Abstract
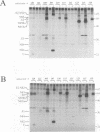
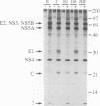
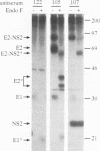
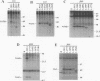
Hepatitis C virus (HCV) is the major cause of transfusion-acquired non-A, non-B hepatitis. HCV is an enveloped positive-sense RNA virus which has been classified as a new genus in the flavivirus family. Like the other two genera in this family, the flaviviruses and the pestiviruses, HCV polypeptides appear to be produced by translation of a long open reading frame and subsequent proteolytic processing of this polyprotein. In this study, a cDNA clone encompassing the long open reading frame of the HCV H strain (3,011 amino acid residues) has been assembled and sequenced. This clone and various truncated derivatives were used in vaccinia virus transient-expression assays to map HCV-encoded polypeptides and to study HCV polyprotein processing. HCV polyproteins and cleavage products were identified by using convalescent human sera and a panel of region-specific polyclonal rabbit antisera. Similar results were obtained for several mammalian cell lines examined, including the human HepG2 hepatoma line. The data indicate that at least nine polypeptides are produced by cleavage of the HCV H strain polyprotein. Putative structural proteins, located in the N-terminal one-fourth of the polyprotein, include the capsid protein C (21 kDa) followed by two possible virion envelope proteins, E1 (31 kDa) and E2 (70 kDa), which are heavily modified by N-linked glycosylation. The remainder of the polyprotein probably encodes nonstructural proteins including NS2 (23 kDa), NS3 (70 kDa), NS4A (8 kDa), NS4B (27 kDa), NS5A (58 kDa), and NS5B (68 kDa). An 82- to 88-kDa glycoprotein which reacted with both E2 and NS2-specific HCV antisera was also identified (called E2-NS2). Preliminary results suggest that a fraction of E1 is associated with E2 and E2-NS2 via disulfide linkages.
Full text
PDF





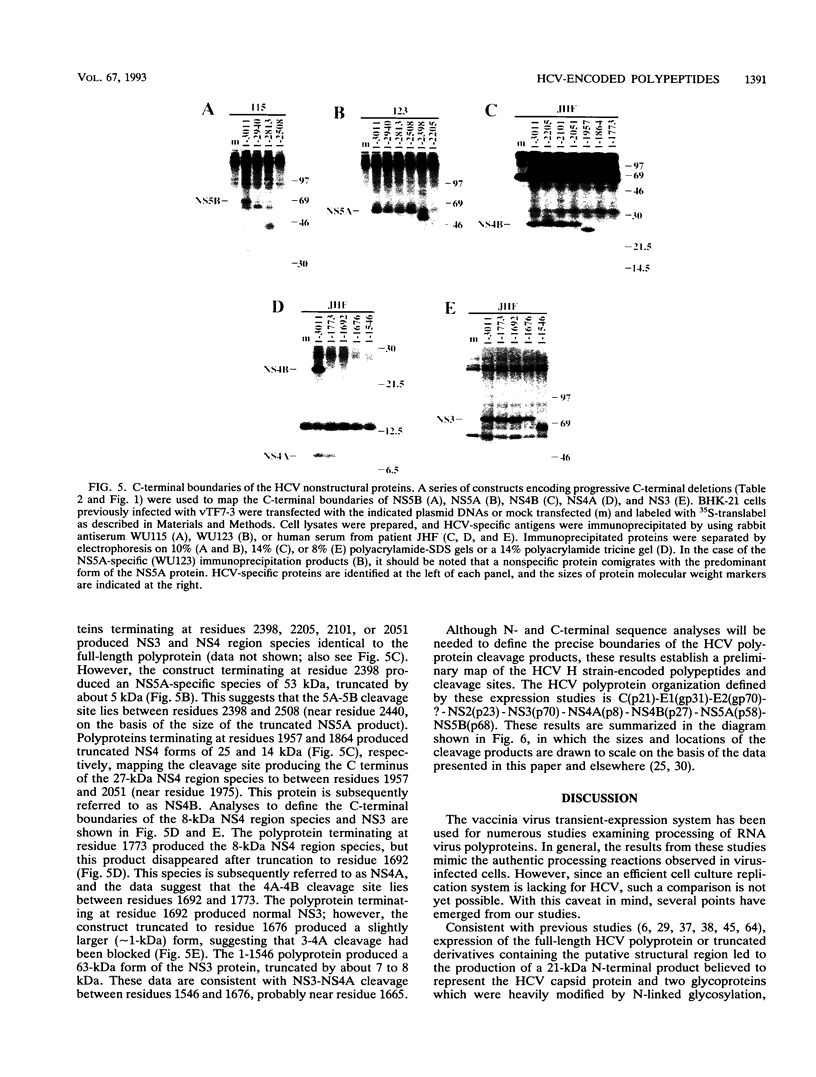




Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Bolivar F., Rodriguez R. L., Greene P. J., Betlach M. C., Heyneker H. L., Boyer H. W., Crosa J. H., Falkow S. Construction and characterization of new cloning vehicles. II. A multipurpose cloning system. Gene. 1977;2(2):95–113. [PubMed] [Google Scholar]
- Brockman W. W., Nathans D. The isolation of simian virus 40 variants with specifically altered genomes. Proc Natl Acad Sci U S A. 1974 Mar;71(3):942–946. doi: 10.1073/pnas.71.3.942. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chambers T. J., Hahn C. S., Galler R., Rice C. M. Flavivirus genome organization, expression, and replication. Annu Rev Microbiol. 1990;44:649–688. doi: 10.1146/annurev.mi.44.100190.003245. [DOI] [PubMed] [Google Scholar]
- Chambers T. J., McCourt D. W., Rice C. M. Yellow fever virus proteins NS2A, NS2B, and NS4B: identification and partial N-terminal amino acid sequence analysis. Virology. 1989 Mar;169(1):100–109. doi: 10.1016/0042-6822(89)90045-7. [DOI] [PubMed] [Google Scholar]
