Abstract
Neogenin is a multifunctional receptor implicated in axon navigation, neuronal differentiation, morphogenesis, and cell death. Very little is known about signaling downstream of neogenin. Because we found that the neogenin intracellular domain (NeICD) interacts with nuclear proteins implicated in transcription regulation, we investigated further whether neogenin signals similarly to the Notch receptor. We show here that neogenin is cleaved by γ-secretase, an event that releases the complete NeICD. We also describe that NeICD is located at the nucleus, a feature regulated through a balance between nuclear import and export. NeICD triggers gene reporter transactivation and associates with nuclear chromatin. Direct transcriptional targets of NeICD were determined and were shown to be up-regulated in the presence of neogenin ligand. Together, we reveal here a novel aspect of neogenin signaling that relies on the direct implication of its intracellular domain in transcriptional regulation.
The seminal work by Fearon and colleagues proposed the 200-kDa-type transmembrane receptor deleted in colorectal cancer (DCC) to be a putative tumor suppressor (12). Even though this role in tumor regulation is still a matter of debate (11, 22), many reports have described the different functions of DCC. More and more attention is now being given to neogenin, a DCC homologue about which very little is currently known (8).
Neogenin displays a large ectodomain homologous to the NCAM protein family members and an intracellular domain with no known specific motifs. Because of the homology between DCC and neogenin, the latter was initially reported to be a receptor for the multifunctional netrin-1 protein (17, 35). Netrin-1 was discovered by Serafini and colleagues as a soluble cue that is key to guiding many axons and neurons during the development of the nervous system (33, 34). Because of the similar nervous system defects in netrin-1 and DCC mutant mice (17, 33), DCC is considered to be the main receptor involved in the role of netrin-1 in neuronal navigation. It has been proposed that neogenin could also be implicated in this guidance effect, although solid evidence is still missing. Netrin-1 and netrin family members are important not only for neuronal navigation but also beyond the brain (6), including in cell adhesion, morphogenesis, cell survival, and angiogenesis. Neogenin has been shown to play a role in mediating part of netrin's role in morphogenesis, especially during breast development (35) and myotube formation (16).
However, the propensity of netrin-1 to act as a ligand for neogenin was recently challenged (9, 18, 19, 32), and it has now been proposed that it is, at best, a low-affinity ligand for neogenin. Rather, neogenin appears to be a high-affinity receptor for the repulsive guidance molecule (RGM). RGM is actually comprised of three family members, RGMa, RGMb, and RGMc (18). Neogenin was proposed to mediate the axon guidance function of RGMa (32), and the RGM/neogenin pair has been shown to play a role in neuronal differentiation (18). Besides this role in neuronal navigation and differentiation, the RGM/neogenin pair also regulates cell survival. Indeed, both DCC and neogenin have been shown to belong to the so-called dependence receptor family (19, 21, 24). Such receptors share the functional property of inducing cell death when disengaged from their ligands, while the presence of their ligands blocks this proapoptotic activity. Such receptors thus create cellular states of dependence on their respective ligands (3, 25) and may consequently act as tumor suppressors by limiting tumor growth to settings of ligand availability (1, 20). This proapoptotic effect of unbound receptors may also be involved in nervous system development: the down-regulation of RGM or overexpression of neogenin in chick embryos is associated with massive cell death in the developing neural tube (19).
The molecular mechanisms that occur downstream of neogenin to promote morphogenesis, neuronal guidance, differentiation, or cell death are unknown. By analogy with its DCC homologue, it is believed that in the absence of RGM, neogenin is cleaved by caspases, exposing a proapoptotic domain, which in turn interacts with proapoptotic proteins (14, 19). On the other hand, in the presence of ligand, neogenin would activate classic positive pathways. Indeed, the RGM/neogenin pair was recently shown to activate the small GTPase RhoA, and it was shown that its role in neuronal navigation required RhoA activation (9). While performing a two-hybrid screen on the intracellular domain of neogenin, we observed that a large fraction of the putative interactors obtained are proteins usually localized within the nucleus and/or involved in transcription regulation. We thus investigated whether neogenin could function similarly to the Notch or amyloid-beta precursor protein (APP) receptors, and here, we present evidence supporting neogenin cleavage by γ-secretase, leading to the release of a neogenin intracellular domain (NeICD) that is imported/exported into/from the nucleus and regulates gene transcription.
MATERIALS AND METHODS
Cell lines, transfection procedures, and reagents.
Human embryonic kidney HEK293T cells, human breast cancer MCF-7 cells, and mouse mammary tumor 67NR cells were cultured in Dulbecco's modified essential medium (DMEM) plus 10% serum and 50 μg/ml gentamicin. Transient transfections of HEK293T cells were performed using FuGENE 6 reagent (Roche Diagnostics) according to the manufacturer's instructions. The γ-secretase inhibitor N-[N-(3,5-difluorophenylacetyl)-l-alanyl]-(S)-phenylglycine t-butyl ester (DAPT) and the proteasome inhibitor Z-Leu-Leu-Leu-al (MG132) were purchased from Sigma and used at 2 μM and 1 μM, respectively. The nuclear export inhibitor leptomycin B (LMB) was purchased from LC Laboratories and used at 1 μM. Lactacystin was purchased from Cayman and used at 10 μM. The α-secretase/TACE/ADAM17 inhibitor TAPI-1 was purchased from Calbiochem and used at 10 μM. Treatments with drugs or the same amount of dimethyl sulfoxide or ethanol vehicle alone were performed at indicated times. Recombinant human RGMa (200 ng/ml) was obtained from Apotech Corp. Mouse embryo brains were rinsed in DMEM-F12, manually dissociated, and incubated for 6 h with the indicated drugs in DMEM-F12 at 37°C.
Two-hybrid analysis.
The two-hybrid system Matchmaker II (Clontech) was used according to the manufacturer's instructions. As bait, we inserted the coding sequence of Neogenin IC (positions 1127 to 1323), amplified from pCDNA3-Neogenin, into plasmid pGBKT7 through the SmaI site. Saccharomyces cerevisiae AH109 cells were cotransformed with the pGBKT7-Neogenin IC construct and a human fetal brain cDNA library. Yeast cells were grown in the absence of leucine, tryptophan, and histidine and in the presence of α-Gal and 5 mM 3-amino-1,2,4-triazole. Putative candidate interactors were further analyzed through DNA sequencing.
Plasmid constructs.
