Abstract
The degree of similarity of two protein three-dimensional structures is usually measured with the root-mean-square distance between equivalent atom pairs. Such a similarity measure depends on the dimension of the proteins, that is, on the number of equivalent atom pairs. The present communication presents a simple procedure to make the root-mean-square distances between pairs of three-dimensional structures independent of their dimensions. This normalization may be useful in evolutionary and fold classification studies as well as in simple comparisons between different structural models.
Keywords: Root-mean-square distance, structure classification, structure comparison, three-dimensional similarity
Quantitative comparison of three-dimensional structures is a fundamental task in structural biology (Carugo and Eisenhaber 1997; Peters-Libeu and Adman 1997), especially in such fields as domain fold classification and structural evolution studies (Domingues et al. 2000; Yang and Honig 2000). A very popular quantity used to express the structural similarity is the root-mean-square distance (rmsd) calculated between equivalent atoms in two structures, defined as
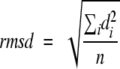 |
1 |
where d is the distance between each of the n pairs of equivalent atoms in two optimally superposed structures. The rmsd is 0 for identical structures, and its value increases as the two structures become more different. Rmsd values are considered as reliable indicators of variability when applied to very similar proteins, like alternative conformations of the same protein. On the other hand, rmsd data calculated for structure pairs of different sizes cannot be directly compared, because the rmsd value obviously depends on the number of atoms included in the structural alignment. Clearly, an rmsd value of, say, 3 Å has a different significance for proteins of 500 residues than for those of 50 residues; accordingly, the structural variability of fold types cannot be easily compared in quantitative terms (Irving et al. 2001). In other words, rmsd is a good indicator for structural identity, but less so for structural divergence.
The present communication aims to define a normalized, size-independent rmsd formula that could help to overcome this problem. In order to derive a formula between rmsd and protein dimension, one would need a database of structural alignments, in which all other parameters, such as secondary structure content and amino acid composition of the protein, are either constant (which is not possible) or are evenly distributed with respect to protein chain length. Such experimental data are presently not available. For example, the FSSP database (Holm and Sander 1996) contains a reasonably high number of structural alignments (about 23,000), but 80% of these have small rmsd values (0–2 Å), which reflects the fact that the percentage of sequence identity is very high (more than 90% residue identity in 60% of the alignments).
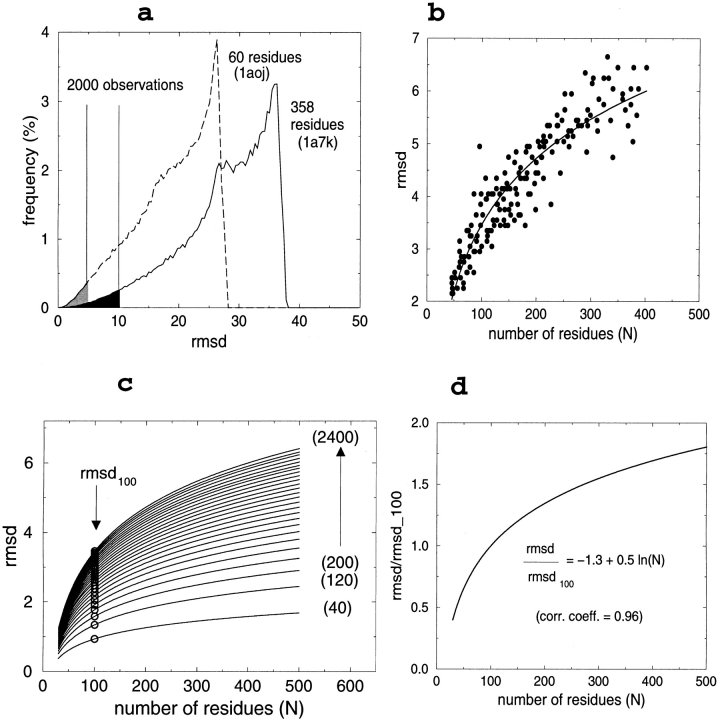
We therefore decided to create a large artificial set of rmsd values via extensive self-comparison of 180 nonhomologous (maximal identity 25%) protein structures, selected from the protein data bank (Berman et al. 2000) using the PDB_SELECT (Hobohm and Sander 1994) algorithm. These proteins were selected so as to represent the largest possible variability of amino acid content, sequence length as well as secondary structure content (Table 1). Each structure was compared, using the algorithm of Kabsch (1976, 1978), with 400,000 of its randomized variants created through random shuffling of the Cα equivalencies. All Cα atoms were included in superposing each structure with all its variants. Overall, we obtained 400,000 rmsd observations in each of the 180 randomization experiments, which corresponds to a database of 72 million structural alignments. As expected, the distribution of rmsd values thus obtained depends on the size of the protein. The rmsd values are not evenly distributed, rather, the histograms are biased toward the high rmsd values (Fig. 1a ▶). Moreover, there are characteristic differences between proteins of different length, illustrated by, for example, the different rmsd limits of the 2000 smallest rmsd values in the two experiments, as shown by the shaded areas in Figure 1a ▶.
