Abstract
Polycomb-group (PcG) genes encode multimeric nuclear protein complexes, PcG complex 1 and 2. PcG complex 2 was proved to induce transcription repression and to further methylate histone H3 at lysine-27 (H3K27). Subsequently PcG complex 1 is recruited through recognition of methylated H3K27 and maintains the transcription silencing by mediating monoubiquitination of histone H2A at lysine-119. Genetic evidence demonstrated a crucial role for PcG complex 1 in stem cells, and Bmi1, a member of PcG complex 1, was shown to sustain adult stem cells through direct repression of the INK4a locus encoding cyclin-dependent kinase inhibitor, p16CKI, and p19ARF. The molecular functions of PcG complex 1, however, remain insufficiently understood. In our study, deficiency of Rae28, a member of PcG complex 1, was found to impair ubiquitin-proteasome-mediated degradation of Geminin, an inhibitor of DNA replication licensing factor Cdt1, and to increase protein stability. The resultant accumulation of Geminin, based on evidence from retroviral transduction experiments, presumably eliminated hematopoietic stem cell activity in Rae28-deficient mice. Rae28 mediates recruiting Scmh1, which provides PcG complex 1 an interaction domain for Geminin. Moreover, PcG complex 1 acts as the E3 ubiquitin ligase for Geminin, as we demonstrated in vivo as well as in vitro by using purified recombinant PcG complex 1 reconstituted in insect cells. Our findings suggest that PcG complex 1 supports the activity of hematopoietic stem cells, in which high-level Geminin expression induces quiescence securing genome stability, by enhancing cycling capability and hematopoietic activity through direct regulation of Geminin.
Keywords: Rae28, Scmh1, HSCs, ubiquitination, DNA replication licensing
Molecular bases supporting the potency of stem cells may be provided by intrinsic and extrinsic signals. Recent studies have started to unveil the molecular nature of bone marrow (BM) niches, which may provide extrinsic signals to support hematopoietic stem cells (HSCs) through cell-to-cell interactions and/or cytokines, but as yet much less is known about the intrinsic signals that maintain HSC property and activity.
The Polycomb-group (PcG) genes may provide an intrinsic signal to support stem cells (1, 2). PcG genes encode multimeric nuclear protein complexes designated as PcG complex 1 or 2. Representative PcG complex 1 consists of M33(Cbx2), Ring1A or Ring1B(Rnf2), Bmi1, Rae28(Edr1, Mph1, or Phc1), and Scmh1. PcG complex 1 is recruited through recognition of histone H3 at lysine-27 (H3K27) methylated by PcG complex 2 and maintains transcription silencing by mediating histone H2A monoubiquitination at lysine-119 (H2AK119) (3). The contribution of PcG proteins to stem cells was indicated by genome-wide profiling of their chromatin binding and H3K27 methylation in ES cells (4) and was further supported by genetic evidence. Findings obtained from mice or ES cells deficient in Rae28, Bmi1, and Ring1B suggested that PcG complex 1 is crucially involved in HSCs (5–8), neural stem cells (9), ES cells (10), and further leukemic stem cells (11). Impaired control of p16 cyclin-dependent kinase inhibitor (p16CKI), p19ARF (7, 12), p21 (13), and E4F1 (14) expression was reported to be at least in part responsible for stem cell defects in Bmi1-deficient mice. Nevertheless, the molecular role of PcG complex 1 in sustaining stem cell activity is believed to be not yet fully understood for the following reasons. (i) Although stem cell defects in Bmi1-deficient mice were genetically compensated for by INK4a deficiency, the recovery was only partial (9). (ii) Overexpression of Bmi1 augments HSC activity even in the absence of the INK4a locus (15). (iii) Although knockdown of E4F1 (14) and p21 (13) was shown to rescue stem cell activity in Bmi1-deficient mice, the recovery was only partial and was not observed in the impaired long-term repopulating (LTR)-HSC activity. In addition, these molecular alterations were not detected in fetal liver cells (FL) of Rae28-deficient mice, which were defective in self-renewal and LTR capacities of HSCs (5, 6). Rae28-deficient mice may thus provide an important clue for clarifying a novel molecular function of PcG complex 1 in sustaining HSC activity.
Because Rae28 recruits Scmh1 (16), which is required for recruitment of Geminin to PcG complex 1 (17), we examined in the study reported here the molecular role of PcG complex 1 in the regulation of HSCs by focusing on Geminin (18), and we demonstrated that PcG complex 1 acts as the E3 ubiquitin ligase for Geminin. It is known that Geminin is absent in the G1 phase and accumulates during progression into the S phase. Geminin directly inhibits the DNA replication licensing factor Cdt1, which recruits the Mcm complex with helicase activity, and prevents rereplication (18). Geminin was also found to maintain undifferentiated states through the regulation of a chromatin-remodeling complex (19). These findings emphasize that Geminin may be a key molecule for the support of stem cells by regulating cellular proliferation and differentiation as well as further genome stability. The findings described in this study disclose a direct role for PcG complex 1 in Geminin regulation and provide important insights into the molecular mechanisms supporting HSCs.
Results
Expression of Geminin, Cdt1, and PcG Complex 1 Members in BM Cells.