- Chiba J., Ohba H., Matsuura Y., Watanabe Y., Katayama T., Kikuchi S., Saito I., Miyamura T. Serodiagnosis of hepatitis C virus (HCV) infection with an HCV core protein molecularly expressed by a recombinant baculovirus. Proc Natl Acad Sci U S A. 1991 Jun 1;88(11):4641–4645. doi: 10.1073/pnas.88.11.4641. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choo Q. L., Kuo G., Weiner A. J., Overby L. R., Bradley D. W., Houghton M. Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science. 1989 Apr 21;244(4902):359–362. doi: 10.1126/science.2523562. [DOI] [PubMed] [Google Scholar]
- Choo Q. L., Richman K. H., Han J. H., Berger K., Lee C., Dong C., Gallegos C., Coit D., Medina-Selby R., Barr P. J. Genetic organization and diversity of the hepatitis C virus. Proc Natl Acad Sci U S A. 1991 Mar 15;88(6):2451–2455. doi: 10.1073/pnas.88.6.2451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Choo Q. L., Weiner A. J., Overby L. R., Kuo G., Houghton M., Bradley D. W. Hepatitis C virus: the major causative agent of viral non-A, non-B hepatitis. Br Med Bull. 1990 Apr;46(2):423–441. doi: 10.1093/oxfordjournals.bmb.a072408. [DOI] [PubMed] [Google Scholar]
- Collett M. S., Anderson D. K., Retzel E. Comparisons of the pestivirus bovine viral diarrhoea virus with members of the flaviviridae. J Gen Virol. 1988 Oct;69(Pt 10):2637–2643. doi: 10.1099/0022-1317-69-10-2637. [DOI] [PubMed] [Google Scholar]
- Collett M. S. Molecular genetics of pestiviruses. Comp Immunol Microbiol Infect Dis. 1992 Jul;15(3):145–154. doi: 10.1016/0147-9571(92)90087-8. [DOI] [PubMed] [Google Scholar]
- Colombo M., Kuo G., Choo Q. L., Donato M. F., Del Ninno E., Tommasini M. A., Dioguardi N., Houghton M. Prevalence of antibodies to hepatitis C virus in Italian patients with hepatocellular carcinoma. Lancet. 1989 Oct 28;2(8670):1006–1008. doi: 10.1016/s0140-6736(89)91016-7. [DOI] [PubMed] [Google Scholar]
- Elder J. H., Alexander S. endo-beta-N-acetylglucosaminidase F: endoglycosidase from Flavobacterium meningosepticum that cleaves both high-mannose and complex glycoproteins. Proc Natl Acad Sci U S A. 1982 Aug;79(15):4540–4544. doi: 10.1073/pnas.79.15.4540. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elroy-Stein O., Fuerst T. R., Moss B. Cap-independent translation of mRNA conferred by encephalomyocarditis virus 5' sequence improves the performance of the vaccinia virus/bacteriophage T7 hybrid expression system. Proc Natl Acad Sci U S A. 1989 Aug;86(16):6126–6130. doi: 10.1073/pnas.86.16.6126. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Falkner F. G., Moss B. Escherichia coli gpt gene provides dominant selection for vaccinia virus open reading frame expression vectors. J Virol. 1988 Jun;62(6):1849–1854. doi: 10.1128/jvi.62.6.1849-1854.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Feinstone S. M., Alter H. J., Dienes H. P., Shimizu Y., Popper H., Blackmore D., Sly D., London W. T., Purcell R. H. Non-A, non-B hepatitis in chimpanzees and marmosets. J Infect Dis. 1981 Dec;144(6):588–598. doi: 10.1093/infdis/144.6.588. [DOI] [PubMed] [Google Scholar]
