Abstract
The gene for the transducer of transforming growth factor-beta/bone morphogenetic protein signalling SMAD4, a potential suppressor of colorectal carcinogenesis, is located at the chromosomal region 18q21. In order to evaluate the clinical relevance of SMAD4 deletion, gene copy alterations were determined by copy dosage using real-time quantitative PCR in 202 colorectal tumour biopsies from a previous randomised study of adjuvant chemotherapy. Patients with normal SMAD4 diploidy turned out to have a three-fold higher benefit of 5-fluorouracil-based adjuvant chemotherapy with a border line significance (overall survival: 3.23, P=0.056; disease-free survival: 2.89, P=0.045). These data are consistent with the previous observation that patients whose cancer had retention of the 18q21 region had a significantly higher benefit from 5-fluorouracil-based therapy. Moreover, these results may provide a refinement at the gene level of the clinical relevance of 18q21 deletion, thereby suggesting SMAD4 as a predictive marker in colorectal cancer. This data also indicate that integrity of this component of the transforming growth factor-beta/bone morphogenetic protein signalling pathway may be a critical factor for benefit of chemotherapy in patients with colorectal cancer.
British Journal of Cancer (2002) 21, 630–634. doi:10.1038/sj.bjc.6600511 www.bjcancer.com
© 2002 Cancer Research UK
Keywords: colorectal cancer, TGFβ, signalling, SMAD, predictive marker
Deletion of the chromosomal region 18q21 is the most frequent cytogenetic alteration observed in colorectal cancer (CRC), suggesting the location of a tumour suppressor locus in this region (Vogelstein et al, 1988; Mitelman et al, 1997). Searches for such candidate genes have led to the identification of a gene designated as Deleted in Pancreatic Cancer locus 4 (DPC4) (Hahn et al, 1996; Thiagalingam et al, 1996). Through its homology with the C. elegans Small (Sma) proteins (Savage et al, 1996) and with the Drosophila protein Mothers against dpp (Mad), initially identified for its genetic interaction with the gene for the BMP-like peptide Decapentaplegic (dpp) (Raftery et al, 1995), DPC4 has been renamed SMAD4 as a merger of Sma and Mad (Derinck et al, 1996).
SMADs form a family of structurally related proteins initially identified for their role in embryonic development of Drosophila (Raftery et al, 1995) and of C. elegans (Savage et al, 1996). Proteins of the SMAD family can be divided into three distinct subtypes that correlate with their respective functions in transforming growth factor beta (TGFβ)/bone morphognetic protein (BMP) signalling (Kretschmar and Massagué, 1999; Newfeld et al, 1999), as depicted in Figure 1: (i) receptor-activated (ra)-SMADs are serine-phosporylated upon binding of the cytokine to its cognate receptor. SMAD2 and SMAD3 are specifically activated by TGFβ-like cytokines, whereas SMAD1, SMAD5 and SMAD8 are exclusively phosphorylated by BMPs; (ii) a co-SMAD, SMAD4 heteropolymerises with activated ra-SMADs. This complex migrates to the nucleus where it associates with tissue specific transcription factors. SMAD4 is the only co-SMAD protein known in mammalians, and therefore is a common signalling mediator to all TGFβ/BMPs; and (iii) among the immediate target genes for SMAD transcription complexes are the genes for anti-SMADs. Thus, the anti-SMADs SMAD6 and SMAD7 prevent activation of ra-SMADs (SMAD1/5/8 and SMAD2/3, respectively), therefore providing a transient cytokine response through a negative feedback loop. Interestingly, SMAD2 (Eppert et al, 1996) and SMAD7 (Nakao et al, 1997) genes have also been assigned to the 18q21 region (Eppert et al, 1996; Röijer et al, 1998), where the SMAD7 gene maps between SMAD2 and SMAD4 genes (Boulay et al, 2001) within four megabases (Venter et al, 2001). Thus, this region encodes the three classes of TGFβ mediators specifically required for the signalling of TGFβ-like cytokines, and one, SMAD4, for both TGFβ and BMP families.