Table 1.
Protein structures examined in the present work
| idcode | n | h | e | t | o | idcode | n | h | e | t | o | idcode | n | h | e | t | o |
| 1a7kA | 358 | 31 | 25 | 25 | 19 | 1agnA | 373 | 30 | 26 | 21 | 23 | 1ak0_ | 264 | 64 | 4 | 20 | 12 |
| 1amx_ | 150 | 8 | 54 | 21 | 17 | 1aojA | 60 | 5 | 45 | 22 | 28 | 1ap8_ | 213 | 21 | 23 | 26 | 29 |
| 1auxA | 292 | 29 | 31 | 20 | 20 | 1avgI | 142 | 8 | 41 | 27 | 25 | 1axn_ | 323 | 79 | 0 | 15 | 6 |
| 1b12A | 239 | 8 | 46 | 20 | 26 | 1b6bA | 168 | 26 | 30 | 21 | 23 | 1b6rA | 349 | 34 | 29 | 19 | 19 |
| 1b87A | 181 | 27 | 30 | 28 | 15 | 1b9lA | 119 | 32 | 39 | 17 | 12 | 1b9xA | 340 | 7 | 46 | 29 | 18 |
| 1bq2_ | 323 | 32 | 28 | 21 | 19 | 1bhu_ | 102 | 4 | 25 | 35 | 36 | 1boeA | 46 | 0 | 24 | 43 | 33 |
| 1bor_ | 56 | 0 | 4 | 55 | 41 | 1bp3A | 186 | 60 | 0 | 22 | 18 | 1bqv_ | 110 | 46 | 0 | 23 | 31 |
| 1bu2A | 229 | 60 | 0 | 27 | 14 | 1buyA | 166 | 57 | 1 | 20 | 22 | 1bxwA | 172 | 0 | 60 | 12 | 28 |
| 1by1A | 209 | 64 | 0 | 25 | 11 | 1bynA | 128 | 5 | 48 | 18 | 29 | 1cl7M | 142 | 84 | 0 | 7 | 9 |
| 1c20A | 128 | 65 | 3 | 15 | 17 | 1ceuA | 51 | 51 | 0 | 25 | 24 | 1cflD | 368 | 7 | 49 | 21 | 24 |
| 1ckv_ | 141 | 16 | 21 | 36 | 27 | 1cn3A | 283 | 7 | 41 | 25 | 27 | 1d0mA | 312 | 49 | 9 | 22 | 20 |
| 1dldA | 220 | 58 | 0 | 25 | 17 | 1d2hA | 252 | 32 | 31 | 19 | 18 | 1de9A | 276 | 27 | 26 | 22 | 25 |
| 1dgvA | 183 | 46 | 2 | 31 | 20 | 1dtjB | 62 | 44 | 32 | 6 | 18 | 1dujA | 187 | 28 | 29 | 18 | 25 |
| 1eus_ | 358 | 3 | 44 | 28 | 25 | 1evtD | 192 | 9 | 52 | 13 | 26 | 1ewiA | 114 | 11 | 25 | 31 | 33 |
| 1gcf_ | 109 | 0 | 44 | 20 | 36 | 1gnhA | 206 | 9 | 44 | 21 | 26 | 1gsa_ | 314 | 35 | 29 | 18 | 18 |
| 1hcd__ | 118 | 3 | 49 | 30 | 19 | 1hoe_ | 74 | 0 | 49 | 26 | 26 | 1hsm_ | 79 | 62 | 0 | 22 | 16 |
| 1ihfA | 96 | 42 | 29 | 9 | 20 | 1iyu_ | 79 | 0 | 47 | 25 | 28 | 1jlyA | 299 | 7 | 45 | 30 | 18 |
| 1ksr_ | 100 | 0 | 43 | 36 | 21 | 1lbd_ | 238 | 66 | 3 | 20 | 12 | 1liaA | 164 | 76 | 0 | 15 | 9 |
| 1mtyB | 384 | 64 | 1 | 19 | 16 | 1nfdA | 203 | 5 | 43 | 22 | 30 | 1oczB | 227 | 30 | 25 | 22 | 23 |
| 1pgs_ | 311 | 5 | 47 | 22 | 26 | 1pho_ | 330 | 2 | 56 | 25 | 16 | 1pslA | 304 | 72 | 0 | 14 | 14 |
| 1pyaA | 81 | 27 | 27 | 22 | 23 | 1qhkA | 47 | 32 | 28 | 21 | 19 | 1qklA | 127 | 20 | 14 | 32 | 33 |
| 1qleC | 273 | 68 | 1 | 16 | 16 | 1qmcA | 52 | 6 | 56 | 21 | 17 | 1qqvA | 67 | 42 | 3 | 30 | 25 |
| 1qrjB | 199 | 57 | 1 | 22 | 20 | 1qslA | 402 | 34 | 29 | 19 | 18 | 1qsoA | 149 | 34 | 30 | 19 | 17 |
| 1qstA | 160 | 35 | 29 | 22 | 14 | 1qu0C | 183 | 3 | 51 | 22 | 24 | 1qu5A | 182 | 10 | 22 | 35 | 33 |
| 1r63_ | 63 | 67 | 0 | 16 | 17 | 1rgs_ | 264 | 34 | 27 | 17 | 21 | 1rip_ | 81 | 0 | 11 | 28 | 60 |
| 1stu_ | 68 | 38 | 31 | 12 | 19 | 1svpA | 160 | 6 | 46 | 26 | 23 | 1tbaA | 67 | 21 | 6 | 39 | 34 |
| 1tig_ | 88 | 35 | 35 | 16 | 14 | 1tiv_ | 86 | 0 | 0 | 50 | 50 | 1tnm_ | 91 | 0 | 46 | 27 | 26 |