- Fuerst T. R., Niles E. G., Studier F. W., Moss B. Eukaryotic transient-expression system based on recombinant vaccinia virus that synthesizes bacteriophage T7 RNA polymerase. Proc Natl Acad Sci U S A. 1986 Nov;83(21):8122–8126. doi: 10.1073/pnas.83.21.8122. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gocke D. J. A prospective study of posttransfusion hepatitis. The role of Australia Antigen. JAMA. 1972 Feb 28;219(9):1165–1170. [PubMed] [Google Scholar]
- Gocke D. J., Greenberg H. B., Kavey N. B. Correlation of Australia antigen with posttransfusion hepatitis. JAMA. 1970 May 4;212(5):877–879. [PubMed] [Google Scholar]
- Han J. H., Shyamala V., Richman K. H., Brauer M. J., Irvine B., Urdea M. S., Tekamp-Olson P., Kuo G., Choo Q. L., Houghton M. Characterization of the terminal regions of hepatitis C viral RNA: identification of conserved sequences in the 5' untranslated region and poly(A) tails at the 3' end. Proc Natl Acad Sci U S A. 1991 Mar 1;88(5):1711–1715. doi: 10.1073/pnas.88.5.1711. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Harada S., Watanabe Y., Takeuchi K., Suzuki T., Katayama T., Takebe Y., Saito I., Miyamura T. Expression of processed core protein of hepatitis C virus in mammalian cells. J Virol. 1991 Jun;65(6):3015–3021. doi: 10.1128/jvi.65.6.3015-3021.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hijikata M., Kato N., Ootsuyama Y., Nakagawa M., Shimotohno K. Gene mapping of the putative structural region of the hepatitis C virus genome by in vitro processing analysis. Proc Natl Acad Sci U S A. 1991 Jul 1;88(13):5547–5551. doi: 10.1073/pnas.88.13.5547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Houghton M., Weiner A., Han J., Kuo G., Choo Q. L. Molecular biology of the hepatitis C viruses: implications for diagnosis, development and control of viral disease. Hepatology. 1991 Aug;14(2):381–388. [PubMed] [Google Scholar]
- Hruby D. E., Guarino L. A., Kates J. R. Vaccinia virus replication. I. Requirement for the host-cell nucleus. J Virol. 1979 Feb;29(2):705–715. doi: 10.1128/jvi.29.2.705-715.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Inchauspe G., Zebedee S., Lee D. H., Sugitani M., Nasoff M., Prince A. M. Genomic structure of the human prototype strain H of hepatitis C virus: comparison with American and Japanese isolates. Proc Natl Acad Sci U S A. 1991 Nov 15;88(22):10292–10296. doi: 10.1073/pnas.88.22.10292. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kato N., Hijikata M., Ootsuyama Y., Nakagawa M., Ohkoshi S., Sugimura T., Shimotohno K. Molecular cloning of the human hepatitis C virus genome from Japanese patients with non-A, non-B hepatitis. Proc Natl Acad Sci U S A. 1990 Dec;87(24):9524–9528. doi: 10.1073/pnas.87.24.9524. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kohara M., Tsukiyama-Kohara K., Maki N., Asano K., Yamaguchi K., Miki K., Tanaka S., Hattori N., Matsuura Y., Saito I. Expression and characterization of glycoprotein gp35 of hepatitis C virus using recombinant vaccinia virus. J Gen Virol. 1992 Sep;73(Pt 9):2313–2318. doi: 10.1099/0022-1317-73-9-2313. [DOI] [PubMed] [Google Scholar]
