Abstract
We present data from antimicrobial assays performed in vitro that pertain to the potential clinical utility of a novel rifamycin-quinolone hybrid antibiotic, CBR-2092, for the treatment of infections mediated by gram-positive cocci. The MIC90s for CBR-2092 against 300 clinical isolates of staphylococci and streptococci ranged from 0.008 to 0.5 μg/ml. Against Staphylococcus aureus, CBR-2092 exhibited prolonged postantibiotic effects (PAEs) and sub-MIC effects (SMEs), with values of 3.2, 6.5, and >8.5 h determined for the PAE (3× MIC), SME (0.12× MIC), and PAE-SME (3× MIC/0.12× MIC) periods, respectively. Studies of genetically defined mutants of S. aureus indicate that CBR-2092 is not a substrate for the NorA or MepA efflux pumps. In minimal bactericidal concentration and time-kill studies, CBR-2092 exhibited bactericidal activity against staphylococci that was retained against rifampin- or intermediate quinolone-resistant strains, with apparent paradoxical cidal characteristics against rifampin-resistant strains. In spontaneous resistance studies, CBR-2092 exhibited activity consistent with balanced contributions from its composite pharmacophores, with a mutant prevention concentration of 0.12 μg/ml and a resistance frequency of <10−12 determined at 1 μg/ml in agar for S. aureus. Similarly, CBR-2092 suppressed the emergence of preexisting rifamycin resistance in time-kill studies undertaken at a high cell density. In studies of the intracellular killing of S. aureus, CBR-2092 exhibited prolonged bactericidal activity that was superior to the activities of moxifloxacin, rifampin, and a cocktail of moxifloxacin and rifampin. Overall, CBR-2092 exhibited promising activity in a range of antimicrobial assays performed in vitro that pertain to properties relevant to the effective treatment of serious infections mediated by gram-positive cocci.
Recent studies of the epidemiology of methicillin-resistant Staphylococcus aureus (MRSA) infections in the United States have confirmed its status as the predominant pathogen associated with invasive skin and soft tissue infections in both the community and health care settings (12, 14). The emergence and spread of invasive MRSA isolates, including those exhibiting multidrug-resistant traits (15), have spurred renewed interest in the potential use of older classes of antistaphylococcal agents for the treatment of serious infections. In an accompanying article (18), we describe detailed mode-of-action studies for CBR-2092, a novel rifampin-quinolone hybrid antibiotic that combines the rifamycin SV and a 4H-4-oxo-quinolizine pharmacophores via a chiral linking group.
Biochemical studies indicate that CBR-2092 exhibits rifampin-like potency as an inhibitor of RNA polymerase and is an equipotent (or balanced) inhibitor of DNA gyrase and DNA topoisomerase IV that retains activity against a prevalent quinolone-resistant variant. Macromolecular biosynthesis studies confirm that CBR-2092 causes rifampin-like effects on RNA and protein synthesis in rifampin-susceptible strains and quinolone-like effects on DNA synthesis in rifampin-resistant strains. Finally, data from studies on the isolation and characterization of mutant strains that exhibit reduced susceptibility to CBR-2092 substantiated that RNA polymerase is the primary cellular target of CBR-2092, with DNA gyrase and DNA topoisomerase IV being secondary and tertiary targets, respectively, in strains exhibiting preexisting rifampin resistance. In contrast to quinolone comparator agents, strains with altered susceptibility to CBR-2092 were not found to exhibit changes consistent with altered efflux properties (18). Overall, the combined data suggest that CBR-2092 may have potential utility as monotherapy for the treatment of S. aureus infections and those mediated by other susceptible cocci. Here we describe the results of microbiology studies conducted to further characterize the in vitro profile of activity of CBR-2092 against staphylococci and streptococci and to compare its activity to the activities of antibiotics of the parental rifamycin and fluoroquinolone classes.
(Portions of this work were previously presented at the 47th Interscience Conference on Antimicrobial Agents and Chemotherapy, 2007 [8a, 12a, 19].)
MATERIALS AND METHODS
Antimicrobial agents.
CBR-2092 was synthesized at Cumbre Pharmaceuticals Inc. Rifampin, ethidium bromide, and reserpine were purchased from Sigma-Aldrich, Inc. (St. Louis, MO). Ciprofloxacin, levofloxacin, gatifloxacin, moxifloxacin, and nadifloxacin were purchased from LKT Laboratories, Inc. (St. Paul, MN).
Bacterial strains.
A total of 300 gram-positive, aerobic, clinical isolates, including methicillin-susceptible S. aureus (MSSA), MRSA, methicillin-susceptible Staphylococcus epidermidis (MSSE), methicillin-resistant S. epidermidis (MRSE), Streptococcus pyogenes, Streptococcus agalactiae, and Streptococcus pneumoniae, were studied. These isolates comprised isolates sourced independently by (i) Focus Technologies Inc. (now Eurofins MediNet Inc.; Herndon, VA) from North American and European contributors to The Surveillance Network collection in years prior to 2004 and (ii) Micromyx LLC (Kalamazoo, MI) from North American and European contributors in years prior to 2006. Other strains employed in these studies are listed in Table 1. The isolation and characterization of derivatives of S. aureus ATCC 29213 (CB190) that exhibit stable resistance to agents of the rifamycin or quinolone classes are described in the accompanying article (18). Derivatives of Streptococcus pyogenes CB1208 that exhibit intermediate resistance (i.e., MIC, 1 μg/ml; strain CB1456) or high-level resistance (i.e., MIC, >32 μg/ml; strain CB1504) to rifampin were isolated sequentially by spontaneous selection on Trypticase soy agar medium supplemented with 5% sheep's blood (TSBA) and 1 or 16 μg/ml rifampin, respectively. Genotypic analysis of these strains was undertaken by standard methods (18).
TABLE 1.
Bacterial strains used in this study
| Strain | Organism | Relevant genotype and/or characteristic | Source or reference |
|---|---|---|---|
| CB190 | S. aureus | ATCC 29213; wild-type parent strain for isogenic strain panel | ATCC |
| CB370 | S. aureus | rpoBH481Y; spontaneous rifampin-resistant variant of CB190 | Robertson et al. (18) |
| CB808 | S. aureus | parCS80F; spontaneous ciprofloxacin-resistant variant of CB190 | Robertson et al. (18) |
| CB809 | S. aureus | rpoBH481Y parCS80F; ciprofloxacin-resistant variant of CB370 | Robertson et al. (18) |
| CB811 | S. aureus | gyrAS84L; spontaneous nadifloxacin-resistant variant of CB190 | Robertson et al. (18) |
| CB812 | S. aureus | rpoBH481Y gyrAS84L; nadifloxacin-resistant variant of CB370 | Robertson et al. (18) |
| CB814 | S. aureus | parCS80FgyrAS84L; higher level ciprofloxacin-resistant variant of CB808 | Robertson et al. (18) |
| CB815 | S. aureus | rpoBH481Y parCS80FgyrAS84L; higher-level ciprofloxacin-resistant variant of CB809 | Robertson et al. (18) |
| CB1406 | S. aureus | ATCC 25923; clinical isolate used for intracellular killing assays | ATCC |
| CB402 | S. aureus | ATCC 12600 (NCTC 8532); clinical isolate, parent strain of CB1927 | ATCC |
| CB1927 | S. aureus | Small colony variant owing to ΔhemB::ermC derived from CB402 | This work |
| CB191 | S. epidermidis | ATCC 35984; methicillin-resistant, gentamicin-resistant clinical isolate | ATCC |
| SA-8325-4 | S. aureus | Parent of SA-K2068 | Kaatz et al. (10) |
| SA-K2068 | S. aureus | MDR mutant of NCTC 8325-4 with elevated efflux activity via MepA | Kaatz et al. (10) |
| CB990 | S. aureus | Mutant of CB190 selected on norfloxacin and ethidium bromide with elevated NorA activity due to a deletion of 4 nucleotides (+3 to +6) in the norA promoter region | This work |
| CB1208 | S. pyogenes | Clinical isolate (Focus Technologies isolate 1166966); rifampin sensitive | Eurofins Medinet Inc. |
| CB1456 | S. pyogenes | rpoBA624D; intermediate rifampin-resistant variant of CB1208 | This work |
| CB1504 | S. pyogenes | rpoBA624D, H486Y; high-level rifampin-resistant variant of CB1456 | This work |
| CU357 | E. coli | DB3.1 host carrying pKOR1 vector for allelic replacement in S. aureus | Bae and Schneewind (2) |
| CU373 | E. coli | GC5 host carrying pKOR1-based ΔhemB::ermC allelic replacement construct | This work |
Deletion-replacement of the hemB locus for the generation of a stable S. aureus small colony variant (17, 25) was accomplished by the pKOR1-mediated allelic replacement methodology (2). Briefly, a linear ΔhemB::ermC amplicon was generated by splicing-overlap extension PCR methods with oligonucleotide primers specific for the 5′ and 3′ flanking regions of hemB or for the ermC coding sequence and promoter region originally obtained from pMP963 (11). Outside primers were engineered to also carry site-specific recombination attB sites for recombination into pKOR1 (2). Deletion-replacement of hemB was accomplished by allelic replacement with a pKOR1-based ΔhemB::ermC construct with selection on hemin (1 μg/ml)-supplemented medium with anhydrotetracycline (1 μg/ml) and erythromycin (2 μg/ml). A single erythromycin-resistant, chloramphenicol-sensitive recombinant (designated CB1927) was identified and was subsequently confirmed to bear the appropriate ΔhemB::ermC deletion-replacement mutation by diagnostic PCR analysis with outside flanking oligonucleotide primers.