| 1upuA | 224 | 39 | 27 | 15 | 18 | 1xikA | 340 | 70 | 4 | 13 | 14 | 1xrc_ | 378 | 32 | 25 | 26 | 17 |
| 2af8_ | 86 | 50 | 0 | 21 | 29 | 2cgpA | 200 | 40 | 30 | 19 | 12 | 2def_ | 146 | 22 | 20 | 25 | 33 |
| 2ezl_ | 99 | 59 | 0 | 19 | 22 | 2jhbA | 143 | 26 | 25 | 24 | 24 | 2myo_ | 118 | 47 | 0 | 34 | 19 |
| 2pcbA | 294 | 49 | 7 | 22 | 21 | 2pcfB | 250 | 8 | 40 | 23 | 28 | 2pii_ | 112 | 26 | 29 | 15 | 29 |
| 2qwc_ | 385 | 3 | 45 | 25 | 26 | 2tbd_ | 134 | 28 | 27 | 21 | 24 | 2tmvP | 154 | 45 | 5 | 26 | 25 |
| 2yfpA | 224 | 9 | 53 | 24 | 14 | 3cla_ | 213 | 30 | 29 | 22 | 20 | 3csmA | 252 | 64 | 0 | 17 | 19 |
| 3sil_ | 378 | 6 | 47 | 25 | 22 | 7prcC | 332 | 42 | 4 | 30 | 24 | 8prn_ | 289 | 4 | 55 | 24 | 17 |
| 8rucI | 123 | 22 | 24 | 27 | 28 | 1a3k_ | 137 | 2 | 58 | 18 | 21 | 1a79A | 171 | 34 | 31 | 20 | 15 |
| 1a7m_ | 180 | 58 | 0 | 22 | 19 | 1a8p_ | 257 | 26 | 34 | 23 | 18 | 1aep_ | 153 | 80 | 0 | 14 | 7 |
| 1aru_ | 336 | 43 | 7 | 24 | 26 | 1avwB | 171 | 2 | 42 | 20 | 36 | 1awj_ | 77 | 0 | 5 | 27 | 68 |
| 1b3kA | 373 | 31 | 31 | 21 | 17 | 1b65A | 363 | 31 | 26 | 18 | 24 | 1bec_ | 238 | 5 | 49 | 20 | 26 |
| 1beg_ | 97 | 61 | 4 | 14 | 21 | 1bmy_ | 107 | 48 | 0 | 27 | 25 | 1bu9A | 168 | 48 | 1 | 27 | 24 |
| 1bw6A | 56 | 55 | 0 | 23 | 21 | 1bx9A | 210 | 56 | 10 | 23 | 11 | 1cd3_ | 294 | 63 | 2 | 20 | 16 |
| 1cby_ | 227 | 30 | 30 | 16 | 24 | 1cczA | 171 | 4 | 51 | 22 | 24 | 1cjkA | 189 | 38 | 30 | 18 | 15 |
| 1cmyA | 141 | 74 | 0 | 15 | 11 | 1cpzA | 68 | 28 | 29 | 28 | 15 | 1d4uA | 111 | 23 | 11 | 38 | 29 |
| 1d8lA | 140 | 39 | 29 | 19 | 14 | 1dipA | 77 | 40 | 0 | 31 | 29 | 1dj7B | 73 | 4 | 44 | 27 | 25 |
| 1dkdA | 146 | 34 | 29 | 17 | 19 | 1dztA | 183 | 10 | 44 | 21 | 25 | 1eioA | 127 | 9 | 58 | 21 | 11 |
| 1ej3A | 187 | 61 | 4 | 18 | 17 | 1eqfA | 267 | 64 | 0 | 21 | 15 | 1exg_ | 110 | 3 | 56 | 22 | 19 |
| 1gdoA | 238 | 25 | 34 | 21 | 20 | 1ghj_ | 79 | 0 | 42 | 25 | 33 | 1hhhB | 100 | 0 | 49 | 21 | 30 |
| 1irl_ | 133 | 52 | 3 | 22 | 23 | 1lkfA | 292 | 3 | 59 | 20 | 18 | 1mrj_ | 247 | 40 | 25 | 21 | 13 |
| 1mut_ | 129 | 12 | 23 | 28 | 37 | 1nflA | 259 | 58 | 0 | 29 | 13 | 1nfa_ | 178 | 2 | 24 | 26 | 49 |
| 1otfA | 59 | 42 | 32 | 10 | 15 | 1ounB | 121 | 28 | 49 | 12 | 11 | 1p32A | 182 | 36 | 26 | 17 | 20 |
| 1pdnC | 123 | 47 | 5 | 24 | 24 | 1pex_ | 192 | 7 | 42 | 33 | 18 | 1qgiA | 259 | 58 | 6 | 21 | 15 |
| 1qhgA | 163 | 56 | 10 | 20 | 15 | 1ghlA | 203 | 37 | 27 | 19 | 16 | 1qj8A | 148 | 0 | 82 | 14 | 5 |
| 1qkfA | 73 | 22 | 21 | 27 | 30 | 1qovM | 302 | 64 | 4 | 14 | 18 | 1qpvA | 133 | 30 | 30 | 23 | 17 |
| 1qu8A | 46 | 0 | 0 | 50 | 50 | 1r2aA | 46 | 63 | 0 | 11 | 26 | 1rof_ | 60 | 12 | 10 | 40 | 38 |
| 1rypL | 212 | 39 | 33 | 16 | 12 | 1sxl_ | 97 | 18 | 13 | 30 | 39 | 1tif_ | 76 | 36 | 30 | 17 | 17 |
| 1tiiD | 98 | 26 | 39 | 21 | 14 | 1tuc_ | 61 | 5 | 44 | 21 | 30 | 1u2fA | 90 | 16 | 18 | 38 | 29 |