The full-length human neogenin-expressing construct pCDNA3-Neogenin was described previously (19). The Flag-Tip60 construct is a kind gift from C. N. Robson (University of Newcastle Upon Tyne, Newcastle Upon Tyne, United Kingdom). The green fluorescent protein (GFP)-neogenin constructs were generated by inserting the corresponding coding sequences, amplified by PCR using pCDNA3-Neogenin as a template, into pEGP-C1 (Clontech) through ApaI-KpnI and XhoI-HindIII sites, respectively. Gal4-neogenin constructs were made with the same method by insertion into plasmid pMst through PstI-HindIII and EcoRI-HindIII sites, respectively. pMst is a modified version of pM (Clontech) and is a kind gift from T. Sudhof. The R1129A/R1130A, K1138A/R1139A, and L1455A neogenin mutations were created using the QuikChange site-directed mutagenesis system (Stratagene). The sequences of the primers that we used are available upon request. The Neo-Gal4 construct was generated in multiple steps by PCR amplification and cloning of neogenin fragments at positions 1 to 1144 and Gal4 at positions 1 to 147 into pCDNA3. Neo-Gal4-IC was generated by inserting the neogenin fragment at positions 1145 to 1462 into Neo-Gal4: the resulting plasmid encodes full-length neogenin with the Gal4 DNA binding domain inserted into the IC domain between positions 1144 and 1145 (more details of the strategy are available upon request). The Flag-NeICD L1455A construct was obtained by PCR amplification using GFP-NeICD L1455A as a template and inserted through NotI-KpnI sites into p3X-Flag (Sigma).
Immunoblotting and immunoprecipitation.
Antibodies against the neogenin intracellular domain (C-20) and against lamin A/C (346) were purchased from Santa Cruz. Antibody against β-actin (C4) was purchased from Chemicon. Coimmunoprecipitations were carried out on cotransfected 293T cells lysed in a solution containing 50 mM Tris (pH 7.5), 150 mM NaCl, 1 mM EDTA, and 0.5% NP-40 in the presence of protease inhibitors using anti-Flag M2 antibody and protein A-Sepharose (Sigma) to pull down Flag-Tip60. Neogenin IC's interaction with Tip60 was detected with anti-Neogenin IC.
Subcellular fractionation.
Cells were harvested and resuspended for 5 min in CLB buffer (10 mM HEPES, 10 mM NaCl, 1 mM KH2PO4, 5 mM NaHCO3, 5 mM EDTA, 1 mM CaCl2, 0.5 mM MgCl2) supplemented with protease inhibitor cocktail (Roche), phosphatase inhibitor cocktails (Sigma), and 1 mM trichostatin A (Calbiochem). Forty strokes were applied using a Dounce grinder, and the homogenate was then centrifuged at 4,000 × g for 5 min. The supernatant was adjusted to 0.5% NP-40 and used as the membrane plus the cytosol fraction. The nuclear pellet was resuspended in TSE (10 mM Tris, 300 mM sucrose, 1 mM EDTA, 0.1% NP-40 [pH 7.5]) plus protease and phosphatase inhibitors and trichostatin A and homogenized with 25 strokes. After centrifugation, the pure nuclear pellet was resuspended in Laemmli buffer and sonicated to obtain the nuclear extract.
Cell imaging.
HEK293T cells seeded onto cell culture slides and transfected with GFP constructs, treated or not with LMB, were fixed with 4% formaldehyde. Nuclei were stained with DAPI (4′,6′-diamidino-2-phenylindole). After mounting with Mowiol, cells were visualized using an Axiovert 200 M epifluorescence Zeiss microscope.
Gene reporter assay.
Plasmid pGLuc, encoding the firefly luciferase reporter gene under the control of the Gal4 upstream activation sequence (UAS), and plasmid pCMV-Renilla, encoding the Renilla luciferase reporter gene under the control of the cytomegalovirus promoter, were provided by T. Sudhof. HEK293T cells were cotransfected with each pMst construct, the same amount of pGLuc, and 1/50 of pCMV-Renilla. Transcriptional activities were determined 48 h after transfection using a dual-luciferase reporter assay according to the manufacturer's (Promega) instructions.
ChIP cloning.
HEK293T cells (1 × 108 cells) were transfected with the Flag-NeICD L1455A construct. Forty-eight hours later, cells were fixed in 2.5 mg/ml dimethyladipimidate (Fluka) in phosphate-buffered saline for 30 min at room temperature and subsequently fixed with 0.8% formaldehyde in phosphate-buffered saline for 10 min at room temperature. Fixation was blocked by the addition of glycine to 125 mM for 5 min at room temperature. Cells were centrifuged at 4°C, and the pellet was resuspended in sonication buffer (10 mM Tris [pH 8.0], 1 mM EDTA, 0.5 mM EGTA plus protease inhibitors). Chromatin was sonicated on ice using a Branson cell disrupter. After centrifugation to remove debris, the lysate was then adjusted to 10 mM Tris (pH 8.0), 1 mM EDTA, 0.5 mM EGTA, 150 mM NaCl, 0.1% Triton X-100, and protease inhibitors and incubated with anti-Flag M2 antibody overnight. The lysate was combined with μMACS protein G microbeads (Miltenyi Biotech) and incubated for 1 h. Lysate and beads were allowed to flow through a MACS column and washed four times with a solution containing 100 mM Tris (pH 8.0), 500 mM LiCl, 1% NP-40, and 1% deoxycholic acid. Chromatin immunoprecipitation (ChIP) products were then eluted in 50 mM NaHCO3-1% sodium dodecyl sulfate buffer. Ten micrograms of RNase A and 0.3 M NaCl were added for elution, and cross-linking was reverted by incubation for 4 h at 67°C. DNA was further purified using QiaQuick columns (Qiagen), digested by MboI, and cloned into BamHI-digested, calf intestinal alkaline phosphatase-dephosphorylated vector pUC19. The clones obtained were sequenced and localized on the human genome with the BLAT search function of the University of California, Santa Cruz, genome browser (http://genome.ucsc.edu).
Quantitative reverse transcription-PCR (Q-RT-PCR).
After RNA extraction using a NucleoSpin RNA II apparatus (Macherey Nagel) and following reverse transcription using an iScript cDNA synthesis kit (Bio-Rad) according to the manufacturer's instructions, quantitative PCR was performed using LightCycler technology according to the manufacturer's instructions. Primers used for GRHL1 amplification were 5′-GCAAAAGAAAAGGCAAGTGTC-3′ and 5′-AGGGAAGCAGTGGCACTTTA-3′, those used for CCM2 amplification were 5′-AGGAGACCTACGAGGTGGAAG-3′ and 5′-ACCACCCACATCCACAGATT-3′, and those used for β2-microglobulin were 5′-ACCCCCACTGAAAAAGATGA-3′ and 5′-ATCTTCAAACCTCCATGATG-3′. Quantification was carried out using the ΔΔCT method.
Quantitative ChIP assay.
Real-time PCR-based quantitative ChIP analysis was conducted essentially as ChIP cloning but using either 1 × 107 HEK293T cells transfected for full-length neogenin or 2 × 107 MCF-7 cells. Before immunoprecipitation, an aliquot (10% input) of sonicated chromatin was removed from each sample, reverse cross-linked, treated with RNase A and proteinase K, and then purified. The rest of the samples were incubated overnight with 4 μg of control goat immunoglobulin G (IgG) or goat anti-NeICD (C-20). After purification using the μMACS system, ChIP products were reverse cross-linked, RNase A and proteinase K treated, purified using QiaQuick columns, and analyzed by real-time PCR (LightCycler). The enrichment of the GRHL1 or CCM2 promoter sequence in chromatin immunoprecipitated with control or anti-neogenin antibody was calculated with the ΔΔCT method using a diluted 10% input sample as a reference and an untreated sample with control IgG as a calibrator. Primers used for the GRHL1 promoter were 5′-CCAGGCACCAGGAACATAAAAATG-3′ and 5′-TGTCTGTCTCCACCAAGTGTGAGC-3′, and those used for the CCM2 promoter were 5′-GCAACATAGTGTGACCTTGTCTCAAC-3′ and 5′-AAGCAATCCTCCCATCTCAGCC-3′.
