Abstract
Transcription of the catalytic subunit of telomerase (hTERT) in keratinocytes can be induced by human papillomavirus type 16 (HPV16) E6/E6AP ubiquitin ligase through degradation of the repressor, NFX1-91. Here, we demonstrate that NFX1-91 interacts with the corepressor complex mSin3A/histone deacetylase (HDAC) at the hTERT promoter. By degrading NFX1-91, E6/E6AP changes the chromatin structure at the hTERT promoter as indicated by enhanced acetylation of histones H3 and H4 as well as dimethylation of H3K4. Knockdown of NFX1-91 by short hairpin RNA (shRNA) mimics the effect of E6 and leads to acetylation of histones H3 and H4. Conversely, knockdown of E6AP by shRNA suppresses histone acetylation at the hTERT promoter. These data demonstrate that targeted degradation of NFX1-91 by E6/E6AP dissociates the mSin3A/HDAC complex from the hTERT promoter and induces hTERT transcription.
Maintenance of telomere length by telomerase is required for sustained cell proliferation, and activation of telomerase is a critical step during cellular immortalization and malignant transformation (7). Expression of the catalytic subunit of telomerase (hTERT) is rate limiting for telomerase activity (28, 30, 32); thus, understanding the regulation of hTERT transcription is critical to elucidate how cells acquire immortality. The hTERT promoter contains several transcription factor-binding sites, including c-Myc (15, 42, 45), Sp1 (18, 37, 43), USF (12), and ER81 (13). Several studies showed that hTERT expression is minimal or absent in most somatic cells through both known transcriptional repressors, such as Mad1, menin, and SIP1 (24), and other endogenous factors demonstrated by chromosome transfer experiments, such as a putative tumor suppressor on chromosome 3p (17). Distinct repression and activation mechanisms likely operate in different tissues. For example, a complex containing BRCA1 inhibits c-Myc-induced hTERT promoter activity in breast cancer cells (23), estrogen upregulates telomerase in both mammary and ovarian epithelial cells (3, 29), and E2F-1 activates the hTERT promoter in some normal human somatic cells (44).
The human papillomavirus type 16 (HPV16) E6 oncoprotein can induce telomerase activity in human primary keratinocytes through induction of hTERT expression (9, 10, 33, 39). Studies on how E6 regulates hTERT have implicated E-box and X-box elements in the hTERT promoter (9, 10, 33, 40). c-Myc's role in directly enhancing transcription of hTERT has been demonstrated in several cell lines (15, 21, 45). However, the mechanism is still controversial and not well defined (9, 33, 39, 40). Although both E-box and c-Myc activities were found to be required for E6-mediated activation of hTERT (33, 39, 40), neither increased c-Myc protein level nor enhanced Myc-Max complex formation was detected with E6 expression (9, 33, 39, 40).
Recently, we identified NFX1-91 as a transcriptional repressor that binds to a consensus X-box element at the hTERT promoter, which overlaps with an E box (10). Two isoforms of NFX1 generated by alternative splicing were characterized and named NFX1-123 and NFX1-91. The short isoform, NFX1-91, is ubiquitinated and destabilized in epithelial cells expressing E6 (10). NFX1-91 is a target of the E6/E6AP ubiquitin ligase complex, and its degradation was proposed to mediate telomerase activation. Knockdown of NFX1-91 by shRNA resulted in derepression of endogenous hTERT expression and elevated the level of telomerase activity (10).
Sin3 is a core component of a large multiprotein corepressor complex with associated histone deacetylase (HDAC) activity (22). The Sin3 protein contains four imperfect repeats of a paired amphipathic helix (PAH) motif (41) that has been shown to mediate protein-protein interactions. The Sin3 protein does not bind to DNA on its own but provides a scaffold for several transcription factors with specific DNA binding activity to recruit HDACs to repress specific target genes. The mammalian Sin3 proteins, mSin3A and mSin3B, were discovered initially through their direct interaction with the transcriptional repressors of the Myc family network, Mad1 and Mxi1 (1, 14). Interaction of mSin3A with many transcription factors has linked the mSin3A corepressor complex to the regulation of diverse signaling pathways and associated biological processes, such as early embryonic development and proliferation and survival of primary, immortalized cells (8).