| 1vcaA | 199 | 5 | 57 | 20 | 19 | 2abd_ | 86 | 60 | 0 | 19 | 21 | 2atcB | 152 | 6 | 5 | 41 | 48 |
| 2aviA | 121 | 2 | 51 | 22 | 24 | 2ayh_ | 214 | 6 | 52 | 19 | 23 | 2bby_ | 69 | 51 | 7 | 10 | 32 |
| 2bidA | 197 | 57 | 0 | 21 | 22 | 2nlrA | 222 | 6 | 50 | 23 | 21 | 2nmbA | 147 | 20 | 20 | 31 | 29 |
| 2shl_ | 48 | 0 | 52 | 27 | 21 | 2trxA | 108 | 33 | 28 | 27 | 12 | 3ncmA | 92 | 4 | 51 | 27 | 17 |
| 3stdA | 162 | 30 | 48 | 14 | 9 | 5daaA | 277 | 27 | 31 | 23 | 19 | 6gsvA | 217 | 49 | 9 | 28 | 14 |
Each entry is identified by its four-letter identification code, followed by the chain identifier. The following features are indicated for each entry: the number of residues (n) and the percentages of residues in helical (h), extended (e), turn (t), and other (o) backbone conformation. The secondary structures, as assigned by DSSP, were simplified as follows: helical if 310-α or π-helix (G, H, and I respectively in DSSP), extended if β-bulge or strand (B and E), turn if bend or reverse turn (S and T), and others in the remaining cases.
Fig. 1.
(a) Typical distributions of rmsd values after 400,000 random superpositions for proteins of different sizes (PDB codes indicated in parentheses); the percentage of observations in each range of 0.4 Å is reported. (b) A typical rmsd-versus-chain-length plot; the upper limits of the smallest 2000 observations (examples indicated by perpendicular lines in a) are plotted for each of the 180 experiments. (c) The dependence of the rmsd-versus-chain-length plots as a function of the different number of smallest observations (indicated in parentheses); the lines were determined by fitting a logarithmic equation of the form y = a + b ln(x) to the data (0.95 < r < 1.00); ◯ is a reference value, corresponding to 100 residues, chosen to normalize the curves. (d) Dividing the rmsd values by the corresponding reference value (indicated with ◯ in c) causes the curves in the previous figure to collapse into one single curve.
In order to check the effect of the uneven distribution of rmsd values, we prepared separate rmsd-versus-chain-length plots for different subsets of the database, selected to represent different rmsd ranges without changing the other parameters (secondary structure content, amino acid composition, etc.). This was achieved by first ordering the structural alignments in growing order of the rmsd values in each of the 180 data sets and then selecting the first n smallest rmsd values from each data set. This procedure guarantees that the data sets will be equal with respect to all parameters; only the range of the rmsd values will be different: that is, gradually increasing the number n of observations in the data sets means not only an increase of the data size but also an inclusion of higher rmsd values. The data of each subset could be fitted with a logarithmic function with correlation