Murine BM cells were sorted for purification of CD34− c-Kit+ Sca1+ lineage marker (Lin)− cells (CD34−KSL) (LTR-HSCs), CD34+KSL [multipotential progenitor cells (MPPs)], c-Kit+ Sca1− Lin− cells (progenitors), and their progeny subpopulations. Total cellular RNA was extracted, and expression of PcG members, Geminin, and Cdt1 was examined by means of quantitative RT-PCR analysis (Fig. 1A). We focused on Ring1B, Bmi1, Rae28, and Scmh1 among the PcG complex 1 members. Expression of Geminin was higher in CD34−KSL and down-regulated in CD34+KSL and progeny except for CD3+ T cells. On the other hand, expression of Cdt1 was up-regulated in CD34+KSL and progenitors. Expression of Geminin and Cdt1 was thus reciprocally regulated during differentiation from CD34−KSL to CD34+KSL and progenitors. Expression of Ring1B, Rae28, and Scmh1 was especially augmented in CD34+KSL, where it was higher than in CD34−KSL and progeny. However, Bmi1 expression was not induced in CD34+KSL and was gradually down-regulated during differentiation, which was consistent with previously reported findings (20). The cell cycle phase of sorted CD34−KSL, CD34+KSL, and progenitors was further examined by using BrdU and 7-aminoactinomycin D. Because Geminin expression oscillates at the protein level in the cell cycle (17), we examined Geminin expression in G0/G1-phase CD34−KSL, CD34+KSL, and progenitors by flow cytometry. The average levels (Fig. 1B) and representative sorting profiles are shown [supporting information (SI) Fig. S1]. Geminin expression is higher in CD34−KSL and is differentially down-regulated. These expressions are schematically represented in Fig. S2.
Fig. 1.
Expression of Geminin, Cdt1, and PcG members. (A) CD34−KSL, CD34+KSL, progenitors, and their progeny subpopulations were isolated from murine BM cells, and mRNA expression was examined by quantitative RT-PCR. The mRNA expression levels are shown as ratios to the level in GAPDH. Lane 1, CD34−KSL (LT-HSCs); lane 2, CD34+KSL (MPPs); lane 3, progenitors; lane 4, Ter119+ cells (erythroid cells); lane 5, Gr1+ cells (granulocytes); lane 6, CD3+ cells (T cells); lane 7, B220+ cells (B cells). (B) Geminin expression in G0/G1-phase CD34−KSL, CD34+KSL, and progenitors was further examined at the protein level by cell sorting. Average intensity of Geminin expression in three independent experiments is shown with SEMs.
Expression and Stability of Geminin in Rae28-Deficient FL.
Rae28 recruits Scmh1, which mediates interaction of PcG complex 1 with Geminin (17), suggesting the possibility that Geminin expression may be affected in Rae28-deficient cells. Immunoblot analysis as well as quantitative RT-PCR analysis of Geminin expression in FL was performed because homozygous Rae28-deficient (Rae−/−) embryos are lethal at the perinatal period. Geminin expression started to increase at the protein level in Rae−/− FL later than 15.5 days postcoitum (Fig. 2A), at which time the clonogenic activity started to decline (5), although it was not affected at the mRNA level (Fig. S3). We then performed pulse–chase analysis of Geminin with [35S]methionine in vivo to examine protein stability in FL (Fig. 2B). This demonstrates that Geminin was stabilized in the absence of Rae28, suggesting that Rae28 was involved in the regulation of protein stability of Geminin. Because HSCs are Mac1+ and CD34+ in FL, we could not sufficiently enrich HSCs in FL by using the conventional HSC-sorting method (data not shown). Thus, further detailed analysis of Geminin expression in the Rae−/− HSC-enriched subpopulation remained untouched in this study.
Fig. 2.
Geminin expression and its effect on LTR activity. (A) Geminin protein expression in FL. FL were examined by immunoblot analysis with an anti-Geminin antibody. β-Actin detection confirmed equal amounts of protein on the filter. Wild type: Rae+/+. (B) Stability of Geminin. FL were pulse-labeled with [35S]methionine in FL. Geminin was immunoprecipitated and was detected by autoradiography. (C) Cdt1 and Mcm2in chromatin fraction of 15.5-days-postcoitum FL. S, soluble fraction; C, chromatin fraction. Histone H2A was detected as a control. (D) Effect of exogeneous Geminin on LTR activity. Retrovirally transduced FL were injected into lethally irradiated syngeneic mice, and the repopulation was examined 1 and 3 months after injection. The number of recipient mice is indicated above each bar. d.p.c., days postcoitum.
Effect of Accumulated Geminin on Licensing of DNA Replication.
We analyzed Cdt1 and Mcm2, a component of the Mcm complex, in the chromatin fraction of 2-day-cultured FL prepared from 15.5-days-postcoitum embryos (Fig. 2C) to examine the effect of accumulated Geminin on DNA replication licensing. Although the cell subpopulation in the G0/G1 phase increased (see below), in which Cdt1 is up-regulated, Cdt1 in the chromatin fraction was severely reduced in Rae−/− FL, suggesting that an increase in Geminin causes a reduction in chromatin-loaded Cdt1. Chromatin-loaded Mcm2, which is essential for prereplication complex formation, was subsequently reduced. In the absence of Rae28, licensing of DNA replication was thus impaired by an increase in Geminin.
Effect of Exogeneous Geminin on Cell Cycle, Apoptosis, and Hematopoiesis.
