Abstract
The plant hormone auxin (indole-3-acetic acid [IAA]) is found both free and conjugated to a variety of carbohydrates, amino acids, and peptides. We have recently shown that IAA could be converted to its methyl ester (MeIAA) by the Arabidopsis (Arabidopsis thaliana) enzyme IAA carboxyl methyltransferase 1. However, the presence and function of MeIAA in vivo remains unclear. Recently, it has been shown that the tobacco (Nicotiana tabacum) protein SABP2 (salicylic acid binding protein 2) hydrolyzes methyl salicylate to salicylic acid. There are 20 homologs of SABP2 in the genome of Arabidopsis, which we have named AtMES (for methyl esterases). We tested 15 of the proteins encoded by these genes in biochemical assays with various substrates and identified several candidate MeIAA esterases that could hydrolyze MeIAA. MeIAA, like IAA, exerts inhibitory activity on the growth of wild-type roots when applied exogenously. However, the roots of Arabidopsis plants carrying T-DNA insertions in the putative MeIAA esterase gene AtMES17 (At3g10870) displayed significantly decreased sensitivity to MeIAA compared with wild-type roots while remaining as sensitive to free IAA as wild-type roots. Incubating seedlings in the presence of [14C]MeIAA for 30 min revealed that mes17 mutants hydrolyzed only 40% of the [14C]MeIAA taken up by plants, whereas wild-type plants hydrolyzed 100% of absorbed [14C]MeIAA. Roots of Arabidopsis plants overexpressing AtMES17 showed increased sensitivity to MeIAA but not to IAA. Additionally, mes17 plants have longer hypocotyls and display increased expression of the auxin-responsive DR5:β-glucuronidase reporter gene, suggesting a perturbation in IAA homeostasis and/or transport. mes17-1/axr1-3 double mutant plants have the same phenotype as axr1-3, suggesting MES17 acts upstream of AXR1. The protein encoded by AtMES17 had a Km value of 13 μm and a Kcat value of 0.18 s−1 for MeIAA. AtMES17 was expressed at the highest levels in shoot apex, stem, and root of Arabidopsis. Our results demonstrate that MeIAA is an inactive form of IAA, and the manifestations of MeIAA in vivo activity are due to the action of free IAA that is generated from MeIAA upon hydrolysis by one or more plant esterases.
Indole-3-acetic acid (IAA), also known as auxin, is a plant hormone involved in many aspects of plant growth and development, such as embryogenesis, vascular differentiation, fruit set and development, and senescence (Woodward and Bartel, 2005; Teale et al., 2006; Delker et al., 2008). Plants utilize a variety of mechanisms to spatially and temporally regulate IAA concentrations and gradients, including de novo synthesis, degradation, transport, and synthesis and hydrolysis of various IAA conjugates (Normanly, 1997; Ljung et al., 2002; Woodward and Bartel, 2005).
IAA is known to be conjugated to sugars, amino acids, and peptides, and some enzymes that catalyze these conjugating reactions have been characterized (Jackson et al., 2001; Staswick et al., 2005). Some conjugates such as IAA-Asp and IAA-Glu are not able to induce auxin responses when applied exogenously and therefore are considered inactive auxin and intermediates in IAA degradation (Ljung et al., 2002; Woodward and Bartel, 2005). Recently, a rice (Oryza sativa) GH3-8 gene encoding IAA-amino acid synthetase has been shown to promote basal immunity in rice by converting active IAA to inactive IAA-Asp and thus reducing the auxin-induced cell wall loosening (Ding et al., 2008). Other IAA conjugates such as IAA-Leu and IAA-Ala induce auxin responses when applied exogenously to plants. However, the hydrolytic cleavage of these compounds parallels the activity (Bartel and Fink, 1995; Ljung et al., 2002). These findings have led to suggestions that these conjugates per se are biologically inactive, and any response obtained in the assay reflected the degree of hydrolysis and the activity of the released free hormone.
A family of hydrolases that acts on IAA-amino acid conjugates and releases IAA from some IAA-amino acid conjugates has been identified in Arabidopsis (Arabidopsis thaliana; LeClere et al., 2002; Rampey et al., 2004). The hydrolyzable IAA conjugates have been proposed to function as storage form to allow the plants to quickly release active IAA when necessary. While the transport of any IAA conjugates has rarely been reported, IAA-inositol has been shown to be transported from the endosperm to shoot in Zea mays at a rate much faster than that of free IAA and there hydrolyzed to yield free IAA (Nowacki and Bandurski, 1980).
We have recently discovered an IAA carboxy methyltransferase (IAMT1) in Arabidopsis and several other species that can methylate IAA to form the ester methyl indole-3-acetate (MeIAA; Zubieta et al., 2003; Qin et al., 2005; Zhao et al., 2008). Furthermore, disruption of the expression levels of IAMT1 led to phenotypes indicative of disruption of IAA homeostasis (Qin et al., 2005). MeIAA has rarely been reported as an endogenous IAA metabolite in plants (Narasimhan et al., 2003), probably due to its low abundance or fast turnover, and therefore its in vivo function remains unknown. MeIAA had been used as a substitute for IAA in physiological studies (Zimmerman and Hitchcock, 1937), and it has been observed that various auxin signaling mutants show decreased sensitivity to exogenously applied MeIAA as well as to IAA (Qin et al., 2005), suggesting that MeIAA and IAA share similar signaling components. Therefore, either MeIAA itself could initiate the auxin signaling pathway, or it must be hydrolyzed to IAA to exert hormonal function. If MeIAA hydrolysis occurs in planta, the reaction is likely to be catalyzed by one or more carboxylesterases.
Carboxylesterases catalyze the hydrolysis of a C-O ester linkage in a wide range of compounds, and structural analyses have shown that such enzymes are all members of the α/β hydrolase “superfamily” (Nardini and Dijkstra, 1999). Carboxylesterases have been extensively studied in animals and microbes. However, the physiological role and substrate specificity of few plant carboxylesterases have been identified. Several putative plant proteins, including those encoded by tobacco (Nicotiana tabacum) hsr203J, the tomato (Solanum lycopersicum) and pea (Pisum sativum) homologs of hsr203J, and PrMC3 from Pinus radiata and pepEST from pepper (Capsicum annuum; Pontier et al., 1994, 1998; Walden et al., 1999; Ichinose et al., 2001; Ko et al., 2005) have been annotated as carboxylesterases based on homology with fungal esterases but with little direct biochemical evidence (Baudouin et al., 1997). Marshall et al. (2003), in turn, searched the Arabidopsis genome, which has several hundred members of the α/β hydrolase superfamily, for genes encoding proteins with the highest similarities to these previously annotated plant carboxylesterases. This bioinformatic search identified a branch of the α/β hydrolase superfamily containing 20 genes, which were collectively named the AtCXE family (Marshall et al., 2003). However, the in vivo substrates of none of the enzymes in the AtCXE family have been experimentally determined. Recently, two proteins belonging to the α/β hydrolase superfamily have been identified in Gentiana triflora and implicated in cold response, but their in vivo substrates remain unknown (Hikage et al., 2007).
Recently, we have demonstrated that a tobacco protein required for development of systemic acquired resistance, SABP2 (originally identified as salicylic acid binding protein 2), is a methyl salicylate (MeSA) esterase (Kumar and Klessig, 2003; Forouhar et al., 2005). The amino acid sequence of SABP2 shares 46% to 56% similarity to two other confirmed methyl esterases from plants, methyl jasmonate (MeJA) esterase (MJE) from tomato and polyneuridine aldehyde esterase (PNAE) from the medicinal plant Rauvolfia serpentina (Dogru et al., 2000; Stuhlfelder et al., 2004; Forouhar et al., 2005). Bioinformatic analysis of the Arabidopsis genome revealed 20 genes encoding proteins with relatively high sequence similarities to SABP2 (Forouhar et al., 2005; Yang et al., 2006a). These proteins are distinct from the group of 20 AtCXE proteins, and their sequence similarity to known methylesterases suggests that they too may be methylesterases and may perhaps be involved in the hydrolysis of MeSA, MeJA, or MeIAA in Arabidopsis (Yang et al., 2006a).
Here, we show that some of the proteins in this group of putative Arabidopsis methylesterases, which we have named MES (for methyl esterases), are able to hydrolyze MeIAA. Analysis of mutants with T-DNA insertions in the AtMES genes indicates that at least one AtMES (AtMES17) is capable of hydrolyzing MeIAA in vivo. We used this mutant to demonstrate that MeIAA itself is an inactive form of IAA.
