Abstract
Viral integrase catalyzes the integration of the linear viral DNA genome into the chromatin of the infected host cell, an essential step in the life cycle of retroviruses. The reaction produces a characteristic small duplication of host sequences at the site of integration, implying that there is a close juxtaposition of the viral DNA ends during a concerted integration event. We have used an in vitro assay to measure the concerted integration of virus-like plasmid DNA into naked lambda DNA catalyzed by virion purified avian integrase. In contrast to in vivo avian integration, which has strong fidelity for a 6-bp duplication, purified avian integrase in the context of this assay produced a distribution of duplication sizes, with the 6-bp size dominating. The metal cofactor Mg2+ induced increased fidelity for the 6-bp duplication relative to that with Mn2+. The immediate sequence of the host site may also influence duplication size in that we found sites that sustained multiple independent integration events producing the same duplication size. Additionally, for each set of cloned integration sites (5, 6, and 7 bp), a unique but similar symmetrical pattern of G/C and A/T sequence biases was found. Using duplex oligonucleotides as target substrates, we tested the significance of the 6-bp G/C and A/T pattern for site selection. In the context of this assay, which is likely dominated by the integration of only one viral end, the 6-bp pattern was not preferred. Instead, integration was predominantly into the 3' ends of the oligonucleotides. The combined results of the lambda and oligonucleotide assays indicated that although host site selection has properties in common with recognition of the viral DNA termini, the nonrandom sequence preferences seen for host site selection were not identical to the sequence requirements for long terminal repeat recognition.
Full text
PDF


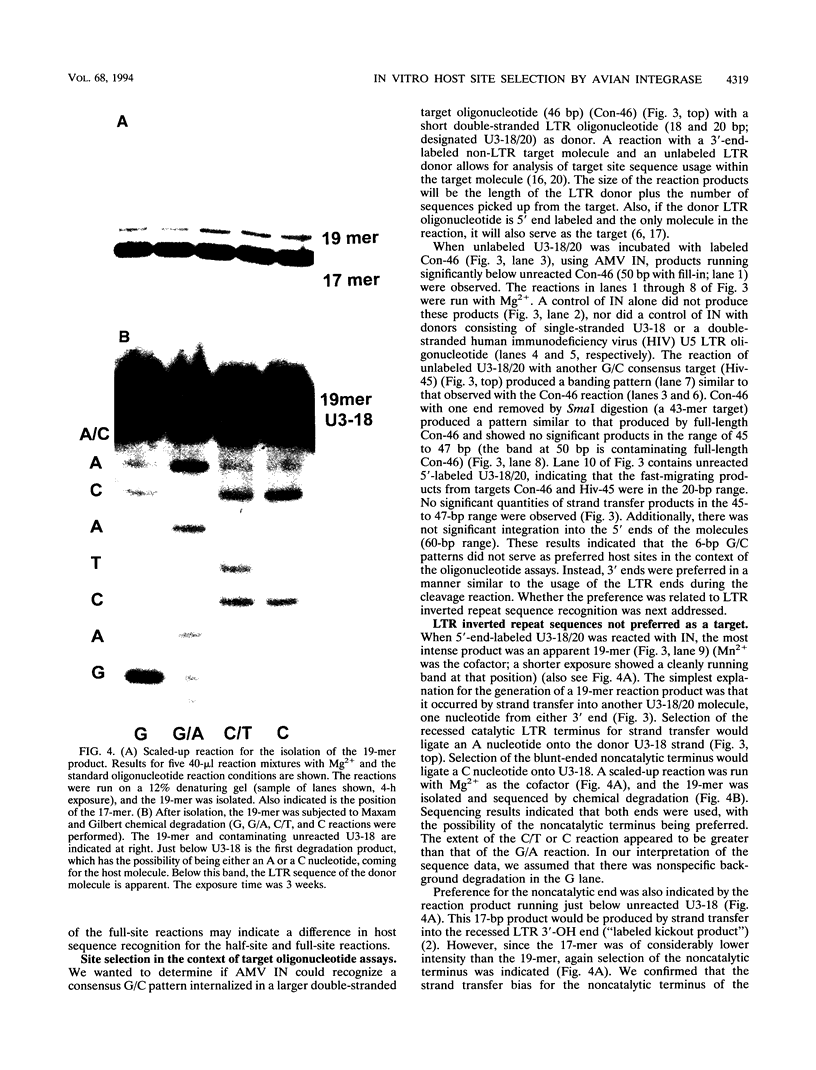


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Brown P. O., Bowerman B., Varmus H. E., Bishop J. M. Correct integration of retroviral DNA in vitro. Cell. 1987 May 8;49(3):347–356. doi: 10.1016/0092-8674(87)90287-x. [DOI] [PubMed] [Google Scholar]
- Bushman F. D., Craigie R. Activities of human immunodeficiency virus (HIV) integration protein in vitro: specific cleavage and integration of HIV DNA. Proc Natl Acad Sci U S A. 1991 Feb 15;88(4):1339–1343. doi: 10.1073/pnas.88.4.1339. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bushman F. D., Craigie R. Integration of human immunodeficiency virus DNA: adduct interference analysis of required DNA sites. Proc Natl Acad Sci U S A. 1992 Apr 15;89(8):3458–3462. doi: 10.1073/pnas.89.8.3458. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bushman F. D., Fujiwara T., Craigie R. Retroviral DNA integration directed by HIV integration protein in vitro. Science. 1990 Sep 28;249(4976):1555–1558. doi: 10.1126/science.2171144. [DOI] [PubMed] [Google Scholar]
- Colicelli J., Goff S. P. Sequence and spacing requirements of a retrovirus integration site. J Mol Biol. 1988 Jan 5;199(1):47–59. doi: 10.1016/0022-2836(88)90378-6. [DOI] [PubMed] [Google Scholar]
- Craigie R., Fujiwara T., Bushman F. The IN protein of Moloney murine leukemia virus processes the viral DNA ends and accomplishes their integration in vitro. Cell. 1990 Aug 24;62(4):829–837. doi: 10.1016/0092-8674(90)90126-y. [DOI] [PubMed] [Google Scholar]
- Engelman A., Bushman F. D., Craigie R. Identification of discrete functional domains of HIV-1 integrase and their organization within an active multimeric complex. EMBO J. 1993 Aug;12(8):3269–3275. doi: 10.1002/j.1460-2075.1993.tb05996.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engelman A., Mizuuchi K., Craigie R. HIV-1 DNA integration: mechanism of viral DNA cleavage and DNA strand transfer. Cell. 1991 Dec 20;67(6):1211–1221. doi: 10.1016/0092-8674(91)90297-c. [DOI] [PubMed] [Google Scholar]
- Fitzgerald M. L., Vora A. C., Grandgenett D. P. Development of an acid-soluble assay for measuring retrovirus integrase 3'-OH terminal nuclease activity. Anal Biochem. 1991 Jul;196(1):19–23. doi: 10.1016/0003-2697(91)90111-6. [DOI] [PubMed] [Google Scholar]