Self-renewal and LTR capabilities were impaired in Rae−/− HSCs, whereas Geminin accumulated in Rae−/− FL, which facilitated examination of the effect of Geminin on HSCs. Geminin contains a destruction box (amino acids 23–31; RRTLKMIQP) that is required for ubiquitination through anaphase-promoting complex/cyclosome (APC/C) and subsequent degradation from the M to the G1 phase (21). We therefore transduced Geminin or Geminin with the destruction box deleted (Geminin-DBD) into FL by using a retrovirus vector, MEP, which contains the gene for an enhanced yellow fluorescence protein (EYFP) as a selectable marker, examined Geminin expression in each phase of the cell cycle (Fig. S4A), and detailed the effect on cell cycle (Fig. S4B), apoptosis (Fig. S4C), and hematopoiesis including LTR activity (Fig. S5 and Fig. 2D). Geminin transduction significantly reduced cells in the S phase but increased cells in the G0/G1 phase (Fig. S4B), probably because of induction of a licensing checkpoint (22). Geminin transduction affected the clonogenic and long-term culture-initiating cell (LTC-IC) activities (Fig. S5). We further examined the effect of Geminin transduction on LTR activity. FL were retrovirally transduced, and the percentage of EYFP+ cells was adjusted to 12.5% by adding FL precultured under the same conditions. Lethally irradiated syngeneic mice were injected with the transduced FL (2 × 106), and their peripheral blood was examined (Fig. 2D). Mice with >0.5% EYFP+ multilineage cells were evaluated as repopulated mice. Three months after injection 62.5% of mice injected with MEP-transduced FL showed repopulation, whereas those with either Geminin or Geminin-DBD did not at all. Thus, Geminin and Geminin-DBD transduction appeared to accurately reproduce cell cycle alterations and apoptosis and further impaired HSC activities in Rae−/− FL (5). It was also confirmed that Geminin transduction reduced chromatin-loaded Cdt1 and Mcm2 to a similar extent in Rae−/− FL (data not shown).
Ubiquitination of Geminin and Scmh1.
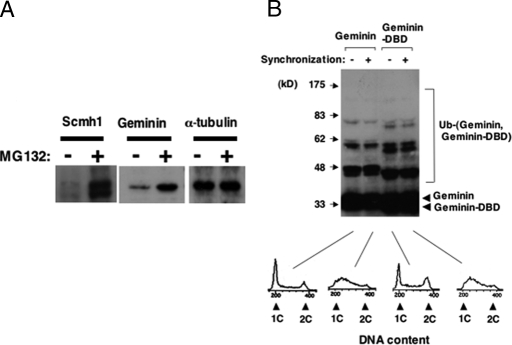
Next, we examined sensitivity to a specific inhibitor of proteasome, MG132. U-2 OS cells, a human osteosarcoma cell line used for analyzing PcG complexes, stably transfected with PcG members, were treated with MG132 and were examined by immunoblot analysis. MG132 treatment markedly augmented protein expression of Scmh1 and Geminin (Fig. 3A), but the effect was only very minor on Ring1B and was undetectable on either Bmi1 or Rae28 (data not shown). We further examined in vivo whether these molecules were subjected to ubiquitination. For this purpose, HEK-293 cells, a cell line with high transfection efficiency derived from human kidney cells, were cotransfected with either PcG members or Geminin combined with ubiquitin and were subjected to immunoblot analysis. Mobility-shifted bands were detected in Scmh1 (Fig. S6) and Geminin (Fig. 3B). These findings showed good consistency with sensitivity of Scmh1 and Geminin to MG132, suggesting that they were regulated by the ubiquitin-proteasome system (UPS). Because Geminin is believed to be ubiquitinated solely by APC/C from the M to the G1 phase, we examined Geminin ubiquitination in synchronized cells (Fig. 3B). HEK-293 cells were cotransfected with Geminin and ubiquitin, treated with thymidine for 24 h, and cultured 3 h after removal of thymidine to synchronize cells in the S phase. Cell synchronization was confirmed by DNA content analysis. Ubiquitinated Geminin was detected even in cells synchronized in the S phase, and intensity of the detected bands was not significantly different for asynchronized and synchronized cells. Ubiquitinated molecules were also detected in HEK-293 cells transfected with Geminin-DBD, which is resistant to APC/C. These findings suggest that Geminin is regulated through UPS even in the S phase and that there exists a ubiquitination mechanism for Geminin other than APC/C. Because PcG complex 1 was reported to act as an E3 ubiquitin ligase for histone H2A and efficiency in Geminin ubiquitination was impaired in the absence of Rae28 (Fig. S7), we hypothesized that PcG complex 1 acts as the E3 ubiquitin ligase for Geminin.
Fig. 3.
Involvement of UPS in the regulation of Geminin and Scmh1. (A) Sensitivity to MG132. The lanes of Geminin and Scmh1 show the effect of MG132 on the endogenous and exogeneous expression in U-2 OS cells, respectively. α-Tubulin detection confirmed equal amounts of protein on the filter. (B) In vivo ubiquitination of Geminin. Either Geminin or Geminin-DBD was cotransfected with ubiquitin into HEK-293 cells and was detected by immunoblot analysis. Cellular synchronization was done by thymidine treatment. DNA content is shown below the blots to confirm cellular synchronization.