RESULTS
Neogenin is cleaved by γ-secretase and interacts with proteins involved in gene transactivation.
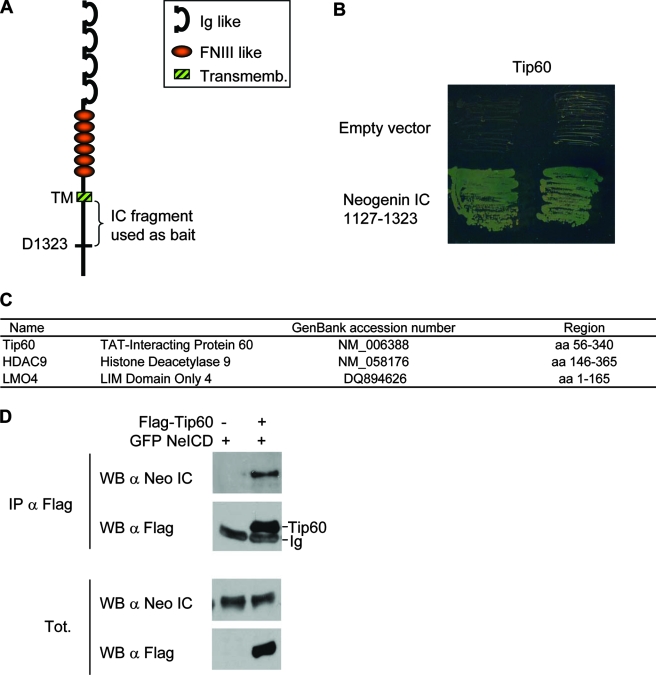
In a search for neogenin interactors, we performed a two-hybrid screen on the N-terminal half of the NeICD (amino acids [aa] 1127 to 1323) (this domain was initially chosen because it was thought to be involved in cell death induction) (19) (Fig. 1A). We were puzzled when we identified putative partners known to be located in the nucleus and to play a role in gene transactivation (Fig. 1B and C). One of these candidates, the TIP60 histone acetyltransferase (5), was further analyzed, and its interaction with neogenin was confirmed by coimmunoprecipitation. Upon forced expression in HEK293T cells, Flag-tagged TIP60 immunoprecipitation pulled down the intracellular domain of neogenin (Fig. 1D).
FIG. 1.
The NeICD interacts with nuclear proteins including TIP60. (A) Scheme of neogenin showing the structure of the protein consisting of Ig-like and fibronectin III-like domains for the extracellular part, a transmembrane domain (TM and Transmemb.), and an intracellular domain (IC). The fragment used as bait for the two-hybrid screen is also shown. (B) Neogenin aa 1127 to 1323 interact with TIP60 in yeast. AH109 yeast cells cotransformed with TIP60 fused to the transactivating domain of Gal4 together with the Gal4 DNA binding domain alone (empty vector) or fused to neogenin aa 1127 to 1323 were grown in selective medium, as described in Materials and Methods. (C) Some of the putative interactors of neogenin aa 1127 to 1323 revealed by the two-hybrid screen and known to display a nuclear activity. (D) Coimmunoprecipitation (IP) performed in HEK293T cells cotransfected with a neogenin-expressing vector together with a Flag-tagged TIP60-expressing construct. Anti-Flag M2 antibody was used for the pull down, which was revealed using a neogenin immunoblot (WB).
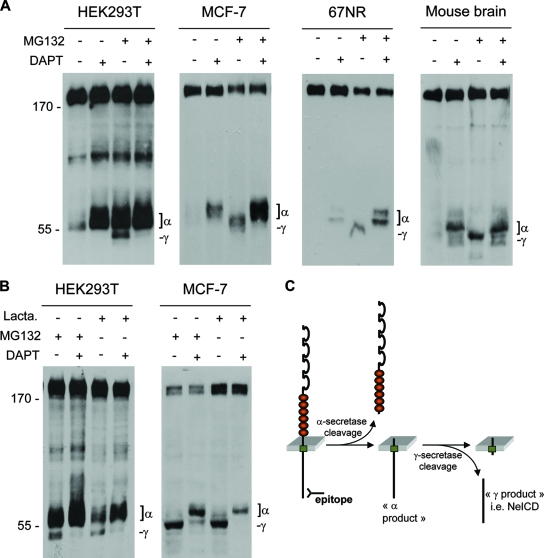
TIP60 was shown to form a complex with the APP intracellular domain and to be required for the gene transactivation activity of APP, which makes it an interesting candidate. The overall view regarding transmembrane receptors that interact with nuclear proteins and/or proteins involved in transcription regulation is that the cleavage of these receptors, e.g., Notch and APP, by a secretase releases an intracellular domain that in turn can be imported into the nucleus. Because DCC was recently shown to be a substrate for γ-secretase (30, 36), we then investigated whether neogenin was also cleaved by secretase. As a first approach, neogenin was expressed in HEK293T cells incubated in the presence or absence of DAPT, a specific γ-secretase inhibitor, with or without the proteasome inhibitor MG132. The resulting neogenin fragments were analyzed by immunoblot using a neogenin C-terminal epitope. As shown in Fig. 2A, in the absence of DAPT, a “γ” fragment of about 45 kDa was detected and disappeared upon DAPT treatment to the benefit of a higher-molecular-mass fragment (55 kDa), which consequently acted as a substrate for γ-secretase. We called this fragment the α-fragment, as it is associated with the cleavage of neogenin by an α-secretase. Indeed, 24 h of treatment with 10 μM TAPI-1, a specific inhibitor of α-secretase/TACE/ADAM17, prevents the apparition of this fragment (see Fig. S1 in the supplemental material). To discard a potential bias of overexpressed neogenin in HEK293T cells, we then assessed whether endogenously expressed neogenin undergoes dual cleavage by secretases. As shown in Fig. 2A, in the mammary tumor cell lines MCF-7 and 67NR, but also in mouse brain extract, neogenin is cleaved by γ-secretase, which leads to the appearance of a fragment encompassing the NeICD. The γ-fragment seems very unstable, as it was observed only in presence of the proteasome inhibitor MG132. To confirm that the γ-fragment is rapidly degraded by the proteasome, and to avoid a possible interference between MG132 and γ-secretase, as suggested previously by some authors (10), we compared the cleavage profiles of transfected or endogenous neogenin in presence of MG132 or lactacystin, another proteasome inhibitor (10 μM for 6 h). A similar profile was observed with both proteasome inhibitors (Fig. 2B) and also with a third one, epoxomicin (10 μM for 6 h) (data not shown). Therefore, based on the results described above and what is known for other secretase-cleaved receptors, such as Notch, neogenin appears to be cleaved by secretases, as depicted in the scenario shown in Fig. 2C.