RESULTS
The Arabidopsis Genome Has 20 MES Genes
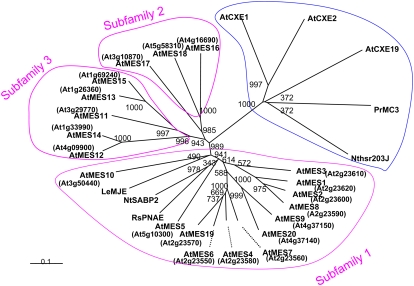
A search of the Arabidopsis genome for genes encoding proteins with the highest identity to tobacco SABP2 (MeSA esterase), tomato MJE, and R. serpentina PNAE identified 20 genes forming a close clade within the α/β hydrolase superfamily, which we named AtMES1 to AtMES20 (Fig. 1; Table I). The amino acid sequences of the AtMES proteins, which range in length from 256 to 444 amino acids (with the exception of the proteins encoded by AtMES19 and 20, which are likely to be pseudogenes; see below), share 30% to 57% similarity with tobacco SABP2, 31% to 42% similarity with tomato MJE, and 29% to 49% similarity with R. serpentina PNAE. The tree topology of the AtMES family shows the presence of three clusters of genes, which we have named subfamilies 1, 2, and 3 (Fig. 1). Members of the previously annotated plant carboxylesterases (CXE) family, including tobacco hsr203J, PrMC3, and three AtCXE genes (AtCXE1, -2, and -19), are more divergent, and they cluster into a clade that is distant from the AtMES family (Fig. 1).
Figure 1.
An unrooted neighbor-joining tree showing the phylogenetic relationships among AtMES proteins and other carboxylesterases. The tree was constructed with protein sequences of 20 Arabidopsis MES members (AtMES1–AtMES20), NtSABP2, R. serpentina PNAE, and LeMJE. Protein sequences of CXE family members AtCXE1, -2, and -19, Nthsr203J, and P. radiata (Pr) MC3 were also included in the phylogenetic analysis. Bootstrap values were calculated from 1,000 replicates. The three clusters of sequences that contain all the AtMES sequences were designated as subfamilies 1, 2, and 3. Analysis using maximum parsimony (not shown) gave a tree with the same four major branches.
Table I.
Substrate specificities of AtMES proteins
AtMES family members (AtMES1–AtMES20) are listed with the respective gene identification numbers. Fifteen heterologously expressed AtMES proteins were assayed for esterase activities with PNPA, MeIAA, MeSA, MeJA, MeGA4, and MeGA9, as described in “Materials and Methods.” +, Active; −, not active; n.d., not determined.
| Name | Gene ID | PNPA | MeIAA | MeSA | MeJA | MeGA4 | MeGA9 |
|---|---|---|---|---|---|---|---|
| AtMES1 | At2g23620 | + | + | + | + | − | − |
| AtMES2 | At2g23600 | + | + | + | + | − | − |
| AtMES3 | At2g23610 | + | + | − | + | − | − |
| AtMES4 | At2g23580 | + | − | + | − | − | − |
| AtMES5 | At5g10300 | − | − | − | − | − | − |
| AtMES6 | At2g23550 | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
| AtMES7 | At2g23560 | + | + | + | − | − | − |
| AtMES8 | At2g23590 | + | − | − | − | − | − |
| AtMES9 | At4g37150 | + | + | + | + | − | − |
| AtMES10 | At3g50440 | − | − | − | + | − | − |
| AtMES11 | At3g29770 | − | − | − | − | − | − |
| AtMES12 | At4g09900 | − | − | − | − | − | − |
| AtMES13 | At1g26360 | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
| AtMES14 | At1g33990 | − | − | − | − | − | − |
| AtMES15 | At1g69240 | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
| AtMES16 | At4g16690 | + | + | − | + | − | − |
| AtMES17 | At3g10870 | + | + | − | − | − | − |
| AtMES18 | At5g58310 | − | + | − | − | − | − |
| AtMES19 | At2g23570 | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
| AtMES20 | At4g37140 | n.d. | n.d. | n.d. | n.d. | n.d. | n.d. |
The sequence alignment of AtMES1 to AtMES20 revealed that the catalytic triad Ser-His-Asp, a characteristic feature of the α/β hydrolase fold family, is conserved in 15 of these proteins (Fig. 2). In the protein sequences of AtMES11, AtMES13, and AtMES15, the conserved Ser in the catalytic triad is replaced by Asp, a substitution previously found in active α/β hydrolases in animals (Holmquist, 2000). AtMES19 and AtMES20 lack part of the N-terminal or C-terminal region, respectively, and are therefore likely to be inactive enzymes.
Figure 2.
Multiple sequence alignment of tobacco SABP2, tomato MJE, and the 20 Arabidopsis AtMES. The sequence alignment was constructed using ClustalX program (Thompson et al., 1997). Identical amino acids at a given position in 17 or more proteins are shown in white letters on black. The catalytic triad residues are indicated by asterisks.
Substrate Specificities of 15 MES Esterases
To examine whether the AtMES genes encode functional esterases, we obtained full-length cDNAs of 15 AtMES genes, expressed the cDNAs in Escherichia coli, and tested the recombinant proteins for esterase activity. Because AtMES19 and AtMES20 were likely to be pseudogenes, they were not tested. Three other AtMES proteins, AtMES6, AtMES13, and AtMES15, were also not tested, because we were not able to obtain full-length cDNAs.
When the 15 AtMES esterases were tested with the chymotryptic synthetic substrate p-nitrophenyl acetate (PNPA), AtMES1, AtMES2, AtMES3, AtMES4, AtMES7, AtMES8, AtMES9, AtMES16, and AtMES17 showed activity (Table I). Because the AtMES proteins are homologs of SABP2 and MJE, esterases that hydrolyze the methylated plant hormones MeSA and MeJA, respectively, we further examined whether the AtMES proteins are active with known methylated plant hormones, including MeIAA, MeSA, MeJA, MeGA4, and MeGA9 (Shulaev et al., 1997; Chen et al., 2003; Qin et al., 2005; Yang et al., 2006a; Varbanova et al., 2007). To assess whether the AtMES proteins are active with any of these substrates, we carried out preliminary assays for each protein with a number of substrates present at a concentration of 1 mm. Reactions that resulted in product formation that was at least 3 times the value found in control assays (using boiled enzyme) were scored “+” as indicating enzymatic activity (Table I).
Among the 15 esterases tested, AtMES1, AtMES2, AtMES3, AtMES7, AtMES9, AtMES16, AtMES17, and AtMES18 displayed hydrolase activity with MeIAA, while AtMES4, AtMES5, AtMES8, AtMES10, AtMES11, AtMES12, and AtMES14 could not hydrolyze MeIAA (Table I). In addition, AtMES1, AtMES2, AtMES4, AtMES7, and AtMES9 displayed MeSA hydrolase activity, while AtMES1, AtMES2, AtMES3, AtMES9, AtMES10, and AtMES16 were active with MeJA. None of the 15 AtMES esterases was active with MeGA4 or MeGA9. AtMES5, AtMES8, AtMES11, AtMES12, and AtMES14 were not active with any of these methylated hormones.
AtMES17 Null Mutant Plants Are More Resistant Than Wild Type to the Root Inhibition Activity of Exogenously Supplied MeIAA and They Are Defective in Hydrolysis of Such MeIAA in Vivo
Because AtMES1, AtMES2, AtMES3, AtMES7, AtMES9, AtMES16, AtMES17, and AtMES18 could all hydrolyze MeIAA in vitro, we examined whether they possess MeIAA hydrolase activity in vivo. It has been previously shown that both IAA and MeIAA inhibit root growth in wild-type Arabidopsis seedlings when applied exogenously (Zimmerman and Hitchcock, 1937; Qin et al., 2005), but it has not been determined if MeIAA itself is active or whether the apparent activity of MeIAA is due to its hydrolysis in planta, giving rise to active IAA.
T-DNA insertional mutants of AtMES1, AtMES9, AtMES16, and AtMES17 were obtained as described in “Materials and Methods,” including two independent mutant lines each for both AtMES16 and AtMES17. There was no T-DNA insertional mutant of AtMES3 reported, and the several T-DNA insertions reported for AtMES2, AtMES7, and AtMES18 turned out upon further examination (described in “Materials and Methods”) not to abolish gene transcriptions (data not shown).