Determination of MICs.
Determination of MICs was done in accordance with the Clinical and Laboratory Standards Institute (CLSI) methodology (3) either by the broth microdilution method with cation-adjusted Mueller Hinton (MHII) medium further supplemented with 0.002% (vol/vol) polysorbate 80 (P-80) (18) or by the agar dilution method with MHII agar medium without P-80 supplementation.
Determination of bactericidal activity.
Bactericidal activity was assessed by determining minimum bactericidal concentration (MBC) endpoints and by time-kill assays (8). The MBC was determined on charcoal agar medium following assessment of the broth microdilution MIC after 18 to 24 h of incubation, with the MBC defined as the lowest consecutive drug concentration that resulted in a ≥99.9% (3-log10) reduction in the starting inoculum in terms of the input CFU. Standard- or high-initial-cell-density time-kill studies employed log-phase cultures of S. aureus which were diluted into prewarmed 37°C MHII broth medium with 0.002% P-80 to yield starting initial cell densities of ≈106 and 108 CFU/ml, respectively. Antimicrobial agents were then added from 100× the final desired dose in dimethyl sulfoxide aliquots to yield the indicated final test concentrations. Growth or killing was measured following serial dilution in physiologic saline and enumeration of the viable CFU recovered after 20 h of incubation on charcoal agar medium. To test for the emergence of resistant subpopulations, samples were also plated on MHII agar further supplemented with either rifampin at 1 μg/ml or ciprofloxacin at 1 or 4 μg/ml. All time-kill assays involving staphylococci were conducted under ambient atmospheric conditions in six-well tissue culture plates at 37°C with aeration to ensure consistent and robust growth. For time-kill studies with Streptococcus pyogenes, cells were harvested following overnight growth on TSBA plates and were suspended for assay in MHII broth with 0.002% P-80 but without lysed horse blood. S. pyogenes was grown statically at 35°C in an atmosphere of 5 to 10% CO2, with viable counts determined on TSBA following dilution in physiologic saline.
Spontaneous resistance selection methods.
Mutant prevention concentrations (MPCs) were determined by plating 1010 viable test organisms on MHII agar medium with twofold doubling concentrations of antibiotic. The MPC was defined as the lowest consecutive drug concentration capable of fully suppressing the emergence of antibiotic-resistant bacterial subpopulations (5). The resistance frequency was similarly determined by plating 1012 viable organisms (CFU) on a series of agar plates containing a fixed dose of 1 μg/ml of the test agent.
In vitro PAEs and SMEs.
Studies were undertaken to measure postantibiotic effects (PAEs) and sub-MIC effects (SMEs) on S. aureus CB190 (ATCC 29213) or S. epidermidis CB191 (ATCC 35984) cultivated in MHII broth supplemented with P-80. The PAE period was determined following the exposure of log-phase bacteria (≈107 to 108 CFU/ml) to 3× MIC for 1 h and 0.5 h for S. aureus CB190 (ATCC 29213) and S. epidermidis CB191, respectively, in which drug was removed by a 10−3 dilution method as described previously (4). The PAE-SME was determined by exposure to a 3×MIC concentration for 1 h or 0.5 h, followed by 10−3 dilution into medium containing the indicated test agents at a final concentration of 0.125× MIC. The SME was determined in parallel by 10−3 dilution of non-drug-exposed control cultures into medium containing the indicated test agents at a final concentration of 0.125×MIC. Viable counts were determined on charcoal agar medium following serial dilution in physiologic saline.
Intracellular killing assays.
Intracellular killing studies employed the adherent, mouse macrophage cell line J774A.1 (ATCC TIB-67) in combination with S. aureus CB1406 (ATCC 25923) or a small colony variant derived from S. aureus ATCC 12600 bearing a deletion-replacement ΔhemB::ermC mutation (CB1927). The bacteria were opsonized with 10% normal mouse serum prior to infection and seeded at a multiplicity of infection of 4:1. To ensure bacterium-macrophage surface contact, the plates were centrifuged briefly at 500 × g and phagocytosis of the bacterial cells was allowed to proceed for 1 h at 37°C in 5% CO2. After phagocytosis, the cell monolayer was washed two times with sterile prewarmed growth medium and replaced with complete growth medium containing an antimicrobial at the indicated test concentration to remove/inactivate any remaining extracellular and adherent nonphagocytosed bacteria. The effect of drug treatment on intracellular bacterial viability was measured by direct viable count measurements following deoxycholate-mediated cell disruption, serial dilution in sterile saline, and enumeration of the viable CFU on charcoal agar following appropriate incubation. In the experiments reported here, the initial intracellular inoculum quantified was typically in the range of 1 × 106 to 3 × 106 CFU per ml. The absence of extracellular S. aureus growth or contamination was monitored in parallel by sampling the cell culture medium at each time point.
RESULTS
Activity of CBR-2092 and comparators against clinical isolates of primary target pathogens.
Table 2 lists the in vitro antimicrobial activity of CBR-2092 against 175 staphylococci and 125 streptococci compared to the activities of rifampin and ciprofloxacin determined in parallel. The broth microdilution CBR-2092 MIC90s for MSSA, MRSA, MSSE, or MRSE isolates were 0.015, 0.015, 0.008, and 0.5 μg/ml, respectively. The activity of CBR-2092 against MSSA, MRSA, and MSSE was equivalent to that of rifampin and was improved over that of ciprofloxacin, whereas the activity of CBR-2092 was improved over the activities of both rifampin and ciprofloxacin against MRSE. The MIC90s of CBR-2092 for S. pyogenes, S. agalactiae, and S. pneumoniae were 0.12, 0.25, and 0.12 μg/ml, respectively, and, again, were equivalent to those of rifampin and improved over those of ciprofloxacin.
TABLE 2.
