Abstract
Xenopus Zic3 is a Xenopus homologue of mouse Zic and Drosophila pair-rule gene, odd-paired. We show here that Zic3 has significant roles both in neural and neural crest development in Xenopus embryo. Expression of Zic3 is first detected in prospective neural plate region at gastrulation. Onset of the expression was earlier than most proneural genes and followed chordin expression. The expression was induced by blockade of BMP4 signal. Overexpression of Zic3 resulted in hyperplastic neural and neural crest derived tissue. In animal cap explant, the overexpression of Zic3 induced expression of all the proneural genes and neural crest marker genes. These findings suggest that Zic3 can determine the ectodermal cell fate and promote the earliest step of neural and neural crest development.
Keywords: Zic, odd-paired, neural induction, proneural gene
During vertebrate neurulation, the original ectoderm gives rise to neural tube, neural crest, and epidermis. Several genes controlling this process have been described (1–3). Among them, recent advances on two issues, neural inducers and the “proneural genes,” have been remarkable.
In Xenopus laevis, the neuroectoderm is induced on the dorsal side by signals such as noggin, chordin, follistatin, and Xnr3 emanating from the Spemann organizer during early gastrulation (4–9). Although BMP4 has an inhibitory effect on ectoderm to differentiate into neuroectoderm (10), the blockade of the signal triggers the neuroectoderm formation (11–13). Organizer signals such as noggin, chordin, follistatin may act as endogenous blockages for BMP4 by direct binding (14–16).
On the other hand, several basic helix-loop-helix transcription factors are shown to participate in vertebrate neural development, as is the case in the Drosophila. Among them, Neurogenin, NeuroD, XASH-3, and XATH-3 are shown to have neurogenic activity, because their overexpression in Xenopus embryos results in enlargement of neural tissue or ectopic neural cell induction (17–22). Although a large number of studies along the above two lines have been made, the link between BMP4 signal and proneural genes is not understood well.
In our previous study (23–27), we investigated about Zic gene family in vertebrate development. The Zic family was originally found as a group of genes encoding zinc finger proteins that are expressed highly restrictedly in mammalian cerebellum (23–26). The Zic genes are the vertebrate homologues of a Drosophila pair-rule gene, odd-paired (opa), which has important roles in the parasegmental subdivision and the visceral mesoderm development of the Drosophila embryo (28, 29). As is the case in Drosophila opa, the expression patterns of mouse Zic genes suggest that Zic genes are involved in ontogenesis (27). In particular, we could detect the Zic expression in the neuroectoderm just before it was formed.
To clarify the role of Zic in the early stage of neural development, we have cloned the Xenopus Zic3. Zic3 expression began at prospective neural plate region as soon as neural induction occurred and was induced by blockade of BMP4 signal in Xenopus embryo. Overexpression of Zic3 caused neural hyperplasia involved with induction of all of the proneural genes reported so far. These data indicate that Zic3 acts in the earliest phase of neural development and that it belongs to a novel class of genes responsible for neural development. In addition, Zic3 overexpression induced neural crest tissue in embryo. These results suggest the essential roles of Zic3 in the induction of neuroectoderm and neural crest.
MATERIALS AND METHODS
Isolation of Xenopus Zic3 cDNA Clone.
Xenopus neurula (stage 17) cDNA was subjected to PCR by following primers. The 5′ primer is a 5′-GAGAACCTCAAGATCCACAA-3′, which derives from ENLKIHK (sequence based on zinc finger domain of the mouse Zic family genes), and the 3′ primer is 5′-TT(C/T)CCATG(A/G)ACCTTCATGTG-3′, which is the reverse translation of HMKVHEE. The fragment was used to screen a λ ZAP cDNA library prepared from Xenopus neurula embryos The isolated Applied Biosystems/PRISM Dye Primer Cycle Sequencing Ready Reaction Kit (Perkin–Elmer).
Embryo Manipulations.
Microinjection was carried out as previously described (30). The Zic3 ORF was cloned into the vector pCS2+ (20). Zic3 mRNA, LacZ mRNA, dominant-negative form of BMP receptor (dnBMPR) mRNA, and noggin mRNA were synthesized by in vitro transcription. Animal cap preparation and dissociation were described (10, 31).
RNA Isolation and Reverse Transcription–PCR (RT-PCR) Assay.
Preparation of total RNA and RT-PCR assay were carried out as described (32). The conditions of RT-PCR assays and primer sequences of Histone H4 and XlPOU 2 were as published (33, 34). Xenopus slug (Xslu) primer sequences were derived from P. A. Wilson (personal communication). Some primer sequences were obtained from The Xenopus Molecular Marker Resource on the internet (http://vize222.zo.utexas.edu/). Our original primer sequences were deposited at the same web site.
Histology and Whole-Mount in Situ Hybridization.
Paraffin sections of Zic3 mRNA injected embryos were prepared essentially as described by Kelly et al. (35). Whole-mount in situ hybridization was performed essentially as described (36) using digoxigenin-labeled antisense probes for Zic3, neural cell adhesion molecule (NCAM) (37), Xslu (38), and Xenopus twist (Xtwi) (39). Whole-mount immunohistochemistry was performed essentially as described using EpA supernatant (40).
RESULTS
Xenopus Zic3 Is Expressed in Prospective Neural Plate in Gastrula and Neural Folds in Neurula.