FIG. 2.
Neogenin is cleaved by γ-secretase. (A) Neogenin immunoblot from lysates of neogenin-transfected HEK293T cells, from lysate of MCF-7 cells or 67NR cells, or from brains from mouse embryos at embryonic day 13.5 and treated for 6 h with 2 μM DAPT and/or the proteasome inhibitor MG132 (1 μM). Positions of the neogenin fragments cleaved by γ-secretase (γ) or α-secretase/ADAM metalloprotease (α), i.e., as this fragment is inhibited by treatment with TAPI-1 (see Fig. S1 in the supplemental material), are indicated. (B) Treatments of neogenin-transfected HEK293T or MCF-7 cells with two proteasome inhibitors show the same neogenin profiles (lactacystin [Lacta.] was used at 10 μM for 6 h). (C) Hypothetical model for the secretases cleavage of neogenin according to our results and the literature on other secretase-cleaved receptors.
The NeICD translocates to the nucleus.
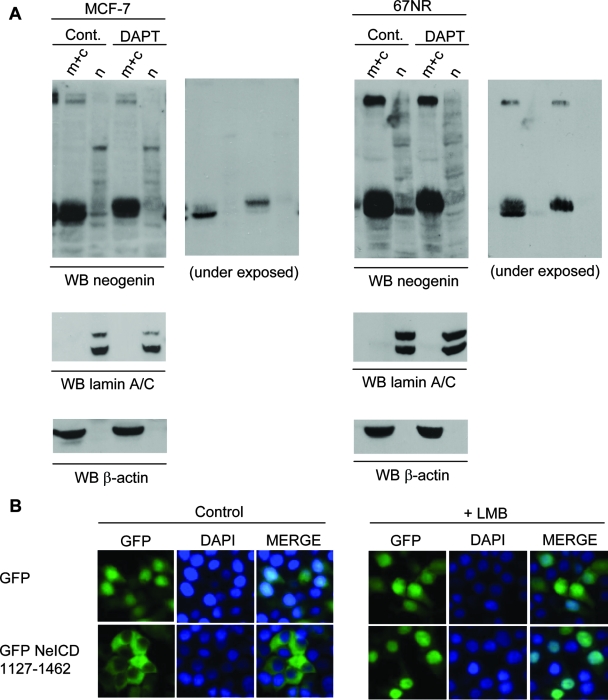
We then searched whether the fragment corresponding to the complete NeICD could translocate to the nucleus. As a first approach, we searched for the location of endogenously expressed neogenin in MCF-7 and 67NR cells following cytosolic/nuclear fractionation. As shown in Fig. 3A, while the majority of neogenin was recovered in the membrane/cytosolic fraction, a 45-kDa neogenin fragment was also detected in the nuclear fraction. Treatment with DAPT is associated with the absence of the neogenin γ-fragment detected in the nuclear extract, indicating that this fragment corresponds specifically to NeICD. Under these conditions, the α-product accumulates in the membrane/cytosolic fraction. Thus, upon γ-secretase cleavage, the NeICD was detected in the nucleus. To further elucidate the nuclear localization of the NeICD, we studied that of a GFP-tagged NeICD upon expression in HEK293T cells. As shown in Fig. 3B, under normal settings, the GFP-tagged NeICD was detected mainly in the cytosol. However, when LMB, an inhibitor of nuclear export, was added to the culture, GFP-NeICD accumulated in the nucleus (Fig. 3B). Thus, the NeICD naturally translocates to the nucleus but is also rapidly exported.
FIG. 3.
The NeICD displays a nuclear localization. (A) Cellular fractionation of MCF-7 or 67NR cells treated or not treated with 2 μM DAPT and/or the proteasome inhibitor MG132 (1 μM) for 6 h. Neogenin immunoblotting (WB) was performed on cellular fractions, the nucleus (n) and the membrane and cytosol (m+c); two exposition times are shown. Please note that a shorter exposition of the blot allows a better visualization of the migration difference between γ and α products in cytosolic fractions. A lamin A/C immunoblot was used as a control for nuclear fraction, while a β-actin immunoblot was used to control the cytosolic fraction. (B) HEK293T cells transfected with GFP-NeICD, further incubated with the inhibitor LMB (1 μM for 1 h), and monitored for fluorescence localization. DAPI staining was used to label the nucleus.
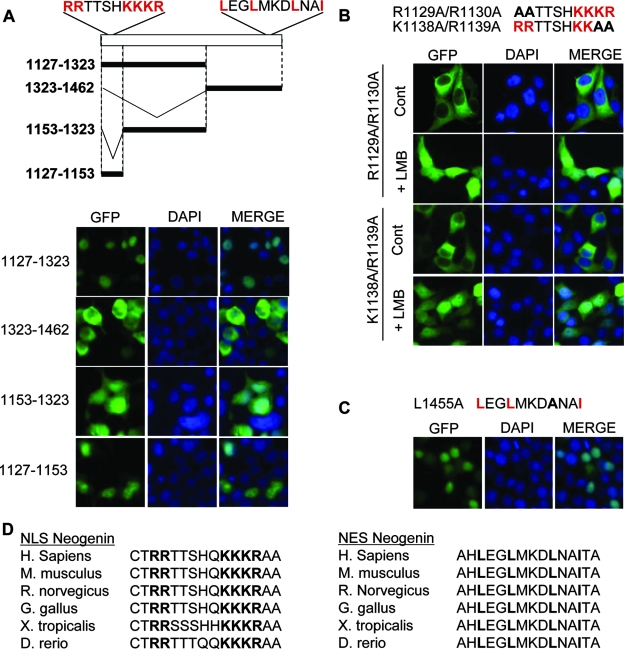
To map the domains responsible for the import and export of NeICD, we first analyzed the location of the N-terminal first half of the NeICD (aa 1127 to 1323) (Fig. 4A). As shown in Fig. 4A, this fragment is constitutively located within the nucleus, while the corresponding C-terminal other half (aa 1323 to 1462) is constitutively detected in the cytosol. Interestingly, in the first amino acids of the fragment at aa 1127 to 1323, a basic-rich domain was detected. As nuclear localization signals (NLSs) are often motifs enriched in basic amino acids (7), we deleted this region and observed that the remaining fragment (aa 1153 to 1323) was no longer specifically located in the nucleus. To further demonstrate that an NLS is present in the region at aa 1127 to 1153, we directly mutated the basic residues (RRTTSHKKKR mutated in AATTSHKKKR or RRTTSHKAA) and assessed the nuclear localization of the NeICD in the presence of LMB. As shown in Fig. 4B, the two mutants failed to adequately locate to the nucleus, demonstrating per se that neogenin displays an NLS located at aa 1127 to 1153. As we observed only nuclear localization of the NeICD in the presence of LMB or the absence of the C-terminal region, we surmised that a nuclear export signal (NES) was present in the C-terminal region of the NeICD. NESs are often characterized by sequences enriched in hydrophobic residues (38). We noted a sequence enriched in leucines in the C terminus of NeICD (L-[X]1-3-L-[X]2-3-L-X-X-I) and mutated one of the leucines (L→A). As shown in Fig. 4C, NeICD L1455A was then located to the nucleus, even in the absence of LMB. Thus, neogenin is cleaved by γ-secretase, which releases a fragment that translocates to the nucleus through an NLS located in its N-terminal end and is exported from the nucleus thanks to an NES located in its C-terminal end. Interestingly, both the NLS and NES appear to be conserved among species (Fig. 4D).