Here, we investigated the mechanism of NFX1-91 repression of the hTERT promoter and demonstrated that NFX1-91 interacts with the corepressor mSin3A/HDAC to maintain the deacetylated status at the hTERT promoter, thus providing a mechanism by which NFX1-91 represses hTERT expression.
MATERIALS AND METHODS
Cell culture, transfection and viral infection.
Primary human foreskin keratinocytes (HFKs) were derived from neonatal foreskin and grown in Epilife medium supplemented with 60 μM calcium chloride and human keratinocyte growth supplement (Cascade Biologics, Portland, OR). 293T cells were grown in Dulbecco's modified Eagle's medium (GIBCO-BRL, Carlsbad, CA) containing 10% fetal bovine serum and penicillin-streptomycin. Retroviruses and lentiviruses were produced either in the established viral producer cell line PA317 or in 293T cells by a transient vesicular stomatitis virus G-pseudotyped virus production protocol as previously described (2, 26). Small interfering RNA (siRNA) was transiently transfected into HFKs using the transfection reagent Lipofectamine RNAiMax (Invitrogen, Carlsbad, CA). Cells were collected at 72 h after transfection.
Plasmid constructs.
A glutathione S-transferase-NFX1-91 N-terminal fusion protein construct (GST-NFX1-91-N) was generated by inserting a DNA fragment encoding amino acids 1 to 356 of NFX1-91 into the EcoRI site of pGEX2T. This DNA fragment of NFX1-91-N was cloned by PCR from the full-length NFX1-91 cDNA. FLAG-tagged NFX1-91 and FLAG-tagged NFX1-123 constructs as well as NFX1-91 shRNA no. 1 in pBabe retroviral vectors were previously described (10). NFX1-91 shRNA and the scrambled shRNA were cloned into the BamHI and EcoRI sites of the FUGW lentiviral vector (26). The NFX1-91 shRNA no. 1 sequence is 5′-TGTGGAACCAGCCCAACTGCCCATCAGTCAA-3′, the NFX1-91 shRNA no. 2 sequence is 5′-AGATACCTGCACACGCATG-3′, and the scrambled shRNA sequence is 5′-CAGATGCTACAGTAGGACCTTACCCCAACGC-3′. E6AP shRNA no. 1 and the scrambled shRNA were cloned into the BamHI and EcoRI sites of the FUGW lentiviral vector. The E6AP shRNA no. 1 sequence is 5′-CTAATAGAACGCTACTACCACCAGTTAAC-3′, and the scrambled shRNA sequence is 5′-GACATAACCTCTACCAACAGTATCGACTA-3′. E6AP siRNA oligonucleotides are as follows: no. 2, ON-TARGETplus SMARTpool L-005137-00-0005 (Dharmacon, Lafayette, CO), and no. 3, CUGUAUACUGGAUGUACAU (sense) and AUGUACAUCCAGUAUACAG (antisense) (Sigma, St. Louis, MO). Control siRNA oligonucleotides are the ON-TARGETplus siControl nontargeting pool (Dharmacon, Lafayette, CO). Myc-tagged mSin3A deletion constructs have been described previously (22).
Pulse-chase assay.
Keratinocytes cultured in 60-mm plates were washed twice with cysteine- and methionine-free 154XP medium (Cascade Biologics, Portland, OR) and incubated for 15 min with the above medium. The medium was removed and pulsed with 1 ml 154XP medium containing 200 μCi/ml [35S]methionine (Easytag NEG772; Amersham Biosciences, Piscataway, NJ) for 10 min. Cells were washed three times with EpiLife keratinocyte medium and chased for various periods of time. Cells were washed twice with cold phosphate-buffered saline (PBS) and lysed with radioimmunoprecipitation assay buffer (10 mM Tris-Cl [pH 8.0], 140 mM NaCl, 1 mM EDTA, 1% Triton X-100, 0.1% sodium dodecyl sulfate [SDS], 0.1% sodium deoxycholate, and Complete protease inhibitor tablet (Roche, Alameda, CA). Lysates were sonicated for 10 seconds and clarified by centrifugation. Lysates were subsequently precleared with protein A-agarose prior to immunoprecipitation with anti-c-Myc (N-262; Santa Cruz Biotechnology, Santa Cruz, CA). After incubation with protein A-agarose beads, immunocomplexes were washed three times with PBS. The beads were boiled in sample buffer and subjected to 10% SDS-polyacrylamide gel electrophoresis (SDS-PAGE). The gels were dried and subjected to PhosphorImager analysis. Band intensities were measured using ImageQuant.