We cloned a Xenopus Zic-related cDNA (2.4 kb) (see Materials and Methods). This cDNA encodes a predicted protein of 441 amino acid residues. A homology search against the current database revealed that mouse Zic3 (24) is the most homologous gene reported previously and that the cDNA is closely related to other Zic and Opa proteins. Therefore, we named the novel gene as Xenopus Zic3.
To clarify the spatial expression pattern of Zic3, we performed a series of whole-mount in situ hybridizations. Zic3 expression was first detected at early gastrula in dorsal lip and prospective neural plate [Fig. 1A, stage 10.25 (41)]. As gastrulation proceeded, the expression decreased in the dorsal lip and increase in the prospective neural plate (Fig. 1 B and C, stage 10.5). In late gastrula, Zic3 expression diminished gradually in the central region (Fig. 1D, stage 12). At the neural plate stage (Fig. 1E, stage 14), Zic3 was expressed strongly in prospective region of mesencephalon and anterior rhombencephalon (42). Following that, Zic3 expression became stronger in the anterior neural folds, whereas that in the trunk neural folds remained weak (Fig. 1F, stage 16). At early tailbud stages (Fig. 1 G and H, stage 20), Zic3 expression was gradually restricted to the dorsal region of forebrain (telencephalon and diencephalon), midbrain, and hindbrain, and weakly to the dorsal region of the trunk. After mid-tailbud stage, Zic3 expression diminished in the diencephalon and an additional expression became clear in the lateral mesoderm of the tailbud region (Fig. 1I, stage 30). The cross section through the head at the same stage, showed that Zic3 expression was restricted in a dorsal part of the neural tube (Fig. 1J).
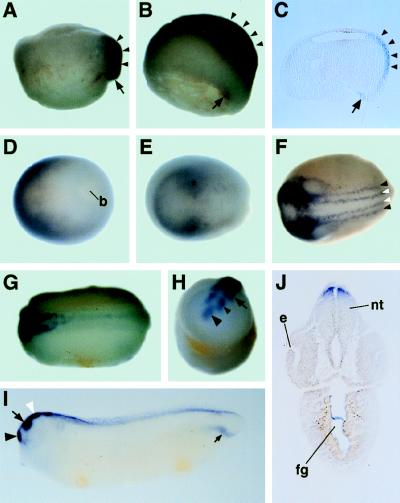
Figure 1.
The spatial expression pattern of Zic3 in Xenopus embryos. Series of embryo were hybridized with digoxigenin-labeled antisense Zic3 RNA. (A) Stage 10.25. (B) Stage 10.5. (C) The expression of Zic3 in dorsal lip (arrow) and prospective neural plate (arrowheads) was confirmed in the cross section of the same embryo in B. (D) Stage 12. (E) Stage 14. Asterisk indicates prospective region of mesencephalon and rhombencephalon. (F) Stage 16. White and black arrowheads indicate the lateral edge of neural plate and neural crest, respectively. (G and H) Stage 20. (I) Stage 30. In H and I, large black arrowhead, small black arrowhead, arrow, and white arrowhead indicate the telencephalon, diencephalon, mesencephalon, and rhombencephalon, respectively. Small arrow in I indicates the staining in a part of lateral mesoderm of tail. (J) Transverse section through the head region of same embryo as in I. Staining in the gut is nonspecific one, because it was detected also by the sense strand probe(data not shown). A, B, and I are the lateral views. The upper side of the panel is the animal side in A and B and the dorsal side in I. D–G are the dorsal views, the anterior side is toward left. H is the anterior view. The upper side is dorsal. b, blastopore; e, eye; nt, neural tube; fg, fore-gut.
Zic3 Is Expressed Earlier Than Any Other Neural Marker Genes in Xenopus Embryo.
Because Zic3 was expressed in prospective neural plate region during gastrulation, we compared the temporal expression profile of Zic3 with other neural marker genes (XASH-3, XATH-3, XlPOU 2, NeuroD, NCAM, N-tubulin) (18–20, 22, 34, 36, 37, 43, 44) by RT-PCR (Fig. 2A). The Zic3 expression was detected from early gastrula (stage 10). On the other hand, other neural marker genes were detected from mid-gastrula (XATH-3, NCAM) or later stage (NeuroD, N-tubulin), except XASH-3 and XlPOU 2. Although XASH-3 and XlPOU 2 transcripts were detected from early gastrula, their expression was extremely limited when compared with Zic3 expression.
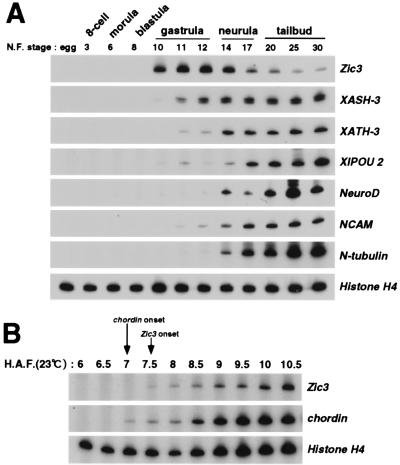
Figure 2.
The temporal expression profiles of Zic3 and neural marker genes during development. (A) RNA was extracted from embryos at the indicated stage of development and the contents of Zic3 and neural marker genes (XASH-3, XATH-3, XlPOU 2, NeuroD, NCAM, N-tubulin) mRNA were measured by RT-PCR. (B) To compare the onset of the expression of Zic3 and neural inducer, chordin, during development, RNA was extracted from embryos at the indicated time after artificial fertilization and the expression level was determined by RT-PCR. The ubiquitous marker, Histone H4 served as a control for both A and B. H.A.F.; hours after fertilization.
