biol. 1996;259:938–946. doi: 10.1006/jmbi.1996.0371. [DOI] [PubMed] [Google Scholar]
- 6.Rini J M, Schulze-Gahmen U, Wilson I A. Science. 1992;255:959–965. doi: 10.1126/science.1546293. [DOI] [PubMed] [Google Scholar]
- 7.Braden B C, Poljak R J. FASEB J. 1995;9:9–16. doi: 10.1096/fasebj.9.1.7821765. [DOI] [PubMed] [Google Scholar]
- 8.Braden B C, Fields B A, Ysern X, Goldbaum F A, Dall’Acqua W, Schwarz F P, Poljak R J, Mariuzza R A. J Mol Biol. 1996;257:889–894. doi: 10.1006/jmbi.1996.0209. [DOI] [PubMed] [Google Scholar]
- 9.Foote J, Milstein C. Nature (London) 1991;352:530–532. doi: 10.1038/352530a0. [DOI] [PubMed] [Google Scholar]
- 10.Foote J, Milstein C. Proc Natl Acad Sci USA. 1994;91:10370–10374. doi: 10.1073/pnas.91.22.10370. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Clackson T, Hoogenboom H R, Griffiths A D, Winter G. Nature (London) 1991;352:624–628. doi: 10.1038/352624a0. [DOI] [PubMed] [Google Scholar]
- 12.Figini M, Marks J D, Winter G, Griffiths A D. J Mol Biol. 1994;239:68–78. doi: 10.1006/jmbi.1994.1351. [DOI] [PubMed] [Google Scholar]
- 13.Jespers L S, Roberts A, Mahler S M, Winter G, Hoogenboom H R. Bio/Technology. 1994;12:899–903. doi: 10.1038/nbt0994-899. [DOI] [PubMed] [Google Scholar]
- 14.Steipe B, Plückthun A, Huber R. J Mol Biol. 1992;225:739–753. doi: 10.1016/0022-2836(92)90398-4. [DOI] [PubMed] [Google Scholar]
- 15.Perisic O, Webb P A, Holliger P, Winter G, Williams R L. Structure. 1994;2:1217–1226. doi: 10.1016/s0969-2126(94)00123-5. [DOI] [PubMed] [Google Scholar]
- 16.Griffiths A D, Malmqvist M, Marks J D, Bye J M, Embleton M J, McCafferty J, Baier M, Holliger K P, Gorick B D, Hughes-Jones N C, Hoogenboom H R, Winter G. EMBO J. 1993;12:725–734. doi: 10.1002/j.1460-2075.1993.tb05706.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Holliger P, Prospero T, Winter G. Proc Natl Acad Sci USA. 1993;90:6444–6448. doi: 10.1073/pnas.90.14.6444. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 18.Nissim A, Hoogenboom H R, Tomlinson I M, Flynn G, Midgley C, Lane D, Winter G. EMBO J. 1994;13:692–698. doi: 10.1002/j.1460-2075.1994.tb06308.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Whitlow M, Filpula D, Rollence M L, Feng S-L, Wood J W. Protein Eng. 1994;7:1017–1026. doi: 10.1093/protein/7.8.1017. [DOI] [PubMed] [Google Scholar]
- 20.Leslie A G W. Joint CCP4 and ESF-EACMB Newsletter on Protein Crystallography. Vol. 26. Warrington, U.K.: Daresbury Lab.; 1992. [Google Scholar]
- 21.CCP4. Acta Crystallogr D. 1994;50:760–763. doi: 10.1107/S0907444994003112. [DOI] [PubMed] [Google Scholar]
- 22.Navaza J. Acta Crystallogr A. 1994;50:157–163. [Google Scholar]
- 23.Lamzin V S, Wilson K S. Acta Crystallogr D. 1993;49:129–147. doi: 10.1107/S0907444992008886. [DOI] [PubMed] [Google Scholar]
- 24.Jones T A, Zou J-Y, Cowan S W, Kjeldgaard M. Acta Crystallogr A. 1991;47:110–119. doi: 10.1107/s0108767390010224. [DOI] [PubMed] [Google Scholar]
- 25.Brünger, A. (1992) x-plor Manual (Yale Univ., New Haven, CT).
- 26.Laskowski R A, MacArthur M W, Moss D S, Thornton J M. J Appl Crystallogr. 1993;26:283–291. [Google Scholar]
- 27.Bennett M J, Schlubegger M P, Eisenberg D. Protein Sci. 1995;4:2455–2468. doi: 10.1002/pro.5560041202. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Jung G, Freiman U, Marschall Z V, Reisfeld R A, Wilmanns W. Eur J Immunol. 1991;21:2431–2435. doi: 10.1002/eji.1830211020. [DOI] [PubMed] [Google Scholar]
- 29.Tutt A, Stevenson G T, Glennie M J. J Immunol. 1991;147:60–69. [PubMed] [Google Scholar]
- 30.Zhu Z, Presta L G, Zapata G, Carter P. Protein Sci. 1997;6:781–788. doi: 10.1002/pro.5560060404. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Mizutani R, Miura K, Nakayama T, Shimada I, Arata Y, Satow Y. J Mol Biol. 1995;254:208–222. doi: 10.1006/jmbi.1995.0612. [DOI] [PubMed] [Google Scholar]
- 32.Stanfield R L, Takimoto-Kamimura M, Rini J M, Profy A T, Wilson I A. Structure. 1993;1:83–93. doi: 10.1016/0969-2126(93)90024-b. [DOI] [PubMed] [Google Scholar]
- 33.Stanfield R L, Wilson I A. Trends Biotechnol. 1994;12:275–279. doi: 10.1016/0167-7799(94)90139-2. [DOI] [PubMed] [Google Scholar]
- 34.Wu S, Cygler M. J Mol Biol. 1993;229:597–601. doi: 10.1006/jmbi.1993.1065. [DOI] [PubMed] [Google Scholar]
- 35.Tiegs S L, Russell D M, Nemazee D. J Exp Med. 1993;177:1009–1020. doi: 10.1084/jem.177.4.1009. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Chen C, Nagy Z, Prak E L, Wiegert M. Immunity. 1995;3:747–755. doi: 10.1016/1074-7613(95)90064-0. [DOI] [PubMed] [Google Scholar]
- 37.Bothwell A L, Paskind M, Reth M, Imanishi-Kari T, Rajewski K, Baltimore D. Cell. 1981;24:625–637. doi: 10.1016/0092-8674(81)90089-1. [DOI] [PubMed] [Google Scholar]
- 38.Kabat E A, Wu T T, Perry H M, Gottesman K S, Foeller C. Sequences of Proteins of Immunological Interest. Bethesda, MD: U.S. Department of Health and Human Services; 1991. [Google Scholar]
- 39.Kraulis P J. J Appl Crystallogr. 1991;24:946–950. [Google Scholar]
- 40.Chothia C, Novotny J, Bruccoleri R, Karplus M. J Mol Biol. 1985;186:651–663. doi: 10.1016/0022-2836(85)90137-8. [DOI] [PubMed] [Google Scholar]