Figure 1.
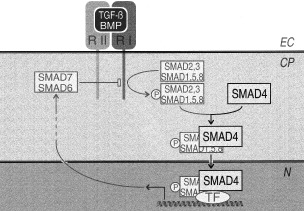
SMADS in the TGFβ/BMP signalling pathway: (i) receptor-activated (ra)-SMADs SMAD2 or SMAD3 are serine-phosporylated upon TGFβ-receptor interaction, whereas SMAD1, SMAD5 or SMAD8 phosphorylation is exclusively induced by BMPs; (ii) SMAD4, the common and unique co-SMAD signalling mediator to all TGFβ/BMP cytokines, heteropolymerises with activated ra-SMADs and migrates to the nucleus where it associates with tissue specific transcription factors (TF); (iii) anti-SMADs produced upon cytokine induction (SMAD7 for TGFβ and SMAD6 for BMPs) block ra-SMAD serine phosphorylation. This inducible negative feedback loop provides a transient response to cytokine activation (see Kretschmar and Massagué, 1999). Extracytoplasmic (EC), cytoplasmic (CP) and nuclear (N) compartments are indicated.
Genetic evidence for the involvement of TGFβ pathway in colon tumour suppression was given by Markowitz et al (1995), who observed frequent frameshift mutations within the TGFβ-receptor II coding sequence in CRC, as a result of microsatellite instability. This observation has been later confirmed in a larger population, where most tumors with microsatellite instability carry this gene mutation (Watanabe et al, 2001). On the other hand, TGFβ has been shown to be a potent cell growth inhibitor (Roberts and Sporn, 1993), and apoptosis inducer on prostatic epithelial cell lines (Hsing et al, 1996), whereas most squamous carcinoma lines are refractory to this function (Reiss et al, 1993; Blobe et al, 2000). Thus, the frequent deletion of the chromosomal region 18q21 in colorectal tumours together with the physiologic functions of TGFβ strongly suggested a role for SMAD4 in the suppression of colorectal carcinogenesis.
For these reasons, we wished to study the influence of the SMAD4 gene on the clinical outcome of patients with CRC, including on benefit of 5FU-based chemotherapy. Indeed, an interaction between markers and treatment responsiveness or lack thereof has led to a separation of these factors into prognostic (independent of treatment) and predictive (interactive with treatment) categories. To do so, we took advantage of archived colorectal tumour biopsies collected in a previous Swiss Association for Clinical Cancer Research (SAKK) study of 5FU-based perioperative adjuvant therapy (SAKK, 1995). Through a strategy based on quantitative real time PCR (Boulay et al, 1999, 2001), we performed genetic analyses of corresponding DNAs by copy dosage of the SMAD4 gene. In order to study on one hand the prognostic value of genotype, and on the other hand its predictive effect on the efficacy of 5FU-based therapy among patients with CRC, we undertook multivariate statistical analysis of SMAD4 gene copy status on survival.
MATERIALS AND METHODS
Patients
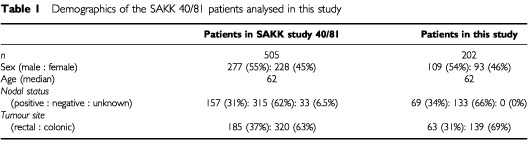
Patients from whom biopsies were isolated, were part of a previous randomised study of the Swiss Association for Clinical Cancer Research (SAKK) on benefit of adjuvant chemotherapy (SAKK study 40/81) (SAKK, 1995). In that study, 533 patients with colorectal cancer about to undergo curative resection were randomly assigned no adjuvant treatment (control group) or an immediate postoperative infusion with 5FU (500 mg m−2) for 7 days, with one single dose of mitomycin (10 mg m−2) on day 1. As a result, patients appeared to significantly benefit from this therapy such that overall survival increased from 55 to 66 months (hazard ratio: 0.74; 95% confidence interval: 0.57–0.97; P=0.026), and disease-free survival, from 48 to 57 months (hazard ratio: 0.79; 95% confidence interval: 0.62–1.00; P=0.051). The relationship between genotypes and clinical outcome was assessed in a subset of 202 patients with genetic data for which we also had clinical and survival data. As shown in Table 1, the subgroup for which genetic and clinical data are available, is closely representative of the patients treated in the SAKK study 40/81 (SAKK, 1995). Our study comprises 164 out of the 233 individuals described in our previous report (Boulay et al, 2001).