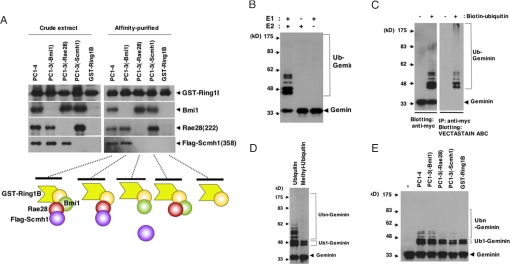
Finally, we reconstituted recombinant PcG complex 1 composed of Ring1B, Bmi1, Rae28, and Scmh1 in Spodoptera frugiperda (Sf9) insect cells. Sf9 cells were coinfected with baculoviruses including glutathione S-transferase (GST)-Ring1B, Bmi1, Rae28, and Flag-Scmh1. Because Rae28 and Scmh1 were unstable in Sf9 cells, a truncated form of Rae28 (amino acids 222-1012), lacking an N-terminal region including serine threonine-rich and glutamine-rich domains [Rae28(222)] (23), and one of Scmh1 (amino acids 358–664), lacking an N-terminal region including the MBT and PEST domains preceded by a Flag tag in the N-terminal portion [Flag-Scmh1(358)], were expressed together with GST-Ring1B and Bmi1 to obtain the stable protein complex (PC1–4). The cell extract was prepared from Sf9 cells expressing PC1–4 and the complex but lacking Bmi1[PC1–3(-Bmi1)], Rae28(222)[PC1–3(-Rae28)], or Flag-Scmh1(358)[PC1–3(-Scmh1)]. The complexes were purified by means of glutathione affinity chromatography. The molecular size of PC1–4 digested with thrombin to release GST was examined by gel filtration fractionation on Superose 6 10/300GL, and the presence of each of the PcG members was confirmed by immunoblot analysis. The molecular mass of the major recombinant complex was close to that of a stoichiometric amount of the members predicted from cDNAs (200 kDa) (Fig. S8A). A pull-down assay clearly showed that Rae28 mediates recruitment of Scmh1 to the complex and further that GST-Ring1B formed a complex with Flag-Scmh1 (358) even in the absence of Bmi1 but not in the absence of Rae28(222) (Fig. 4A), which may be because of direct interaction of GST-Ring1B with Rae28(222) (data not shown).
Fig. 4.
Reconstitution of PcG complex 1 in Sf9 cells and the E3 ubiquitin ligase activity for Geminin. (A) Pull-down assay of the recombinant PcG complex 1. Schematic representation of the complex is shown below the blots. Note that Rae28 and Scmh1 interact with GST-Ring1B even in the absence of Bmi1. Affinity-purified recombinant complex was subjected to in vitro ubiquitination reaction (myc-Geminin+E1+E2+ubiquitin) with indicated modifications. B–E shows E3 ubiquitin ligase activity for Geminin of the recombinant complexes. (B) E1 and E2 dependency. (C) Reaction with biotin-tagged ubiquitin (Biotin-ubiquitin). The reaction products were immunoblotted with an anti-myc antibody (Right) and were also immunoprecipitated with an anti-myc antibody, and molecules modified with biotin-ubiquitin were detected with the VECTASTAIN ABC kit (Left). (D) Effect of methyl-ubiquitin. Mobility-shifted bands other than the lower two may represent polyubiquitinated Geminin. Mono- and polyubiquitinated molecules are indicated by Ub1 and Ubn, respectively. (E) Comparison of E3 ubiquitin ligase activity among PC1–4, PC1–3(-Bmi1), PC1–3(-Rae28), PC1–3(-Scmh1), and GST-Ring1B. *, no reaction. One microgram of PC1–4 was subjected to the reaction, and the amount of GST-Ring1B in the other complexes was adjusted to that in 1 μg of PC1–4. E3 ubiquitin ligase activity was reduced in PC1–3 and GST-Ring1B. Note that polyubiquitinated Geminin was hardly detectable in PC1–3(-Rae28), PC1–3(-Scmh1), and GST-Ring1B.
Recombinant myc-tagged Geminin was subjected to an in vitro ubiquitination assay with the affinity-purified PcG complex. The reaction product was analyzed by immunoblot analysis using an anti-myc monoclonal antibody. PC1–4 was used to detect mobility-shifted Geminin bands in the reaction products (Fig. 4B), but they were undetectable in the absence of either E1 or E2 (Fig. 4B). Intensity of the bands increased and the mobility shifted further depending on reaction time and quantity of PC1–4 (Fig. S8B). Weak mobility-shifted bands detected in the reaction product with GST (Fig. S8B) were also detectable in the reaction product solely with E1 and E2 (data not shown). We examined with biotin-tagged ubiquitin whether the shifted bands corresponded to ubiquitinated Geminin. The reaction product was immunoprecipitated with an anti-myc polyclonal antibody, and the exactly same mobility-shifted bands were detected through biotin-streptavidin interaction (Fig. 4C), confirming that these represented ubiquitinated Geminin. Mobility-shifted bands for ubiquitinated Geminin may represent either formation of an elongated ubiquitin chain at a single lysine residue in Geminin or attachment of a single ubiquitin molecule to multiple lysine residues in Geminin. To determine the validity of these possibilities, we examined Geminin ubiquitination with methyl-ubiquitin, which is incapable of serving as a substrate for further elongation of the ubiquitin chain. The lower two mobility-shifted bands may correspond to monoubiquitinated Geminin (Fig. 4D), but the other more mobility-shifted bands may represent Geminin with elongated ubiquitin chains. PC1–4 may thus mediate the formation of polyubiquitin chains at specific lysine residues in Geminin. To further confirm that PC1–4 directly exerts its E3 ubiquitin ligase activity for Geminin, we prepared mutant GST-Ring1B containing a single amino acid substitution (R70C) in Ring1B (GST-mut-Ring1B), which is defective in the E3 ubiquitin ligase activity for histone H2A (3). R70C mutation did not affect the formation of complex PC1–4 (data not shown), but the ubiquitination activity was abrogated (Fig. S8C), suggesting that the E3 ubiquitin ligase activity of PC1–4 for Geminin was mediated by the RING finger domain providing an interaction domain with E2.