Zic3 was first detected in prospective neural plate region just after neural induction (Fig. 1A). We therefore compared the onset of expression between Zic3 and a neural inducer, chordin, precisely (6) (Fig. 2B). Zic3 expression began 7.5 h after fertilization, which was 30 min later than chordin expression. These data led us to hypothesize that the Zic3 expression is induced at the initial step of neural induction.
Zic3 Is Induced by Blockade of BMP4-Mediated Signaling.
Xenopus ectoderm (animal cap) is neuralized by prolonged culture in dispersal (45, 46). To examine whether Zic3 is induced in animal cap explant under such a condition, we cultured animal caps in dispersal to the time point equivalent to mid-neurula and the expression of Zic3 and other genes was tested by RT-PCR (Fig. 3A). Zic3 was expressed in dispersed animal caps with neural marker NCAM, whereas it was not detected in intact caps in which epidermal marker, epidermal keratin, expressed. Interestingly, Xtwi and Xslu, neural crest markers (38, 39), were not expressed in dispersed animal caps (see Discussion). The neuralization that occurred in the dispersed animal caps is considered to be due to the attenuation of BMP4-mediated signal (10).
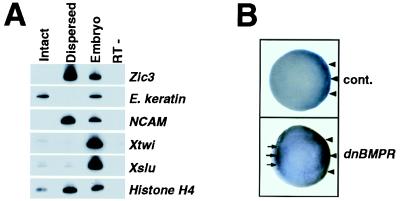
Figure 3.
Zic3 was induced by blockade of BMP4 signaling. (A) Expression of Zic3 and other marker genes in the dispersed animal cap explants were measured by RT-PCR. The dispersed caps (Dispersed) differentiated into neural cells expressing NCAM whereas not dissociated caps (Intact) differentiated into epidermal cells expressing epidermal keratin (E. keratin). Zic3 expression was detected only in the dispersed animal caps. Sibling control embryos served as a positive control (Embryo) and PCR on the same RNA without reverse transcription was done to check the absence of genomic DNA (RT−). (B) Zic3 was induced by the injection of dnBMPR mRNA. Zic3 expression was detected by whole-mount in situ hybridization of early gastrula (stage 10.25) embryos injected with control LacZ mRNA (Upper) or dnBMPR mRNA (Lower) into the ventromarginal region at two-cell stage. The embryo injected with dnBMPR mRNA showed an ectopic Zic3 expression in the ventromarginal zone (arrows in Lower) in addition to the proper site (arrowheads in Lower), which was also found in the control embryo (arrowheads in Upper).
We therefore considered if the blockade of BMP4-mediated signal actually can induce the Zic3 expression in vivo. A dominant negative form of BMP receptor (dnBMPR) (47) or noggin (5) was overexpressed in a ventral region by injection into two-cell stage embryo. The embryos were subjected to in situ hybridization with Zic3 probe at the early gastrula stage. Both dnBMPR mRNA (Fig. 3B) and noggin mRNA (data not shown), which were shown to induce neuroectoderm by inhibition of BMP4 signal (14), were able to induce ectopic Zic3 mRNA in ventromarginal zone in gastrula. Therefore, blockade of BMP4 signaling is sufficient to induce Zic3 in vivo.
Overexpression of Zic3 in Xenopus Embryos Leads to Hyperplasia of Neural and Neural Crest Tissue.
The expression pattern of Zic3 in Xenopus embryos and its regulation in neural induction suggest that one role of Zic3 is in the initial step of neural development. Therefore, we decided to examine the function of Zic3 by the overexpression experiments in embryos. First, we injected Zic3 mRNA into one blastomere of two-cell stage embryos to overexpress Zic3 in a hemilateral body.
In almost all cases, the sides of the heads injected with Zic3 mRNA were enlarged and coarse showing poorly formed eyes(Fig. 4 B and C), whereas uninjected control side were normal (Fig. 4A). The sections through the head region of injected embryo showed that neural walls were considerably thickened in the injected side (Fig. 4 D–F). In addition to the change in neural wall, presumptive mesenchymal tissue, which may derive from the neural crest in the cephalic region, showed a remarkable hyperplasia. In most cases, neural retinas were considerably distorted and, less frequently, hyperplastic (Fig. 4D). On the other hand, retinal pigment cells diminished variably and, in particular, lenses were not induced at all (Fig. 4 C and D).
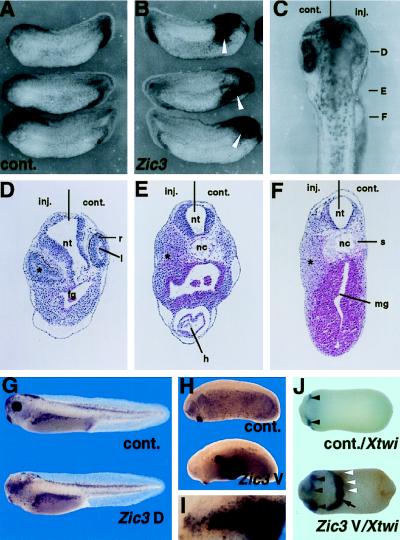
Figure 4.