Western blotting.
HFK whole-cell lysates were prepared by trypsinizing cells, washing with cold PBS, and resuspending in WE16 lysis buffer (50 mM Tris-HCl [pH 7.5], 250 mM NaCl, 1% NP-40, 0.1% SDS, 20% glycerol, 10 μM zinc chloride, 80 mM β-glycerophosphate, 50 mM sodium chloride, 1 mM sodium orthovanadate, and Complete protease inhibitor tablet (Roche, Alameda, CA). Lysates were sonicated with a cup sonicator and clarified by centrifugation. Proteins were subjected to SDS-PAGE and transferred to Immunobilon-P membranes (Millipore, Billerica, MA). Western blotting was performed with mouse anti-c-Myc (C-33; Santa Cruz Biotechnology, Santa Cruz, CA), mouse anti-GAPDH (glyceraldehyde-3-phosphate dehydrogenase) (Abcam, Cambridge, MA), mouse anti-E6AP (Abnova, Taipei, Taiwan), and mouse antinucleolin (MS-3; Santa Cruz Biotechnology, Santa Cruz, CA). Rabbit anti-NFX1 and -NFX1-91 antibodies were described previously (10).
Real-time PCR.
RNA was isolated from HFK cells using the RNeasy kit (Qiagen, Valencia, CA). cDNA was synthesized using random hexamers and the Superscript II reverse transcriptase system (Invitrogen, Carlsbad, CA). Expression of hTERT, NFX1-91, E6AP, and 36B4 was detected by real-time PCR. The sequences of the hTERT, NFX1-91, and 36B4 primers were described previously (9). E6AP primers (GenBank accession no. NM_130838) were 5′-TTTTGGTGGTGATAGTGTCCCA-3′ (forward) and 5′-AGCCCACATGTCCCCAATAA-3′ (reverse).
ChIP.
Chromatin-cross-linking immunoprecipitation (ChIP) assays were performed as previously described (34) with minor modifications. HFKs grown in 150-mm plates were cross-linked for 10 min at room temperature with 10 ml EpiLife keratinocyte medium containing 1% formaldehyde. Cells were scraped and washed once with Paro Rinse I buffer (10 mM Tris-Cl [pH 8.0], 10 mM EDTA, 0.5 mM EGTA, 0.25% Triton X-100, 10 mM sodium butyrate) and once with Paro Rinse II buffer (10 mM Tris-Cl [pH 8.0], 1 mM EDTA, 0.5 mM EGTA, 0.2 M sodium chloride, 10 mM sodium butyrate). Cell pellets were resuspended in sonication buffer (10 mM Tris-Cl [pH 8.0], 1 mM EDTA, 0.5 mM EGTA, 10 mM sodium butyrate) and sonicated 10 for 15 seconds each to achieve an average chromatin size of about 600 bp. All buffers were supplemented with Complete protease inhibitor (Roche, Alameda, CA). Chromatin was solubilized by adding 1/20 of the volume of 20% sarcosyl and incubated at room temperature for 20 min. Lysates were clarified by centrifugation, diluted with 2× radioimmunoprecipitation assay buffer and precleared with protein A-agarose beads for 1 h at 4°C. Immunoprecipitations were performed overnight at 4°C with 25 μl rabbit anti-c-Myc (N-262; Santa Cruz Biotechnology, Santa Cruz, CA), 10 μl rabbit anti-acetyl-H3 (06-599; Upstate, Lake Placid, NY), 10 μl rabbit anti-acetyl-H4 (06-866; Upstate, Lake Placid, NY), 10 μl rabbit anti-dimethyl-H3K4 (07-030; Upstate, Lake Placid, NY), 15 μl rabbit anti-NFX1-91, and 15 μl rabbit anti-mSin3A. The same amounts of normal rabbit immunoglobulin G (IgG) and preimmune (pre-I) serum were used as negative control. Lysate from 1 × 107 cells was used for c-Myc immunoprecipitation, and lysates from 2 × 107 cells were used for acetyl-H3, acetyl-H4, dimethyl-H3K4, NFX1-91, and mSin3A immunoprecipitation. After incubation with protein A-agarose beads, immunocomplexes were washed twice with wash buffer 2 (20 mM Tris-Cl [pH 8.1], 2 mM EDTA, 500 mM NaCl, 0.1% SDS, 1% Triton X-100), three