- Kumar U., Cheng D., Thomas H., Monjardino J. Cloning and sequencing of the structural region and expression of putative core gene of hepatitis C virus from a British case of chronic sporadic hepatitis. J Gen Virol. 1992 Jun;73(Pt 6):1521–1525. doi: 10.1099/0022-1317-73-6-1521. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Laskey R. A., Mills A. D. Quantitative film detection of 3H and 14C in polyacrylamide gels by fluorography. Eur J Biochem. 1975 Aug 15;56(2):335–341. doi: 10.1111/j.1432-1033.1975.tb02238.x. [DOI] [PubMed] [Google Scholar]
- Mackett M., Smith G. L. Vaccinia virus expression vectors. J Gen Virol. 1986 Oct;67(Pt 10):2067–2082. doi: 10.1099/0022-1317-67-10-2067. [DOI] [PubMed] [Google Scholar]
- Martell M., Esteban J. I., Quer J., Genescà J., Weiner A., Esteban R., Guardia J., Gómez J. Hepatitis C virus (HCV) circulates as a population of different but closely related genomes: quasispecies nature of HCV genome distribution. J Virol. 1992 May;66(5):3225–3229. doi: 10.1128/jvi.66.5.3225-3229.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Mason P. W. Maturation of Japanese encephalitis virus glycoproteins produced by infected mammalian and mosquito cells. Virology. 1989 Apr;169(2):354–364. doi: 10.1016/0042-6822(89)90161-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Matsuura Y., Harada S., Suzuki R., Watanabe Y., Inoue Y., Saito I., Miyamura T. Expression of processed envelope protein of hepatitis C virus in mammalian and insect cells. J Virol. 1992 Mar;66(3):1425–1431. doi: 10.1128/jvi.66.3.1425-1431.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers G., Tautz N., Stark R., Brownlie J., Dubovi E. J., Collett M. S., Thiel H. J. Rearrangement of viral sequences in cytopathogenic pestiviruses. Virology. 1992 Nov;191(1):368–386. doi: 10.1016/0042-6822(92)90199-Y. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Miller R. H., Purcell R. H. Hepatitis C virus shares amino acid sequence similarity with pestiviruses and flaviviruses as well as members of two plant virus supergroups. Proc Natl Acad Sci U S A. 1990 Mar;87(6):2057–2061. doi: 10.1073/pnas.87.6.2057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Moss B., Elroy-Stein O., Mizukami T., Alexander W. A., Fuerst T. R. Product review. New mammalian expression vectors. Nature. 1990 Nov 1;348(6296):91–92. doi: 10.1038/348091a0. [DOI] [PubMed] [Google Scholar]
- Negro F., Pacchioni D., Shimizu Y., Miller R. H., Bussolati G., Purcell R. H., Bonino F. Detection of intrahepatic replication of hepatitis C virus RNA by in situ hybridization and comparison with histopathology. Proc Natl Acad Sci U S A. 1992 Mar 15;89(6):2247–2251. doi: 10.1073/pnas.89.6.2247. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ogata N., Alter H. J., Miller R. H., Purcell R. H. Nucleotide sequence and mutation rate of the H strain of hepatitis C virus. Proc Natl Acad Sci U S A. 1991 Apr 15;88(8):3392–3396. doi: 10.1073/pnas.88.8.3392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Okamoto H., Kojima M., Okada S., Yoshizawa H., Iizuka H., Tanaka T., Muchmore E. E., Peterson D. A., Ito Y., Mishiro S. Genetic drift of hepatitis C virus during an 8.2-year infection in a chimpanzee: variability and stability. Virology. 1992 Oct;190(2):894–899. doi: 10.1016/0042-6822(92)90933-g. [DOI] [PubMed] [Google Scholar]