Zic3 overexpression induces neural tissue hyperplasia and ectopic pigment cell expression. A total of 100 pg of Zic3 mRNA (A–F) was injected into one blastomere of two-cell stage embryos. (A) Uninjected control side (stage 27). (B) Zic3 mRNA injected side (stage 27). Clusters of pigment cells appeared in the anterior region (arrow). (C) Dorsal view of anterior region of the injected embryo (stage 36). (D–F) Transverse sections of the embryo shown in C. The levels of sections are indicated in C. The injected side was characterized by the hyperplastic neural walls, distorted eye, and unidentified tissue, probably derived from the neural crest (asterisk in D, E, and F, see Results). A total of 100 pg of Zic3 mRNA or control LacZ mRNA was injected into dorsoanimal (G) or ventroanimal (H–J) two blastomeres of eight-cell stage embryo, and cultured to stage 36 (G), stage 25 (H and I), or stage 20 (J). Lateral view of control LacZ mRNA injected embryos (G and H, Upper) and Zic3 mRNA injected embryos (G and H, Lower). The eye abnormalities were observed in the embryos injected with Zic3 mRNA into dorsal blastomeres (G). Remarkable clusters of ectopic pigment cells appeared in the embryos injected with Zic3 mRNA into ventral blastomeres (H). (I) Higher magnification of the pigment cell clusters in (H). (J) Xtwi expression in the embryos (ventral view) injected with LacZ mRNA (Upper) or Zic3 mRNA (Lower) into ventroanimal two blastomeres at eight-cell stage. The Xtwi expression was observed in the head neural crest of the control embryo (Upper, arrowhead). However, the Zic3 mRNA injection induced the ectopic Xtwi expression (Lower, arrow) near the ectopic clusters of pigment cells (white arrowheads; the brown staining indicates the remnants of the pigments) in the ventral side of embryo. The expansion of the Xtwi expressing cephalic neural crest (Lower, black arrowheads) was also observed in this embryo.
In addition to the phenotypes in the head, the ectopic clusters of pigment-containing cells were found in the head region (Fig. 4B). However, the extent and position of the clusters varied in each case. We speculated that these variations reflected the difference of the sites where Zic3 was expressed. We therefore injected Zic3 mRNA into the dorsoanimal or ventroanimal two blastomeres of the eight-cell stage embryo to express Zic3 restrictedly in dorsal or ventral side (Fig. 4 G–J). When Zic3 mRNA were injected into dorsoanimal blastomeres, heads of the embryos were enlarged and the eyes showed the same change as observed in the embryos injected at two-cell stage (58/80 embryos tested) (Fig. 4G), and neural tube closure in the anterior region was delayed (data not shown). The pigment cells were found in the dorsal head (58/80 embryo tested). In contrast, if the mRNA was injected into ventroanimal blastomeres, clusters of ectopic pigment cells appeared in the ventral epidermis (90/111 embryos tested) (Fig. 4 H and I). The clusters of the ventral pigment cells were arrayed remarkably on the ridge of the hyperplastic tissue which transverses the ventral side. These pigment cells were considered to be melanocytes, which derived from neural crest. Therefore, we performed in situ hybridization with the probes of neural crest marker, Xtwi, in Zic3 mRNA injected embryo (Fig. 4J, data not shown). Ectopic Xtwi expression was observed near the ectopically appearing clusters of pigment cells in addition to the expansion of the Xtwi-expressing region of the cephalic neural crest.
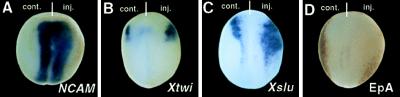
To determine whether Zic3 overexpression cause an early alteration in cell fate, we examined the expression of NCAM, Xtwi, Xslu, and EpA, an epidermal antigen (40), at an early neurula (Fig. 5 A–D). Zic3 mRNA was injected into a blastomere of two-cell stage embryos and the expression patterns were examined at stage 14. It became clear that the NCAM-expressing region (31/45 embryos tested) increased markedly in the anterior neural plate region of injected side(Fig. 5A). Xtwi and Xslu expression in neural crest cells were also expanded in injected side(Fig. 5 B and C) (42/45 embryos tested by Xtwi probe and 12/12 embryos tested by Xslu probe). If the epidermal fate change into a neural and neural crest fate, epidermis should be reduced in Zic3 mRNA-injected side. To test this possibility, we next determined the expression of EpA in the injected embryos. On the Zic3-injected side, expression of EpA was significantly reduced (Fig. 5D). These facts suggest that Zic3 altered epidermal cell fate into neural and neural crest cell fate.
Figure 5.
Zic3 induced NCAM, Xtwi, and Xslu expression but reduced epidermis in early stage embryo. A total of 100 pg of Zic3 mRNA was injected into one blastomere of two-cell stage embryos (A–D). In situ hybridization was performed with NCAM (A), Xtwi (B), and Xslu (C) probe, and immunohistochemistry was performed with EpA mAb (D). (A–D) Dorsal view of a stage 14 embryo. NCAM, Xtwi, and Xslu expressing regions expand laterally in the injected side (A–C). EpA staining in the epidermis is reduced in the injected side (D).
Zic3 Induces Neural Marker Genes Including Proneural Genes and Neural Crest Marker Genes Without Mesoderm Induction in Animal Cap Explant.
The above studies indicate that Zic3 has important roles in early neural and neural crest development. How does Zic3 act in this process? To address this question, we examined the expression of several marker genes by RT-PCR in the animal cap explants injected with Zic3 mRNA (Fig. 6).
Figure 6.
