Abstract
Mouse hepatitis virus (MHV), a coronavirus, has been shown to undergo a high frequency of RNA recombination both in tissue culture and in animal infection. So far, RNA recombination has been demonstrated only between genomic RNAs of two coinfecting viruses. To understand the mechanism of RNA recombination and to further explore the potential of RNA recombination, we studied whether recombination could occur between a replicating MHV RNA and transfected RNA fragments. We first used RNA fragments which represented the 5' end of genomic-sense sequences of MHV RNA for transfection. By using polymerase chain reaction amplification with two specific primers, we were able to detect recombinant RNAs which incorporated the transfected fragment into the 5' end of the viral RNA in the infected cells. Surprisingly, even the anti-genomic-sense RNA fragments complementary to the 5' end of MHV genomic RNA could also recombine with the MHV genomic RNAs. This observation suggests that RNA recombination can occur during both positive- and negative-strand RNA synthesis. Furthermore, the recombinant RNAs could be detected in the virion released from the infected cells even after several passages of virus in tissue culture cells, indicating that these recombinant RNAs represented functional virion RNAs. The crossover sites of these recombinants were detected throughout the transfected RNA fragments. However, when an RNA fragment with a nine-nucleotide (CUUUAUAAA) deletion immediately downstream of a pentanucleotide (UCUAA) repeat sequence in the leader RNA was transfected into MHV-infected cells, most of the recombinants between this RNA and the MHV genome contained crossover sites near this pentanucleotide repeat sequence. In contrast, when exogenous RNAs with the intact nine-nucleotide sequence were used in similar experiments, the crossover sites of recombinants in viral genomic RNA could be detected at more-downstream sites. This study demonstrated that recombination can occur between replicating MHV RNAs and RNA fragments which do not replicate, suggesting the potential of RNA recombination for genetic engineering.
Full text
PDF
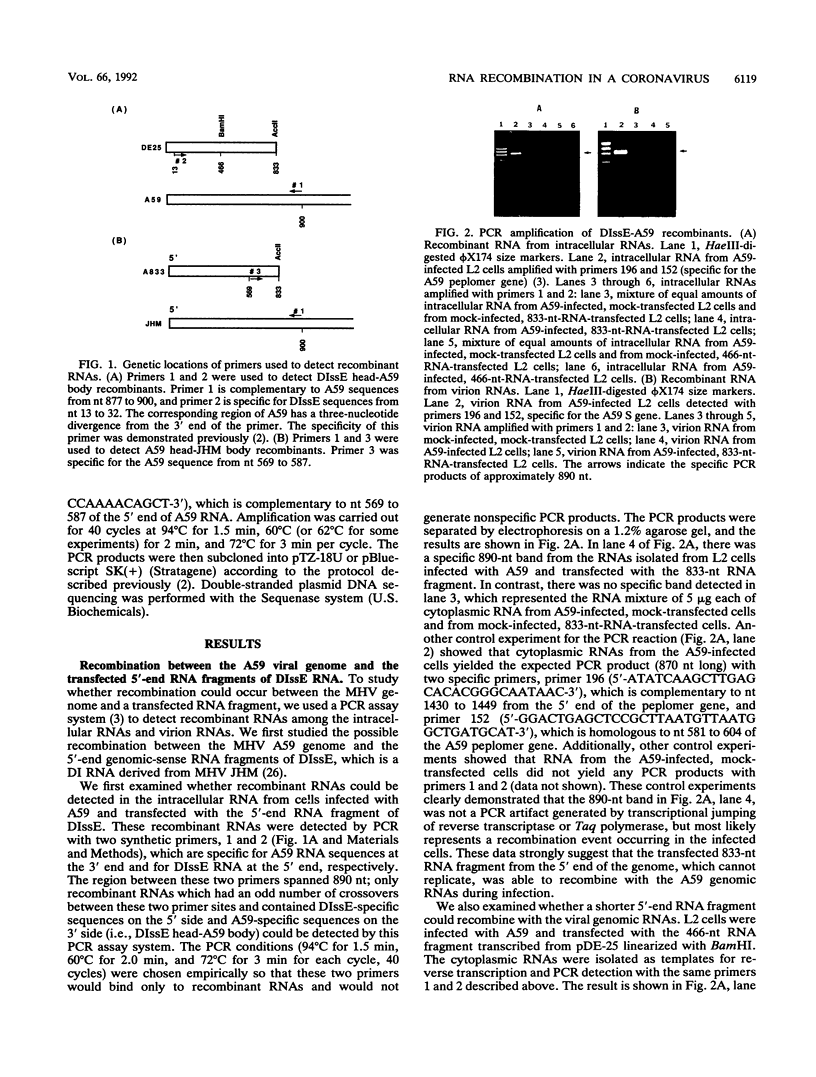







Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Allison R., Thompson C., Ahlquist P. Regeneration of a functional RNA virus genome by recombination between deletion mutants and requirement for cowpea chlorotic mottle virus 3a and coat genes for systemic infection. Proc Natl Acad Sci U S A. 1990 Mar;87(5):1820–1824. doi: 10.1073/pnas.87.5.1820. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baker S. C., Lai M. M. An in vitro system for the leader-primed transcription of coronavirus mRNAs. EMBO J. 1990 Dec;9(12):4173–4179. doi: 10.1002/j.1460-2075.1990.tb07641.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Banner L. R., Lai M. M. Random nature of coronavirus RNA recombination in the absence of selection pressure. Virology. 1991 Nov;185(1):441–445. doi: 10.1016/0042-6822(91)90795-D. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baric R. S., Fu K., Schaad M. C., Stohlman S. A. Establishing a genetic recombination map for murine coronavirus strain A59 complementation groups. Virology. 1990 Aug;177(2):646–656. doi: 10.1016/0042-6822(90)90530-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baric R. S., Shieh C. K., Stohlman S. A., Lai M. M. Analysis of intracellular small RNAs of mouse hepatitis virus: evidence for discontinuous transcription. Virology. 1987 Feb;156(2):342–354. doi: 10.1016/0042-6822(87)90414-4. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baric R. S., Stohlman S. A., Razavi M. K., Lai M. M. Characterization of leader-related small RNAs in coronavirus-infected cells: further evidence for leader-primed mechanism of transcription. Virus Res. 1985 Jul;3(1):19–33. doi: 10.1016/0168-1702(85)90038-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bujarski J. J., Dzianott A. M. Generation and analysis of nonhomologous RNA-RNA recombinants in brome mosaic virus: sequence complementarities at crossover sites. J Virol. 1991 Aug;65(8):4153–4159. doi: 10.1128/jvi.65.8.4153-4159.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bujarski J. J., Kaesberg P. Genetic recombination between RNA components of a multipartite plant virus. 1986 May 29-Jun 4Nature. 321(6069):528–531. doi: 10.1038/321528a0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Copper P. D., Steiner-Pryor A., Scotti P. D., Delong D. On the nature of poliovirus genetic recombinants. J Gen Virol. 1974 Apr;23(1):41–49. doi: 10.1099/0022-1317-23-1-41. [DOI] [PubMed] [Google Scholar]