FIG. 4.
The NeICD displays an NLS and an NES to regulate its nuclear localization. HEK23T cells were transfected with the indicated constructs, further incubated or not with the inhibitor LMB (1 μM for 1 h), and monitored for fluorescence localization. DAPI staining was used to label the nucleus. (A, top) Scheme representing the different mutants of neogenin IC used in the following panels. (Bottom) Same as in Fig. 3B but with the indicated deletion mutants. (B) The neogenin NLS is in the N-terminal region of NeICD. Data are the same as described above (A) but with the indicated point mutants. (C) The NES is the C-terminal region of NeICD. Data are the same as described above (A), with the neogenin IC L1455A mutant. (D) Sequence alignments showing conservation of NESs and NLSs in the different species indicated. H. sapiens, Homo sapiens; M. musculus, Mus musculus; R. norvegicus, Rattus norvegicus; G. gallus, Gallus gallus; X. tropicalis, Xenopus tropicalis; D. rerio, Danio rerio.
The NeICD triggers gene transcription.
To monitor the role of this fragment, we investigated whether the NeICD was able to transactivate gene transcription. To assay this, the NeICD was fused to a Gal4 DNA binding domain, and HEK293T cells were forced to express Gal4-NeICD together with a construct encoding a luciferase reporter gene under the control of the Gal4 UAS. As shown in Fig. 5A, Gal4-NeICD is able to transactivate the Gal4 UAS (more than 12-fold over the Gal4 DNA binding domain alone), thereby demonstrating that this fragment displays a gene transactivation activity. To further demonstrate that this gene transactivation activity depends on the γ-secretase cleavage of the full-length neogenin, a Gal4 DNA binding domain was inserted within the intracellular domain of full-length neogenin (Fig. 5B). HEK293T cells were then forced to express Neo-Gal4ICD together with a construct encoding a reporter gene under the control of the Gal4 UAS and further treated (or not) with the γ-secretase inhibitor. As shown in Fig. 5B, in the absence of DAPT treatment, Neo-Gal4ICD transactivates the Gal4 UAS, whereas such transactivation is strongly inhibited by DAPT treatment. Thus, neogenin is cleaved by γ-secretase, which releases the NeICD that displays gene transactivation activity.
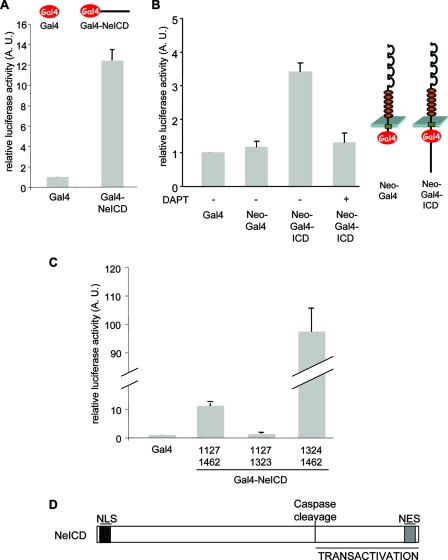
FIG. 5.
The NeICD displays a gene-transactivating activity. (A) The NeICD promotes reporter gene transcription. The Gal4-expressing construct or Gal4 DNA binding domain fused to the NeICD as indicated (top) was cotransfected with a Gal4-lucerifase reporter plasmid into HEK293T cells, and luminescence signals were recorded. The relative luciferase activity index is shown as the ratio between the neogenin construct and the control Gal4 vector. Standard errors of the means are indicated. (B) γ-Secretase-dependent NeICD release from full-length neogenin promotes reporter gene transcription. The Gal4 DNA binding domain was inserted into full-length neogenin. The resulting construct, NeoGal4ICD, was cotransfected with a Gal4 luciferase reporter plasmid into HEK293T cells. The NeoGal4 construct lacking the intracellular domain was used as a control. DAPT treatment (2 μM) was performed for 24 h. (C) Same as described above (A), with the indicated deletion mutants of NeICD-Gal4. (D) Scheme representing different functional domains of the NeICD. A.U., arbitrary units.
This activity is located in the C-terminal second half of NeICD (aa 1324 to 1462), as the deletion of this region was sufficient to fully abrogate this activity (Fig. 5C). In the same line, the expression of this C-terminal region only (fused to the Gal4 DNA binding domain) is sufficient to transactivate the Gal4 UAS (Fig. 5C). Intriguingly, the NeICD at aa 1324 to 1462 displays a stronger activity than does the NeICD, suggesting that the N-terminal region of NeICD also contains an inhibitory domain.
To formally prove that this cleaved fragment is associated with DNA complexes and triggers/enhances specific gene transcription, ChIPs were performed using NeICD L1455A for the pull down (the L1455A mutant was chosen for the immunoprecipitation to increase the level of the NeICD in the nucleus). DNA was extracted from the pull down and further vector cloned, demonstrating that the NeICD forms a complex with DNA. DNA sequencing of a small amount of cloned DNA (the goal here was not to describe new target genes but rather to demonstrate that the NeICD regulates the transcription of some of them) revealed a list of putative target genes that may potentially be regulated by neogenin signaling (Fig. 6A). We next assessed whether some of these genes were regulated by neogenin signaling. Two candidate genes, GRHL1 and CCM2, were selected on the basis of their relative known roles and expression statuses with regard to those of neogenin (CCM2 encodes a scaffold for mitogen-activated protein kinase [MAPK] [37], a kinase involved in DCC/neogenin signaling [13], and GRHL1 is a transcription factor implicated during nervous system development). We then assessed the presence of GRHL1 and CCM2 mRNA by Q-RT-PCR in HEK293T cells forced to express full-length neogenin and further incubated or not with RGMa. As shown in Fig. 6B, while neogenin expression by itself failed to increase GRHL1 and CCM2 levels, RGMa treatment triggers a time-dependent increased expression of GRHL1 and CCM2. Of interest, this increased expression is inhibited by DAPT treatment, thus suggesting that RGMa triggers gene transcription via γ-secretase cleavage of neogenin. To further analyze whether the same effect would be seen with endogenous neogenin, RGMa was added to MCF-7 cells, and as shown in Fig. 6C, RGMa does indeed trigger a GRHL1 mRNA increase, and this up-regulation is inhibited by DAPT.