Table 1. Demographics of the SAKK 40/81 patients analysed in this study.
Gene copy status scoring
Genomic samples were tested for gene dosage using the TaqMan® system on an ABI Prism® 7700 sequence detector (PE Applied Biosystems, Foster, USA). All reactions were made in triplicate. For each individual, the Ct value (calculated by the built-in software) obtained for the gene 36B4 (Masiakowski et al, 1982) on normal tissue was subtracted from that of tumour tissue thus defining ΔCt36B4. A similar calculation was made for the SMAD4 gene (ΔCtSMAD4). Gene copy status is indicated by the ΔCt value (ΔCt36B4–ΔCtSMAD4) as following. ΔCt>−0.45: no deletion; ΔCt<−0.55: hemizygous. Primers: SMAD4, gca gac aga aac tgg att aaa aca att and gaa tgt gtt tct cct aat ctt caa gct; 36B4: agc aag tgg gaa ggt gta atc c and cca ttc tat cat caa cgg gta caa. Probes: SMAD4, tgt tgt ggt ccc tat ggc tgt tta cta tcc a; 36B4: tct cca cag aca agg cca gga ctc g.
Statistical analysis
Cox proportional hazard modelling was undertaken to assess the impact of genotype on overall survival and on disease-free survival after controlling for possible confounding. All analyses were performed using S Plus. The relationship between genotypes and survival/disease-free survival was assessed in a subset of 202 patients with genetic data for whom we also had clinical and survival data.
RESULTS
We aimed to test whether the deletion of the SMAD4 gene would have a significant influence on the outcome of patients with CRC. Within the individual tumours in this statistical analysis, the frequency of SMAD4 gene deletion was 67% (135 out of 202). This deletion frequency was similar to that previously reported (Boulay et al, 2001). For each individual, the associations between gene copy dosage and clinical data was investigated in multivariate statistical analyses that included age, sex, stage, tumour location, grade, nodal status and chemotherapy as covariates. Hazard ratios (HR) for death and relapse associated with SMAD4 gene deletion were close to one (1.22 and 1.16, respectively) with non-significant P values (0.43 and 0.56, respectively, Table 2, top). Thus we concluded that deletion of SMAD4 gene has no significant influence on the outcome of patients with CRC.
Table 2. Association of SMAD4 status with patient outcome.

The original clinical study from which tumour samples were derived had shown the benefit of 5FU-based perioperative adjuvant chemotherapy in colorectal cancer (SAKK, 1995). Thus, a similar multivariate statistical analysis was performed to evaluate the deletion of the SMAD4 gene as a potential marker for a predictive effect on 5FU treatment. Regarding disease-free survival, after controlling for confounding in the multivariate models, the HR associated with 5FU treatment among patients with normal diploidy for SMAD4 was 0.32, whereas the HR associated with 5FU treatment in patients with SMAD4 deletion was 2.89 times as large (Table 2, bottom). The difference between these two HRs (i.e. the statistical interaction between gene deletion and the effect of 5FU chemotherapy) was of borderline statistical significance (P=0.045). A similar result was found for overall survival, the HR associated with 5FU treatment being 0.25 among patients with normal SMAD4 genotype and 3.23 times as great among patients with SMAD4 deletion (Table 2, bottom; Figure 2). As with disease-free survival, the difference was of borderline statistical significance (P=0.056). This suggests SMAD4 as a predictive marker for 5FU/mitomycin adjuvant chemotherapy
Figure 2.