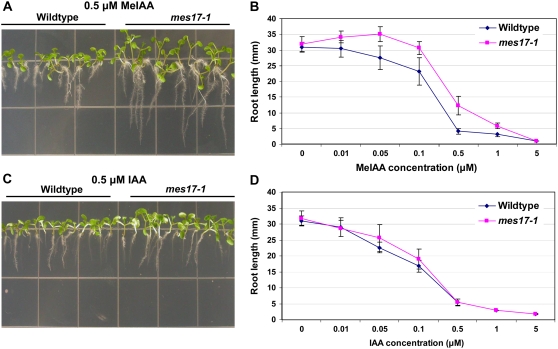
All mutant lines as well as wild-type Arabidopsis plants were next grown on one-half-strength Murashige and Skoog (MS) medium containing various concentrations of MeIAA or no MeIAA, and their root lengths were measured after 7 d. While in unsupplemented medium, root length of an AtMES17 T-DNA mutant mes17-1 (SALK_092550) seedlings were similar to that of wild type; in the presence of MeIAA concentrations ranging from 0.01 to 1 μm, root length of mutant seedlings was consistently longer than the root length of wild-type seedlings (Fig. 3B). For example, at 0.5 μm MeIAA, a concentration that inhibits the root growth of wild-type Arabidopsis by 85% on average, wild-type seedlings had an average root length of 4.1 mm and mes17-1 plants had an average root length of 12 mm, 3 times as long as that of the wild type (Fig. 3, A and B). Similar results were obtained with a second independent AtMES17 T-DNA mutant, mes17-2 (SAIL-503-c03; data not shown). The root lengths of AtMES1, AtMES9, and AtMES16 mutant lines grown on MeIAA were the same as wild type. All mutant plants, including the mes17-1 and mes17-2, when grown on one-half-strength MS medium containing different concentrations of IAA, showed no statistically significant difference in root length from that of wild-type plants, although the mes17 mutants appeared to have a slightly diminished response to IAA (Fig. 3, C and D).
Figure 3.
Plants with a null mutation in AtMES17 are more resistant to MeIAA but not to IAA. A, Seedlings were grown on one-half-strength MS medium containing 0.5 μm MeIAA for 7 d. B, Root length (mean ± sd, n ≥ 20) of wild-type and mes17-1 plants grown for 7 d on one-half-strength MS medium containing various concentrations of MeIAA. C, Seedlings were grown on one-half-strength MS medium containing 0.5 μm IAA for 7 d. D, Root length (mean ± sd, n ≥ 20) of wild-type and mes17-1 plants grown for 7 d on one-half-strength MS medium containing various concentrations of IAA.
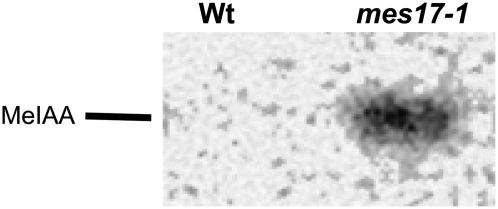
To examine directly the fate of exogenously added MeIAA in wild-type and Atmes17 mutant plants, we soaked plants in a 0.5 μm solution of [14C]MeIAA and examined the total amount of [14C]label taken up by the plant and the relative amounts of [14C]MeIAA remaining in the plant tissues. After 30 min of incubation, wild-type plants had no [14C]MeIAA left, but Atmes17-1 plants still contained 58.5% ± 16.5% of the [14C]label taken up in the form of MeIAA (Fig. 4).
Figure 4.
Plants with a null mutation in AtMES17 hydrolyze exogenously supplied MeIAA at a much lower efficiency than wild-type plants. Wild-type and mes17-1 seedlings were incubated in a 0.5 μm solution of [14C]MeIAA for 30 min. [14C]MeIAA absorbed by the plants was extracted and analyzed by radio-TLC. The position of an authentic MeIAA standard is also shown.
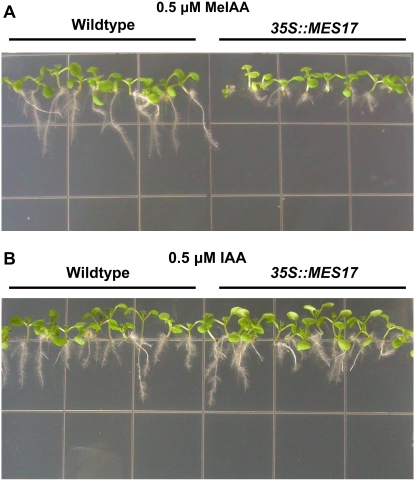
We also obtained several lines that overexpress AtMES17 under the control of the 35S promoter and tested them for sensitivity to MeIAA and IAA treatments. When AtMES17-overexpressing plants of three independent lines were grown in the presence of 0.5 μm MeIAA for 7 d, their root growth was more severely inhibited than that of wild-type seedlings (see Fig. 5A for one of the lines). However, both types of seedlings had a similar root length in the presence of 0.5 μm IAA (Fig. 5B), suggesting that the increased root inhibition of MeIAA on MES17-overexpressing plants was caused by increased auxin concentration derived from increased rate of MeIAA hydrolysis.
Figure 5.
Plants overexpressing AtMES17 are more sensitive to MeIAA treatment but not to IAA. Wild-type plants and plants overexpressing AtMES17 were grown on one-half-strength MS medium containing 0.5 μm MeIAA (A) or IAA (B) for 7 d.
Atmes17 Null Mutants Have a Longer Hypocotyl
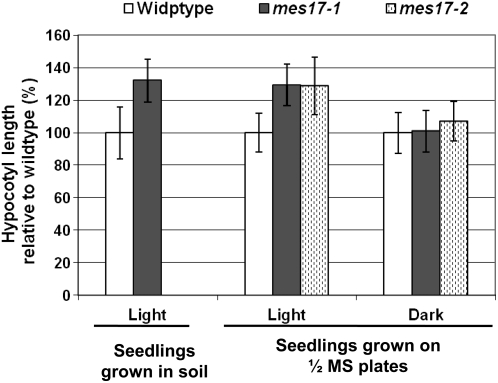
We observed that mes17-1 mutant plants grown in soil had, in general, longer hypocotyls than wild-type plants. The hypocotyls of 4-week-old mes17-1 plants were on average 32% longer than that of the wild type (Fig. 6). Statistical analysis performed by Student's t test returned P values <1 × 10−9, indicating that the differences are significant. When grown on one-half-strength MS medium under continuous light for 8 d, mes17-1 and mes17-2 mutants had hypocotyls longer on average by 29% than that of wild-type seedlings (Fig. 6), with statistical analysis indicating that the differences are significant (P values <1 × 10−5). When grown on one-half-strength MS medium in the dark, however, the hypocotyl lengths of mes17-1 and mes17-2 mutants were the same as that of wild type (Fig. 6). With the exception of hypocotyl length, mes17 mutants grown under normal conditions did not display any obvious phenotypic differences compared to wild type.
Figure 6.
Plants carrying a null mutation in AtMES17 have longer hypocotyls. Left, Analysis of hypocotyl lengths of 4-week-old wild-type and mes17-1 Arabidopsis plants grown in soil. Right, Analysis of the hypocotyl length of wild-type, mes17-1, and mes17-2 seedlings grown on one-half-strength MS medium under continuous light for 8 d or in the dark for 4 d. Hypocotyl lengths of the seedlings grown on one-half-strength MS plates were measured with Image J as described in “Materials and Methods.” The mean value and sd were calculated from 20 samples.
The DR5:GUS Reporter Gene Is More Highly Expressed in Atmes17 Null Mutants Compared with Wild-Type Plants
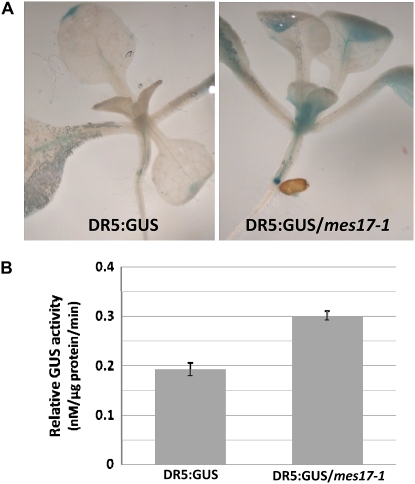
DR5 is a synthetic auxin response element, and the DR5:GUS reporter has been widely used as a marker to study the endogenous distribution of auxin (Ulmasov et al., 1997; Ottenschlager et al., 2003). We constructed mes17 null mutant plants carrying the DR5:GUS reporter gene and tested them for GUS activity. mes17 null plants had much stronger GUS staining overall than wild-type plants, including in the shoot apex and in the root primordia and leaf tip (Fig. 7).