Antibacterial activities of CBR-2092 against clinical isolates of gram-positive bacteria
| Organism (no. of strains) | Drug | MICa (μg/ml)
|
||
|---|---|---|---|---|
| Range | 50% | 90% | ||
| S. aureus | ||||
| MSSA (51)b | CBR-2092 | ≤0.004-0.03 | 0.008 | 0.015 |
| Rifampin | ≤0.008-0.06 | 0.015 | 0.015 | |
| Ciprofloxacin | 0.06->4 | 0.25 | 1 | |
| MRSA (54)b | CBR-2092 | ≤0.004-2 | 0.015 | 0.015 |
| Rifampin | ≤ 0.008->4 | 0.015 | 0.015 | |
| Ciprofloxacin | 0.12->4 | >4 | >4 | |
| S. epidermidis | ||||
| MSSE (35)b | CBR-2092 | ≤0.004-0.015 | ≤0.008 | 0.008 |
| Rifampin | ≤0.004-0.06 | 0.015 | 0.015 | |
| Ciprofloxacin | 0.12-> 4 | 0.25 | >4 | |
| MRSE (35)b | CBR-2092 | ≤0.004-1 | 0.008 | 0.5 |
| Rifampin | ≤0.004-> 4 | 0.015 | >4 | |
| Ciprofloxacin | 0.12-> 4 | >4 | >4 | |
| S. pyogenes (35)c | CBR-2092 | 0.008-0.12 | 0.03 | 0.12 |
| Rifampin | ≤0.008-0.25 | 0.06 | 0.12 | |
| Ciprofloxacin | 0.12-2 | 0.5 | 1 | |
| S. agalactiae(35)c | CBR-2092 | 0.06-2 | 0.12 | 0.25 |
| Rifampin | 0.03-2 | 0.12 | 0.12 | |
| Ciprofloxacin | 0.25-1 | 0.5 | 1 | |
| S. pneumoniae (55)c | CBR-2092 | 0.06-0.25 | 0.12 | 0.12 |
| Rifampin | 0.12-1 | 0.25 | 0.25 | |
| Ciprofloxacin | 0.5->4 | 2 | >4 | |
Determined by the broth microdilution method.
Tested in MHII broth with 0.002% (vol/vol) P-80.
Tested in MHII broth with 5% (vol/vol) LHB and 0.002% (vol/vol) P-80.
While enterococci were not primary target pathogens, the activity of CBR-2092 against enterococci was evaluated in studies with limited numbers of clinical isolates (n = 10) and yielded MIC90s for rifampin and CBR-2092 of 8 and 2 μg/ml, respectively, for Enterococcus faecalis and 8 and 1 μg/ml, respectively, for Enterococcus faecium (data not shown). The mechanistic basis underlying the lower susceptibility of enterococci to rifamycins and, to a lesser degree, CBR-2092 compared to those of other gram-positive cocci is not well understood but may reflect lower cell surface penetration.
Correlation between MIC and MBCs.
As shown in Table 3, the MBCs for CBR-2092 and the comparators were evaluated by using S. aureus CB190 (ATCC 29213) and otherwise isogenic derivatives bearing a high-level rifamycin resistance mutation in rpoB (rpoBH481Y) and/or quinolone resistance mutations in both gyrA (gyrAS84L) and parC (parCS80F). The MICs for rifampin against the rifamycin-susceptible strains were 0.008 μg/ml; however, rifampin exhibited only bacteriostatic activity in these assays, and no MBC endpoints were determined in the concentration range tested here (i.e., up to 4 μg/ml). Ciprofloxacin, in contrast, exhibited bactericidal activity against quinolone-susceptible strains, with equivalent MICs and MBCs of 0.24 μg/ml. The MIC determined for CBR-2092 against a rifampin- and quinolone-susceptible strain was 0.008 μg/ml, and thus, CBR-2092 had potency equivalent to that of rifampin. However, unlike rifampin, an MBC of 1 μg/ml was determined, indicating that CBR-2092 exhibits bactericidal activity that is not accounted for by its rifamycin pharmacophore alone (Table 3). The CBR-2092 MIC for rifampin-resistant strain CB370 (rpoBH481Y) was 0.12 μg/ml and, in this context, was bactericidal in nature, with an equivalent MBC of 0.12 μg/ml. The notion that the quinolone pharmacophore of CBR-2092 is responsible for the apparent bactericidal activity exhibited against strain CB370 (rpoBH481Y) is substantiated by the observation that the MBC determined for strain CB815 (rpoBH481Y gyrAS84L parCS80F) is >4 μg/ml.
TABLE 3.
Influence of rifamycin or quinolone resistance phenotype on CBR-2092 MBCs against S. aureus
| Strain (relevant genotype) | MBC/MIC (μg/ml)
|
||
|---|---|---|---|
| CBR-2092 | Rifampin | Ciprofloxacin | |
| CB190 (ATCC 29213; wild-type parent) | 1/0.008 | >4/0.008 | 0.24/0.24 |
| CB814 (gyrAS84LparCS80F) | >4/0.008 | >4/0.008 | >16/16 |
| CB370 (rpoBH481Y) | 0.12/0.12a | >4/>4 | 0.24/0.24 |
| CB815 (rpoBH481YgyrAS84L parCS80F) | >4/1 | >4/>4 | >16/16 |
Cidality was considered paradoxical (i.e., the drug is less cidal at higher doses; Fig. 1).
Bactericidal activity in time-kill assays.
In the standard time-kill assay format, when ciprofloxacin was dosed at ≥1 μg/ml, it was rapidly cidal and promoted a 3-log10 (99.9%) reduction in the CFU of wild-type S. aureus CB190 (ATCC 29213) within 2 to 3 h (Fig. 1C). In contrast, rifampin dosed in the same fashion failed to achieve a 3-log10 cidal endpoint within the 24-h time period (Fig. 1B) and, as such, behaved as a bacteriostatic agent. CBR-2092 exhibited time-kill properties that were distinct from and intermediate between those of ciprofloxacin and rifampin. At fixed drug doses of 0.1, 1, or 10 μg/ml of CBR-2092, 3-log10 CFU reductions in wild-type S. aureus CB190 (ATCC 29213) were recorded in 24-h, 10-h, and 3-h increments, respectively (Fig. 1A); and as such, CBR-2092 exhibited attributes of both time- and concentration-dependent killing in vitro.
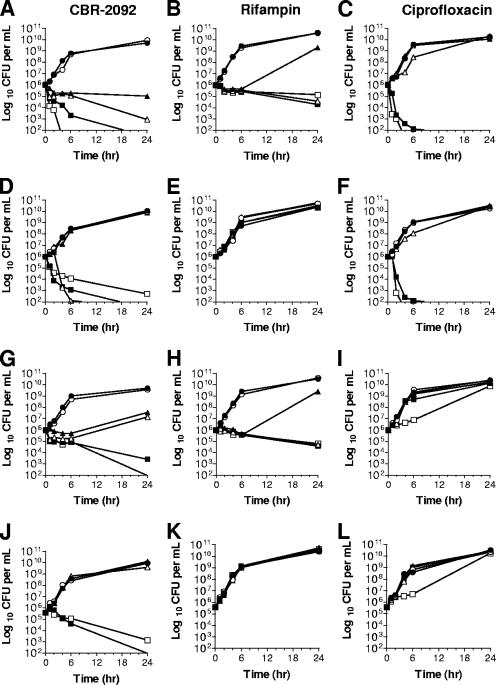
FIG. 1.