- Hirano N., Fujiwara K., Hino S., Matumoto M. Replication and plaque formation of mouse hepatitis virus (MHV-2) in mouse cell line DBT culture. Arch Gesamte Virusforsch. 1974;44(3):298–302. doi: 10.1007/BF01240618. [DOI] [PubMed] [Google Scholar]
- Holland J., Spindler K., Horodyski F., Grabau E., Nichol S., VandePol S. Rapid evolution of RNA genomes. Science. 1982 Mar 26;215(4540):1577–1585. doi: 10.1126/science.7041255. [DOI] [PubMed] [Google Scholar]
- Keck J. G., Matsushima G. K., Makino S., Fleming J. O., Vannier D. M., Stohlman S. A., Lai M. M. In vivo RNA-RNA recombination of coronavirus in mouse brain. J Virol. 1988 May;62(5):1810–1813. doi: 10.1128/jvi.62.5.1810-1813.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keck J. G., Soe L. H., Makino S., Stohlman S. A., Lai M. M. RNA recombination of murine coronaviruses: recombination between fusion-positive mouse hepatitis virus A59 and fusion-negative mouse hepatitis virus 2. J Virol. 1988 Jun;62(6):1989–1998. doi: 10.1128/jvi.62.6.1989-1998.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Keck J. G., Stohlman S. A., Soe L. H., Makino S., Lai M. M. Multiple recombination sites at the 5'-end of murine coronavirus RNA. Virology. 1987 Feb;156(2):331–341. doi: 10.1016/0042-6822(87)90413-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King A. M., McCahon D., Saunders K., Newman J. W., Slade W. R. Multiple sites of recombination within the RNA genome of foot-and-mouth disease virus. Virus Res. 1985 Nov;3(4):373–384. doi: 10.1016/0168-1702(85)90437-X. [DOI] [PMC free article] [PubMed] [Google Scholar]
- King A. M., McCahon D., Slade W. R., Newman J. W. Recombination in RNA. Cell. 1982 Jul;29(3):921–928. doi: 10.1016/0092-8674(82)90454-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kirkegaard K., Baltimore D. The mechanism of RNA recombination in poliovirus. Cell. 1986 Nov 7;47(3):433–443. doi: 10.1016/0092-8674(86)90600-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Koetzner C. A., Parker M. M., Ricard C. S., Sturman L. S., Masters P. S. Repair and mutagenesis of the genome of a deletion mutant of the coronavirus mouse hepatitis virus by targeted RNA recombination. J Virol. 1992 Apr;66(4):1841–1848. doi: 10.1128/jvi.66.4.1841-1848.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai M. M., Baric R. S., Makino S., Keck J. G., Egbert J., Leibowitz J. L., Stohlman S. A. Recombination between nonsegmented RNA genomes of murine coronaviruses. J Virol. 1985 Nov;56(2):449–456. doi: 10.1128/jvi.56.2.449-456.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai M. M., Brayton P. R., Armen R. C., Patton C. D., Pugh C., Stohlman S. A. Mouse hepatitis virus A59: mRNA structure and genetic localization of the sequence divergence from hepatotropic strain MHV-3. J Virol. 1981 Sep;39(3):823–834. doi: 10.1128/jvi.39.3.823-834.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lai M. M. Coronavirus: organization, replication and expression of genome. Annu Rev Microbiol. 1990;44:303–333. doi: 10.1146/annurev.mi.44.100190.001511. [DOI] [PubMed] [Google Scholar]
- Lai M. M. RNA recombination in animal and plant viruses. Microbiol Rev. 1992 Mar;56(1):61–79. doi: 10.1128/mr.56.1.61-79.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lee H. J., Shieh C. K., Gorbalenya A. E., Koonin E. V., La Monica N., Tuler J., Bagdzhadzhyan A., Lai M. M. The complete sequence (22 kilobases) of murine coronavirus gene 1 encoding the putative proteases and RNA polymerase. Virology. 1991 Feb;180(2):567–582. doi: 10.1016/0042-6822(91)90071-I. [DOI] [PMC free article] [PubMed] [Google Scholar]