Figure 7.
Histochemical staining of GUS activity in DR5:GUS and DR5:GUS/mes17-1 seedlings. A, DR5:GUS seedlings and DR5:GUS/mes17-1 seedlings were stained for GUS activity for 16 h. B, Quantitative GUS assay of DR5:GUS and DR5:GUS/mes17-1 seedlings. The mean value and sd of GUS activity were calculated from three replicates and represented as nanomoles of 4-methyl umbelliferone per milligram protein per minute, as described in “Materials and Methods.”
mes17-1/axr1-3 Double Mutant Plants Have the Same Phenotype as axr1-3
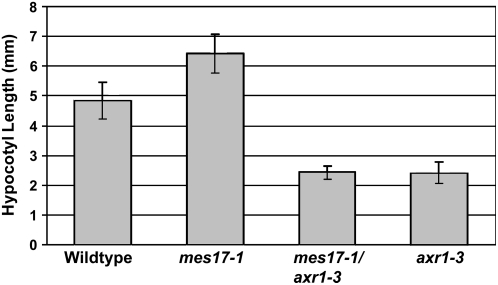
Plants homozygous for the allele axr1-3, which carries a missense mutation in the AXR1 gene, display resistance to exogenous auxin, as well as a variety of morphological defects due to compromised auxin signaling (Lincoln et al., 1990; Leyser et al., 1993). We have previously shown that axr1-3 mutants also have reduced sensitivity to MeIAA (Qin et al., 2005), suggesting that MeIAA shares similar signaling components as IAA. We therefore constructed plants that were homozygous for both mes17-1 and axr1-3 and tested them for their phenotype and response to IAA and MeIAA. While mes17-1 mutant had a longer hypocotyl than wild type when grown in one-half-strength MS medium in light, axr1-3 has a shorter hypocotyl than wild type (Fig. 8; Jensen et al., 1998). The hypocotyl length of mes17-1/axr1-3 was shorter than that of wild type, similar with the hypocotyl length of axr1-3 mutant (Fig. 8). In addition, mes17-1/axr1-3 mutant plants had the same morphological phenotype as the axr1-3 mutant line, which includes irregular rosette leaves, reduced height, and reduced fertility (data not shown). When plants were tested for their responses to the root growth inhibition activity of IAA and MeIAA, the mes17-1/axr1-3 double mutant displayed reduced sensitivity to both IAA and MeIAA, as did axr1-3 (data not shown).
Figure 8.
The mes17/axr1-3 double mutant has the same hypocotyl length as axr1-3. Plants were grown on one-half-strength MS medium under continuous light for 9 d. The hypocotyl lengths of the seedlings were measured with Image J as described in “Materials and Methods.” The mean value and sd were calculated from 20 samples.
Biochemical Characterization of AtMES17
Because AtMES17 displays MeIAA hydrolase activity in vitro and likely does so in vivo, we performed a more detailed in vitro kinetic analysis of the E. coli-expressed and purified AtMES17. AtMES17 displayed hydrolase activity toward MeIAA but not MeJA, MeSA, or MeGAs (Table I). AtMES17 displayed the highest MeIAA hydrolase activity at pH 8.5 and about 60% of the highest activity at pH 6.5 or 9.5. However, at pH 8.0 or higher, nonenzymatic hydrolysis of MeIAA was also observed. We therefore used buffers with pH 7.5, which gave 93% of the maximal enzymatic activity and no observable nonenzymatic hydrolysis. Under these conditions, AtMES17 had a Km value of 13 μm and a Kcat value of 0.18 s−1 for MeIAA. The hydrolase activity was strongly inhibited (44%–75%) by 5 mm Fe2+, Fe3+, Zn2+, and Cu2+, and mildly inhibited by 5 mm Ca2+ and Mn2+ (20% and 34%, respectively). At 5 mm concentration, Na+, Mg2+, K+, and NH4+ had no effects on the hydrolase activity.
Expression Pattern of AtMES17
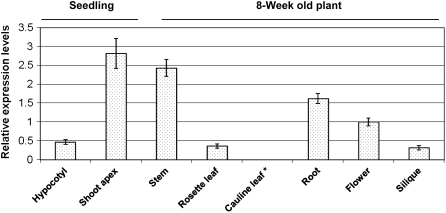
Real-time reverse transcription (RT)-PCR analysis showed that the expression of AtMES17 in 10-d-old seedlings is approximately 5-fold higher in the region of the shoot apex than in the rest of the hypocotyl (Fig. 9). When AtMES17 transcript levels in the 8-week mature plants were examined, the highest expression levels were observed in stems, followed by roots, flowers, rosette leaves, and siliques, and no AtMES17 transcripts were detectable in cauline leaves (Fig. 9).
Figure 9.
Real-time RT-PCR analysis of AtMES17 transcript levels in different plant organs at different developmental stages. AtMES17 transcript levels were normalized to the levels of ubiquitin gene expression in respective samples. The levels of AtMES17 transcript in flowers were arbitrarily set to 1.0. Data are plotted as means ± sd. *, Below the detection limit.
DISCUSSION
The Arabidopsis MES Methylesterase Family
We have identified a family of 20 Arabidopsis proteins that we have designated the AtMES family, based on the sequence similarity of these proteins to experimentally identified methyl esterases, including the tobacco MeSA esterase SABP2 (NtSABP2) and the tomato MJE (LeMJE). This AtMES family is distinct from the previously annotated AtCXE family (Fig. 1), yet both families belong to the α/β hydrolase superfamily, which is characterized by the structurally conserved “canonical” α/β hydrolase fold and catalytic residues (Nardini and Dijkstra, 1999).
Most members of the AtMES family encode proteins of approximately 250 amino acids, similarly to NtSABP2 and LeMJE. AtMES11, AtMES12, AtMES13, AtMES14, and AtMES15 contain an extra region of 90 to 190 amino acids at their N termini (Fig. 2). This N-terminal extension does not appear to constitute a targeting signal peptide, suggesting that these AtMES proteins are localized in the cytosol like the rest of the members in the family. The AtMES11, AtMES12, AtMES13, AtMES14, and AtMES15 proteins also cluster into a close clade within the AtMES family (subfamily 3, Fig. 1), and so far we have not been able to ascribe any enzymatic activity to any of the proteins in this subfamily.
Because the AtMES proteins are closely related to LeMJE and NtSABP2, we hypothesized that members of the AtMES family could encode MeIAA esterase(s). Of the members of the AtMES family that we tested, eight AtMES proteins were found to be active with MeIAA, and they all belong to subfamilies 1 and 2. Some of these proteins also hydrolyze other methyl esters under the experimental conditions used in this study, and it is likely that many, and perhaps all, of the AtMES proteins would be found to use multiple substrates upon a more extensive survey of substrates.
AtMES17 Encodes an Esterase Capable of Hydrolyzing MeIAA, Which Is Not Itself Active
We further showed that AtMES17 encodes an esterase that efficiently and specifically hydrolyzes MeIAA to IAA in vitro and is likely to do so in vivo as well. The kinetic parameters of AtMES17 are comparable to previously characterized esterases (Forouhar et al., 2005), indicating that MeIAA is likely to be a relevant substrate for AtMES17 in planta. As pointed out above, although MES17 showed activity with only MeIAA in our limited survey, we cannot yet conclude that MeIAA is its only substrate.