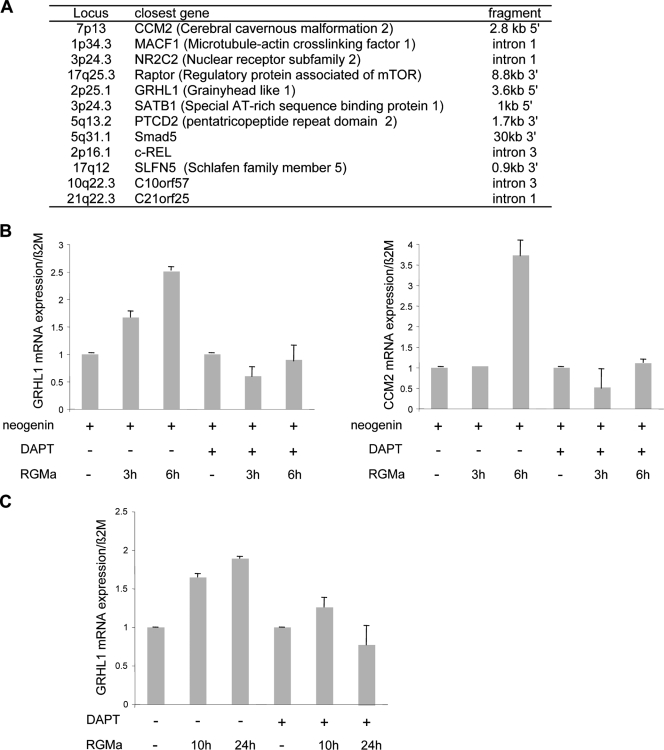
FIG. 6.
The NeICD behaves as a transactivator of gene transcription. (A) Different genes in which the NeICD has been shown to interact within their proximities (12 of over 25 clones analyzed). ChIP was performed on the NeICD, and immunoprecipitated DNA was cloned and sequenced. (B) GRHL1 and CCM2 Q-RT-PCR performed on mock- versus neogenin-transfected HEK293T cells treated or not treated for 3 or 6 h with RGMa in the presence or absence of DAPT. CCM2 and GRHL1 mRNA levels are regulated by neogenin/RGM signaling in neogenin-transfected HEK293T cells. Normalization was done on β2-microglobulin. The CCM2 or GRHL1 level is presented for each condition as a ratio between the normalized neogenin sample to the mock-transfected sample. (C) Same as described above (B), but the GRHL1 mRNA level was determined in RGMa/DAPT-treated MCF7 cells. Normalization was also done with β2-microglobulin, and the GRHL1 level is presented as a ratio between GRHL1 and β2-microglobulin.
To more formally demonstrate that RGMa regulates the nuclear signaling of the NeICD, which in turn directly enhances the transcription of GRHL1 and CCM2, as opposed to an indirect effect on RGMa/neogenin alternative signaling, we investigated whether the NeICD could be detected within GRHL1 and CCM2 promoter sequences and whether this detection could be increased upon RGMa treatment. To do so, ChIPs were performed using an anti-NeICD antibody for the pull down in HEK293T cells forced to express neogenin or in MCF-7 cells (which express endogenous neogenin) treated or not treated with RGMa. NeICD binding to GRHL1 and CCM2 promoter sequences was then quantitated by quantitative PCR of GRHL1 and CCM2 promoter sequences. As shown in Fig. 7, while RGMa treatment failed to enhance NeICD binding when the ChIP assay was performed with a control antibody, when the anti-NeICD antibody was used for the ChIP, we detected an increase in NeICD binding to both promoter sequences in response to RGMa. Together, these data support the view that the nuclear translocation and transactivation activities of the NeICD are part of the RGM signaling that occurs through the regulation of the transcription of a specific set of genes.
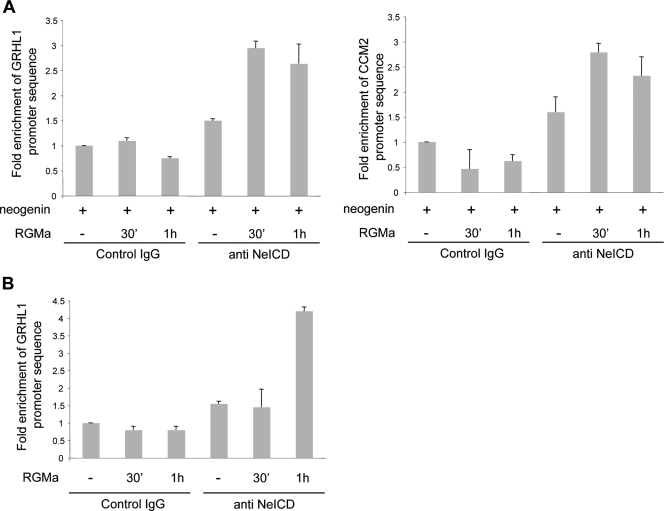
FIG. 7.
The NeICD binds target promoters after RGMa stimulation. (A) The NeICD binds GRHL1 and CCM2 promoters upon RGMa stimulation in neogenin-transfected HEK293T cells. Cross-linked chromatin was isolated from untreated and RGMa-treated neogenin-transfected HEK293T cells and precipitated with an NeICD-specific antibody or a control antibody. The precipitated chromatin was used as a template for quantitative PCR. (B) The endogenous NeICD binds the GRHL1 promoter upon RGMa stimulation. Data are the same as described above (A), but the experiment was performed with MCF7 cells untreated or treated with RGMa.
DISCUSSION
We propose here that neogenin is cleaved at the membrane by γ-secretase, an event that releases an intracellular domain, which migrates to the nucleus to regulate the transcription of a specific set of genes. Thus, functionally, neogenin resembles the APP or Notch receptors. Interestingly, the Notch intracellular domain is known to recruit histone acetyltransferases but also a transcriptional regulator, RBP-J/CBF1, which is the factor that activates gene transcription per se (2). As shown here, the NeICD also interacts with a histone acetyltransferase, but the factor present in the NeICD/DNA complex that activates transcription remains to be determined.
Another puzzling question is the role of the ligand in this novel neogenin signaling cascade. It has been shown that presence of the ligand stimulates the γ-secretase cleavage of the Notch or Erbb-4 receptors (28, 29). This has not been shown for other receptors such as the growth hormone receptor (10) or APP, even though in the latter case, this is due to the fact that no functional ligand is known for APP. Here, the functional data strongly support the initiating role of the ligand (RGMa) in this signaling: (i) neogenin-mediated transcription of target genes is stimulated in the presence of RGMa, a mechanism dependent on γ-secretase activity, and (ii) in response to RGMa, the NeICD shows an increased binding to target gene promoters. Thus, it is tempting to speculate that RGM controls this signaling. However, at this stage, we have not detected a change in the γ-secretase cleavage of neogenin dependent on the presence of RGM (not shown). This could either suggest that the ligand affects a downstream effect (e.g., nuclear import) or highlight technical difficulties in detecting a subtle change in the amount of a fragment that appears to be tightly regulated (e.g., the proteasome inhibitors MG132 and lactacystin were used to detect the neogenin γ-fragment indicated in Fig. 2).