We also compared the ubiquitin ligase activity in PC1–4, PC1–3(-Bmi1), PC1–3(-Rae28), PC1–3(-Scmh1), and GST-Ring1B. Intensity of the bands, especially those for polyubiquitinated Geminin, was reduced in PC1–3(-Rae28), PC1–3(-Scmh1), and GST-Ring1B (Fig. 4E). The intensity was decreased in PC1–3(-Bmi1), but to a lesser extent, so that it was reproducible. The polyubiquitinated bands were relatively conserved, probably because GST-Ring1B formed a complex directly with Rae28(222) and Flag-Scmh1(358) even in the absence of Bmi1 (Fig. 4A). Rae28 and Scmh1 appeared to be required for an effective induction of polyubiquitination. These findings may represent a molecular mechanism underlying the affected Geminin ubiquitination in Rae28-deficient cells. To eliminate the possibility that Geminin was ubiquitinated by contaminated APC/C, we confirmed that the similar activity was detected in Geminin-DBD (Fig. S8D).
Polyubiquitinated GST-Ring1B, Bmi1, and Rae28(222) were detected in the reaction product with PC1–4, but not with PC1–4 containing R70C mutation in Ring1B (Fig. S8E), suggesting self-ubiquitination of these members. However, it remains unclear whether ubiquitination of PcG complex 1 is required for regulating the stability of PcG complex 1 itself through UPS or for inducing the E3 ubiquitin ligase activity as suggested previously (24).
Discussion
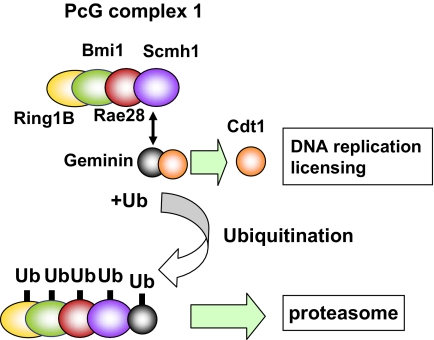
Based on the findings presented in this study, we propose here a model for a novel role of PcG complex 1, which acts as the E3 ubiquitin ligase for Geminin to regulate protein stability (Fig. 5). This model may help to deepen conceptual understanding of the molecular function of PcG complex 1, which has long been believed to be involved solely in transcription silencing (25). Some light has been shed on the molecular role of PcG complex 1 in transcription silencing by evidence of the E3 ubiquitin ligase activity for H2AK119 of PcG complex 1 (3, 26). H2AK119 monoubiquitination correlates well with transcription silencing and is presumed to inhibit RNA polymerase II transcriptional elongation (27). The polyubiquitin chain, on the other hand, which is attached to Geminin through PcG complex 1, may serve as a recognition signal for the proteasome to regulate protein stability of Geminin. In addition, Pc2 (Cbx4), an isoform member of PcG complex 1, is reportedly involved in SUMO E3 ligase (28). Previously reported biochemical evidence indicates that PcG complex 1 competitively inhibits activity of SWI/SNF in Drosophila (29). Thus, it might be reasonable to assume that PcG complex 1 regulates protein stability and activity of a target protein in SWI/SNF through ubiquitination or sumoylation. This makes it tempting to speculate that PcG complex 1 plays various regulatory roles by mediating ubiquitination and/or sumoylation for a variety of target molecules.
Fig. 5.
PcG complex 1 directly regulates Geminin by acting as the E3 ubiquitin ligase. Cdt1 is loaded on chromatin and licenses the chromatin for DNA replication, whereas Geminin inhibits the licensing through the direct interaction with Cdt1. PcG complex 1 induces polyubiquitination of Geminin by acting as the E3 ubiquitin ligase, and the polyubiquitin chain may serve as a recognition signal for the proteasome to regulate stability of Geminin. Ub, ubiquitin.