The growth of roots of two independent mes17 mutants was much less inhibited by MeIAA than was wild-type root growth (Fig. 3A). However, both mes17 null mutants responded similarly as did wild type to the root inhibition activity of IAA (Fig. 3B), indicating normal auxin signaling in these mutants. Incubating seedlings in the presence of [14C]MeIAA also revealed that mes17 mutants were much less efficient in hydrolyzing [14C]MeIAA than wild-type plants (Fig. 4). We thus conclude that the response of Arabidopsis seedlings to the root inhibition activity of MeIAA is at least partly due to the hydrolytic activity of AtMES17. The observations that some hydrolysis of [14C]MeIAA occurred in the mes17 mutant line and that the root growth of mes17 mutants retained some response to the inhibitory activity of MeIAA also indicate that there are other esterases in addition to AtMES17 that participate in MeIAA hydrolysis in Arabidopsis, consistent with our finding that other AtMES proteins could hydrolyze MeIAA in vitro. These proteins include AtMES16 and AtMES18, which have the highest sequence similarities to AtMES17. However, we obtained two independent T-DNA insertional lines of AtMES16 and observed that both of these null mutants responded to MeIAA similarly to wild type, including in the root growth assay. Microarray data indicates that transcript levels of AtMES17 in roots are more than 10 times higher than those of AtMES16 (AtGenExpress Visualization Tool; Schmid et al., 2005). In addition, the double mutant mes16/mes17-1 was phenotypically indistinguishable from the mes17-1 mutant in all organs or developmental stages (data not shown). We were unable to obtain a T-DNA insertion in AtMES18. It remains to be determined whether AtMES16 or other AtMES can hydrolyze MeIAA in vivo.
The much-reduced (although not completely abolished) response of roots of mes17 mutants in the root inhibition assay with exogenously supplied MeIAA coupled with the observation that these mutants are much less efficient in the hydrolysis of MeIAA also suggest that the inhibition is due to IAA and not MeIAA. Consistent with this interpretation, when AtMES17 was overexpressed, its roots were even more sensitive to MeIAA than wild-type plants (Fig. 5), likely because MeIAA was hydrolyzed in these plants even faster than in wild type. The observation that the mes17-1/axr1-3 double mutant plants have the same phenotype as axr1-3 is also consistent with the putative role of MES17 in producing IAA, which acts upstream of AXR1. A similar albeit more extensive analysis of the effects of MeIAA treatment on axr1 and other auxin response mutants has also concluded that MeIAA is likely to be inactive by itself (Li et al., 2008). In addition, the recently solved structure of the auxin receptor TIR1 supports the notion that MeIAA is not an active auxin, because it was shown that the carboxyl group of the IAA molecule interacts with two residues in the binding pocket of TIR1, docking IAA to the bottom of the pocket (Tan et al., 2007). MeIAA has a methyl ester group instead of a carboxyl group and therefore is not likely to be accommodated in the binding site of TIR1 to initiate TIR1-mediated auxin signaling. Analogous to our finding, it was recently shown that silencing of a tobacco MJE (NaMJE) reduced MeJA- but not JA-induced herbivore resistance, indicating that the resistance elicited by MeJA treatment is directly elicited not by MeJA but by its demethylated product, JA (Wu et al., 2008).
Possible Role of MeIAA in Arabidopsis
mes17-1 mutants have longer hypocotyls than wild-type plants. The regulation of hypocotyl length is a complex process that is under the influence of many factors, including light, nutrients, and hormones such as IAA, ethylene, and brassinosteroids (Jensen et al., 1998; Collett et al., 2000; Vandenbussche et al., 2005). Earlier physiological studies have shown that auxin promotes the growth of excised hypocotyl segments from various plant species (Evans, 1985). Several Arabidopsis mutants that accumulate increased overall auxin levels also have longer hypocotyls than wild type, although the auxin gradient may be more important than its actual concentration (Boerjan et al., 1995; Zhao et al., 2001, 2002). In addition, Jensen et al. (1998) have demonstrated that auxin transport from the shoot apex to the root is required for hypocotyl elongation in Arabidopsis during development in the light.
In seedlings, AtMES17 is expressed at highest levels in the shoot apex, but it is also expressed at lower levels elsewhere (Fig. 9). mes17-1 plants display a stronger auxin response in the shoot apex as well as in other parts of the plant (as assessed by the DR5:GUS reporter system; Fig. 7). Although it seems paradoxical that mes17 mutants appear to have higher levels of IAA, it may be that the higher GUS staining in this line indicates a higher rate of transport of IAA rather than a higher level of IAA concentration, brought about by higher but transient and localized concentrations of MeIAA due to the decrease in (but not complete absence of) overall MeIAA esterase activity. Methylation of IAA to enhance its transport (and subsequent hydrolysis by MES enzymes) would be analogous to the transport of SA as biologically inactive MeSA from the site of infection to distal tissue for development of systemic acquired resistance (Park et al., 2007). For this explanation to be valid, however, it would appear necessary to postulate that there are distinct types of cells, probably in close proximity to each other, that contain either IAA methylation activity or MeIAA esterase activity.
A high-resolution spatial map depicting auxin concentration as well as activities of MES17 and IAMT is needed to validate this hypothesis. Recently, a cell type-specific microarray analysis has shown that in a given area of roots, genes involved in auxin biosynthesis are expressed in different cells than genes regulating auxin homeostasis or auxin transport (Brady et al., 2007). Transient methylation of IAA and small local differences in expression of MES17 (and IAMT) among populations of cells may also explain our failure to detect differences in MeIAA concentration between wild-type, mes17, and 35S∷IAMT lines, which all contain MeIAA levels that are barely detectable (Y. Yang, unpublished data). The expression of YUCCA genes, involved in biosynthesis of auxin, are localized to small populations of cells, and the actual differences in IAA concentrations in yucca mutants are also difficult to demonstrate, despite strong defects in organ formation in these mutants (Cheng et al., 2007). Alternatively, it is possible that other auxin biosynthetic pathways are induced in response to the loss of MES17 activity in the shoot apex as well as in other parts of the plants.
In conclusion, our results suggest that MES17 functions in auxin homeostasis in vivo and that MeIAA itself is not an active auxin. Because MeIAA is more nonpolar than IAA, MeIAA could more easily diffuse across membranes, and it is therefore possible that transport of IAA (in the form of MeIAA) to neighboring cells or even to more distant targets could be enhanced, where it could be hydrolyzed back to the active auxin IAA by esterases belonging to the MES family.
MATERIALS AND METHODS
Plant Material and Growth Conditions
Wild-type Arabidopsis (Arabidopsis thaliana) ecotype Columbia was used in all experiments. The AtMES17 full-length cDNA was ligated into pCHF3 vector (Varbanova et al., 2007) using the Gateway system (Hartley et al., 2000). The resulting 35S∷AtMES17 construct was then introduced into wild-type Arabidopsis using Agrobacterium-mediated transformation by the floral dip method (Clough and Bent, 1998). Three independent homozygous transgenic lines were selected by examining the pattern of kanamycin resistance in T2 and T3 generations, and overexpression of AtMES17 in these homozygous lines was confirmed by northern blot (D'Auria et al., 2002).
Arabidopsis plants grown in soil were under 16-h-light/8-h-dark cycles at 22°C. Arabidopsis plants grown on one-half-strength MS medium (Murashige and Skoog, 1962) were subjected to constant light at 22°C.
Chemicals
All chemicals were purchased from Sigma. MeIAA and IAA were dissolved in 95% ethanol to make stock solutions of different concentrations. Stock solutions were then diluted 1:1,000 into one-half-strength MS medium, and the medium was poured into square plates. Plates containing chemicals were wrapped in aluminum foil and stored at 4°C before use.
Protein Expression and Purification
Isolation of AtMES cDNAs and construction of Escherichia coli expression vectors of all AtMES genes, except AtMES11, AtMES12, and AtMES18, are described elsewhere (A.C. Vlot and D.F. Klessig, unpublished data). Full-length cDNA of AtMES11 (U22904), AtMES12 (U15905), and AtMES18 (U50042) were obtained from the Arabidopsis Biological Resource Center (ABRC), and subcloned into pENTR/D-TOPO (Invitrogen) and subsequently p-His-9 vector (a Gateway adapted derivative of pET28a). The plasmid containing the respective AtMES cDNA was transformed into E. coli and expressed as previously described (Nam et al., 1999), with the following minor modifications. All expression constructs in this study were transformed into the E. coli cell line BL21 Codon plus. E. coli cells were grown to an OD600 of 0.4, then induced with 0.4 mm isopropylthio-β-galactoside and grown at 18°C overnight. The cell lysate used in esterase enzyme assays or protein purification was first examined by SDS-PAGE to ensure that the protein encoded by the cDNA was expressed.