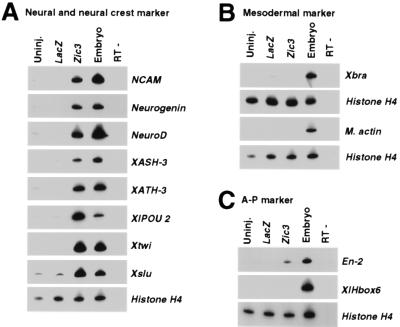
Zic3 induced neural and neural crest marker genes without mesoderm induction in animal cap explant. Embryos were injected with 100 pg of Zic3 or control LacZ mRNA at the two-cell stage. Animal caps were explanted at stage 9 and cultured. (A) Zic3 induced all the neural and neural crest marker genes tested. When the sibling embryos reached stage 20, the expression of neural marker (NCAM, Neurogenin, NeuroD, XASH-3, XATH-3, XlPOU 2) and neural crest marker (Xtwi, Xslu) were examined by RT-PCR. Although uninjected (Uninj.) or LacZ injected (LacZ) caps do not express any of these markers, animal caps injected with Zic3 mRNA (Zic3) express all of the neural markers and neural crest markers we tested. (B) Zic3 did not induce mesodermal marker. An early mesodermal marker, Xbra, and a dorsal mesodermal marker, M. actin, expression were examined by RT-PCR when the sibling embryos became stage 10.5 and stage 20, respectively. (C) Zic3 induced anterior neural character. Expression of the neural markers, which is expressed differentially along the anteroposterior axis, was examined by RT-PCR when the sibling embryos became stage 27. Anterior neural marker, En-2, but not the posterior neural marker, XlHbox6, was expressed in the Zic3 overexpressed animal cap. In each experiment, sibling control embryos served as a positive control (Embryo) and PCR on the same RNA without reverse transcription was done to check the absence of genomic DNA (RT−).
As expected, a neural marker (NCAM) was induced in the explant by the Zic3 overexpression (Fig. 6A). Interestingly, neural crest markers (Xtwi and Xslu) were also induced in this case (Fig. 6A). This is in contrast to the finding that Xtwi and Xslu were not induced in dispersed animal caps although Zic3 is expressed (Fig. 3) (see Discussion). An early mesodermal marker [Xenopus brachyury (Xbra)] and a dorsal mesodermal marker [muscle actin (M. actin)] were not induced (Fig. 6B). These results demonstrate that Zic3 is able to generate neural tissue without mesoderm induction and that Zic3 can directly convert epidermal fate to neural and neural crest fate.
As for the anteroposterior neural properties, the molecular markers which are expressed in anterior neural plate were induced [En-2 (48)] by Zic3 overexpression, whereas the posterior marker [XlHbox6 (49)] was not (Fig. 6C). This result is consistent with previous finding that anterior neuroectoderm is induced in animal cap explant by a BMP4 signal blockade which also induces Zic3 expression (7, 11, 50).
Finally, we considered if Zic3 can induce so called proneural genes, because Zic3 is expressed earlier than these genes. (In this paper, we tentatively use the words “proneural gene” for the gene which is shown to have neurogenic activity and to be expressed in neural development.) Surprisingly, all the proneural genes (Neurogenin, NeuroD, XASH-3, XATH-3, and XlPOU 2) which we tested were induced by Zic3 overexpression, whereas uninjected or LacZ injected animal cap explant did not show any expression (Fig. 6A). The fact suggests that Zic3 may act upstream of proneural genes reported so far.
DISCUSSION
Zic3 Expression Is Induced by Blockade of BMP4 Signal.
Zic3 expressed in the prospective neural plate in early gastrula. The expression begins just after the chordin expression. These expression patterns led us to hypothesize that Zic3 is regulated directly by the neural tissue-inducing signals (1, 3).
Recent studies showed that the factors antagonizing BMP4, which represses differentiation into neural tissue and promotes epidermal differentiation (10), have important roles in neural induction (14–16). These factors, such as noggin, chordin, and follistatin, are expressed in the organizer and have neural tissue inducing activities.
We showed that Zic3 mRNA level is elevated in the dispersed animal cap explant and that Zic3 expression in gastrula is ectopically induced by the dnBMPR or noggin overexpression. Both results suggest that blockade of BMP4 signal can induce the Zic3 expression. Based on these findings, we consider that Zic3 expression may be regulated negatively by the BMP4 signal from nonneural ectoderm and that the neural inducers may relieve the Zic3 from the repression by the BMP4 signal.
Zic3 Functions in Neural Development Upstream of Proneural Genes.
Zic3 overexpression in the developing embryos results in the expansion of neural plate and the hyperplasia of neural tissue. Moreover, several neural markers are induced in the Zic3 overexpressed animal caps. These data indicate that Zic3 has a neurogenic activity. Our study strongly indicates that Zic3 acts in the earliest phase of neural development and that Zic3 belongs to a novel class of genes involved in neural development based on the following three points.
First, the initiation of the Zic3 expression is in the late blastula, earlier than most proneural genes. Although Neurogenin transcripts are detected earlier than Zic3 expression, as a maternal message in our assay (data not shown), the function of Neurogenin as a proneural gene seems to be activated after neural induction occurs.
Second, Zic3 is first expressed in almost entire prospective neural plate region. This spatial expression pattern is considerably different from those of proneural genes. Of the proneural genes reported so far, the Neurogenin, NeuroD, and XATH-3 are expressed in the three longitudinal regions where the primary neurons are generated (17, 18, 22). As for XASH-3 and XlPOU 2, the restricted expression in these regions has not been reported. However, XASH-3 expression is detected in a restricted region of the neural plate particularly from its expression onset (stage 11.5) (20, 43). The XASH-3 expression pattern is in contrast to that of Zic3, which is first expressed in almost entire prospective neural plate. XlPOU 2 is detected as early as stage 10 in the involuting mesoderm, but not in prospective neural plate (34) where Zic3 is expressed (Fig. 1 B and C).