- MANAKER R. A., PICZAK C. V., MILLER A. A., STANTON M. F. A hepatitis virus complicating studies with mouse leukemia. J Natl Cancer Inst. 1961 Jul;27:29–51. [PubMed] [Google Scholar]
- Makino S., Fujioka N., Fujiwara K. Structure of the intracellular defective viral RNAs of defective interfering particles of mouse hepatitis virus. J Virol. 1985 May;54(2):329–336. doi: 10.1128/jvi.54.2.329-336.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino S., Keck J. G., Stohlman S. A., Lai M. M. High-frequency RNA recombination of murine coronaviruses. J Virol. 1986 Mar;57(3):729–737. doi: 10.1128/jvi.57.3.729-737.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino S., Lai M. M. High-frequency leader sequence switching during coronavirus defective interfering RNA replication. J Virol. 1989 Dec;63(12):5285–5292. doi: 10.1128/jvi.63.12.5285-5292.1989. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino S., Shieh C. K., Keck J. G., Lai M. M. Defective-interfering particles of murine coronavirus: mechanism of synthesis of defective viral RNAs. Virology. 1988 Mar;163(1):104–111. doi: 10.1016/0042-6822(88)90237-1. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino S., Soe L. H., Shieh C. K., Lai M. M. Discontinuous transcription generates heterogeneity at the leader fusion sites of coronavirus mRNAs. J Virol. 1988 Oct;62(10):3870–3873. doi: 10.1128/jvi.62.10.3870-3873.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino S., Taguchi F., Hirano N., Fujiwara K. Analysis of genomic and intracellular viral RNAs of small plaque mutants of mouse hepatitis virus, JHM strain. Virology. 1984 Nov;139(1):138–151. doi: 10.1016/0042-6822(84)90335-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Makino S., Yokomori K., Lai M. M. Analysis of efficiently packaged defective interfering RNAs of murine coronavirus: localization of a possible RNA-packaging signal. J Virol. 1990 Dec;64(12):6045–6053. doi: 10.1128/jvi.64.12.6045-6053.1990. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meyers G., Tautz N., Dubovi E. J., Thiel H. J. Viral cytopathogenicity correlated with integration of ubiquitin-coding sequences. Virology. 1991 Feb;180(2):602–616. doi: 10.1016/0042-6822(91)90074-L. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pachuk C. J., Bredenbeek P. J., Zoltick P. W., Spaan W. J., Weiss S. R. Molecular cloning of the gene encoding the putative polymerase of mouse hepatitis coronavirus, strain A59. Virology. 1989 Jul;171(1):141–148. doi: 10.1016/0042-6822(89)90520-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soe L. H., Shieh C. K., Baker S. C., Chang M. F., Lai M. M. Sequence and translation of the murine coronavirus 5'-end genomic RNA reveals the N-terminal structure of the putative RNA polymerase. J Virol. 1987 Dec;61(12):3968–3976. doi: 10.1128/jvi.61.12.3968-3976.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steinhauer D. A., Holland J. J. Direct method for quantitation of extreme polymerase error frequencies at selected single base sites in viral RNA. J Virol. 1986 Jan;57(1):219–228. doi: 10.1128/jvi.57.1.219-228.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Surratt C. K., Milan S. C., Chamberlin M. J. Spontaneous cleavage of RNA in ternary complexes of Escherichia coli RNA polymerase and its significance for the mechanism of transcription. Proc Natl Acad Sci U S A. 1991 Sep 15;88(18):7983–7987. doi: 10.1073/pnas.88.18.7983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weiss B. G., Schlesinger S. Recombination between Sindbis virus RNAs. J Virol. 1991 Aug;65(8):4017–4025. doi: 10.1128/jvi.65.8.4017-4025.1991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yokomori K., Banner L. R., Lai M. M. Heterogeneity of gene expression of the hemagglutinin-esterase (HE) protein of murine coronaviruses. Virology. 1991 Aug;183(2):647–657. doi: 10.1016/0042-6822(91)90994-M. [DOI] [PMC free article] [PubMed] [Google Scholar]