For protein purification, nickel-nitrilotriacetic acid agarose (Qiagen) was loaded into a column and washed with 10 bed volumes of water followed by 10 bed volumes of lysis buffer (50 mm Tris-HCl, pH 8.0, 500 mm NaCl, 20 mm imidazole, pH 8.0, 20 mm β-mercaptoethanol, 10% [v/v] glycerol, and 1% [v/v] Tween 20). Ten bed volumes of cell lysate was passed over the column and subsequently washed with 10 bed volumes of lysis buffer, and 20 bed volumes of wash buffer (50 mm Tris-HCl, pH 8.0, 500 mm NaCl, 20 mm imidazole, pH 8.0, 20 mm β-mercaptoethanol, and 10% [v/v] glycerol). The protein was eluted with elution buffer (50 mm Tris-HCl, pH 8.0, 500 mm NaCl, 250 mm imidazole, pH 8.0, 20 mm β-mercaptoethanol, and 10% [v/v] glycerol) and collected in 0.5-mL fractions. After being examined by SDS-PAGE, elution fractions containing the most abundant purified proteins were pooled and concentrated by centrifugation in the Amicon Ultra-4 centrifugal filter (Millipore). Concentrated proteins were finally resuspended in a buffer containing 50 mm Tris-HCl, pH 8.0, 10 mm NaCl, 20 mm β-mercaptoethanol, and 10% (v/v) glycerol. All purification procedures were performed at 4°C.
Esterase Enzyme Assay
The chymotryptic substrate PNPA was dissolved in acetonitrile to make a stock solution of 100 mm. An assay was prepared containing 50 mm Tris-HCl, pH 7.5, 0.05% Triton X-100, 1 mm PNPA, and 200 μL expression lysate. Control assays were set up in parallel with denatured protein. Esterase activity was estimated by the rate of hydrolysis determined spectrophometrically at 410 nm. The assay was carried out at room temperature, and OD410 values were measured at 2-min intervals up to 30 min. All assays were performed in duplicate. An AtMES protein was considered active when the reaction product determined by OD410 was at least 3 times that of the control assay.
Esterase assays with MeIAA, MeSA, MeJA, MeGA4, and MeGA9 as substrates were performed using the coupled methyltransferase assay, as previously described (Forouhar et al., 2005). All assays were performed in triplicate.
For kinetic analysis of AtMES17, the amount of IAA generated from the esterase assay was quantified by HPLC analysis on a Waters 2690 Separations Module. HPLC separation of MeIAA and IAA was achieved over a Waters Nova-Pak C18 column, using an 8-min linear gradient from 65% acetonitrile in 1.5% phosphoric acid to 90% acetonitrile, with the flow rate set at 1 mL/min and the column temperature set to 30°C. In-line UV light spectra (200–450 nm) were obtained using an attached Waters 996 photodiode array detector. Eluting compounds were identified by comparison of both UV light spectra and elution volume with authentic MeIAA and IAA. IAA peak area detected at 278.4 nm (the maximum absorption wavelength for IAA) was plotted onto a standard curve created at identical parameters to calculate the product of each reaction.
[14C]MeIAA Uptake and in Vivo Hydrolysis Assays
[14C]MeIAA was produced by incubating IAA with [14C]SAM and IAMT under assay conditions described previously (Zubieta et al., 2003). Seedlings (8 d old) were incubated in a 100-μL solution containing 0.5 μm [14C]MeIAA and 50 mm Tris-HCl, pH 7.5. After 30 min of incubation, the solution was removed, the seedlings were washed with 1 mL of distilled, deionized water three times, and then ground with a pestle in 100 μL of Tris-HCl buffer. [14C]MeIAA in the plants was then extracted with ethyl acetate and analyzed by radio-TLC and in a scintillation counter as previously described (Fridman et al., 2005). [14C]MeIAA was loaded on the same TLC plate to show the position of MeIAA. Each experiment was repeated three times and results calculated on per fresh weight basis.
Characterization of AtMES17 Kinetic Parameters
Appropriate enzyme concentrations and incubation time were chosen so that the reaction velocity was linear over time with no more than 10% of the substrate consumed during the time period. The determination of kinetic parameters was as described (Yang et al., 2006b), except that 50 mm BisTris propane, pH 7.5, was used to examine all steady-state kinetics, because control assays prepared with denatured enzyme indicated that nonenzymatic hydrolysis occurs at pH 8.0 and increases as pH increases.
Screening of T-DNA Insertional Mutants
The following T-DNA insertional mutants were obtained from ABRC: Salk_006044 (AtMES1), Salk_030442 (AtMES9), Salk_151578 (AtMES16), Salk_139756 (AtMES16), Salk_092550 (AtMES17), and SAIL-503-c03 (AtMES17). The T-DNA insertion sites in these AtMES genes were verified first by PCR using T-DNA-specific primer SALKLBb1 (5′-GCGTGGACCGCTTGCTGCAACT-3′, for SALK lines) or SAILLB3 (5′-TAGCATCTGAATTTCATAACCAATCT-3′, for SAIL lines) and the genomic primers designed for each T-DNA insertional line as follows: Salk_006044 forward (5′-CACCGAACACTCACCATCCTTCG-3′) and reverse (5′-TTAAACGAATTTGTCCGCGATTTTCAG-3′); Salk_030442 forward (5′-ATGAAGCATTATGTGCTAGTTCACGGAGGC-3′) and reverse (5′-TTAGGGATATTTATCAGCAATCTTTAGAAG-3′); Salk_151578 forward (5′-TTACTAACTCACCTCTCTTCTTCTTCG-3′) and reverse (5′-ATACGCTAAGGCATCGAAGGG-3′); Salk_139756 forward (5′CTCTCTTGTCCGATCTCCCTCC-3′) and reverse (5′-CCCTGGATTGCTTCGCATG-3′); Salk_092550 forward (5′-GCGTTTGACAAATGTGACAAGGC-3′) and reverse (5′-GGTTTGATAATAGCACTGGTGGG-3′); and SAIL-503-c03 forward (5′-ATGGCGGAGGAGAATC-3′) and reverse (5′-TTAGATAGAACCGACGGAAACGGC-3′). PCR results were verified by sequencing. All homozygous T-DNA insertional lines were confirmed by PCR with specific primers and subsequent Southern blot. RT-PCR was done with RNA extracted from homozygous lines to ensure absence of the respective gene transcript (see Supplemental Fig. S1 for mes17 mutants). Homozygous T-DNA insertional lines were also obtained for AtMES2 (Salk_050266), AtMES7 (Salk_054303, Salk_036791), and AtMES18 (CS826062). However, full-length gene transcripts were detectable in these mutants.
Measurement of Root Length and Hypocotyl Length
The root length of seedlings was measured with a ruler, and at least 20 measurements were taken to calculate the mean and sd values. Hypocotyl lengths of 4-week-old plants grown in soil were measured with a ruler. To measure hypocotyl length of seedlings grown on plates, seedlings were gently lifted with forceps from plates onto acetate sheets and digitized with a flat-bed scanner at a resolution of 1,200 dpi. Seedling scans were analyzed by ImageJ 1.37v software (National Institutes of Health), through which the hypocotyl lengths of seedling were measured. Twenty seedlings were analyzed for each measurement to calculate mean and sd values.
Real-Time RT-PCR Analysis
RNA extraction, purification, and real-time RT-PCR were performed as described (Varbanova et al., 2007). Shoot apex and hypocotyl were collected from 10-d-old plants grown in soil. Flowers, siliques, stems, rosette leaves, cauline leaves, and roots were collected from 8-week-old flowering plants grown in soil. AtMES17 gene-specific primers were designed as follows: forward 5′-GTTTTGGTCTAGGACCGGAGAATC-3′ and reverse 5′-CCAAGGAACATTCCTGTTGAGG-3′.
DR5:GUS Reporter Analysis
DR5:GUS/mes17 plants were obtained by crossing the DR5:GUS line into the mes17-1 mutant line. Plants homozygous for both DR5:GUS and mes17-1 were analyzed for GUS activity and compared to that of wild-type DR5:GUS. Seedlings were grown on one-half-strength MS medium for 8 d and incubated in GUS staining solution (100 mm sodium phosphate buffer, pH 6.8, 10 mm EDTA, 0.2% Triton X-100, and 0.2 mg/mL 5-bromo-4-chloro-3-indolyl-β-d-glucuronide) for 16 h, after which chlorophyll were extracted with 75% ethanol for 24 h. Quantitative GUS assay was carried out as described by Nakamura et al. (2003) except that 1.3 mm 4-methylumbelliferyl-β-d-glucuronide was used and the assay was carried out for 32 min.