Even though the elucidation of molecular mechanisms will require further investigation, it is tempting to speculate that ligand-dependent nuclear signaling is important in the context of the different functions of neogenin. For example, it may be important for the guidance activity of RGM. However, one could wonder how relevant the nuclear translocation of the intracellular domain of a membrane-associated receptor located in the growth cone of neurons would be in the context of RGM-mediated axon guidance. Indeed, neogenin would be cleaved by γ-secretase at the level of the growth cone and then transported to the soma, where the NeICD could enter the nucleus and trigger specific gene transcription and subsequent translation before being transported back to the growth cone, where axon navigation occurs. The requirement of neotranscription in axon guidance has already been supported: for example, transcription factors like NFaT are activated in response to netrin-1 and are required for the netrin-1 response (15). Here, it is interesting that several of the target genes pulled down with the NeICD could turn out to be important in RGM-mediated axon guidance. As an example, CCM2 appears to act as a scaffold for MAPK (37), and the importance of such a scaffold and MAPK activation has been demonstrated in various systems of axon guidance, including netrin-1-mediated axon guidance (4, 13, 27). Future studies should reveal whether direct target genes for the NeICD are important for RGM signaling.
Another putative role of the RGM/neogenin interaction is the inhibition of neogenin's proapoptotic activity. This RGM-dependent nuclear signaling could antagonize the prodeath signaling of neogenin. Even though we have not detected a target gene with a clear antiapoptotic effect at this stage, an interesting hypothesis is that RGM could inhibit neogenin-mediated cell death via a signaling loop that would take neogenin to the nucleus and possibly involve the neogenin-mediated transcription of antiapoptotic genes. In this respect, it is intriguing that neogenin undergoes two antagonistic cleavages in its intracellular domain. The γ-secretase cleavage appears to promote nuclear signaling speculated to be antiapoptotic, i.e., as being ligand mediated, while the caspase cleavage promotes the proapoptotic activity neogenin (19). Of interest, the caspase cleavage not only promotes apoptosis via the exposition of a proapoptotic domain (19) but also separates the NLS domain of neogenin from the neogenin transactivating domain (Fig. 5D). This should theoretically kill the gene transactivation activity of neogenin. Therefore, the engagement toward neogenin-mediated apoptosis should be associated not only with apoptosis promotion by the exposure of the neogenin proapoptotic domain but also with the inhibition of NeICD-mediated transcription.
Future studies will probably stress the importance of this nuclear signaling in neogenin functions. It is then intriguing that neogenin NLSs and NESs appear to be conserved in the different species in which neogenin exists. Similarly, an in silico analysis revealed that the NLS and NES of neogenin are also conserved in DCC (not shown), thus suggesting a conserved role of nuclear signaling in these receptors. We have preliminary results showing that DCC also interacts with TIP60 (not shown). Thus, even though the intracellular domains of DCC and neogenin are only modestly conserved (37%) (26), except in three specific domains called P1, P2, and P3 (23), and taking into account that Notch nuclear signaling is one of the few signaling pathways that has been conserved throughout evolution and is key to controlling cell fate (31), it is tempting to speculate that the neogenin/DCC nuclear signaling pathway is a conserved and important signaling pathway as well.
Supplementary Material
Acknowledgments
We thank C. N. Robson, T. Sudhof, and S. Douc-Rasy for materials; J. Fitamant and C. Guenebeaud for their advice on breast cancer cell lines; and H. Bilak for text correction.
This work was supported by an institutional grant for CNRS and Centre Léon Bérard, by the Ligue Contre le Cancer, by the NIH (NS45093), by the ARC, by the Region Rhone-Alpes, by the Agence Nationale de la Recherche, and by the Institut National du Cancer. D.G. was supported by fellowships from the Ligue Contre le Cancer and from the Schlumberger Foundation.
Footnotes
Published ahead of print on 7 April 2008.
Supplemental material for this article may be found at http://mcb.asm.org/.
REFERENCES
- 1.Bernet, A., L. Mazelin, M. M. Coissieux, N. Gadot, S. L. Ackerman, J. Y. Scoazec, and P. Mehlen. 2007. Inactivation of the UNC5C Netrin-1 receptor is associated with tumor progression in colorectal malignancies. Gastroenterology 1331840-1848. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bray, S. J. 2006. Notch signalling: a simple pathway becomes complex. Nat. Rev. Mol. Cell Biol. 7678-689. [DOI] [PubMed] [Google Scholar]
- 3.Bredesen, D. E., P. Mehlen, and S. Rabizadeh. 2005. Receptors that mediate cellular dependence. Cell Death Differ. 121031-1043. [DOI] [PubMed] [Google Scholar]
- 4.Campbell, D. S., and C. E. Holt. 2003. Apoptotic pathway and MAPKs differentially regulate chemotropic responses of retinal growth cones. Neuron 37939-952. [DOI] [PubMed] [Google Scholar]
- 5.Cao, X., and T. C. Südhof. 2001. A transcriptively [sic] active complex of APP with Fe65 and histone acetyltransferase Tip60. Science 293115-120. [DOI] [PubMed] [Google Scholar]
- 6.Cirulli, V., and M. Yebra. 2007. Netrins: beyond the brain. Nat. Rev. Mol. Cell Biol. 8296-306. [DOI] [PubMed] [Google Scholar]
- 7.Cokol, M., R. Nair, and B. Rost. 2000. Finding nuclear localization signals. EMBO Rep. 1411-415. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Cole, S. J., D. Bradford, and H. M. Cooper. 2007. Neogenin: a multi-functional receptor regulating diverse developmental processes. Int. J. Biochem. Cell Biol. 391569-1575. [DOI] [PubMed] [Google Scholar]
- 9.Conrad, S., H. Genth, F. Hofmann, I. Just, and T. Skutella. 2007. Neogenin-RGMa signaling at the growth cone is bone morphogenetic protein-independent and involves RhoA, ROCK, and PKC. J. Biol. Chem. 28216423-16433. [DOI] [PubMed] [Google Scholar]
- 10.Cowan, J. W., X. Wang, R. Guan, K. He, J. Jiang, G. Baumann, R. A. Black, M. S. Wolfe, and S. J. Frank. 2005. Growth hormone receptor is a target for presenilin-dependent gamma-secretase cleavage. J. Biol. Chem. 28019331-19342. [DOI] [PubMed] [Google Scholar]