coefficients higher than 0.95 (an example is shown in Fig. 1b ▶). The fitted curves are different as higher rmsd data are included in the calculation, which results in the series of curves shown in Figure 1c ▶. This observation therefore confirms that the uneven distribution of rmsd values would bias the parameters obtained by simple curve fitting. Interestingly, dividing the rmsd values with a reference value, chosen here as the value of the fitted rmsd curve at 100 residues, rmsd100 (Fig. 1c ▶), makes the curves collapse into one single logarithmic curve (Fig. 1d ▶) that is described by the following equation:
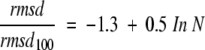 |
2 |
where N is the number of amino acid residues. This curve is accordingly independent of both the number n of observations included in the calculation and the magnitude of rmsd values; a statistical bias is therefore not likely. Given that −1.3 ≅ 1 − ln(10), the equation can be rearranged to give
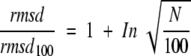 |
3 |
It is interesting to note that the value 100, the residue number corresponding to the chosen reference value, rmsd100, appears in the equation. We repeated the normalization procedure on the entire data set with residue numbers of 50, 75, 150, and 200, respectively, and in fact found that a generalized equation is valid with correlation coefficients 0.96–0.99:
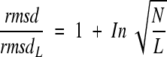 |
4 |
where L is the number of residues chosen as a reference. In other words, the relative root-mean-square distance rmsd/rmsdL is a simple function of the relative dimension N/L. Equation 3 can be simply rearranged to give a formula for a normalized rmsd value:
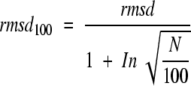 |
5 |
The chain length of 100 residues was primarily chosen because this is the mean number of amino acids per domain (Xu and Nussinov 1998). rmsd100 is therefore an rmsd value that would be observed for a pair of structures of 100 residues exhibiting the same degree of similarity as the structures actually compared. In other words, the rmsd100 value can be considered as a normalized, size-independent indicator of structural variability. For example, suppose that the Cα atoms of two pairs of protein structures, 50 and 200 residues long, respectively, can be superposed to give a final rmsd value of 1.0 Å. For the first pair of sequences sharing N = 50 equivalent residues, the corresponding rmsd100 value will be 1.524 Å The second pair of structures (N = 200) is considerably more similar to each other (rmsd100 = 0.741 Å) despite the fact that the crude rmsd values are the same. In other words, the normalized rmsd100 qualitatively reflects the intuitive view that larger structures have a higher probability to differ one from the other. Because the data were derived from proteins with more than 40 residues we suggest that the rmsd100 formula should be applied to alignments that include more than 40 residues. On the other hand, it follows from the mathematical form of the equation that the formula can be applied only for structural alignments with more than 14 residues; for smaller N values the ratio in equation 2 would be negative.
