Abstract
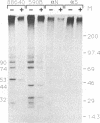
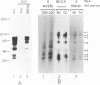
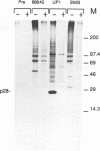
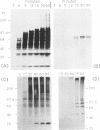
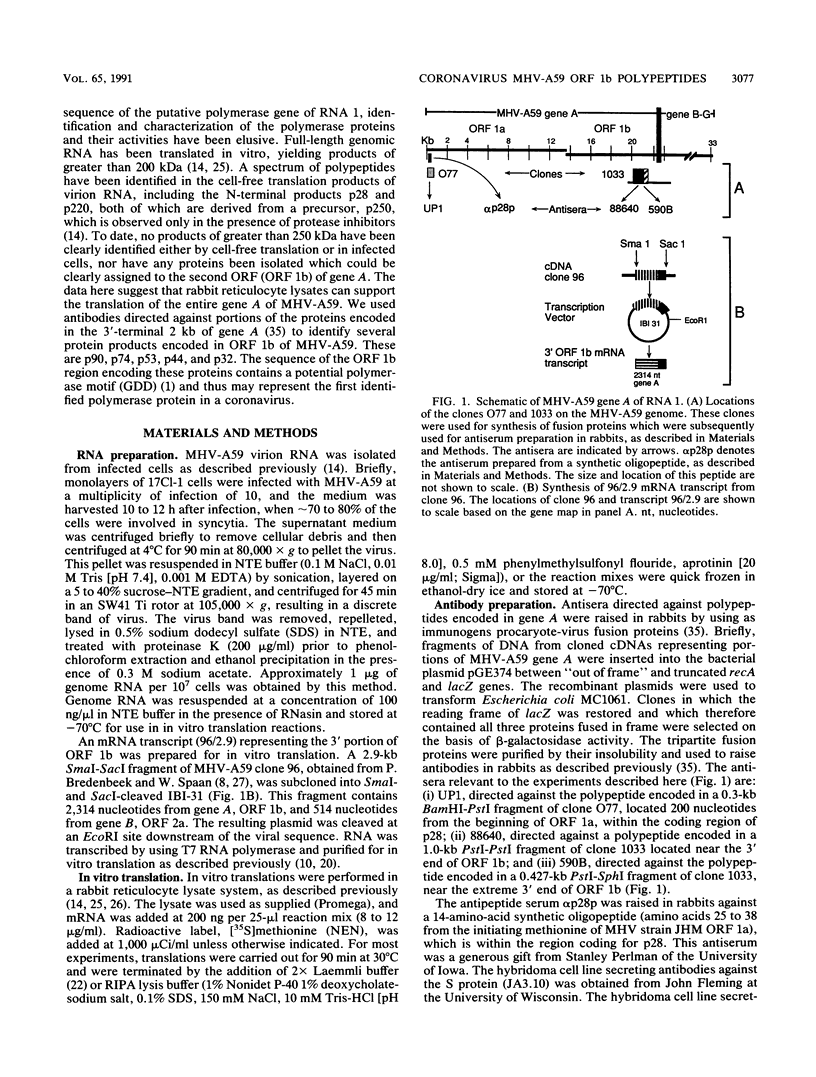
The polypeptides encoded in open reading frame (ORF) 1b of the mouse hepatitis virus A59 putative polymerase gene of RNA 1 were identified in the products of in vitro translation of genome RNA. Two antisera directed against fusion proteins containing sequences encoded in portions of the 3'-terminal 2.0 kb of ORF 1b were used to immunoprecipitate p90, p74, p53, p44, and p32 polypeptides. These polypeptides were clearly different in electrophoretic mobility, antiserum reactivity, and partial protease digestion pattern from viral structural proteins and from polypeptides encoded in the 5' end of ORF 1a, previously identified by in vitro translation. The largest of these polypeptides had partial protease digestion patterns similar to those of polypeptides generated by in vitro translation of a synthetic mRNA derived from the 3' end of ORF 1b. The polypeptides encoded in ORF 1b accumulated more slowly during in vitro translation than polypeptides encoded in ORF 1a. This is consistent with the hypothesis that translation of gene A initiates at the 5' end of ORF 1a and that translation of ORF 1b occurs following a frameshift at the ORF 1a-ORF 1b junction. The use of in vitro translation of genome RNA and immunoprecipitation with antisera directed against various regions of the polypeptides encoded in gene A should make it possible to study synthesis and processing of the putative coronavirus polymerase.