Finally, the overexpression of Zic3 induces the expression of all the proneural genes in the animal cap. It is not likely that a gene other than Zic3 can induce all of the other proneural genes based on their spatio-temporal expression patterns.
Zic3 Functions in Neural Crest Formation.
Zic3 is expressed in the neural folds including the neural crest, and strongly in the cephalic region at stage 12–16. Furthermore, Zic3 overexpression induces ectopic pigment cells, which derive from the neural crest and neural crest markers, Xtwi and Xslu, in embryo (38, 39, 51, 52). In addition, Xtwi and Xslu are induced in the Zic3 overexpressed animal cap explant. All these findings suggest that Zic3 has an essential role in the neural crest induction.
Xtwi and Xslu are induced, not in dispersed animal cap, but in Zic3 mRNA injected animal caps, although Zic3 is expressed in both cases. This fact indicates that factors which are present only in the nondissociated animal cap explants, possibly the secretory factors, can induce the Xtwi and Xslu expression in cooperation with Zic3. As a candidate of such factors, BMP4 and BMP7 might be appropriate because both factors are expressed in nondissociated animal caps (13, 33, 53, 54) and can induce neural crest markers in chicken neural plate explants (55, 56).
Zic3 as an Initial Regulator of the Ectodermal Cell Fate Decision.
The findings in this paper suggest that Zic3 plays a part in the vertebrate ectodermal cell fate decision. The role of Zic3 is peculiar when the results of Zic3 overexpression are compared with those of the proneural genes. First, Neurogenin, NeuroD, XATH-3, and XlPOU 2 induce ectopic neuronal differentiation in the epidermis (17, 18, 22, 34) whereas Zic3 does not. In the course of neural development, Zic3 may cooperate with other factors restricted to the dorsal ectoderm and/or Zic3 is more sensitive to inhibitors that may be present in the epidermis. The difference suggests that the role of Zic3 in “neuronal differentiation,” occurring in a relatively late phase of neural development, is not so definitive as with the other genes. The role of Zic3 may be confined to an early phase of neural development. Second, NeuroD, XASH-3, or XATH-3 overexpression abolishes the expression of Xtwi, a neural crest marker (18–20, 22). On the other hand, the expression is enhanced by Zic3. This difference may be elucidated by the fact that Zic3 is capable of inducing not only neural but also neural crest tissues, possibly cooperating with diffusible factors from epidermis. The neural tissue induced by XASH-3, NeuroD, or XATH-3 overexpression, and which may represent the neural tissue of later stages, would be insensitive to these factors. Therefore, the Xtwi was not induced by the overexpression of the three genes.
These features of Zic3 seem to correspond closely with the ideas that Zic3 functions as an initial regulator of ectodermal cell fate decision. Understanding the mechanism involved in the regulation of Zic3 expression, and how Zic3 regulates the proneural genes, would be helpful for the understanding this process.
Acknowledgments
We thank Dr. H. Okano for his critical reading of the manuscript; Dr. N. Ueno for helpful advice and dnBMPR expression vector; Dr. R. Kageyama for helpful advice and preliminary data on XATH-3; Drs. D. Melton, D. Turner, R. Harland, J. B. Gurdon, and A. Muto for plasmids; Drs. M. Asashima, H. Okamoto, P. A. Wilson, A. Suzuki, and S. Kume for helpful advice; Mr. K. Yamanaka for technical assistance; and Dr. C. S. Langham for helpful comments on the manuscript. This work was supported by Special Coordination Funds for Promoting Science and Technology, grants from the Japanese Ministry of Education, Science and Culture to J.A. and K.M., CREST (Core Research for Evolutional Science and Technology) of Japan Science and Technology Cooperation to J.A., the Japanese Brain Science Foundation to J.A., and the Japan Society for Promotion of Science to K.N. and T.N.
Note Added in Proof
The Xenopus Zic family has also been independently characterized by both Hazel Sive’s group and Ariel Ruiz i Altaba’s group (personal communications).
Footnotes
This paper was submitted directly (Track II) to the Proceedings Office.
Abbreviations: RT-PCR, reverse transcription–PCR; NCAM, neural cell adhesion molecule, Xtwi, Xenopus twist; Xslu, Xenopus slug.
Data deposition: The sequence reported in this paper has been deposited in the GenBank database (accession no. AB005292).