- 11.Fazeli, A., S. L. Dickinson, M. L. Hermiston, R. V. Tighe, R. G. Steen, C. G. Small, E. T. Stoeckli, K. Keino-Masu, M. Masu, H. Rayburn, J. Simons, R. T. Bronson, J. I. Gordon, M. Tessier-Lavigne, and R. A. Weinberg. 1997. Phenotype of mice lacking functional Deleted in colorectal cancer (Dcc) gene. Nature 386796-804. [DOI] [PubMed] [Google Scholar]
- 12.Fearon, E. R., K. R. Cho, J. M. Nigro, S. E. Kern, J. W. Simons, J. M. Ruppert, S. R. Hamilton, A. C. Preisinger, G. Thomas, K. W. Kinzler, et al. 1990. Identification of a chromosome 18q gene that is altered in colorectal cancers. Science 24749-56. [DOI] [PubMed] [Google Scholar]
- 13.Forcet, C., E. Stein, L. Pays, V. Corset, F. Llambi, M. Tessier-Lavigne, and P. Mehlen. 2002. Netrin-1-mediated axon outgrowth requires deleted in colorectal cancer-dependent MAPK activation. Nature 417443-447. [DOI] [PubMed] [Google Scholar]
- 14.Forcet, C., X. Ye, L. Granger, V. Corset, H. Shin, D. E. Bredesen, and P. Mehlen. 2001. The dependence receptor DCC (deleted in colorectal cancer) defines an alternative mechanism for caspase activation. Proc. Natl. Acad. Sci. USA 983416-3421. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Graef, I. A., F. Wang, F. Charron, L. Chen, J. Neilson, M. Tessier-Lavigne, and G. R. Crabtree. 2003. Neurotrophins and netrins require calcineurin/NFAT signaling to stimulate outgrowth of embryonic axons. Cell 113657-670. [DOI] [PubMed] [Google Scholar]
- 16.Kang, J. S., M. J. Yi, W. Zhang, J. L. Feinleib, F. Cole, and R. S. Krauss. 2004. Netrins and neogenin promote myotube formation. J. Cell Biol. 167493-504. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Keino-Masu, K., M. Masu, L. Hinck, E. D. Leonardo, S. S. Chan, J. G. Culotti, and M. Tessier-Lavigne. 1996. Deleted in Colorectal Cancer (DCC) encodes a netrin receptor. Cell 87175-185. [DOI] [PubMed] [Google Scholar]
- 18.Matsunaga, E., H. Nakamura, and A. Chedotal. 2006. Repulsive guidance molecule plays multiple roles in neuronal differentiation and axon guidance. J. Neurosci. 266082-6088. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Matsunaga, E., S. Tauszig-Delamasure, P. P. Monnier, B. K. Mueller, S. M. Strittmatter, P. Mehlen, and A. Chedotal. 2004. RGM and its receptor neogenin regulate neuronal survival. Nat. Cell Biol. 6749-755. [DOI] [PubMed] [Google Scholar]
- 20.Mazelin, L., A. Bernet, C. Bonod-Bidaud, L. Pays, S. Arnaud, C. Gespach, D. E. Bredesen, J. Y. Scoazec, and P. Mehlen. 2004. Netrin-1 controls colorectal tumorigenesis by regulating apoptosis. Nature 43180-84. [DOI] [PubMed] [Google Scholar]
- 21.Mehlen, P., and D. E. Bredesen. 2004. The dependence receptor hypothesis. Apoptosis 937-49. [DOI] [PubMed] [Google Scholar]
- 22.Mehlen, P., and E. R. Fearon. 2004. Role of the dependence receptor DCC in colorectal cancer pathogenesis. J. Clin. Oncol. 223420-3428. [DOI] [PubMed] [Google Scholar]
- 23.Mehlen, P., and C. Furne. 2005. Netrin-1: when a neuronal guidance cue turns out to be a regulator of tumorigenesis. Cell. Mol. Life Sci. 622599-2616. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Mehlen, P., S. Rabizadeh, S. J. Snipas, N. Assa-Munt, G. S. Salvesen, and D. E. Bredesen. 1998. The DCC gene product induces apoptosis by a mechanism requiring receptor proteolysis. Nature 395801-804. [DOI] [PubMed] [Google Scholar]
- 25.Mehlen, P., and C. Thibert. 2004. Dependence receptors: between life and death. Cell. Mol. Life Sci. 611854-1866. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Meyerhardt, J. A., A. T. Look, S. H. Bigner, and E. R. Fearon. 1997. Identification and characterization of neogenin, a DCC-related gene. Oncogene 141129-1136. [DOI] [PubMed] [Google Scholar]
- 27.Ming, G. L., S. T. Wong, J. Henley, X. B. Yuan, H. J. Song, N. C. Spitzer, and M. M. Poo. 2002. Adaptation in the chemotactic guidance of nerve growth cones. Nature 417411-418. [DOI] [PubMed] [Google Scholar]
- 28.Mumm, J. S., E. H. Schroeter, M. T. Saxena, A. Griesemer, X. Tian, D. J. Pan, W. J. Ray, and R. Kopan. 2000. A ligand-induced extracellular cleavage regulates gamma-secretase-like proteolytic activation of Notch1. Mol. Cell 5197-206. [DOI] [PubMed] [Google Scholar]
- 29.Ni, C. Y., M. P. Murphy, T. E. Golde, and G. Carpenter. 2001. γ-Secretase cleavage and nuclear localization of ErbB-4 receptor tyrosine kinase. Science 2942179-2181. [DOI] [PubMed] [Google Scholar]
- 30.Parent, A. T., N. Y. Barnes, Y. Taniguchi, G. Thinakaran, and S. S. Sisodia. 2005. Presenilin attenuates receptor-mediated signaling and synaptic function. J. Neurosci. 251540-1549. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Pires-daSilva, A., and R. J. Sommer. 2003. The evolution of signalling pathways in animal development. Nat. Rev. Genet. 439-49. [DOI] [PubMed] [Google Scholar]
- 32.Rajagopalan, S., L. Deitinghoff, D. Davis, S. Conrad, T. Skutella, A. Chedotal, B. K. Mueller, and S. M. Strittmatter. 2004. Neogenin mediates the action of repulsive guidance molecule. Nat. Cell Biol. 6756-762. [DOI] [PubMed] [Google Scholar]
- 33.Serafini, T., S. A. Colamarino, E. D. Leonardo, H. Wang, R. Beddington, W. C. Skarnes, and M. Tessier-Lavigne. 1996. Netrin-1 is required for commissural axon guidance in the developing vertebrate nervous system. Cell 871001-1014. [DOI] [PubMed] [Google Scholar]
- 34.Serafini, T., T. E. Kennedy, M. J. Galko, C. Mirzayan, T. M. Jessell, and M. Tessier-Lavigne. 1994. The netrins define a family of axon outgrowth-promoting proteins homologous to C. elegans UNC-6. Cell 78409-424. [DOI] [PubMed] [Google Scholar]
- 35.Srinivasan, K., P. Strickland, A. Valdes, G. C. Shin, and L. Hinck. 2003. Netrin-1/neogenin interaction stabilizes multipotent progenitor cap cells during mammary gland morphogenesis. Dev. Cell 4371-382. [DOI] [PubMed] [Google Scholar]
- 36.Taniguchi, Y., S. H. Kim, and S. S. Sisodia. 2003. Presenilin-dependent “gamma-secretase” processing of deleted in colorectal cancer (DCC). J. Biol. Chem. 27830425-30428. [DOI] [PubMed] [Google Scholar]
- 37.Uhlik, M. T., A. N. Abell, N. L. Johnson, W. Sun, B. D. Cuevas, K. E. Lobel-Rice, E. A. Horne, M. L. Dell'Acqua, and G. L. Johnson. 2003. Rac-MEKK3-MKK3 scaffolding for p38 MAPK activation during hyperosmotic shock. Nat. Cell Biol. 51104-1110. [DOI] [PubMed] [Google Scholar]
- 38.Wen, W., J. L. Meinkoth, R. Y. Tsien, and S. S. Taylor. 1995. Identification of a signal for rapid export of proteins from the nucleus. Cell 82463-473. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.