- Fitzgerald M. L., Vora A. C., Zeh W. G., Grandgenett D. P. Concerted integration of viral DNA termini by purified avian myeloblastosis virus integrase. J Virol. 1992 Nov;66(11):6257–6263. doi: 10.1128/jvi.66.11.6257-6263.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Craigie R. Integration of mini-retroviral DNA: a cell-free reaction for biochemical analysis of retroviral integration. Proc Natl Acad Sci U S A. 1989 May;86(9):3065–3069. doi: 10.1073/pnas.86.9.3065. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandgenett D. P., Inman R. B., Vora A. C., Fitzgerald M. L. Comparison of DNA binding and integration half-site selection by avian myeloblastosis virus integrase. J Virol. 1993 May;67(5):2628–2636. doi: 10.1128/jvi.67.5.2628-2636.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hong T., Murphy E., Groarke J., Drlica K. Human immunodeficiency virus type 1 DNA integration: fine structure target analysis using synthetic oligonucleotides. J Virol. 1993 Feb;67(2):1127–1131. doi: 10.1128/jvi.67.2.1127-1131.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Katz R. A., Merkel G., Kulkosky J., Leis J., Skalka A. M. The avian retroviral IN protein is both necessary and sufficient for integrative recombination in vitro. Cell. 1990 Oct 5;63(1):87–95. doi: 10.1016/0092-8674(90)90290-u. [DOI] [PubMed] [Google Scholar]
- Khan E., Mack J. P., Katz R. A., Kulkosky J., Skalka A. M. Retroviral integrase domains: DNA binding and the recognition of LTR sequences. Nucleic Acids Res. 1991 Feb 25;19(4):851–860. doi: 10.1093/nar/19.4.851. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kitamura Y., Lee Y. M., Coffin J. M. Nonrandom integration of retroviral DNA in vitro: effect of CpG methylation. Proc Natl Acad Sci U S A. 1992 Jun 15;89(12):5532–5536. doi: 10.1073/pnas.89.12.5532. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leavitt A. D., Rose R. B., Varmus H. E. Both substrate and target oligonucleotide sequences affect in vitro integration mediated by human immunodeficiency virus type 1 integrase protein produced in Saccharomyces cerevisiae. J Virol. 1992 Apr;66(4):2359–2368. doi: 10.1128/jvi.66.4.2359-2368.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mizuuchi K. Polynucleotidyl transfer reactions in transpositional DNA recombination. J Biol Chem. 1992 Oct 25;267(30):21273–21276. [PubMed] [Google Scholar]
- Pryciak P. M., Sil A., Varmus H. E. Retroviral integration into minichromosomes in vitro. EMBO J. 1992 Jan;11(1):291–303. doi: 10.1002/j.1460-2075.1992.tb05052.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sandmeyer S. B., Hansen L. J., Chalker D. L. Integration specificity of retrotransposons and retroviruses. Annu Rev Genet. 1990;24:491–518. doi: 10.1146/annurev.ge.24.120190.002423. [DOI] [PubMed] [Google Scholar]
- Vincent K. A., Ellison V., Chow S. A., Brown P. O. Characterization of human immunodeficiency virus type 1 integrase expressed in Escherichia coli and analysis of variants with amino-terminal mutations. J Virol. 1993 Jan;67(1):425–437. doi: 10.1128/jvi.67.1.425-437.1993. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vora A. C., Fitzgerald M. L., Grandgenett D. P. Removal of 3'-OH-terminal nucleotides from blunt-ended long terminal repeat termini by the avian retrovirus integration protein. J Virol. 1990 Nov;64(11):5656–5659. doi: 10.1128/jvi.64.11.5656-5659.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Gent D. C., Oude Groeneger A. A., Plasterk R. H. Identification of amino acids in HIV-2 integrase involved in site-specific hydrolysis and alcoholysis of viral DNA termini. Nucleic Acids Res. 1993 Jul 25;21(15):3373–3377. doi: 10.1093/nar/21.15.3373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- van Gent D. C., Vink C., Groeneger A. A., Plasterk R. H. Complementation between HIV integrase proteins mutated in different domains. EMBO J. 1993 Aug;12(8):3261–3267. doi: 10.1002/j.1460-2075.1993.tb05995.x. [DOI] [PMC free article] [PubMed] [Google Scholar]