References
- 1.Simpson P. Neuron. 1995;15:739–742. doi: 10.1016/0896-6273(95)90163-9. [DOI] [PubMed] [Google Scholar]
- 2.Calof A L. Curr Opin Neurobiol. 1995;5:19–27. doi: 10.1016/0959-4388(95)80082-4. [DOI] [PubMed] [Google Scholar]
- 3.Tanabe Y, Jessell T M. Science. 1996;274:1115–1123. doi: 10.1126/science.274.5290.1115. [DOI] [PubMed] [Google Scholar]
- 4.Spemann H, Mangold H. Roux’ Arch f Entw mech. 1924;100:599–638. [Google Scholar]
- 5.Smith W C, Harland R M. Cell. 1992;70:829–840. doi: 10.1016/0092-8674(92)90316-5. [DOI] [PubMed] [Google Scholar]
- 6.Sasai Y, Lu B, Steinbeisser H, Geissert D, Gont L K, De Robertis E M. Cell. 1994;79:779–790. doi: 10.1016/0092-8674(94)90068-x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 7.Hemmati-Brivanlou A, Kelly O G, Melton D A. Cell. 1994;77:283–295. doi: 10.1016/0092-8674(94)90320-4. [DOI] [PubMed] [Google Scholar]
- 8.Smith W C, McKendry R, Ribisi S, Harland R M. Cell. 1995;82:37–46. doi: 10.1016/0092-8674(95)90050-0. [DOI] [PubMed] [Google Scholar]
- 9.Hansen C S, Marion C D, Steele K, George S, Smith W C. Development (Cambridge, UK) 1997;124:483–492. doi: 10.1242/dev.124.2.483. [DOI] [PubMed] [Google Scholar]
- 10.Wilson P A, Hemmati-Brivanlou A. Nature (London) 1995;376:331–333. doi: 10.1038/376331a0. [DOI] [PubMed] [Google Scholar]
- 11.Sasai Y, Lu B, Steinbeisser H, De Robertis E M. Nature (London) 1995;376:333–336. doi: 10.1038/376333a0. [DOI] [PubMed] [Google Scholar]
- 12.Xu R H, Kim J, Taira M, Zhan S, Sredni D, Kung H F. Biochem Biophys Res Commun. 1995;212:212–219. doi: 10.1006/bbrc.1995.1958. [DOI] [PubMed] [Google Scholar]
- 13.Hawley S H B, Wünnenberg-Stapleton K, Hashimoto C, Laurent M N, Watabe T, Blumberg B W, Cho K W Y. Genes Dev. 1995;9:2923–2935. doi: 10.1101/gad.9.23.2923. [DOI] [PubMed] [Google Scholar]
- 14.Zimmerman L B, De Jesús-Escobar J M, Harland R M. Cell. 1996;86:599–606. doi: 10.1016/s0092-8674(00)80133-6. [DOI] [PubMed] [Google Scholar]
- 15.Piccolo S, Sasai Y, Lu B, De Robertis E M. Cell. 1996;86:589–598. doi: 10.1016/s0092-8674(00)80132-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Fainsod A, Deißler K, Yelin R, Marom K, Epstein M, Pillermer G, Steinbeisser H, Blum M. Mech Dev. 1997;63:39–50. doi: 10.1016/s0925-4773(97)00673-4. [DOI] [PubMed] [Google Scholar]
- 17.Ma Q, Kintner C, Anderson D J. Cell. 1996;87:43–52. doi: 10.1016/s0092-8674(00)81321-5. [DOI] [PubMed] [Google Scholar]
- 18.Lee J E, Hollenberg S M, Snider L, Turner D L, Lipnick N, Weintraub H. Science. 1995;268:836–844. doi: 10.1126/science.7754368. [DOI] [PubMed] [Google Scholar]
- 19.Ferreiro B, Kintner C, Zimmerman K, Anderson D, Harris W A. Development (Cambridge, UK) 1994;120:3649–3655. doi: 10.1242/dev.120.12.3649. [DOI] [PubMed] [Google Scholar]
- 20.Turner D L, Weintraub H. Genes Dev. 1994;8:1434–1447. doi: 10.1101/gad.8.12.1434. [DOI] [PubMed] [Google Scholar]
- 21.Chitnis A, Kintner C. Development (Cambridge, UK) 1996;122:2295–2301. doi: 10.1242/dev.122.7.2295. [DOI] [PubMed] [Google Scholar]
- 22.Takebayashi K, Takahashi S, Yokota C, Tsuda H, Nakanishi S, Asashima M, Kageyama R. EMBO J. 1997;16:384–395. doi: 10.1093/emboj/16.2.384. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Aruga J, Yokota N, Hashimoto M, Furuichi T, Fukuda M, Mikoshiba K. J Neurochem. 1994;63:1880–1890. doi: 10.1046/j.1471-4159.1994.63051880.x. [DOI] [PubMed] [Google Scholar]
- 24.Aruga J, Nagai T, Tokuyama T, Hayashizaki Y, Okazaki Y, Chapman V M, Mikoshiba K. J Biol Chem. 1996;271:1043–1047. doi: 10.1074/jbc.271.2.1043. [DOI] [PubMed] [Google Scholar]
- 25.Aruga J, Yozu A, Hayashizaki Y, Okazaki Y, Chapman V M, Mikoshiba K. Gene. 1996;172:291–294. doi: 10.1016/0378-1119(96)00111-4. [DOI] [PubMed] [Google Scholar]
- 26.Yokota N, Aruga J, Takai S, Yamada K, Hamazaki M, Iwase T, Sugimura H, Mikoshiba K. Cancer Res. 1996;56:377–383. [PubMed] [Google Scholar]
- 27.Nagai T, Aruga J, Takada S, Günther T, Spörle R, Schughart K, Mikoshiba K. Dev Biol. 1997;182:299–313. doi: 10.1006/dbio.1996.8449. [DOI] [PubMed] [Google Scholar]