Retroviral transduction of Geminin into Rae+/+ FL reduced the cells in the S phase, whereas that of Geminin-DBD further increased the cells in the G2/M phase and induced apoptosis. These findings are reminiscent of those for Rae−/− FL and may result from reduced chromatin-loaded Cdt1 and Mcm2. The resultant reduction in active replication origins is presumed to give rise to a licensing checkpoint, which may reduce the cells in the S phase, and/or replication fork stalling, which may trigger G2/M checkpoint and apoptosis (22). This may constitute, at least in part, the mechanism for the reduction of clonogenic and LTR activities seen in Rae−/− FL. Because derepression of the INK4a locus and altered p21 expression in Bmi1-deficient mice were reported (7, 13), we examined expression of p16CKI, p19ARF, and p21 in Geminin-transduced FL. p16CKI and p19ARF mRNAs increased by <50% of those in control cells, whereas the induction could not be detected by immunoblot analysis (data not shown). And expression of p21 mRNA was not altered at all (data not shown). Therefore, transcriptional alterations of these genes can hardly be assumed to be responsible for HSC deficiency in Geminin- or Geminin-DBD-transduced FL, which is consistent with our previously reported findings for Rae−/− FL (5). An increase in Geminin may directly impair HSC activity in Rae−/− FL through reduced chromatin-loaded Cdt1 and Mcm2, as was further evidenced by the finding of a genetic complementation experiment; i.e., retroviral transduction of Cdt1 partially complemented a reduction of clonogenic activity of Rae−/− FL (S.Y. and Y.T., unpublished observation). Because Geminin was found to be subjected to nonproteolytic inactivation through polyubiquitination (30), Geminin may also be regulated at the activity level as well as at the protein level by PcG complex 1. Additional roles for Geminin have been reported, such as its repression of transcription by interaction with PcG complex 1, Hox (17), Six3 (31), SMRT, and AP4 (32). It has also been proposed that Geminin maintains undifferentiated cell states by antagonizing activity of Brg1, a subunit of SWI/SNF, which is reportedly crucial for neural and myeloid differentiation from stem and progenitor cells (19, 33). This suggests the possible existence of an additional function, namely, that an increase in Geminin further affects cellular differentiation capacity through direct interaction with these factors.
Geminin expression is higher in LTR-HSCs than in MPPs and its progeny. This higher Geminin expression may help induce quiescence probably through a mechanism similar to the licensing checkpoint and may also firmly secure genome stability after entering the S phase to self-renew in LTR-HSCs. Otherwise, LTR-HSCs may be susceptible to rereplication in the same cell cycle, which could lead to genome instability. This assumption is supported by recent reports that lack of Geminin causes premature endoreduplication and abrogated development of pluripotent cells of the inner cell mass (34, 35). Because Rae28 deficiency resulted in deficient self-renewal and LTR capability of HSCs (5), it is possible that PcG complex 1 balanced growth potential and genome stability in LTR-HSCs through direct regulation of Geminin. Geminin expression is lower in MPPs and progenitors than in LTR-HSCs, and the reverse is true for Cdt1, which may help the efficient triggering DNA replication licensing in MPPs and progenitors to induce cell cycling. On the other hand, the mRNA levels of PcG complex 1 members Ring1B, Rae28, and Scmh1 are higher in MPPs than in LTR-HSCs, and these PcG members were reported to be well colocalized in hematopoietic cells including HSCs (36). Because PC1–4 was shown to induce ubiquitination of Geminin in a dose-dependent manner at least in the in vitro ubiquitination assay, PcG complex 1 may affect down-regulation of Geminin at the protein level to provide sufficient growth potential during transition from LTR-HSCs to MPPs and progenitors. Alternatively, the high level of Geminin in LTR-HSCs may help maintain undifferentiated states through the direct inhibition of Brg1 as described above, whereas in MPPs and progenitors down-regulation of Geminin may be required for inducing their differentiation into more mature progeny.
Materials and Methods
Rae28-Deficient Mice.
The generation of Rae28-deficinet mice with the C57BL/6 genetic background was described previously (37).
Quantitative RT-PCR.
RNA was extracted by using the Mini RNA isolation kit (ZYMO Research), reverse-transcribed with TaqMan reverse transcription agents (Applied Biosystems), and subjected to real-time quantitative PCR analysis with ABI Prism 7700 sequence detection system (Applied Biosystems).
Transfection and Cellular Analyses.
Transfection experiments and analyses of cell cycle and apoptosis were performed with standard methods as detailed in SI Materials and Methods.
Immunoblot, Chromatin Fraction, and Protein Stability Analyses.
Immunoblot and chromatin fraction analyses were performed as detailed in SI Materials and Methods, and pulse–chase analysis with [35S]methionine was done as described previously (38).
Retrovirus-Mediated Gene Transduction.
Retrovirus-mediated gene transduction was done as described previously (39), and preparation of high-titer retrovirus is detailed in SI Materials and Methods.
Flow Cytometry, HSC, and Progenitor Analyses.
Flow cytometry analysis and cell sorting were performed on FACSCalibur and FACSAria (BD Biosciences Immunocytometry Systems) by using antibodies listed in Table S1, and methylcellulose, LTC-IC, and competitive repopulation assays were performed as described previously (5).
Reconstitution of PcG Complex 1 in Sf9 Cells and Purification.
Recombinant PcG complex was composed in Sf9 cells by the baculovirus-mediated gene transfection method and was purified by affinity column chromatography as described previously (40). The details are in SI Materials and Methods.
In Vitro Ubiquitination Analysis.
E3 ubiquitin ligase activity of the complex was examined in vitro with recombinant myc-tagged Geminin as a substrate as detailed in SI Materials and Methods.
Antibodies.
Primary and secondary antibodies used are listed in Table S1.
Statistical Analysis.
More than three independent experiments were performed, and the findings were statistically analyzed. The results are shown with SEM.
Supplementary Material
Acknowledgments.