times with wash buffer 3 (10 mM Tris-Cl [pH 8.0], 1 mM EDTA, 0.25% LiCl, 1% NP-40, 1% sodium deoxycholate) and twice with Tris-EDTA (TE) buffer (pH 8.0). Extracted DNA from each immunoprecipitation was resuspended in 25 μl TE. In parallel, the input DNA prepared from 150 μl of lysate was resuspended in 150 μl TE. Duplex PCR was performed in 25-μl volumes with the inclusion of 0.1 μl [32P]dCTP by using the Failsafe PCR PreMix Selection kit, PreMix G (Epicenter Biotechnologies, Madison, WI). The primers for hTERT promoter were 5′-GGCCGGGCTCCCAGTGGATTCG-3′ (forward) and 5′-CAGCGGGGAGCGCGCGGCATCG-3′ (reverse), as previously described (46). A pair of primers for the β-globin promoter, 5′-GACAGGTACGGCTGTCATCA-3′ (forward) and 5′-AACGGCAGACTTCTCCTCAG-3′ (reverse), was used as an internal control, as previously described (11). The products were visualized by autoradiography and quantified by PhosphorImager analysis. For real-time PCR, extracted DNA from each immunoprecipitation was resuspended in 10 μl TE. In parallel, the input DNA prepared from 100 μl (10%) of lysate was resuspended in 100 μl TE. Real-time PCR was performed in 25-μl volumes by using Power Sybr green PCR master mix (Applied Biosystems, Foster City, CA). The primers for the hTERT promoter were 5′-CCGTCCAGGGAGCAAT-3′ (forward) (from position −575 to −560) and 5′-ACCACGAATGCCGGACGTGA-3′ (from position −509 to −490) (reverse).
In vitro binding assay.
35S-labeled mSin3A protein was made by in vitro translation reactions using the TNT Quick coupled transcription-translation system (Promega, Madison, WI). Cell lysates containing endogenous NFX1-91 were obtained from 293T cells treated with 10 μM MG-132 (Calbiochem, San Diego, CA) for 2 h prior to harvest as described previously (10), and 2.5 mg lysate was used in each immunoprecipitation. After lysates were precleared with protein A-agarose, 8 μl anti-NFX1-91 or pre-I serum and 5 μl in vitro-translated [35S]mSin3A were added in lysates and incubated for 2 h at 4°C. After incubation with protein A-agarose, the beads were washed four times with lysis buffer. The beads were boiled in 1× sample buffer, and proteins were subjected to 6% SDS-PAGE. The gels were dried and subjected to PhosphorImager analysis. Band intensities were measured using ImageQuant.
In vivo coimmunoprecipitation.
293T cells were cotransfected with pCS2-MT-mSin3A and pCMVg-FLAG-NFX1-91 using FuGENE 6 (Roche, Alameda, CA). Twenty-four hours later, cells were treated with 10 μM MG-132 for 2 h prior to harvest, and lysates were prepared as described previously (10). Cell lysates (2,500 μg) were precleared with protein A-agarose and then incubated with rabbit anti-FLAG antibody (Sigma, F7425, St. Louis, MO) overnight at 4°C. After incubation with protein A-agarose for 1 h, the beads were washed twice with lysis buffer, twice with high salt buffer (lysis buffer with 250 mM NaCl), and twice with lysis buffer again. Immunoprecipitates were separated by SDS-PAGE and immunoblotted with anti-Myc (9E10; Santa Cruz Biotechnology, Santa Cruz, CA) or rabbit anti-FLAG (F7425; Sigma, St. Louis, MO) antibody. For endogenous coimmunoprecipitation of NFX1-91 and mSin3A, cell lysates (1,500 μg) from 293T cells were immunoprecipitated with 15 μl rabbit anti-NFX1-91 antibody (15 μl pre-I serum was used as a control). Rabbit anti-mSin3A antibody (K-20; Santa Cruz Biotechnology) and rabbit anti-NFX1-91 antibody were used for Western blotting.