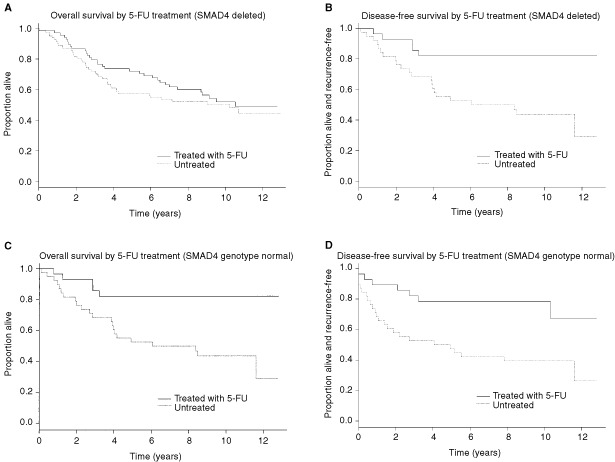
Kaplan–Meier plotting of survival in response to 5FU therapy in patients (n=202) with SMAD4 deletion (top) and with no loss of SMAD4 (bottom). Overall survival (left): HR=3.23, 95% CI=0.97–10.8, P=0.056. Disease-free survival (right): HR=2.89, 95% CI=1.02–8.12, P=0.045.
DISCUSSION
We established that among the patients with colorectal cancer involved in this study, SMAD4 was deleted in 67% of cases. These results, obtained by gene copy dosage, are consistent with those deduced from earlier cytogenetic (Vogelstein et al, 1988; Mitelman et al, 1997) and loss of hetereozygocity (LOH) studies on the 18q21 region (Laurent-Puig et al, 1992; Jen et al, 1994; Martínez-López et al, 1998; Jernwall et al, 1999; Watanabe et al, 2001). However, these LOH studies are frequently based on microsatellite markers that span several centiMorgans (cM), and therefore several megabases (Mb). In contrast to that approach, our strategy allows for a more refined analysis by targeting an individual gene rather than a wide chromosomal region that certainly contains a number of important genes. Thus, it is likely that an analysis at the single gene level will give a more accurate image of eventual clinical implications of genetic alterations.
We observed that colorectal cancer patients with normal SMAD4 gene copy status had a three-fold higher benefit of 5FU-based therapy than those with SMAD4 deletion. This result is consistent with the previous observation by Watanabe et al (2001), that patients with retention of 18q21 alleles had a benefit of 5FU-based chemotherapy of the same order as was found in this study. Moreover, refinement of deletion studies from the chromosomal band level to the gene level may provide a clue to possible mechanisms through which 18q21 deletion influences the outcome of patients with CRC. Therefore, our results reinforce the hypothesis that TGFβ and its signalling components have a role in tumour suppression. This result also suggests the definition of SMAD4 as a predictive marker for benefit of 5FU-based chemotherapy in patients with colorectal cancer. Finally, these findings suggests a mode of action of this cytostatic compound that is SMAD4-dependent. Thus, the integrity of this component of the TGFβ/BMP pathway is not only required for cytokine signalling, but may also be an important factor for 5FU-mediated apoptosis. In addition to the requirement of functional apoptotic pathways such as CD95/Fas (Houghton et al, 1997), bax (Rampino et al, 1997) and p53 (Vogelstein et al, 2000) for drug sensitivity in colorectal tumour cells, this suggests that integrity of the TGFβ pathway may be an additional condition for efficiency of 5FU treatment. Thus, our results provide an additional clue to the genetic basis of drug resistance in cancer.
Acknowledgments
Supported by the Swiss Cancer League, the Krebsliga beider Basel, the Sander Stiftung and by the Novartis (formerly Ciba-Geigy) Foundation.