Time-kill studies with rifampin-resistant and/or fluoroquinolone-resistant S. aureus strains. The influence of the rifamycin or quinolone resistance phenotype of S. aureus on the bactericidal properties of CBR-2092 (A, D, G, and J), rifampin (B, E, H, and K), and ciprofloxacin (C, F, I, and L) were measured by the standard time-kill method with wild-type S. aureus CB190 (ATCC 29213) (A to C), rifamycin-resistant derivative CB370 (rpoBH481Y) (D to F), quinolone-resistant derivative CB814 (parCS80F gyrAS84L) (G to I), and quinolone- plus rifamycin-resistant derivative CB815 (parCS80F gyrAS84L rpoBH481Y) (J to L). The compounds were dosed at fixed concentrations, as indicated. Symbols: closed circles, control (no drug); open circles, 0.001 μg/ml drug; closed triangles, 0.01 μg/ml drug; open triangles, 0.1 μg/ml drug; closed squares, 1 μg/ml drug; open squares, 10 μg/ml drug.
As expected, rifampin lacked any appreciable antimicrobial activity against the rifamycin-resistant strain CB370 or CB815 (Fig. 1E and K), whereas ciprofloxacin was similarly ineffective againt strain CB814 or CB815 bearing quinolone resistance owing to specific mutations in parCS80F and gyrAS84L (Fig. 1I and L). In contrast, CBR-2092 exhibited markedly better activity than ciprofloxacin (compare Fig. 1G and I) and better overall cidality than equivalently dosed rifampin against quinolone-resistant strain CB814 (compare Fig. 1G and H). Similarly, CBR-2092 exhibited activity superior to that of rifampin against high-level rifampin-resistant strain CB370 (compare Fig. 1D and E) and achieved cidal 3-log10-CFU reductions at CBR-2092 concentrations of 10, 1, and 0.1 μg/ml within 24, 6, and 4 h, respectively. Such paradoxical cidality has been reported previously for quinolone agents in studies with a variety of gram-positive cocci. As this paradoxical killing is observed with CBR-2092 only with highly rifampin-resistant strains, it seems likely that only the quinolone pharmacophore of CBR-2092 is necessary and that ongoing RNA synthesis is a contributory factor. Finally, CBR-2092 exhibited bactericidal activity against rifamycin-resistant and quinolone-resistant strain CB815 that was superior to the activities of both rifampin and ciprofloxacin (compare Fig. 1J to K and L). Less overall killing was again observed at 10 μg/ml, indicating a possible paradoxical bactericidal effect of CBR-2092 in this genetic background.
A comparison of the time-kill properties exhibited by CBR-2092, rifampin, and ciprofloxacin against S. epidermidis CB191 (ATCC 35984) and S. aureus CB190 (ATCC 29213) is shown in Fig. 2, which shows the results of studies in which the compounds were dosed at multiples of their respective MICs. As expected from precedents described in the literature, rifampin exhibited more rapid bactericidal characteristics against S. epidermidis CB191 (ATCC 35984) than against S. aureus CB190 (ATCC 29213). However, for both S. aureus and S. epidermidis, rifampin resistance emerged at either 1× or 2×MIC (data not shown), and the rebound of viable cells was observed in the 2× to 8×MIC range with S. epidermidis (Fig. 2E). CBR-2092 also exhibited more pronounced cidality against S. epidermidis but was distinguished from rifampin in that no resistance development was observed and no rebound of the treated cultures occurred (Fig. 2D).
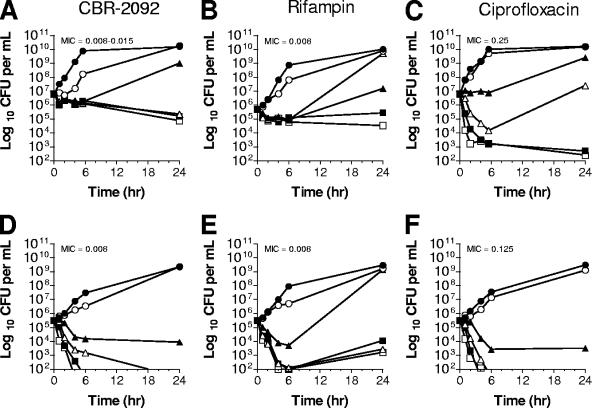
FIG. 2.
Time-kill studies with wild-type S. aureus and S. epidermidis. The bactericidal properties of CBR-2092 (A and D), rifampin (B and E), and ciprofloxacin (C and F) were measured by the standard time-kill method with wild-type S. aureus CB190 (ATCC 29213) (A to C) or S. epidermidis CB191 (ATCC 35984) (D to F) as the test organisms. The compounds were dosed at multiples of the MIC, as indicated. Symbols: closed circles, control (no drug); open circles, 0.25× MIC; closed triangles, 1× MIC; open triangles, 2× MIC; closed squares, 4× MIC; open squares, 8× MIC.
The activities of CBR-2092 against a clinical isolate of Streptococcus pyogenes (CB1208) and two otherwise isogenic strains bearing intermediate or high-level resistance to rifampin are shown in Fig. 3. Against the rifampin-sensitive parent strain (CB1208), CBR-2092 exhibited time-dependent cidal properties but failed to achieve a 3-log10 killing following 24 h of incubation (Fig. 3A). However, against strain CB1456, a derivative of CB1208 that exhibits intermediate-level resistance to rifampin (MIC of 1 μg/ml) and that bears the rpoBA624D mutation, CBR-2092 exhibited bactericidal activity, with elements of both time and concentration dependence, and achieved a killing of ≥99.9% (or ≥3 log10) at ≥2× MIC following ≤24 h of exposure (Fig. 3B). This effect was further pronounced in strain CB1504, a derivative of S. pyogenes CB1456 that exhibits high-level resistance to rifampin (MIC > 32 μg/ml) and that bears the rpoBH486Y mutation, in addition to rpoBA624D. In this background, CBR-2092 achieved a killing of ≥99.9% (or ≥3 log10) at ≥4× MIC following ≤6 h of exposure, suggesting that the bactericidal properties of CBR-2092 are further enhanced against this strain that exhibits high-level resistance to rifampin (Fig. 3C). This effect might be anticipated if the relative contribution of the fluoroquinolone pharmacophore is in effect further elevated owing to the more complete genetic silencing of the target of the rifamycin pharmacophore. The resulting effect is a shift in the relative time-kill properties of CBR-2092 toward a more fluoroquinolone-like activity with bactericidal activity that, again, has elements of both time and concentration dependence. Overall, these combined data are consistent with the anticipated properties of a rifamycin-fluoroquinolone hybrid antibacterial agent with antimicrobial contributions from both of its composite pharmacophores.
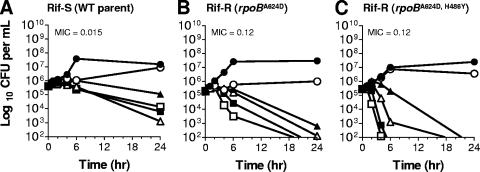
FIG. 3.
Time-kill studies with isogenic Streptococcus pyogenes rifampin-sensitive, -intermediate, or -resistant strains. The influence of the rifamycin resistance phenotype of Streptococcus pyogenes rifampin-sensitive (Rif-S) wild-type (WT) strain CB1208 (A), rifampin-intermediate strain CB1456 (B), and rifampin-resistant (Rif-R) strain CB1504 (C) on the bactericidal properties of CBR-2092 was determined. Time-kill studies were undertaken with static incubation at 35°C in 5 to 10% CO2 in MHII broth medium with 0.002% (vol/vol) P-80 in the absence of further supplements. Compounds were dosed at multiples of the MIC, as indicated. Symbols: closed circles, no drug; open circles, 0.25× MIC; closed triangles, 1× MIC; open triangles, 2× MIC; closed squares, 4× MIC; open squares, 8× MIC.
Single-step resistance studies.