GST pull-down assay.
GST-NFX1-91-N and GST control protein were expressed in E. coli strain BL21-CodonPlus (Stratagene, La Jolla, CA). Proteins were purified with immobilized glutathione-Sepharose 4B (Amersham Biosciences, Piscataway, NJ). For assay of binding of GST-NFX1-91-N with endogenous mSin3A, nuclear extracts of 293T cells were dialyzed against PBS plus 0.1% NP-40 and then incubated with the same amount of GST-NFX1-91N or GST protein for 2 h at 4°C. Glutathione-Sepharose beads were added and left for 1 h. Beads were washed four times with PBS, boiled in sample buffer, and analyzed by SDS-PAGE. Immunoblotting was performed with rabbit anti-mSin3A antibody. For assay of binding of GST-NFX1-91-N with in vitro-translated mSin3A mutants, 20 μl GST-NFX1-91-N or GST protein was incubated with 5 μl in vitro-translated [35S]mSin3A mutants in 75 μl binding buffer (PBS, 1% NP-40, 2 mM dithiothreitol, 10 μM ZnCl, and protease inhibitors) for 2 h at 4°C. Twenty microliters of glutathione-Sepharose beads was added, left for 1 h, and washed three times with binding buffer, and bound proteins were subjected to SDS-PAGE. The gels were dried and subjected to PhosphorImager analysis for quantitation.
HDAC assay.
293T cells were transiently transfected with cytomegalovirus (CMV)-FLAG-NFX1-91, CMV-FLAG-NFX1-123, CMV-FLAG-HDAC2, and CMV-FLAG control vector using FuGENE 6. Immunoprecipitations were performed as described for the coimmunoprecipitation assay. Beads were washed three times with coimmunoprecipitation lysis buffer and once with HDAC assay buffer. HDAC activities were assayed by using the HDAC fluorescent activity assay/drug discovery kit (Biomol Research Laboratories, Plymouth Meeting, PA).
RESULTS AND DISCUSSION
E6 expression induces changes of chromatin modifications at the hTERT promoter without changing the c-Myc level.
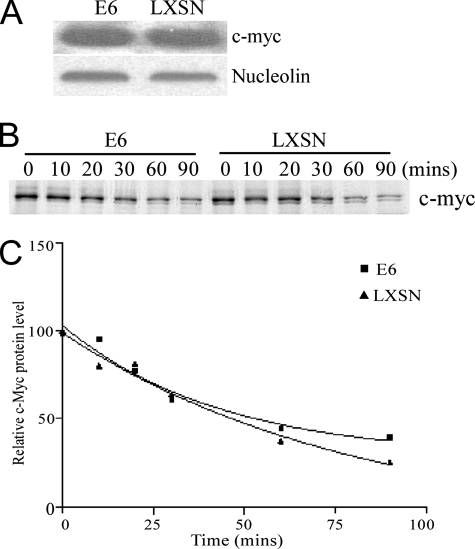
To examine whether c-Myc plays a role in E6-mediated hTERT transcription, we first investigated whether E6 has an effect on c-Myc protein stability. As shown in Fig. 1A, the expression of E6 in HFKs (E6-HFKs) did not detectably affect the total protein level of c-Myc compared to that in HFKs expressing the LXSN vector control (LXSN-HFKs), which is consistent with our previous research and other reports (9, 39). Also, no significant difference in the turnover of c-Myc protein was observed in E6-HFKs and LXSN-HFKs as determined by a paired t test (Fig. 1B and C). Immunofluorescence experiments did not show a difference in the subcellular location of c-Myc or the intensity of c-Myc staining in E6-HFKs compared to LXSN-HFKs (data not shown).
FIG. 1.
c-Myc steady state and half-life in E6- and LXSN-expressing HFKs. (A) Western blot of whole-cell lysates from HPV16 E6-HFK and LXSN-HFK cells. c-Myc was detected with mouse anti-c-Myc antibody (with nucleolin as a loading control). (B) c-Myc pulse-chase analysis in LXSN-HFKs and HPV16 E6-HFKs. Cells were pulse-labeled with [35S]methionine for 10 min and chased for the indicated times. c-Myc was immunoprecipitated with rabbit anti-c-Myc antibody and analyzed by SDS-PAGE. (C) The signal in panel B was visualized and quantitated by a PhosphorImager analysis. Nonlinear regression was performed with Prism software. The r2 values of the E6 and LXSN samples are 0.9689 and 0.9738, respectively. A paired t test was performed (P = 0.18).