- 28.Benedyk M J, Mullen J R, DiNardo S. Genes Dev. 1994;8:105–117. doi: 10.1101/gad.8.1.105. [DOI] [PubMed] [Google Scholar]
- 29.Cimbora D M, Sakonju S. Dev Biol. 1995;169:580–595. doi: 10.1006/dbio.1995.1171. [DOI] [PubMed] [Google Scholar]
- 30.Moon R T, Christian J L. Technique. 1989;1:76–89. [Google Scholar]
- 31.Asashima M, Nakano H, Shimada K, Kinoshita K, Ishii K, Shibai H, Ueno N. Roux’s Arch Dev Biol. 1990;198:330–335. doi: 10.1007/BF00383771. [DOI] [PubMed] [Google Scholar]
- 32.Suzuki A, Nagai T, Nishimatsu S, Sugino H, Eto Y, Shibai H, Murakami K, Ueno N. Biochem J. 1994;298:275–280. doi: 10.1042/bj2980275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Fainsod A, Steinbeisser H, De Robertis E M. EMBO J. 1994;13:5015–5025. doi: 10.1002/j.1460-2075.1994.tb06830.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Witta S E, Agarwal V R, Sato S M. Development (Cambridge, UK) 1995;121:721–730. doi: 10.1242/dev.121.3.721. [DOI] [PubMed] [Google Scholar]
- 35.Kelly G M, Eib D W, Moon R T. Methods Cell Biol. 1991;36:389–417. doi: 10.1016/s0091-679x(08)60289-7. [DOI] [PubMed] [Google Scholar]
- 36.Chitnis A, Henrique D, Lewis J, Ish-Horowicz D, Kintner C. Nature (London) 1995;375:761–766. doi: 10.1038/375761a0. [DOI] [PubMed] [Google Scholar]
- 37.Kintner C R, Melton D A. Development (Cambridge, UK) 1987;99:311–325. doi: 10.1242/dev.99.3.311. [DOI] [PubMed] [Google Scholar]
- 38.Mayor R, Morgan R, Sargent M G. Development (Cambridge, UK) 1995;121:767–777. doi: 10.1242/dev.121.3.767. [DOI] [PubMed] [Google Scholar]
- 39.Hopwood N D, Pluck A, Gurdon J B. Cell. 1989;59:893–903. doi: 10.1016/0092-8674(89)90612-0. [DOI] [PubMed] [Google Scholar]
- 40.Jones E A, Woodland H R. Cell. 1986;44:345–355. doi: 10.1016/0092-8674(86)90769-5. [DOI] [PubMed] [Google Scholar]
- 41.Nieuwkoop P D, Faber J. Normal Table of Xenopus laevis. North Holland, Amsterdam: Daudin; 1967. [Google Scholar]
- 42.Eagleson G W, Harris W A. J Neurobiol. 1989;21:427–440. doi: 10.1002/neu.480210305. [DOI] [PubMed] [Google Scholar]
- 43.Zimmerman K, Shih J, Bars J, Collazo A, Anderson D J. Development (Cambridge, UK) 1993;119:221–232. doi: 10.1242/dev.119.1.221. [DOI] [PubMed] [Google Scholar]
- 44.Oschwald R, Richter K, Grunz H. Int J Dev Biol. 1991;35:399–405. [PubMed] [Google Scholar]
- 45.Gruntz H, Tacke L. Cell Differ Dev. 1989;28:211–218. doi: 10.1016/0922-3371(89)90006-3. [DOI] [PubMed] [Google Scholar]
- 46.Godsave S F, Slack J M. Dev Biol. 1989;134:486–490. doi: 10.1016/0012-1606(89)90122-x. [DOI] [PubMed] [Google Scholar]
- 47.Suzuki A, Shinoda N, Ueno N. Growth Differ. 1995;37:581–588. doi: 10.1046/j.1440-169X.1995.t01-3-00013.x. [DOI] [PubMed] [Google Scholar]
- 48.Hemmati-Brivanlou A, de la Torre J R, Holt C, Harland R M. Development (Cambridge, UK) 1991;111:715–724. doi: 10.1242/dev.111.3.715. [DOI] [PubMed] [Google Scholar]
- 49.Wright C V E, Morita E A, Wilkin D J, De Robertis E M. Development (Cambridge, UK) 1990;109:225–234. doi: 10.1242/dev.109.1.225. [DOI] [PubMed] [Google Scholar]
- 50.Lamb T M, Knecht A K, Smith W C, Stachel S E, Economides A N, Stahl N, Yancopolous D, Harland R M. Science. 1993;262:713–718. doi: 10.1126/science.8235591. [DOI] [PubMed] [Google Scholar]
- 51.Le Douarin NM. The Neural Crest. Cambridge: Cambridge Univ. Press; 1982. [Google Scholar]
- 52.Mancilla A, Mayor R. Dev Biol. 1996;177:580–589. doi: 10.1006/dbio.1996.0187. [DOI] [PubMed] [Google Scholar]
- 53.Schmidt J E, Suzuki A, Ueno N, Kimelman D. Dev Biol. 1995;169:37–50. doi: 10.1006/dbio.1995.1124. [DOI] [PubMed] [Google Scholar]
- 54.Hemmati-Brivanlou A, Thomsen G H. Dev Genet. 1995;17:78–89. doi: 10.1002/dvg.1020170109. [DOI] [PubMed] [Google Scholar]
- 55.Liem K F, Jr, Tremml G, Roelink H, Jessell T M. Cell. 1995;82:969–979. doi: 10.1016/0092-8674(95)90276-7. [DOI] [PubMed] [Google Scholar]
- 56.Bang A G, Papalopulu N, Kintner C, Goulding M D. Development (Cambridge, UK) 1997;124:2075–2085. doi: 10.1242/dev.124.10.2075. [DOI] [PubMed] [Google Scholar]