MPCs and spontaneous resistance frequencies were determined for CBR-2092, rifampin, and ciprofloxacin against S. aureus CB190 (ATCC 29213). In these studies, concentrations of rifampin up to 32 μg/ml were insufficient to prevent the emergence of rifamycin resistance following the plating of 1010 viable CFU, and the resistance frequency for rifampin at the 1 μg/ml concentration was determined to be 2.9 × 10−8 (Table 4). In contrast, the MPCs for ciprofloxacin and CBR-2092 were 2 μg/ml and 0.12 μg/ml, respectively. These data allow the calculation of mutant selection windows (MSWs), which are the MPC/MIC ratios (26), and yield much smaller MSW values (i.e., eightfold) for ciprofloxacin and CBR-2092 than that apparent for rifampin (i.e., >4,000). These data are in accordance with those from studies with S. aureus RN4220, for which MSW values of 13 and 160,000 were determined for ciprofloxacin and rifampin, respectively (26).
TABLE 4.
Results of single-step resistance selection studies with S. aureus ATCC 29213
| Compound | MICa (μg/ml) | MPCb (μg/ml) | MSWc | Resistance freq.d |
|---|---|---|---|---|
| CBR-2092 | 0.015 | 0.12 | 8 | <10−12 |
| Rifampin | 0.008 | >32 | >4,000 | 2.9 × 10−8 |
| Ciprofloxacin | 0.24 | 2 | 8 | 5.0 × 10−9 |
Determined by the broth microdilution method in MHII broth with 0.002% (vol/vol) P-80.
The MPC is the drug dose at which resistant mutants (1010 CFU) were no longer obtained.
MSW is the ratio of the MPC to the MIC.
Resistance freq., resistance frequency, which is the number of mutants obtained divided by the total number of input cells at a fixed drug dose of 1 μg/ml.
Finally, ciprofloxacin dosed at 1 μg/ml exhibited an observed resistance frequency of 5 × 10−9, whereas no resistant mutants were recovered for CBR-2092 at this concentration, indicating an apparent resistance frequency of <10−12. These combined data are consistent with the proposed multitargeting nature of CBR-2092 and hold promise for a low potential for spontaneous resistance development.
In vitro PAEs and SMEs.
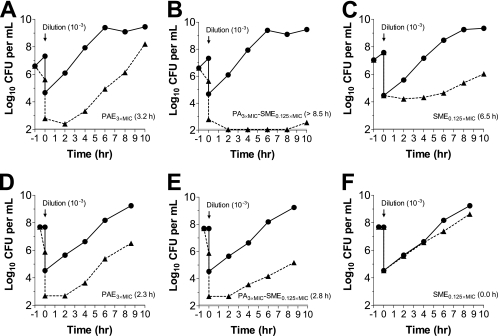
CBR-2092 exhibited in vitro PAEs against S. aureus CB190 (ATCC 29213) and S. epidermidis CB191 (ATCC 35984) that were similar to those of rifampin and improved over those of ciprofloxacin (Fig. 4A and D; Table 5). However, unlike rifampin, CBR-2092 exhibited prolonged postantibiotic SMEs (PA-SME) and SMEs against S. aureus CB190 (ATCC 29213) (Fig. 4B and C) that were consistent with the efficient cell penetration and/or retention of CBR-2092. Interestingly, such dramatic SMEs were not similarly observed for S. epidermidis (ATCC 35984) (Fig. 4E and F), possibly reflecting the improved cidality of CBR-2092 in the latter case.
FIG. 4.
In vitro PAEs and SMEs. The in vitro PAEs (A and D), PA-SMEs (B and E), and SMEs alone (C and F) of CBR-2092 against S. aureus CB190 (ATCC 29213) (A to C) and S. epidermidis CB191 (ATCC 35984) (D to F) were determined. The bacteria were exposed to drug at 0× or 3× MIC for 1 h (S. aureus) or 0.5 h (S. epidermidis) and then diluted (10−3) into fresh medium containing drug at 0 or 0.125× MIC. Symbols: closed circles, control (vehicle only); closed triangles, CBR-2092.
TABLE 5.
In vitro MICs, PAEs, PA-SMEs, and SMEsa
| Antimicrobial |
S. aureus ATCC 29213
|
S. epidermidis ATCC 35984
|
||||||
|---|---|---|---|---|---|---|---|---|
| MICb (μg/ml) | PAE (h) | PA-SME (h) | SME (h) | MICb (μg/ml) | PAE (h) | PA-SME (h) | SME (h) | |
| CBR-2092 | 0.015 | 3.2 | >8.5 | 6.5 | 0.008 | 2.3 | 2.8 | —c |
| Rifampin | 0.008 | 3.0 | 3.8 | — | 0.008 | 2.8 | 3.9 | — |
| Ciprofloxacin | 0.24 | — | — | — | 0.12 | — | 0.5 | — |
PAEs were determined with 3× MIC for 1 h, PA-SMEs were determined with 3× MIC for 1 h for the PAE and 0.125× MIC for the SME, and SMEs were determined with 0.125× MIC.
Determined by the broth microdilution method in MHII broth with 0.002% (vol/vol) P-80.
—, no measured effect was observed.
Impact of intrinsic or mutationally elevated efflux systems.
In order to determine the possible effect of the NorA and/or MepA efflux system on CBR-2092, MIC assays were conducted with S. aureus strains that exhibit mutationally activated NorA or MepA efflux activity (strains CB990 and SA-K2068, respectively) or basal NorA or MepA efflux activity (CB190 and SA-8325-4, respectively) (Table 6). MICs for rifampin were equivalent between otherwise isogenic strain pairs (strains CB990 and CB190 and strains SA-K2068 and SA-8325-4), and it was concluded that rifampin is not a substrate for the NorA or MepA efflux system. In contrast, and consistent with precedents described in the literature, the quinolone agents tested were differentially affected by the mutationally elevated NorA- and/or MepA-mediated efflux activity in S. aureus (strains CB990 versus CB190 and strains SA-K2068 versus SA-8325-4 in Table 6). In contrast, CBR-2092, like rifampin, did not appear to be a substrate for either the NorA or the MepA efflux system, and as such, its MIC was equivalent between otherwise isogenic pairs of strains (strains CB990 and CB190 and strains SA-K2068 and SA-8325-4) that exhibit mutationally activated or intrinsic basal efflux pump activity. The conclusion that CBR-2092 is not a substrate for NorA- or MepA-mediated efflux in S. aureus is substantiated by the observation that the mutational activation of efflux systems does not appear to be a contributory factor in the stepwise development of resistance to CBR-2092 (18).
TABLE 6.
Impact of elevated NorA or MepA efflux resistance pumps on the antibacterial activity of CBR-2092
| Antimicrobial | MICa (μg/ml)
|
Fold change (norAup)b | MICa (μg/ml)
|
Fold change (mepAup)c | ||
|---|---|---|---|---|---|---|
| Wild type (CB190) | norAup (CB990) | Wild type (SA-8325-4) | mepAup (SA-K2068) | |||
| CBR-2092 | 0.015 | 0.015 | 1 | 0.004 | 0.004 | 1 |
| Rifampin | 0.008 | 0.008 | 1 | 0.004 | 0.004 | 1 |
| Ciprofloxacin | 0.24 | 2 | 8 | 0.24 | 2 | 8 |
| Levofloxacin | 0.24 | 0.5 | 2 | 0.24 | 1 | 4 |
| Gatifloxacin | 0.12 | 0.24 | 2 | 0.12 | 0.5 | 4 |
| Moxifloxacin | 0.06 | 0.06 | 1 | 0.06 | 0.24 | 4 |
Determined by the broth microdilution method in MHII broth with 0.002% (vol/vol) P-80.
Fold change in MIC measured for CB990 (norAup) over that for CB190 (ATCC 29213 [wild-type parent]).
Fold change in MIC measured for SA-K2068 (mepAup) over that for SA-8325-4 (wild-type parent).
Suppression of rifampin resistance emergence in high-cell-density time-kill assays.