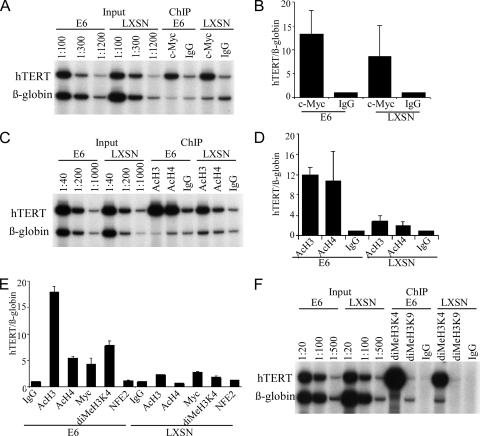
We therefore explored the possibility that c-Myc accessibility to the hTERT promoter is modulated by E6. ChIP experiments indicated that c-Myc bound to the hTERT promoter in both E6-HFKs and LXSN-HFKs (Fig. 2A). Results from seven experiments are summarized in Fig. 2B. Although the amount of c-Myc binding at the hTERT promoter appears to be greater in E6-HFKs, no significant difference was found (t test, P = 0.15). In contrast, ChIP experiments with anti-pan-acetyl-histone 3 (AcH3) and histone 4 (AcH4) indicated that acetylation levels of the hTERT promoter were increased approximately sixfold in E6-HFKs versus LXSN-HFKs (Fig. 2C and D). We repeated these experiments using real-time PCR to quantitate the immunoprecipitated sequences. Similar results were seen, with sixfold-increased acetylation in E6-expressing cells (Fig. 2E). Another marker of transcriptionally active chromatin, dimethylation of histone H3 lysine 4, was elevated fourfold in the E6-HFKs (Fig. 2E and F). Both the dimethyl- and trimethyl-H3K4 modifications have been shown to be enriched at actively transcribed genes (4, 31, 36). Recently, in yeast, methylation of H3K4 was shown to be an anchor for chromatin-remodeling complexes containing histone acetylases (HATs) (35). However, not all chromatin modifications were altered by E6 expression, as observed by ChIP with antibody to dimethylated histone 3 lysine 9 (Fig. 2F). In addition to IgG as a negative control, immunoprecipitation with an antibody to another transcription factor, NFE2, which does not bind to the hTERT promoter, showed only a background level of pulldown of hTERT promoter sequences and was unchanged by E6 expression (Fig. 2E).
FIG. 2.
c-Myc occupancy and histone modifications at the hTERT promoter in HFKs with or without E6. (A) ChIP analysis with anti-c-Myc antibody from LXSN-HFKs and HPV16 E6-HFKs. The first six lanes represent diluted DNAs from the inputs to indicate the linearity of the PCRs. (B) The enrichment of the hTERT promoter DNA relative to the β-globin gene as a control was quantitated. Error bars represent standard deviations from seven independent experiments. (C and D) ChIP analysis using anti-acetylated histone H3 and H4 (anti-acH3 and anti-acH4) antibodies from LXSN-HFKs and HPV16 E6-HFKs. Error bars represent standard deviations from two independent experiments. (E) ChIP analysis using anti-acH3, anti-acH4, anti-dimeH3K4, anti-c-Myc, and anti-NFE2 antibodies from LXSN-HFKs and E6-HFKs, with quantitation by real-time PCR. NFE2 is a nuclear factor that binds to the β-globin gene promoter, used as a negative control. The enrichment of hTERT promoter DNA is relative to the β-globin gene as control. Error bars represent standard deviations of multiple PCRs from two independent experiments. (F) ChIP analysis with anti-dimethyl-H3K4 and anti-dimethyl-H3K9 antibodies from LXSN-HFKs and E6-HFKs. The first six lanes represent diluted DNAs from the inputs to indicate the linearity of the PCRs. The β-globin gene was used as a control.