Supplemental Data
The following materials are available in the online version of this article.
Supplemental Figure S1. mes17-1 and mes17-2 are null mutants of AtMES17.
Supplementary Material
Acknowledgments
We thank Dr. Mark Estelle at the University of Indiana for providing the axr1-3 line. We thank Dr. Yunde Zhao at the University of California at San Diego for providing the DR5:GUS line.
This work was supported by the National Science Foundation (Arabidopsis 2010 project grant no. MCB–0312466 to E.P., and grant no. IOB–0525360 to D.F.K.).
The author responsible for distribution of materials integral to the findings presented in this article in accordance with the policy described in the Instructions for Authors (www.plantphysiol.org) is: Eran Pichersky (lelx@umich.edu).
The online version of this article contains Web-only data.
Open Access articles can be viewed online without a subscription.
References
- Bartel B, Fink G (1995) ILR1, an amidohydrolase that releases active indole-3-acetic acid from conjugates. Science 268 1745–1748 [DOI] [PubMed] [Google Scholar]
- Baudouin E, Charpenteau M, Roby D, Marco Y, Ranjeva R, Ranty B (1997) Functional expression of a tobacco gene related to the serine hydrolase family-esterase activity towards short-chain dinitrophenyl acylesters. Eur J Biochem 248 700–706 [DOI] [PubMed] [Google Scholar]
- Boerjan W, Cervera MT, Delarue M, Beeckman T, Dewitte W, Bellini C, Caboche M, Onckelen HV, Montagu MV, Inze D (1995) superroot, A recessive mutation in Arabidopsis, confers auxin overproduction. Plant Cell 7 1405–1419 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brady SM, Orlando DA, Lee JY, Wang JY, Koch J, Dinneny JR, Mace D, Ohler U, Benfey PN (2007) A high-resolution root spatiotemporal map reveals dominant expression patterns. Science 318 801–806 [DOI] [PubMed] [Google Scholar]
- Chen F, D'Auria JC, Tholl D, Ross JR, Gershenzon J, Noel JP, Pichersky E (2003) An Arabidopsis thaliana gene for methylsalicylate biosynthesis, identified by a biochemical genomics approach, has a role in defense. Plant J 36 577–588 [DOI] [PubMed] [Google Scholar]
- Cheng Y, Dai X, Zhao Y (2007) Auxin synthesized by the YUCCA flavin monooxygenases is essential for embryogenesis and leaf formation in Arabidopsis. Plant Cell 19 2430–2439 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Clough SJ, Bent AF (1998) Floral dip: a simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J 16 735–743 [DOI] [PubMed] [Google Scholar]
- Collett CE, Harberd NP, Leyser O (2000) Hormonal interactions in the control of Arabidopsis hypocotyl elongation. Plant Physiol 124 553–562 [DOI] [PMC free article] [PubMed] [Google Scholar]
- D'Auria JC, Chen F, Pichersky E (2002) Characterization of an acyltransferase capable of synthesizing benzylbenzoate and other volatile esters in flowers and damaged leaves of Clarkia breweri. Plant Physiol 130 466–476 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Delker C, Raschke A, Quint M (2008) Auxin dynamics: the dazzling complexity of a small molecule's message. Planta 227 929–941 [DOI] [PubMed] [Google Scholar]
- Ding X, Cao Y, Huang L, Zhao J, Xu C, Li X, Wang S (2008) Activation of the indole-3-acetic acid amido synthetase GH3-8 suppresses expansin expression and promotes salicylate- and jasmonate-independent basal immunity in rice. Plant Cell 20 228–240 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dogru E, Warzecha H, Seibel F, Haebel S, Lottspeich F, Stöckigt J (2000) The gene encoding polyneuridine aldehyde esterase of monoterpenoid indole alkaloid biosynthesis in plants is an ortholog of the alpha/beta hydrolase super family. Eur J Biochem 267 1397–1406 [DOI] [PubMed] [Google Scholar]
- Evans ML (1985) The action of auxin on plant-cell elongation. Crc Crit Rev Plant Sci 2 317–365 [DOI] [PubMed] [Google Scholar]
- Forouhar F, Yang Y, Kumar D, Chen Y, Fridman E, Park SW, Chiang Y, Acton TB, Montelione GT, Pichersky E, et al (2005) Structural and biochemical studies identify tobacco SABP2 as a methyl salicylate esterase and implicate it in plant innate immunity. Proc Natl Acad Sci USA 102 1773–1778 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fridman E, Wang J, Iijima Y, Froehlich JE, Gang DR, Ohlrogge J, Pichersky E (2005) Metabolic, genomic, and biochemical analyses of glandular trichomes from the wild tomato species Lycopersicon hirsutum identify a key enzyme in the biosynthesis of methylketones. Plant Cell 17 1252–1267 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hartley JL, Temple GF, Brasch MA (2000) DNA cloning using in vitro site-specific recombination. Genome Res 10 1788–1795 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hikage T, Saitoh Y, Tanaka-Saito C, Hagami H, Satou F, Shimotai Y, Nakano Y, Takahashi M, Takahata Y, Tsutsumi K (2007) Structure and allele-specific expression variation of novel alpha/beta hydrolase fold proteins in gentian plants. Mol Genet Genomics 278 95–104 [DOI] [PubMed] [Google Scholar]
- Holmquist M (2000) Alpha beta-hydrolase fold enzymes structures, functions and mechanisms. Curr Protein Pept Sci 1 209–235 [DOI] [PubMed] [Google Scholar]
- Ichinose Y, Hisayasu Y, Sanematsu S, Ishiga Y, Seki H, Toyoda K, Shiraishi T, Yamada T (2001) Molecular cloning and functional analysis of pea cDNA E86 encoding homologous protein to hypersensitivity-related hsr203J. Plant Sci 160 997–1006 [DOI] [PubMed] [Google Scholar]
- Jackson RG, Lim EK, Li Y, Kowalczyk M, Sandberg G, Hoggett J, Ashford DA, Bowles DJ (2001) Identification and biochemical characterization of an Arabidopsis indole-3-acetic acid glucosyltransferase. J Biol Chem 276 4350–4356 [DOI] [PubMed] [Google Scholar]
- Jensen PJ, Hangarter RP, Estelle M (1998) Auxin transport is required for hypocotyl elongation in light-grown but not dark-grown Arabidopsis. Plant Physiol 116 455–462 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ko MK, Jeon WB, Kim KS, Lee HH, Seo HH, Kim YS, Oh BJ (2005) A Colletotrichum gloeosporioides-induced esterase gene of nonclimacteric pepper (Capsicum annuum) fruit during ripening plays a role in resistance against fungal infection. Plant Mol Biol 58 529–541 [DOI] [PubMed] [Google Scholar]
- Kumar D, Klessig DF (2003) High-affinity salicylic acid-binding protein 2 is required for plant innate immunity and has salicylic acid-stimulated lipase activity. Proc Natl Acad Sci USA 100 16101–16106 [DOI] [PMC free article] [PubMed] [Google Scholar]
- LeClere S, Tellez R, Rampey RA, Matsuda SPT, Bartel B (2002) Characterization of a family of IAA-amino acid conjugate hydrolases from Arabidopsis. J Biol Chem 277 20446–20452 [DOI] [PubMed] [Google Scholar]
- Leyser HM, Lincoln CA, Timpte C, Lammer D, Turner J, Estelle M (1993) Arabidopsis auxin-resistance gene AXR1 encodes a protein related to ubiquitin-activating enzyme E1. Nature 364 161–164 [DOI] [PubMed] [Google Scholar]
- Li L, Hou X, Tsuge T, Ding M, Aoyama T, Oka A, Gu H, Zhao Y, Qu LJ (2008) The possible action mechanisms of indole-3-acetic acid methyl ester in Arabidopsis. Plant Cell Rep 27 575–584 [DOI] [PubMed] [Google Scholar]
- Lincoln C, Britton JH, Estelle M (1990) Growth and development of the Axr1 mutants of Arabidopsis. Plant Cell 2 1071–1080 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ljung K, Hull AK, Kowalczyk M, Marchant A, Celenza J, Cohen JD, Sandberg G (2002) Biosynthesis, conjugation, catabolism and homeostasis of indole-3-acetic acid in Arabidopsis thaliana. Plant Mol Biol 50 309–332 [DOI] [PubMed] [Google Scholar]