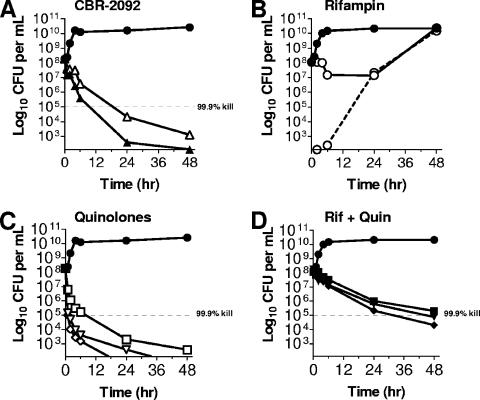
Time-kill assays were also conducted with seed cultures with a high initial cell density (≈108 CFU per ml) that allowed the outgrowth of spontaneous rifampin-resistant subpopulations present in the starter population. As shown in Fig. 5B, dosing with rifampin at 1 μg/ml (∼125× MIC) resulted in an initial drop in the numbers of viable CFU by 4 h but failed to suppress the emergence of rifampin-resistant subpopulations, such that by 48 h, essentially 100% of the recovered cells exhibit resistance to rifampin. The quinolone agents (i.e., ciprofloxacin, levofloxacin, and moxifloxacin) dosed here were tested alone at their respective CLSI resistance breakpoints; they were appreciably bactericidal and did not select for measured quinolone resistance (Fig. 5C). The combination of rifampin and ciprofloxacin, levofloxacin, or moxifloxacin suppressed the emergence of a rifampin-resistant subpopulation (data not shown) but was markedly less cidal than the individual quinolone agents when they were tested alone (compare Fig. 5C and D). The in vitro antagonism of quinolone cidality by rifamycins and translation inhibitors has been well documented in studies of gram-positive cocci and is best explained by the requirement for de novo RNA and/or protein synthesis as a key step in the process of quinolone-mediated, bacterial cell killing (7, 13). In contrast, CBR-2092, dosed here at 0.25 μg/ml or 4 μg/ml, retained excellent cidal activity and also suppressed the emergence of rifampin-resistant subpopulations (Fig. 5A). Such results are wholly consistent with the multifunctional activity of CBR-2092 and an apparent lack of self-antagonism.
FIG. 5.
Time-kill studies conducted with a high initial cell density. The influence of the S. aureus CB190 (ATCC 29213) cell density on the antimicrobial activities and resistance development potential of CBR-2092, rifampin, quinolones, or cocktails of rifampin (Rif) plus quinolones (Quin) was determined in time-kill assays. All quinolones were dosed at their respective CLSI resistance breakpoints, rifampin was dosed at 1 μg/ml, and CBR-2092 was dosed at 0.25 and 4 μg/ml. Symbols: closed circles, control (no drug); open circles, rifampin (1 μg/ml); open triangles, CBR-2092 (0.25 μg/ml); closed triangles, CBR-2092 (4 μg/ml); open squares, ciprofloxacin (4 μg/ml); open downward triangles, levofloxacin (4 μg/ml); open diamonds, moxifloxacin (2 μg/ml); closed squares, rifampin (1 μg/ml) plus ciprofloxacin (4 μg/ml); closed downward triangles, rifampin (1 μg/ml) plus levofloxacin (4 μg/ml); closed diamonds, rifampin (1 μg/ml) plus moxifloxacin (2 μg/ml); open circles, dashed line, rifampin-resistant CFU recovered.
Intracellular activity against wild-type or genetically defined small colony variants of S. aureus.
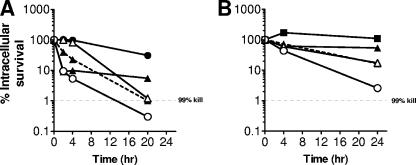
Staphylococci are capable of stably residing within eukaryotic cells such as macrophages and monocytes, and it is postulated that such organisms represent reservoirs of sequestered bacteria that may contribute to persistent and recurrent infections (16). In light of the utility of the rifamycin class of agents for the treatment of persistent intracellular infections, we investigated the ability of CBR-2092 to eradicate S. aureus cells growing in cultured J774A.1 murine macrophages. During the first 2 h, CBR-2092 exhibited excellent intracellular activity against wild-type S. aureus CB1406 (ATCC 25923) that was similar to that seen for the potent quinolone moxifloxacin when it was tested against this quinolone-sensitive S. aureus strain (Fig. 6A). However, unlike moxifloxacin, CBR-2092 also exhibited protracted rifampin-like bactericidal activity that persisted through 24 h, resulting in a >99% cell reduction in ∼14 h (Fig. 6A). Such results are consistent with the effective accumulation and retention of CBR-2092 in subcellular compartments occupied by reservoirs of viable intracellular staphylococci. In contrast, gentamicin, which exhibits poor intracellular accumulation, exhibited little effect on the viability of the intracellular staphylococci. The intracellular killing activity of CBR-2092 (dosed at 4 μg/ml) was also improved over that of a rifampin-plus-moxifloxacin cocktail (dosed at 4 μg/ml of each agent), and evidence of rifampin-mediated antagonism of early rapid moxifloxacin cidality was again apparent for the cocktail combination (Fig. 6A).
FIG. 6.
In vitro intracellular killing activity. Changes in the number of intracellular S. aureus CB1406 (ATCC 25923) (A) or stable small colony variant CB1927 with a ΔhemB::ermC deletion-replacement mutation (B) in infected J774A.1 murine macrophages with treatment with fixed doses of indicated antibiotics were determined. Viable CFU were recovered following deoxycholate-mediated cell disruption, serial dilution, and plating on charcoal agar medium. Survival or killing was expressed as the percent survival relative to the number of viable intracellular CFU at the start of the study. Symbols: closed squares, oxacillin (4 μg/ml); closed circles, gentamicin (4 μg/ml); open circles, CBR-2092 (4 μg/ml); open triangles, rifampin (4 μg/ml); closed triangles, moxifloxacin (4 μg/ml); closed triangles, dashed line, rifampin (4 μg/ml) plus moxifloxacin (4 μg/ml).
Finally, CBR-2092 exhibited markedly improved intracellular killing activity relative to that of the comparators tested when their activities against an engineered ΔhemB::ermC mutant (CB1927) which exhibits hemin auxotrophy and a small colony variant phenotype (Fig. 6B) were assayed (24). This observation is quite striking, as clinical isolates that exhibit a small colony variant phenotype have been implicated in a variety of persistent bacterial infections, including those of indwelling medical devices, and are typically recalcitrant to current antimicrobial therapies (17).
DISCUSSION
CBR-2092 exhibits potent activity against prevalent staphylococcal and streptococcal pathogens associated with skin and soft tissue infections, hospital- and community-acquired pneumonia, bloodstream infections, and biofilm-associated infections of indwelling medical devices. Importantly, activity is retained against the MRSA and MRSE populations prevalent among the staphylococci. For the MRSA and MRSE isolates, the MIC ranges determined for CBR-2092 and rifampin were ≤0.004 to 2 and ≤0.004 to >4 μg/ml, respectively, and for a number of isolates the rifampin MICs were highly elevated (data not shown). As quinolone resistance is more prevalent in MRSA and MRSE strains, it will be important to understand and monitor the incidence of coresistance to rifamycins and quinolones in these key pathogens to elucidate the impact of such on the potential future clinical utility of CBR-2092.
Data from a series of assays indicate that CBR-2092 exhibits bactericidal properties against staphylococci that are consistent with those expected of a rifamycin-quinolone hybrid agent with contributions from both of its composite pharmacophores. Against rifampin-susceptible strains, CBR-2092 exhibited bactericidal activity in vitro similar to that of rifampin, but with altered time and concentration dependence. Direct comparison of this time-kill behavior with the behaviors of equivalently dosed rifampin or ciprofloxacin suggests that CBR-2092 exhibits activity against fully susceptible strains at concentrations of ≥0.1 μg/ml via its fluoroquinolone pharmacophore. Against rifampin-resistant strains, CBR-2092 exhibits rapid, fluoroquinolone-like killing activity that has elements of both time and concentration dependence. However, in this case, a paradoxical bactericidal effect was evident, with decreased killing observed at higher CBR-2092 concentrations. Paradoxical bactericidal effects are apparent for a number of quinolone agents and have been explained by a number of alternate theories (8). Further studies will be necessary to address the mechanistic features underlying the apparent paradoxical killing properties observed with CBR-2092 and its potential clinical manifestations.