We thank Drs. K. R. Humphries (Terry Fox Laboratory, Vancouver) and R. Hawley (George Washington University, Washington, DC) for MEP; Drs. A. Kingsman and T. Friedmann (University of California at San Diego School of Medicine, La Jolla, CA) for expression plasmids for gag, pol, and VSV-G envelope; Dr. T. Kitamura (Institute of Medical Science, University of Tokyo, Tokyo, Japan) for PlatE; the Analysis Center of Life Science in Hiroshima University for helping with cell sorting; Ms. R. Miyachi-Hatasako and R. Tokimoto for technical assistance; Ms. A. Harada for secretarial assistance; Dr. Y. Masuda for technical advices in a gel-filtration analysis; and Drs. T. W. Mak and T. Nakano and the Leukemia Program Project of Hiroshima University for encouragement. This work was supported by Grants-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science and Technology of Japan, the Uehara Memorial Foundation, the Yamanouchi Foundation for Research on Metabolic Disorders, the Japan Leukaemia Research Fund, the Mitsubishi Pharma Research Foundation, the Novartis Foundation for the Promotion of Science, and the Daiwa Securities Health Foundation.
Footnotes
The authors declare no conflict of interest.
This article is a PNAS Direct Submission.
This article contains supporting information online at www.pnas.org/cgi/content/full/0800672105/DCSupplemental.
References
- 1.Takihara Y, Hara J. Polycomb-group genes and hematopoiesis. Int J Hematol. 2000;72:165–172. [PubMed] [Google Scholar]
- 2.Takihara Y. Role of Polycomb-group genes in normal and malignant stem cells. Int J Hematol. 2008;87:25–34. doi: 10.1007/s12185-007-0006-y. [DOI] [PubMed] [Google Scholar]
- 3.Wang H, et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature. 2004;431:873–878. doi: 10.1038/nature02985. [DOI] [PubMed] [Google Scholar]
- 4.Boyer LA, et al. Polycomb complexes repress developmental regulators in murine embryonic stem cells. Nature. 2006;441:349–353. doi: 10.1038/nature04733. [DOI] [PubMed] [Google Scholar]
- 5.Ohta H, et al. Polycomb group gene rae28 is required for sustaining activity of hematopoietic stem cells. J Exp Med. 2002;195:759–770. doi: 10.1084/jem.20011911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Kim JY, et al. Defective long-term repopulating ability in hematopoietic stem cells lacking the Polycomb-group gene rae28. Eur J Haematol. 2004;73:75–84. doi: 10.1111/j.1600-0609.2004.00268.x. [DOI] [PubMed] [Google Scholar]
- 7.Park IK, et al. Bmi-1 is required for maintenance of adult self-renewing haematopoietic stem cells. Nature. 2003;423:302–305. doi: 10.1038/nature01587. [DOI] [PubMed] [Google Scholar]
- 8.Calés C, et al. Inactivation of the Polycomb group protein Ring1B unveils an antiproliferative role in hematopoietic cell expansion and cooperation with tumorigenesis associated to Ink4a deletion. Mol Cell Biol. 2008;28:1018–1028. doi: 10.1128/MCB.01136-07. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Molofsky AV, et al. Bmi-1 dependence distinguishes neural stem cell self-renewal from progenitor proliferation. Nature. 2003;425:962–967. doi: 10.1038/nature02060. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Leeb M, Wutz A. Ring1B is crucial for the regulation of developmental control genes and PRC1 proteins but not X inactivation in embryonic cells. J Cell Biol. 2007;178:219–229. doi: 10.1083/jcb.200612127. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Lessard J, Sauvageau G. Bmi-1 determines the proliferative capacity of normal and leukaemic stem cells. Nature. 2003;423:255–260. doi: 10.1038/nature01572. [DOI] [PubMed] [Google Scholar]
- 12.Kotake Y, et al. pRB family proteins are required for H3K27 trimethylaion and Polycomb repression complexes binding to and silencing p16INK4a tumor suppressor gene. Genes Dev. 2007;21:49–54. doi: 10.1101/gad.1499407. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Fasano CA, et al. shRNA knockdown of Bmi-1 reveals a critical role for p21-Rb pathway in NSC self-renewal during development. Cell Stem Cell. 2007;1:87–99. doi: 10.1016/j.stem.2007.04.001. [DOI] [PubMed] [Google Scholar]
- 14.Chagraoui J, et al. E4F1: A novel candidate factor for mediating BMI1 function in primitive hematopoietic cells. Genes Dev. 2006;20:2110–2120. doi: 10.1101/gad.1453406. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Oguro H, et al. Differential impact of Ink4a and Arf on hematopoietic stem cells and their bone marrow microenvironment in Bmi1-deficient mice. J Exp Med. 2006;203:2247–2253. doi: 10.1084/jem.20052477. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Tomotsune D, et al. A novel member of murine Polycomb-group proteins, Sex comb on midleg homolog protein, is highly conserved, and interacts with RAE28/mph1 in vitro. Differentiation. 1999;65:229–239. doi: 10.1046/j.1432-0436.1999.6540229.x. [DOI] [PubMed] [Google Scholar]
- 17.Luo L, Yang X, Takihara Y, Knoetgen H, Kessel M. The cell-cycle regulator geminin inhibits Hox function through direct and polycomb-mediated interactions. Nature. 2004;427:749–753. doi: 10.1038/nature02305. [DOI] [PubMed] [Google Scholar]