Full text
PDF


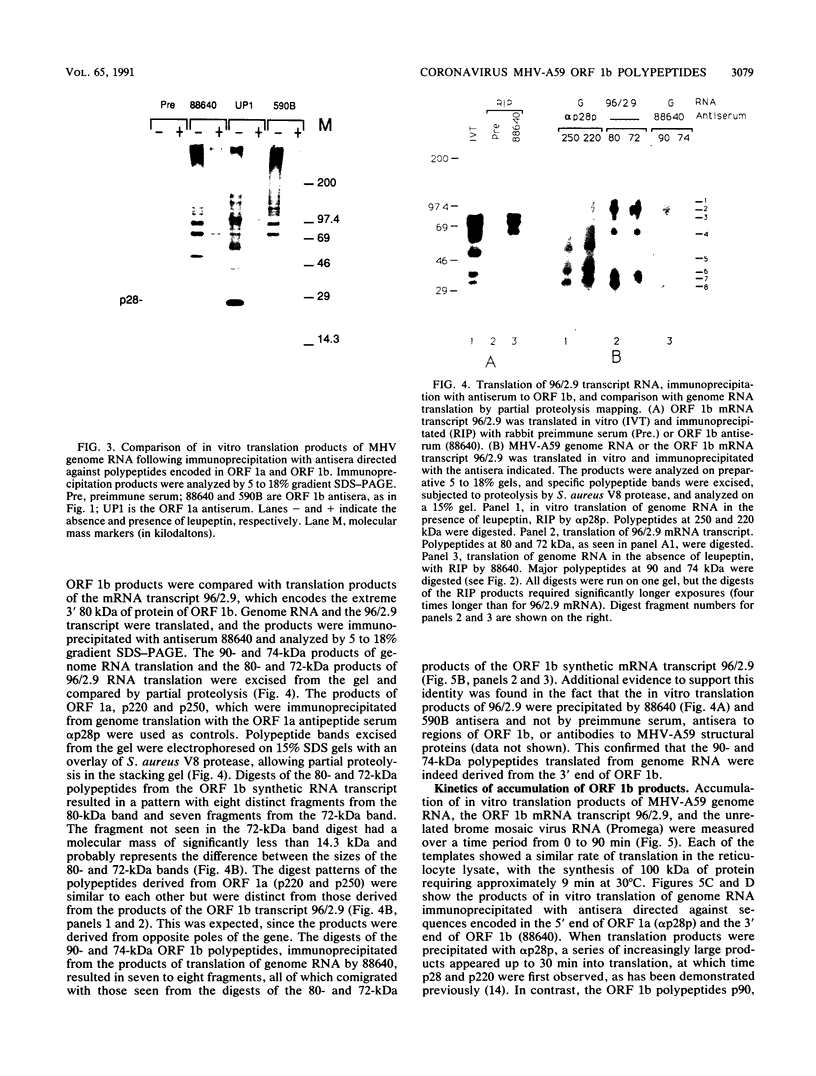
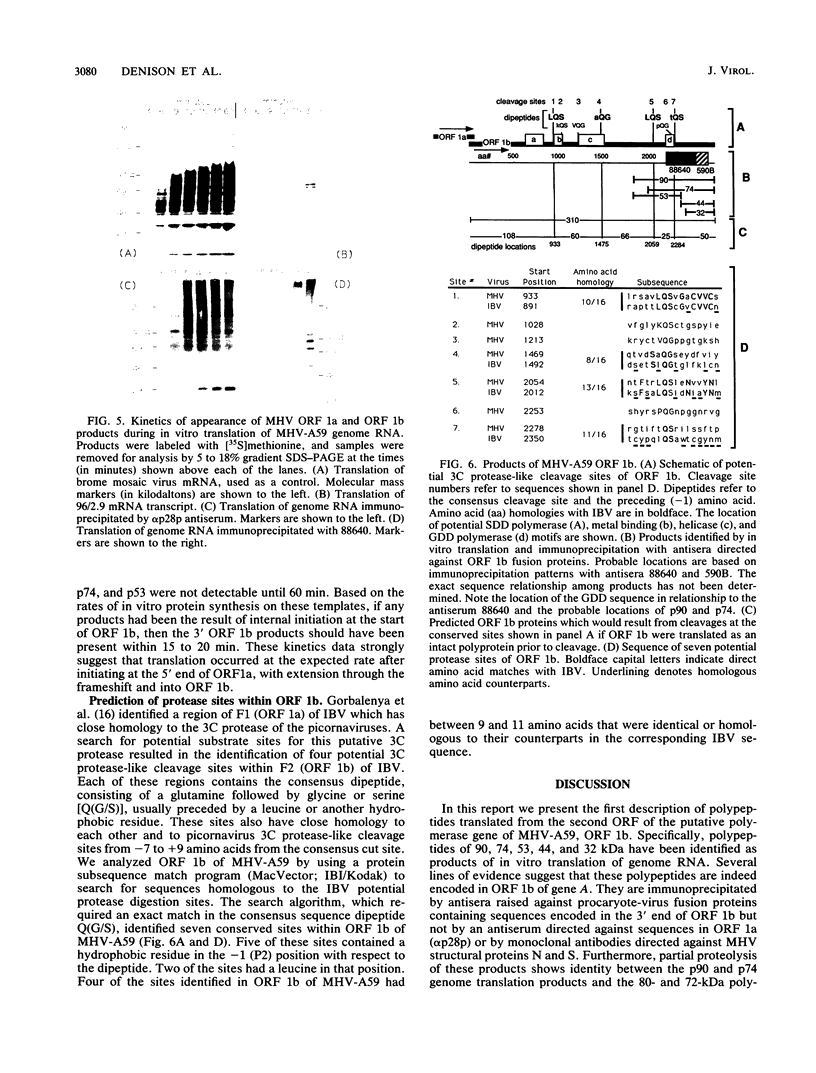


Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Argos P. A sequence motif in many polymerases. Nucleic Acids Res. 1988 Nov 11;16(21):9909–9916. doi: 10.1093/nar/16.21.9909. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker S. C., Shieh C. K., Soe L. H., Chang M. F., Vannier D. M., Lai M. M. Identification of a domain required for autoproteolytic cleavage of murine coronavirus gene A polyprotein. J Virol. 1989 Sep;63(9):3693–3699. doi: 10.1128/jvi.63.9.3693-3699.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baric R. S., Shieh C. K., Stohlman S. A., Lai M. M. Studies into the mechanism of MHV transcription. Adv Exp Med Biol. 1987;218:137–149. doi: 10.1007/978-1-4684-1280-2_16. [DOI] [PubMed] [Google Scholar]
- Baric R. S., Stohlman S. A., Razavi M. K., Lai M. M. Characterization of leader-related small RNAs in coronavirus-infected cells: further evidence for leader-primed mechanism of transcription. Virus Res. 1985 Jul;3(1):19–33. doi: 10.1016/0168-1702(85)90038-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Boursnell M. E., Brown T. D., Foulds I. J., Green P. F., Tomley F. M., Binns M. M. Completion of the sequence of the genome of the coronavirus avian infectious bronchitis virus. J Gen Virol. 1987 Jan;68(Pt 1):57–77. doi: 10.1099/0022-1317-68-1-57. [DOI] [PubMed] [Google Scholar]
- Brayton P. R., Lai M. M., Patton C. D., Stohlman S. A. Characterization of two RNA polymerase activities induced by mouse hepatitis virus. J Virol. 1982 Jun;42(3):847–853. doi: 10.1128/jvi.42.3.847-853.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brayton P. R., Stohlman S. A., Lai M. M. Further characterization of mouse hepatitis virus RNA-dependent RNA polymerases. Virology. 1984 Feb;133(1):197–201. doi: 10.1016/0042-6822(84)90439-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bredenbeek P. J., Pachuk C. J., Noten A. F., Charité J., Luytjes W., Weiss S. R., Spaan W. J. The primary structure and expression of the second open reading frame of the polymerase gene of the coronavirus MHV-A59; a highly conserved polymerase is expressed by an efficient ribosomal frameshifting mechanism. Nucleic Acids Res. 1990 Apr 11;18(7):1825–1832. doi: 10.1093/nar/18.7.1825. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brierley I., Boursnell M. E., Binns M. M., Bilimoria B., Blok V. C., Brown T. D., Inglis S. C. An efficient ribosomal frame-shifting signal in the polymerase-encoding region of the coronavirus IBV. EMBO J. 1987 Dec 1;6(12):3779–3785. doi: 10.1002/j.1460-2075.1987.tb02713.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Budzilowicz C. J., Weiss S. R. In vitro synthesis of two polypeptides from a nonstructural gene of coronavirus mouse hepatitis virus strain A59. Virology. 1987 Apr;157(2):509–515. doi: 10.1016/0042-6822(87)90293-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Budzilowicz C. J., Wilczynski S. P., Weiss S. R. Three intergenic regions of coronavirus mouse hepatitis virus strain A59 genome RNA contain a common nucleotide sequence that is homologous to the 3' end of the viral mRNA leader sequence. J Virol. 1985 Mar;53(3):834–840. doi: 10.1128/jvi.53.3.834-840.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chou P. Y., Fasman G. D. Prediction of the secondary structure of proteins from their amino acid sequence. Adv Enzymol Relat Areas Mol Biol. 1978;47:45–148. doi: 10.1002/9780470122921.ch2. [DOI] [PubMed] [Google Scholar]
- Cleveland D. W., Fischer S. G., Kirschner M. W., Laemmli U. K. Peptide mapping by limited proteolysis in sodium dodecyl sulfate and analysis by gel electrophoresis. J Biol Chem. 1977 Feb 10;252(3):1102–1106. [PubMed] [Google Scholar]
- Denison M. R., Perlman S. Translation and processing of mouse hepatitis virus virion RNA in a cell-free system. J Virol. 1986 Oct;60(1):12–18. doi: 10.1128/jvi.60.1.12-18.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Engelman D. M., Steitz T. A., Goldman A. Identifying nonpolar transbilayer helices in amino acid sequences of membrane proteins. Annu Rev Biophys Biophys Chem. 1986;15:321–353. doi: 10.1146/annurev.bb.15.060186.001541. [DOI] [PubMed] [Google Scholar]