- Okamoto H., Okada S., Sugiyama Y., Kurai K., Iizuka H., Machida A., Miyakawa Y., Mayumi M. Nucleotide sequence of the genomic RNA of hepatitis C virus isolated from a human carrier: comparison with reported isolates for conserved and divergent regions. J Gen Virol. 1991 Nov;72(Pt 11):2697–2704. doi: 10.1099/0022-1317-72-11-2697. [DOI] [PubMed] [Google Scholar]
- Poch O., Sauvaget I., Delarue M., Tordo N. Identification of four conserved motifs among the RNA-dependent polymerase encoding elements. EMBO J. 1989 Dec 1;8(12):3867–3874. doi: 10.1002/j.1460-2075.1989.tb08565.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Purcell R. H., Walsh J. H., Holland P. V., Morrow A. G., Wood S., Chanock R. M. Seroepidemiological studies of transfusion-associated hepatitis. J Infect Dis. 1971 Apr;123(4):406–413. doi: 10.1093/infdis/123.4.406. [DOI] [PubMed] [Google Scholar]
- Rice C. M., Lenches E. M., Eddy S. R., Shin S. J., Sheets R. L., Strauss J. H. Nucleotide sequence of yellow fever virus: implications for flavivirus gene expression and evolution. Science. 1985 Aug 23;229(4715):726–733. doi: 10.1126/science.4023707. [DOI] [PubMed] [Google Scholar]
- Rice C. M., Strauss J. H. Association of sindbis virion glycoproteins and their precursors. J Mol Biol. 1982 Jan 15;154(2):325–348. doi: 10.1016/0022-2836(82)90067-5. [DOI] [PubMed] [Google Scholar]
- Rümenapf T., Stark R., Meyers G., Thiel H. J. Structural proteins of hog cholera virus expressed by vaccinia virus: further characterization and induction of protective immunity. J Virol. 1991 Feb;65(2):589–597. doi: 10.1128/jvi.65.2.589-597.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Saiki R. K., Gelfand D. H., Stoffel S., Scharf S. J., Higuchi R., Horn G. T., Mullis K. B., Erlich H. A. Primer-directed enzymatic amplification of DNA with a thermostable DNA polymerase. Science. 1988 Jan 29;239(4839):487–491. doi: 10.1126/science.2448875. [DOI] [PubMed] [Google Scholar]
- Saito I., Miyamura T., Ohbayashi A., Harada H., Katayama T., Kikuchi S., Watanabe Y., Koi S., Onji M., Ohta Y. Hepatitis C virus infection is associated with the development of hepatocellular carcinoma. Proc Natl Acad Sci U S A. 1990 Sep;87(17):6547–6549. doi: 10.1073/pnas.87.17.6547. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schägger H., von Jagow G. Tricine-sodium dodecyl sulfate-polyacrylamide gel electrophoresis for the separation of proteins in the range from 1 to 100 kDa. Anal Biochem. 1987 Nov 1;166(2):368–379. doi: 10.1016/0003-2697(87)90587-2. [DOI] [PubMed] [Google Scholar]
- Spaete R. R., Alexander D., Rugroden M. E., Choo Q. L., Berger K., Crawford K., Kuo C., Leng S., Lee C., Ralston R. Characterization of the hepatitis C virus E2/NS1 gene product expressed in mammalian cells. Virology. 1992 Jun;188(2):819–830. doi: 10.1016/0042-6822(92)90537-y. [DOI] [PubMed] [Google Scholar]