References
- BlobeGCSchiemanWPLodishHF2000Role of the transformiong growth factor β in human disease N Engl J Med 34213501358 [DOI] [PubMed] [Google Scholar]
- BoulayJLReuterJRitschardRTerraccianoLHerrmannRRochlitzC1999Gene dosage by quantitative real-time PCR Biotechniques 27228232 [DOI] [PubMed] [Google Scholar]
- BoulayJLMildGReuterJLagrangeMTerraccianoLLowyALafferUOrthBMetzgerUStammBMartinoliSHerrmannRRochlitzC2001Combined copy status of 18q21 genes in colorectal cancer shows frequent retention of SMAD7 Genes Chrom Cancer 31240247 [DOI] [PubMed] [Google Scholar]
- DerinckRGelbartWMHarlandRMHeldinCHKernSEMassaguéJMeltonDAMlodzikMPadgettRWRobertsABSmithJThomsenGHVogelsteinBWangXF1996Nomenclature: vertebrate mediators of TGFβ family signals Cell 87173. [DOI] [PubMed] [Google Scholar]
- EppertKSchererSWOzcelikHPironeRHoodlessPKimHTsuiLCBapatBGallingerSAndrulisILThomsenGHWranaJLAttisanoL1996MADR2 maps to 18q21 and encodes a TGFbeta-regulated MAD-related protein that is functionally mutated in colorectal carcinoma Cell 86543552 [DOI] [PubMed] [Google Scholar]
- HahnSASchutteMShamsul HoqueATMMoskalukCAda CostaLTRozenblumEWeinsteinCLFischerAYeoCJHrubanRHKernSE1996DPC4, a candidate tumour suppressor gene at human chromsome 18q21.1 Science 271350353 [DOI] [PubMed] [Google Scholar]
- HoughtonJAHarwoodFGTillmanDM1997Thymineless death in colon carcinoma cells is mediated via Fas signalling Proc Natl Acad Sci USA 9481448149 [DOI] [PMC free article] [PubMed] [Google Scholar]
- HsingAYKadomatsuKBonhamMJDanielpourD1996Regulation of apoptosis induced transforming growth factor-beta1 in nontumorigenic and tumorigenic rat prostatic epithelial cell lines Cancer Res 5651465149 [PubMed] [Google Scholar]
- JenJKimHPiantadosiSLiuAFLevittRCSistonenPKinzlerKWVogelsteinBHamiltonSR1994Allelic loss of chromosome 18q and prognosis in colorectal cancer N Engl J Med 331213221 [DOI] [PubMed] [Google Scholar]
- JernwallPMäkinenMJKarttunenTJMäkeläJVihkoP1999Loss of Heterozygocity at 18q21 is indicative of recurrence and therefore poor prognosis in a subset of colorectal cancers Br J Cancer 79903908 [DOI] [PMC free article] [PubMed] [Google Scholar]
- KretschmarMMassaguéJ1999SMADs: mediators and regulators of TGF-beta signalling Curr Opin Genet Dev 8103111 [DOI] [PubMed] [Google Scholar]
- Laurent-PuigPOlschwangSDelattreORemvikosYAsselainBMelotTValidirePMulerisMGirodetJSalmonRJ1992Survival and acquired genetic alterations in colorectal cancer Gastroenterology 10211361141 [PubMed] [Google Scholar]
- MarkowitzSWangJMyeroffLParsonsRSunLLutterbaughJFanRSZborowskaEKinzlerKWVogelsteinBBrattainMWillsonJKV1995Inactivation of the type II TGF-beta receptor in colon cancer cells with microsatellite instability Science 26813361338 [DOI] [PubMed] [Google Scholar]
- Martínez-LópezEAbadAFontAMonzóMOjangurenIPifarréASánchezJJMartínCRosellR1998Allelic loss on chromosome 18q as a prognostic marker in stage II colorectal cancer Gastroenterology 11411801187 [DOI] [PubMed] [Google Scholar]
- MasiakowskiPBreathnachRBlochJGannonFKrustAChambonP1992Cloning of cDNA sequences of hormone-regulated genes from the MCF-7 human breast cancer cell line Nucleic Acids Res 1078957903 [DOI] [PMC free article] [PubMed] [Google Scholar]