- 18.Saxena S, Dutta A. Geminin-Cdt1 balance is critical for genetic stability. Mutat Res. 2005;569:111–121. doi: 10.1016/j.mrfmmm.2004.05.026. [DOI] [PubMed] [Google Scholar]
- 19.Seo S, et al. Geminin regulates neuronal differentiation by antagonizing Brg1 activity. Genes Dev. 2005;19:1723–1734. doi: 10.1101/gad.1319105. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Lessard J, et al. Functional antagonism of the Polycomb-group genes eed and Bmi1 in hemopoietic cell proliferation. Genes Dev. 1999;13:2691–2703. doi: 10.1101/gad.13.20.2691. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.McGarry TJ, Kirschner MW. Geminin, an inhibitor of DNA replication, is degraded during mitosis. Cell. 1998;93:1043–1053. doi: 10.1016/s0092-8674(00)81209-x. [DOI] [PubMed] [Google Scholar]
- 22.Shreeram S, Sparks A, Lane DP, Blow JJ. Cell type-specific responses of human cells to inhibition of replication licensing. Oncogene. 2002;21:6624–6632. doi: 10.1038/sj.onc.1205910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Nomura M, Takihara Y, Shimada K. Isolation and characterization of retinoic acid-inducible cDNA clones in F9 cells: One of the early inducible clones encodes a novel protein sharing several highly homologous regions with a Drosophila polyhomeotic protein. Differentiation. 1994;57:39–50. doi: 10.1046/j.1432-0436.1994.5710039.x. [DOI] [PubMed] [Google Scholar]
- 24.Ben-Saadon R, Zaaroor D, Ziv T, Ciechanover A. The Polycomb protein Ring1B generates self atypical mixed ubiquitin chains required for its in vitro histone H2A ligase activity. Mol Cell. 2006;24:701–711. doi: 10.1016/j.molcel.2006.10.022. [DOI] [PubMed] [Google Scholar]
- 25.Schuettengruber B, Chourrout D, Vervoort M, Leblanc B, Cavalli G. Genome regulation by polycomb and trithorax proteins. Cell. 2007;128:735–745. doi: 10.1016/j.cell.2007.02.009. [DOI] [PubMed] [Google Scholar]
- 26.de Napoles M, et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev Cell. 2004;7:663–676. doi: 10.1016/j.devcel.2004.10.005. [DOI] [PubMed] [Google Scholar]
- 27.Zhou W, et al. Histone H2A monoubiquitination represses transcription by inhibiting polymerase II transcriptional elongation. Mol Cell. 2008;29:69–80. doi: 10.1016/j.molcel.2007.11.002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 28.Kagey MH, Melhuish TA, Wotton D. The Polycomb protein Pc2 is a SUMO E3. Cell. 2003;113:127–137. doi: 10.1016/s0092-8674(03)00159-4. [DOI] [PubMed] [Google Scholar]
- 29.Shao Z, et al. Stabilization of chromatin structure by PRC1, a Polycomb complex. Cell. 1999;98:37–46. doi: 10.1016/S0092-8674(00)80604-2. [DOI] [PubMed] [Google Scholar]
- 30.Li A, Blow JJ. Non-proteolytic inactivation of geminin requires CDK-dependent ubiquitination. Nat Cell Biol. 2004;6:260–267. doi: 10.1038/ncb1100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 31.Del Bene F, Tessmar-Raible K, Wittbrodt J. Direct interaction of geminin and Six3 in eye development. Nature. 2004;427:745–749. doi: 10.1038/nature02292. [DOI] [PubMed] [Google Scholar]
- 32.Kim MY, et al. A repressor complex, AP4 transcription factor and geminin, negatively regulates expression of target genes in nonneuronal cells. Proc Natl Acad Sci USA. 2006;103:13074–13079. doi: 10.1073/pnas.0601915103. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 33.Vradii D, et al. Brg1, the ATPase subunit of the SWI/SNF chromatin remodeling complex, is required for myeloid differentiation to granulocytes. J Cell Physiol. 2006;206:112–118. doi: 10.1002/jcp.20432. [DOI] [PubMed] [Google Scholar]
- 34.Gonzalez MA, et al. Geminin is essential to prevent endoreduplication and to form pluripotent cells during mammalian development. Genes Dev. 2006;20:1880–1884. doi: 10.1101/gad.379706. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Hara K, Nakayama K-I, Nakayama K. Geminin is essential for the development of preimplantation mouse embryos. Genes Cells. 2006;11:1281–1293. doi: 10.1111/j.1365-2443.2006.01019.x. [DOI] [PubMed] [Google Scholar]
- 36.Kato Y, Koseki H, Vidal M, Nakauchi H, Iwama A. Unique composition of Polycomb repressive complex 1 in hematopoietic stem cells. Int J Hematol. 2007;85:179–181. doi: 10.1532/IJH97.06235. [DOI] [PubMed] [Google Scholar]
- 37.Takihara Y, et al. Targeted disruption of the mouse homologue of the Drosophila polyhomeotic gene leads to altered anteroposterior patterning and neural crest defects. Development. 1997;124:3673–3682. doi: 10.1242/dev.124.19.3673. [DOI] [PubMed] [Google Scholar]
- 38.Ohtsubo M, Okazaki H, Nishimoto T. The RCC1 protein, a regulator for the onset of chromosome condensation locates in the nucleus and binds to DNA. J Cell Biol. 1989;109:1389–1397. doi: 10.1083/jcb.109.4.1389. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Sawada A, et al. A congenital mutation of the novel gene LRRC8 causes agammaglobulinemia in humans. J Clin Invest. 2003;112:1707–1713. doi: 10.1172/JCI18937. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Kikuchi A, Williams LT. The post-translational modification of ras p21 is important for Raf-1 activation. J Biol Chem. 1994;269:20054–20059. [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.