- Marshall SG, Putterill J, Plummer K, Newcomb R (2003) The carboxylesterase gene family from Arabidopsis thaliana. J Mol Evol V57 487–500 [DOI] [PubMed] [Google Scholar]
- Murashige T, Skoog F (1962) A revised medium for rapid growth and bioassays with tobacco tissue cultures. Physiol Plant 15 473–497 [Google Scholar]
- Nakamura A, Higuchi K, Goda H, Fujiwara MT, Sawa S, Koshiba T, Shimada Y, Yoshida S (2003) Brassinolide induces IAA5, IAA19, and DR5, a synthetic auxin response element in Arabidopsis, implying a cross talk point of brassinosteroid and auxin signaling. Plant Physiol 133 1843–1853 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nam KH, Dudareva N, Pichersky E (1999) Characterization of benzylalcohol acetyltransferases in scented and non-scented Clarkia species. Plant Cell Physiol 40 916–923 [DOI] [PubMed] [Google Scholar]
- Narasimhan K, Basheer C, Bajic VB, Swarup S (2003) Enhancement of plant-microbe interactions using a rhizosphere metabolomics-driven approach and its application in the removal of polychlorinated biphenyls. Plant Physiol 132 146–153 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nardini M, Dijkstra BW (1999) Alpha/beta hydrolase fold enzymes: the family keeps growing. Curr Opin Struct Biol 9 732–737 [DOI] [PubMed] [Google Scholar]
- Normanly J (1997) Auxin metabolism. Physiol Plant 100 431–442 [Google Scholar]
- Nowacki J, Bandurski RS (1980) Myoinositol esters of indole-3-acetic-acid as seed auxin precursors of Zea Mays L. Plant Physiol 65 422–427 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ottenschlager I, Wolff P, Wolverton C, Bhalerao RP, Sandberg G, Ishikawa H, Evans M, Palme K (2003) Gravity-regulated differential auxin transport from columella to lateral root cap cells. Proc Natl Acad Sci USA 100 2987–2991 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Park SW, Kaimoyo E, Kumar D, Mosher S, Klessig DF (2007) Methyl salicylate is a critical mobile signal for plant systemic acquired resistance. Science 318 113–116 [DOI] [PubMed] [Google Scholar]
- Pontier D, Godiard L, Marco Y, Roby D (1994) Hsr203j, A tobacco gene whose activation is rapid, highly localized and specific for incompatible plant/pathogen interactions. Plant J 5 507–521 [DOI] [PubMed] [Google Scholar]
- Pontier D, Tronchet M, Rogowsky P, Lam E, Roby D (1998) Activation of hsr203, a plant gene expressed during incompatible plant-pathogen interactions, is correlated with programmed cell death. Mol Plant Microbe Interact 11 544–554 [DOI] [PubMed] [Google Scholar]
- Qin G, Gu H, Zhao Y, Ma Z, Shi G, Yang Y, Pichersky E, Chen H, Liu M, Chen Z, et al (2005) Regulation of Arabidopsis leaf development by an indole-3-acetic acid carboxyl methyltransferase in Arabidopsis. Plant Cell 17 2693–2704 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rampey RA, LeClere S, Kowalczyk M, Ljung K, Sandberg G, Bartel B (2004) A family of auxin-conjugate hydrolases that contributes to free indole-3-acetic acid levels during Arabidopsis germination. Plant Physiol 135 978–988 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schmid M, Davison TS, Henz SR, Pape UJ, Demar M, Vingron M, Scholkopf B, Weigel D, Lohmann JU (2005) A gene expression map of Arabidopsis thaliana development. Nat Genet 37 501–506 [DOI] [PubMed] [Google Scholar]
- Shulaev V, Silverman P, Raskin I (1997) Airborne signalling by methyl salicylate in plant pathogen resistance. Nature 385 718–721 [Google Scholar]
- Staswick PE, Serban B, Rowe M, Tiryaki I, Maldonado MT, Maldonado MC, Suza W (2005) Characterization of an Arabidopsis enzyme family that conjugates amino acids to indole-3-acetic acid. Plant Cell 17 616–627 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Stuhlfelder C, Mueller MJ, Warzecha H (2004) Cloning and expression of a tomato cDNA encoding a methyl jasmonate cleaving esterase. Eur J Biochem 271 2976–2983 [DOI] [PubMed] [Google Scholar]
- Tan X, Calderon-Villalobos LIA, Sharon M, Zheng C, Robinson CV, Estelle M, Zheng N (2007) Mechanism of auxin perception by the TIR1 ubiquitin ligase. Nature 446 640–645 [DOI] [PubMed] [Google Scholar]
- Teale WD, Paponov IA, Palme K (2006) Auxin in action: signalling, transport and the control of plant growth and development. Nat Rev Mol Cell Biol 7 847–859 [DOI] [PubMed] [Google Scholar]
- Thompson JD, Gibson TJ, Plewniak F, Jeanmougin F, Higgins DG (1997) The clustalx windows interface: flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res 24 4876–4882 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Ulmasov T, Murfett J, Hagen G, Guilfoyle TJ (1997) Aux/IAA proteins repress expression of reporter genes containing natural and highly active synthetic auxin response elements. Plant Cell 9 1963–1971 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Vandenbussche F, Verbelen P, Van Der Straeten D (2005) Of light and length: regulation of hypocotyl growth in Arabidopsis. Bioessays 27 275–284 [DOI] [PubMed] [Google Scholar]
- Varbanova M, Yamaguchi S, Yang Y, McKelvey K, Hanada A, Borochov R, Yu F, Jikumaru Y, Ross J, Cortes D, et al (2007) Methylation of gibberellins by Arabidopsis GAMT1 and GAMT2. Plant Cell 19 32–45 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Walden AR, Walter C, Gardner RC (1999) Genes expressed in Pinus radiata male cones include homologs to anther-specific and pathogenesis response genes. Plant Physiol 121 1103–1116 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Woodward AW, Bartel B (2005) Auxin: regulation, action, and interaction. Ann Bot (Lond) 95 707–735 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wu J, Wang L, Baldwin I (2008) Methyl jasmonate-elicited herbivore resistance: does MeJA function as a signal without being hydrolyzed to JA? Planta 227 1161–1168 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang Y, Varbanova M, Ross J, Wang G, Cortes D, Fridman E, Shulaev V, Noel JP, Pichersky E (2006. a) Methylation and demethylation of plant signaling molecules. In JT Romeo, ed, Recent Advances in Phytochemistry, Ed 1, Vol 40. Elsevier Science, Oxford, p 253
- Yang Y, Yuan JS, Ross J, Noel JP, Pichersky E, Chen F (2006. b) An Arabidopsis thaliana methyltransferase capable of methylating farnesoic acid. Arch Biochem Biophys 448 123–132 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao N, Ferrer J, Ross J, Guan J, Yang Y, Pichersky E, Noel JP, Chen F (2008) Structural, biochemical and phylogenetic analyses suggest that indole-3-acetic acid methyltransferase is an evolutionarily ancient member of the SABATH family. Plant Physiol 146 455–467 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao Y, Hull AK, Gupta NR, Goss KA, Alonso J, Ecker JR, Normanly J, Chory J, Celenza JL (2002) Trp-dependent auxin biosynthesis in Arabidopsis: involvement of cytochrome P450s CYP79B2 and CYP79B3. Genes Dev 16 3100–3112 [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zhao YD, Christensen SK, Fankhauser C, Cashman JR, Cohen JD, Weigel D, Chory J (2001) A role for flavin monooxygenase-like enzymes in auxin biosynthesis. Science 291 306–309 [DOI] [PubMed] [Google Scholar]
- Zimmerman P, Hitchcock AE (1937) Comparative effectiveness of acids, esters, and salts as growth substances and methods of evaluating them. Contrib. Boyce Thompson Inst 8 337–350 [Google Scholar]
- Zubieta C, Ross JR, Koscheski P, Yang Y, Pichersky E, Noel JP (2003) Structural basis for substrate recognition in the salicylic acid carboxyl methyltransferase family. Plant Cell 15 1704–1716 [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.