- Studier F. W., Rosenberg A. H., Dunn J. J., Dubendorff J. W. Use of T7 RNA polymerase to direct expression of cloned genes. Methods Enzymol. 1990;185:60–89. doi: 10.1016/0076-6879(90)85008-c. [DOI] [PubMed] [Google Scholar]
- Takamizawa A., Mori C., Fuke I., Manabe S., Murakami S., Fujita J., Onishi E., Andoh T., Yoshida I., Okayama H. Structure and organization of the hepatitis C virus genome isolated from human carriers. J Virol. 1991 Mar;65(3):1105–1113. doi: 10.1128/jvi.65.3.1105-1113.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Tanaka T., Kato N., Nakagawa M., Ootsuyama Y., Cho M. J., Nakazawa T., Hijikata M., Ishimura Y., Shimotohno K. Molecular cloning of hepatitis C virus genome from a single Japanese carrier: sequence variation within the same individual and among infected individuals. Virus Res. 1992 Apr;23(1-2):39–53. doi: 10.1016/0168-1702(92)90066-i. [DOI] [PubMed] [Google Scholar]
- Thiel H. J., Stark R., Weiland E., Rümenapf T., Meyers G. Hog cholera virus: molecular composition of virions from a pestivirus. J Virol. 1991 Sep;65(9):4705–4712. doi: 10.1128/jvi.65.9.4705-4712.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiland E., Stark R., Haas B., Rümenapf T., Meyers G., Thiel H. J. Pestivirus glycoprotein which induces neutralizing antibodies forms part of a disulfide-linked heterodimer. J Virol. 1990 Aug;64(8):3563–3569. doi: 10.1128/jvi.64.8.3563-3569.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiner A. J., Brauer M. J., Rosenblatt J., Richman K. H., Tung J., Crawford K., Bonino F., Saracco G., Choo Q. L., Houghton M. Variable and hypervariable domains are found in the regions of HCV corresponding to the flavivirus envelope and NS1 proteins and the pestivirus envelope glycoproteins. Virology. 1991 Feb;180(2):842–848. doi: 10.1016/0042-6822(91)90104-j. [DOI] [PubMed] [Google Scholar]
- Weiner A. J., Kuo G., Bradley D. W., Bonino F., Saracco G., Lee C., Rosenblatt J., Choo Q. L., Houghton M. Detection of hepatitis C viral sequences in non-A, non-B hepatitis. Lancet. 1990 Jan 6;335(8680):1–3. doi: 10.1016/0140-6736(90)90134-q. [DOI] [PubMed] [Google Scholar]
- Wengler G., Wengler G. Cell-associated West Nile flavivirus is covered with E+pre-M protein heterodimers which are destroyed and reorganized by proteolytic cleavage during virus release. J Virol. 1989 Jun;63(6):2521–2526. doi: 10.1128/jvi.63.6.2521-2526.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wengler G., Wengler G. The carboxy-terminal part of the NS 3 protein of the West Nile flavivirus can be isolated as a soluble protein after proteolytic cleavage and represents an RNA-stimulated NTPase. Virology. 1991 Oct;184(2):707–715. doi: 10.1016/0042-6822(91)90440-m. [DOI] [PubMed] [Google Scholar]
- Wiskerchen M., Collett M. S. Pestivirus gene expression: protein p80 of bovine viral diarrhea virus is a proteinase involved in polyprotein processing. Virology. 1991 Sep;184(1):341–350. doi: 10.1016/0042-6822(91)90850-b. [DOI] [PubMed] [Google Scholar]