Both rifampin and CBR-2092 were observed to exhibit improved cidality against a susceptible S. epidermidis strain compared with that observed against S. aureus. Similar results have been reported from studies of the activities of rifamycins against S. epidermidis and other coagulase-negative staphylococci. However, under the conditions employed in the present study, CBR-2092 was found to be superior to rifampin in that no resistance development or rebound of treated cultures was observed in time-kill studies at doses above the MIC, and this differentiation presumably reflects the activity of the quinolone pharmacophore of CBR-2092. Overall, the reasons underlying the increased cidalities that both rifampin and CBR-2092 exhibit against S. epidermidis compared to those observed against S. aureus are unknown and are worthy of further investigation.
CBR-2092 exhibits PAEs and SMEs that are, overall, distinct from those previously described for members of the rifamycin and fluoroquinolone antibiotic classes. Under the conditions employed here, neither rifampin nor ciprofloxacin exhibited an SME. In striking contrast, the SME measured for CBR-2092 against S. aureus was 6.5 h, and its PA-SME (>8.5 h) was significantly prolonged over that observed with rifampin (3.9 h). Further studies will be necessary to address the mechanistic basis underlying the distinct difference between rifampin and CBR-2092 with regard to these parameters. However, the activity of efflux systems is known to be an important factor in determining PAEs for fluoroquinolones against S. aureus (1) and rifampin against Escherichia coli (22). Hence, the apparent nonsusceptibility of CBR-2092 to either the intrinsic or the mutationally activated efflux systems of S. aureus that are active against fluoroquinolones may be one factor underlying the prolonged SMEs and PA-SMEs observed. As up to ∼50% of fluoroquinolone-resistant clinical isolates of S. aureus exhibit an enhanced quinolone efflux phenotype (4, 20, 21, 23), the prolonged SMEs and PA-SMEs observed with CBR-2092 may contribute to an activity profile that is distinct from that of rifamycin-fluoroquinolone cocktail combinations.
Aside from the aforementioned differentiation of CBR-2092 from rifamycin-fluoroquinolone cocktail combinations at the level of efflux, the studies reported here and elsewhere (19) highlight some additional instances in which the covalent linkage of these two pharmacophores appears to translate to an improved activity profile. First, CBR-2092 exhibits bactericidal activity (≥99.9% at 24 h) at 0.25 or 4 μg/ml in time-kill assays initiated at a high cell density and does not cause the development of either CBR-2092 or rifampin resistance (data not shown). As expected from past studies, a variety of fluoroquinolones dosed at concentrations corresponding to their individual CLSI resistance breakpoints similarly exhibit rapid bactericidal characteristics in this context, whereas rifampin alone (at 1 μg/ml) is bacteriostatic and fails to suppress the outgrowth of a rifampin-resistant population. However, use of the combination of rifampin (at 1 μg/ml) and the fluoroquinolones dosed at their CLSI resistance breakpoints is observed to have a dramatic effect on their cidality, with no combination achieving a killing of ≥99.9% at 24 h. In a related example, rifampin-mediated antagonism of the effective cidality of moxifloxacin is apparent in the intracellular killing assays undertaken with S. aureus CB1406 (Fig. 6A). These data are consistent with past reports in the literature of the antagonism of the quinolone-mediated killing of staphylococci by rifamycins and translation inhibitors (6-8) and are consistent with the aforementioned mode of killing that requires the de novo synthesis of a quinolone lethality factor (8).
Finally, CBR-2092 was found to exhibit intracellular killing activity against staphylococci, including an engineered small colony variant, that was improved over the killing activities of various comparator agents and a cocktail made up of rifampin plus moxifloxacin. The excellent activity of rifamycins against a number of intracellular pathogens has been well documented and presumably reflects the effective penetration of rifamycins into mammalian cells and relevant subcellular compartments and the essential nature of the transcription process in the propagation and/or persistence of bacteria in the intracellular environment (9). Small colony variants of S. aureus (and other pathogens) have metabolic defects that confer a small colony phenotype in vitro and have been associated with a variety of persistent and recurrent diseases, including chronic osteomyelitis, cystic fibrosis, and infections of indwelling medical devices (17). Aside from the increased expression of surface adhesion factors (17, 24), staphylococcal small colony variants have been shown to exhibit improved intracellular persistence characteristics through protection from host immune defenses and reduced susceptibility to many classes of antibiotics (17, 25). Indeed, one explanation for the recurrent nature of certain device-related infections is that S. aureus isolates persisting as small colony variants inside host cells both evade host immune factors and antibiotics and act as a seed reservoir for repeat infections (17). The improved intracellular killing activity of CBR-2092 compared to that of rifampin from the in vitro studies described here, including the retention of activity against a prevalent auxotrophic class of small colony variant, holds promise for the potential utility of CBR-2092 in the treatment of recurrent and/or persistent infections mediated by susceptible gram-positive cocci for which intracellular reservoirs represent an important clinical manifestation.
Acknowledgments
We thank Melissa Peterson, Susan Poppe, Debora Sweeney, Dean Shinabarger, and Gary Zurenko (Micromyx LLC) plus Dan Sahm, Deborah Draghi, Nina Brown, and Parveen Grover (Eurofins MediNet Inc.) for the design and execution of the MIC studies with clinical isolates; Glenn Kaatz (Wayne State University) for providing S. aureus strains SA-8325 and SA-K2068; and Olaf Schneewind (University of Chicago) for the pKOR1 allelic replacement system. We also acknowledge past contributors to the rifamycin-quinolone program at Cumbre Pharmaceuticals, including Donghui Bao, Keith Combrink, Jing Li, Zhenkun Ma, and Paul Renick, and the contributions of current colleagues Charles Ding, Steve Madden, and William Weiss.
Footnotes
Published ahead of print on 28 April 2008.
REFERENCES
- 1.Aeschlimann, J. R., L. D. Dresser, G. W. Kaatz, and M. J. Rybak. 1999. Effects of NorA inhibitors on in vitro antibacterial activities and postantibiotic effects of levofloxacin, ciprofloxacin, and norfloxacin in genetically related strains of Staphylococcus aureus. Antimicrob. Agents Chemother. 43:335-340. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 2.Bae, T., and O. Schneewind. 2006. Allelic replacement in Staphylococcus aureus with inducible counter-selection. Plasmid 55:58-63. [DOI] [PubMed] [Google Scholar]
- 3.Clinical and Laboratory Standards Institute. 2006. Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically, approved standard, 7th ed. CLSI document M7-A7, vol. 26, no. 2. Clinical and Laboratory Standards Institute, Wayne, PA.
- 4.DeMarco, C. E., L. A. Cushing, E. Frempong-Manso, S. M. Seo, T. A. A. Jaravaza, and G. W. Kaatz. 2007. Efflux-related resistance to norfloxacin, dyes, and biocides in bloodstream isolates of Staphylococcus aureus. Antimicrob. Agents Chemother. 51:3235-3239. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 5.Drlica, K. 2003. The mutant selection window and antimicrobial resistance. J. Antimicrob. Chemother. 52:11-17. [DOI] [PubMed] [Google Scholar]
- 6.Drlica, K., and D. C. Hooper. 2003. Mechanisms of quinolone action, p. 19-40. In D. C. Hooper and E. Rubinstein (ed.), Quinolone antimicrobial agents, 3rd ed. ASM Press, Washington, DC.