- MitelmanFMertensFJohanssonB1997A breakpoint map of recurrent chromosomal rearrangements in human neoplasia Nature Genetics 15417474 [DOI] [PubMed] [Google Scholar]
- NakaoAAfrakhteMMorenANakayamaTChristianJLHeuchel RItohSKawabataMHeldinNEHeldinCHten DijkeP1997Identification of Smad7, a TGFbeta-inducible antagonist of TGF-beta signalling Nature 389631635 [DOI] [PubMed] [Google Scholar]
- NewfeldSJWisotzkeyRGKumarS1999Molecular evolution of a developmental pathway: phylogenetic analyses of transforming growth factor-β family ligands, receptors and smad signal transducers Genetics 152783795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- RafteryLATwomblyVWhartonKGelbartWM1995Genetic screens to identify elements of the decapentaplegic signalling pathway in Drosophila Genetics 139241254 [DOI] [PMC free article] [PubMed] [Google Scholar]
- RampinoNYamamotoHIonovYLiYSawaiHReedJCPeruchoM1997Somatic frameshift mutations in the BAX gene in colon cancers of the microsatellite mutator phenotype Science 275967969 [DOI] [PubMed] [Google Scholar]
- ReissMMuñoz-AntoniaTCowanJMWilkinsPCZhouZLVellucciVF1993Resistance of human squamous carcinoma cells to transforming growth factor beta1 is a recessive trait Proc Natl Acad Sci USA 9062806284 [DOI] [PMC free article] [PubMed] [Google Scholar]
- RobertsABSpornMB1993The transforming growth factor-betasIn:Handbook of experimental pharmacology: peptide growth factors and their receptorsSporn MB, Roberts AB (eds)Berlin-Heidelberg: Springer-Verlagpp419472 [Google Scholar]
- RöijerEMorénAten DijkePStenmanG1998Assignment of the SMAD-7 gene (MADH7) to human chromosome 18q21.1 by fluorescence in situ hybridization Cytogenet Cell Genet 81189190 [DOI] [PubMed] [Google Scholar]
- SavageCDasPFinelliALTownsendSRSunCYBairdSEPadgettRW1996Caenorhabditis elegans genes sma-2, sma-3 and sma-4 define a conserved family of transforming growth factor beta pathway components Proc Natl Acad Sci USA 93790794 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swiss Group for Clinical Cancer Research SAKK1995Long-term results of single course of adjuvant intraportal chemotherapy for colorectal cancer The Lancet 345349353 [PubMed] [Google Scholar]
- ThiagalingamSLengauerCLeachFSSchutteMHahnSAOverhauserJWillsonJKVMarkowitzSHamiltonSRKernSEKinzlerKWVogelsteinB1996Evaluation of candidate tumour suppressor genes on chromosome 18 in colorectal cancers Nature Genet 13343346 [DOI] [PubMed] [Google Scholar]
- VenterCet al2001The sequence of the human genome Science 29113041345 [DOI] [PubMed] [Google Scholar]
- VogelsteinBFearonERHamiltonSRKernSEPreisingerACLeppertMNakamuraYWhiteRSmitsAMBosJL1988Genetic alterations during colorectal-tumour development N Engl J Med 319525532 [DOI] [PubMed] [Google Scholar]
- VogelsteinBLaneDLevineAJ2000Surfing the p53 network Nature 408307310 [DOI] [PubMed] [Google Scholar]
- WatanabeTWuTTCatalanoPJUekiTSatrianoRHallerDGBensonIIIABHamiltonSR2001Molecular predictors of survival after adjuvant chemotherapy for colon cancer N Engl J Med 34411961206 [DOI] [PMC free article] [PubMed] [Google Scholar]