We think that the normalized rmsd can be useful in estimating the quality of an NMR ensemble of models, in applying multivariate statistical techniques to structural bioinformatic problems, as well as in comparing limited sets of protein three-dimensional structures.
Acknowledgments
We thank János Murvai and Alessandro Pintar for helpful discussions.
The publication costs of this article were defrayed in part by payment of page charges. This article must therefore be hereby marked "advertisement" in accordance with 18 USC section 1734 solely to indicate this fact.
Article and publication are at www.proteinscience.org/cgi/doi/10.1110/
References
- Berman, H., Westbrook, J., Feng, Z., Gilliland, G., Bhat, T., Weissig, H., Shindyalov, I., and Bourne, P. 2000. The Protein Data Bank. Nucleic Acid Res. 28 235–242. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Carugo, O. and Eisenhaber, F. 1997. Probabilistic evaluation of similarity between pairs of three-dimensional protein structures utilizing temperature factors. J. Appl. Cryst. 30 547–549. [Google Scholar]
- Domingues, F.S., Koppensteiner, W.A., and Sippl, M.J. 2000. The role of protein structure in genomics. FEBS Lett. 476 98–102. [DOI] [PubMed] [Google Scholar]
- Hobohm, U. and Sander, C. 1994. Enlarged representative set of protein structures. Protein Sci. 3 522–531. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Holm, L. and Sander, C. 1996 Mapping the protein universe. Science 273595–602. [DOI] [PubMed]
- Irving J.A., Whisstock J.C., and Lesk A.M. 2001. Protein structural alignments and functional genomics. Proteins 42378–382. [DOI] [PubMed] [Google Scholar]
- Kabsch, W. 1976. A solution for the best rotation to relate two sets of vectors. Acta Crystallogr. A 32 922–923. [Google Scholar]
- ———. 1978. A discussion of the solution for the best rotation to relate two sets of vectors. Acta Crystallogr. A 34 827–828. [Google Scholar]
- Peters-Libeu, C. and Adman, E.T. 1997. Displacement-parameter weighted coordinate comparison: I. Detection of significant structural differences between oxidation states. Acta Crystallogr. D 53 56–76. [DOI] [PubMed] [Google Scholar]
- Xu, D. and Nussinov, R. 1998. Favorable domain size in proteins. Fold. Des. 3 11–17. [DOI] [PubMed] [Google Scholar]
- Yang, A.-S. and Honig, B. 2000. An integrated approach to the analysis and modeling of protein sequences and structures. I. Protein structural alignment and a quantitative measure for protein structural distance. J. Mol. Biol. 301 665–678. [DOI] [PubMed] [Google Scholar]