- 7.Drlica, K., and M. Malik. 2003. Fluoroquinolones: action and resistance. Curr. Top. Med. Chem. 3:249-282. [DOI] [PubMed] [Google Scholar]
- 8.Drlica, K., M. Malik, R. J. Kerns, and X. Zhao. 2008. Quinolone-mediated bacterial death. Antimicrob. Agents Chemother. 52:385-392. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8a.Du, Q., E. J. Bonventre, T. B. Doyle, L. Duncan, G. T. Robertson, E. D. Roche, and A. S. Lynch. 2007. In vitro microbiology profiling studies of CBR-2092, a novel rifamycin-quinolone hybrid antibiotic, abstr. F1-2103. Abstr. 47th Intersci. Conf. Antimicrob. Agents Chemother. American Society for Microbiology, Washington, DC.
- 9.Garzoni, C., P. Francois, A. Huyghe, S. Couzinet, C. Tapparel, Y. Charbonnier, A. Renzoni, S. Lucchini, D. P. Lew, P. Vaudaux, W. L. Kelley, and J. Schrenzel. 2007. A global view of Staphylococcus aureus whole genome expression upon internalization in human epithelial cells. BMC Genomics 8:171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Kaatz, G. W., V. V. Moudgal, and S. M. Seo. 2002. Identification and characterization of a novel efflux-related multidrug resistance phenotype in Staphylococcus aureus. J. Antimicrob. Chemother. 50:833-838. [DOI] [PubMed] [Google Scholar]
- 11.Katayama, Y., H. Z. Zhang, and H. F. Chambers. 2003. Effect of disruption of Staphylococcus aureus PBP4 gene on resistance to beta-lactam antibiotics. Microb. Drug Resist. 9:329-336. [DOI] [PubMed] [Google Scholar]
- 12.Klevens, R. M., M. A. Morrison, J. Nadle, S. Petit, K. Gershman, S. Ray, L. H. Harrison, R. Lynfield, G. Dumyati, J. M. Townes, A. S. Craig, E. R. Zell, G. E. Fosheim, L. K. McDougal, R. B. Carey, S. K. Fridkin, and Active Bacterial Core Surveillance (ABCs) MRSA Investigators. 2007. Invasive methicillin-resistant Staphylococcus aureus infections in the United States. JAMA 298:1763-1771. [DOI] [PubMed] [Google Scholar]
- 12a.Lynch, A. S., E. J. Bonventre, T. B. Doyle, Q. Du, L. Duncan, G. T. Robertson, and E. D. Roche. 2007. In vitro mode-of-action studies of CBR-2092: a novel rifamycin-quinolone hybrid antibiotic, abstr. F1-2101. Abstr. 47th Intersci. Conf. Antimicrob. Agents Chemother. American Society for Microbiology, Washington, DC.
- 13.Malik, M., T. Lu, X. Zhao, A. Singh, C. M. Hattan, J. Domagala, R. Kerns, and K. Drlica. 2005. Lethality of quinolones against Mycobacterium smegmatis in the presence or absence of chloramphenicol. Antimicrob. Agents Chemother. 49:2008-2014. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Moran, G. J., A. Krishnadasan, R. J. Gorwitz, G. E. Fosheim, L. K. McDougal, R. B. Carey, D. A. Talan, and the Emergency ID Net Study Group. 2006. Methicillin-resistant S. aureus infections among patients in the emergency department. N. Engl. J. Med. 355:666-674. [DOI] [PubMed] [Google Scholar]
- 15.Pillar, C. M., D. C. Draghi, D. J. Sheehan, and D. F. Sahm. 2007. Prevalence of multidrug-resistant, methicillin-resistant Staphylococcus aureus in the United States: findings of the stratified analysis of the. 2004 to 2005 LEADER Surveillance Programs. Diagn. Microbiol. Infect. Dis. 60:221-224. [DOI] [PubMed] [Google Scholar]
- 16.Plouin-Gaudon, I., S. Clement, E. Huggler, C. Chaponnier, P. François, D. Lew, J. Schrenzel, P. Vaudaux, and J. S. Lacroix. 2006. Intracellular residency is frequently associated with recurrent Staphylococcus aureus rhinosinusitis. Rhinology 44:249-254. [PubMed] [Google Scholar]
- 17.Proctor, R. A., C. von Eiff, B. C. Kahl, K. Becker, P. McNamara, M. Herrmann, and G. Peters. 2006. Small colony variants: a pathogenic form of bacteria that facilitates persistent and recurrent infections. Nat. Rev. Microbiol. 4:295-305. [DOI] [PubMed] [Google Scholar]
- 18.Robertson, G. T., E. J. Bonventre, T. B. Doyle, Q. Du, L. Duncan, T. W. Morris, E. D. Roche, D. Yan, and A. S. Lynch. 2008. In vitro evaluation of CBR-2092, a novel rifamycin-quinolone hybrid antibiotic: studies of the mode of action in Staphylococcus aureus. Antimicrob. Agents Chemother. 52:2313-2323. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Robertson, G. T., E. J. Bonventre, T. B. Doyle, Q. Du, L. Duncan, E. D. Roche, and A. S. Lynch. 2007. Comparative in vitro activity of CBR-2092, a novel rifamycin-quinolone hybrid antibiotic, and rifampin + quinolone combinations, abstr. F1-2102. Abstr. 47th Intersci. Conf. Antimicrob. Agents Chemother. American Society for Microbiology, Washington, DC.
- 20.Schmitz, F., A. Fluit, M. Luckefahr, B. Engler, B. Hofmann, J. Verhoef, H. Heinz, U. Hadding, and M. Jones. 1998. The effect of reserpine, an inhibitor of multidrug efflux pumps, on the in-vitro activities of ciprofloxacin, sparfloxacin and moxifloxacin against clinical isolates of Staphylococcus aureus. J. Antimicrob. Chemother. 42:807-810. [DOI] [PubMed] [Google Scholar]
- 21.Schmitz, F. J., A. C. Fluit, S. Brisse, J. Verhoef, K. Kohrer, and D. Milatovic. 1999. Molecular epidemiology of quinolone resistance and comparative in vitro activities of new quinolones against European Staphylococcus aureus isolates. FEMS Immunol. Med. Microbiol. 26:281-287. [DOI] [PubMed] [Google Scholar]
- 22.Stubbings, W., J. Bostock, E. Ingham, and I. Chopra. 2005. Deletion of the multiple-drug efflux pump AcrAB in Escherichia coli prolongs the postantibiotic effect. Antimicrob. Agents Chemother. 49:1206-1208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Tanaka, M., T. Wang, Y. Onodera, Y. Uchida, and K. Sato. 2000. Mechanism of quinolone resistance in Staphylococcus aureus. J. Infect. Chemother. 6:131-139. [DOI] [PubMed] [Google Scholar]
- 24.Vaudaux, P., P. Francois, C. Bisognano, W. L. Kelley, D. P. Lew, J. Schrenzel, R. A. Proctor, P. J. McNamara, G. Peters, and C. Von Eiff. 2002. Increased expression of clumping factor and fibronectin-binding proteins by hemb mutants of Staphylococcus aureus expressing small colony variant phenotypes. Infect. Immun. 70:5428-5437. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.von Eiff, C., C. Heilmann, R. A. Proctor, C. Woltz, G. Peters, and F. Gotz. 1997. A site-directed Staphylococcus aureus hemB mutant is a small-colony variant which persists intracellularly. J. Bacteriol. 179:4706-4712. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Zhao, X., and K. Drlica. 2002. Restricting the selection of antibiotic-resistant mutant bacteria: measurement and potential use of the mutant selection window. J. Infect. Dis. 185:561-565. [DOI] [PubMed] [Google Scholar]