- Gorbalenya A. E., Koonin E. V., Donchenko A. P., Blinov V. M. Coronavirus genome: prediction of putative functional domains in the non-structural polyprotein by comparative amino acid sequence analysis. Nucleic Acids Res. 1989 Jun 26;17(12):4847–4861. doi: 10.1093/nar/17.12.4847. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hopp T. P., Woods K. R. Prediction of protein antigenic determinants from amino acid sequences. Proc Natl Acad Sci U S A. 1981 Jun;78(6):3824–3828. doi: 10.1073/pnas.78.6.3824. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Jacks T., Varmus H. E. Expression of the Rous sarcoma virus pol gene by ribosomal frameshifting. Science. 1985 Dec 13;230(4731):1237–1242. doi: 10.1126/science.2416054. [DOI] [PubMed] [Google Scholar]
- Janin J., Wodak S. Conformation of amino acid side-chains in proteins. J Mol Biol. 1978 Nov 5;125(3):357–386. doi: 10.1016/0022-2836(78)90408-4. [DOI] [PubMed] [Google Scholar]
- Krieg P. A., Melton D. A. Functional messenger RNAs are produced by SP6 in vitro transcription of cloned cDNAs. Nucleic Acids Res. 1984 Sep 25;12(18):7057–7070. doi: 10.1093/nar/12.18.7057. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kyte J., Doolittle R. F. A simple method for displaying the hydropathic character of a protein. J Mol Biol. 1982 May 5;157(1):105–132. doi: 10.1016/0022-2836(82)90515-0. [DOI] [PubMed] [Google Scholar]
- Laemmli U. K. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970 Aug 15;227(5259):680–685. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- Lai M. M., Patton C. D., Stohlman S. A. Further characterization of mRNA's of mouse hepatitis virus: presence of common 5'-end nucleotides. J Virol. 1982 Feb;41(2):557–565. doi: 10.1128/jvi.41.2.557-565.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leibowitz J. L., DeVries J. R., Rodriguez M. Increased hepatotropism of mutants of MHV, strain JHM, selected with monoclonal antibodies. Adv Exp Med Biol. 1987;218:321–331. doi: 10.1007/978-1-4684-1280-2_41. [DOI] [PubMed] [Google Scholar]
- Leibowitz J. L., Weiss S. R., Paavola E., Bond C. W. Cell-free translation of murine coronavirus RNA. J Virol. 1982 Sep;43(3):905–913. doi: 10.1128/jvi.43.3.905-913.1982. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merrick W. C. Translation of exogenous mRNAs in reticulocyte lysates. Methods Enzymol. 1983;101:606–615. doi: 10.1016/0076-6879(83)01041-1. [DOI] [PubMed] [Google Scholar]
- Pachuk C. J., Bredenbeek P. J., Zoltick P. W., Spaan W. J., Weiss S. R. Molecular cloning of the gene encoding the putative polymerase of mouse hepatitis coronavirus, strain A59. Virology. 1989 Jul;171(1):141–148. doi: 10.1016/0042-6822(89)90520-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Palmenberg A. C. Picornaviral processing: some new ideas. J Cell Biochem. 1987 Mar;33(3):191–198. doi: 10.1002/jcb.240330306. [DOI] [PubMed] [Google Scholar]
- Sawicki S. G., Sawicki D. L. Coronavirus minus-strand RNA synthesis and effect of cycloheximide on coronavirus RNA synthesis. J Virol. 1986 Jan;57(1):328–334. doi: 10.1128/jvi.57.1.328-334.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sawicki S. G., Sawicki D. L. Coronavirus transcription: subgenomic mouse hepatitis virus replicative intermediates function in RNA synthesis. J Virol. 1990 Mar;64(3):1050–1056. doi: 10.1128/jvi.64.3.1050-1056.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sethna P. B., Hung S. L., Brian D. A. Coronavirus subgenomic minus-strand RNAs and the potential for mRNA replicons. Proc Natl Acad Sci U S A. 1989 Jul;86(14):5626–5630. doi: 10.1073/pnas.86.14.5626. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soe L. H., Shieh C. K., Baker S. C., Chang M. F., Lai M. M. Sequence and translation of the murine coronavirus 5'-end genomic RNA reveals the N-terminal structure of the putative RNA polymerase. J Virol. 1987 Dec;61(12):3968–3976. doi: 10.1128/jvi.61.12.3968-3976.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spaan W. J., Rottier P. J., Horzinek M. C., van der Zeijst B. A. Isolation and identification of virus-specific mRNAs in cells infected with mouse hepatitis virus (MHV-A59). Virology. 1981 Jan 30;108(2):424–434. doi: 10.1016/0042-6822(81)90449-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Spaan W., Cavanagh D., Horzinek M. C. Coronaviruses: structure and genome expression. J Gen Virol. 1988 Dec;69(Pt 12):2939–2952. doi: 10.1099/0022-1317-69-12-2939. [DOI] [PubMed] [Google Scholar]
- Zoltick P. W., Leibowitz J. L., DeVries J. R., Weinstock G. M., Weiss S. R. A general method for the induction and screening of antisera for cDNA-encoded polypeptides: antibodies specific for a coronavirus putative polymerase-encoding gene. Gene. 1989 Dec 28;85(2):413–420. doi: 10.1016/0378-1119(89)90434-4. [DOI] [PMC free article] [PubMed] [Google Scholar]