A role for histone modifications in the regulation of hTERT expression is consistent with previous studies indicating that inhibition of HDAC by trichostatin A in telomerase-negative cells can alter chromatin structure and induce hTERT expression (6, 19, 38). Thus, we hypothesized that in normal HFKs a repressor is responsible for inhibiting histone acetylation and that the presence of E6 interferes with this proposed repressor. E6 expression leads to multiple chromatin modifications at the hTERT promoter, as shown here (Fig. 2C to E), and to activation of hTERT transcription, as shown previously (9, 10, 33, 39, 40). The levels of c-Myc at the hTERT promoter are not commensurate with the changes seen in histone modifications or hTERT transcription (Fig. 2A, B, and E) (40).
E6-induced changes in histone modifications at the hTERT promoter are E6AP dependent.
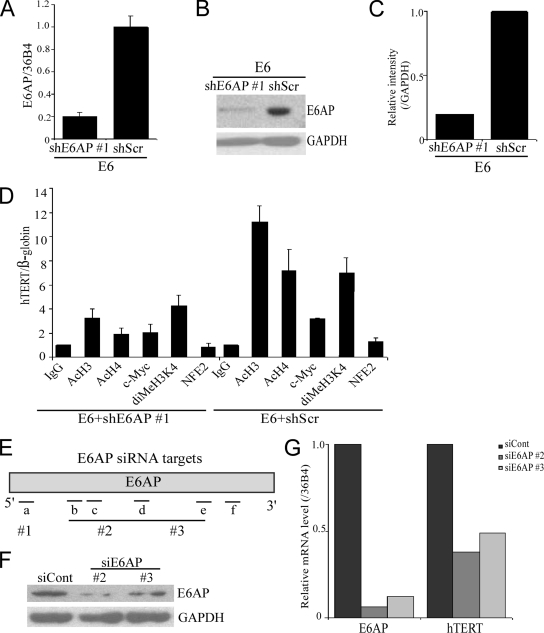
We previously reported that activation of telomerase by HPV16 E6 was dependent on E6AP (10), which has been confirmed recently (20, 25). We showed that knockdown of E6AP in E6-HFKs greatly reduced the levels of endogenous hTERT mRNA and telomerase activity measured by TRAP assay (10). Thus, if acetylation of the hTERT promoter is induced by E6/E6AP, then knockdown of E6AP expression with a specific shRNA to E6AP should block the acetylation of histone H3 and H4 at the hTERT promoter induced by E6. E6AP mRNA levels in E6-HFKs transduced with E6AP shRNA no. 1 were decreased to 20% compared to those in E6-HFKs with the scrambled shRNA (Fig. 3A). E6AP protein levels were determined by Western blotting (Fig. 3B), and the signal intensity was quantitated (Fig. 3C). Indeed, knockdown of E6AP in E6-HFKs showed that levels of AcH3, AcH4, and dimethyl-H3K4 at the hTERT promoter were reduced ∼2- to 4-fold relative to those in control E6-HFKs (Fig. 3D). These data indicate that E6-induced changes in histone modifications at the hTERT promoter are E6AP dependent. A 30% decrease in the level of c-Myc at the hTERT promoter was also observed in E6-HFKs with knockdown of E6AP compared to the control cells (Fig. 3D). We attribute this to the restoration of p53 levels by E6AP, which may reduce the level of c-Myc by both inhibiting cell proliferation and perhaps affecting c-Myc levels directly (16). To exclude off-target effects of E6AP shRNA, we have tested two more E6AP siRNAs. The target sites of these E6AP siRNAs are shown in Fig. 3E. E6AP expression was knocked down by E6AP siRNAs compared to control siRNA, as determined by Western blot analysis (Fig. 3F). As expected, knockdown of E6AP by different siE6APs in E6-expressing cells reduced the hTERT mRNA level (Fig. 3G).
FIG. 3.
Knockdown of E6AP induces changes of histone modifications at the hTERT promoter in E6-HFKs. (A) E6AP mRNA levels in E6-expressing cells with E6AP shRNA no. 1 (shE6AP no. 1) or the scrambled shRNA (shScr) were monitored by real-time PCR analysis of cDNAs using primers specific for E6AP and 36B4 (36B4 is a ribosomal protein gene used as a control). (B) Western blot analysis of E6AP in E6-expressing cells with knockdown of E6AP by specific shRNA or the scrambled shRNA as control (with GAPDH as a loading control). (C) The relative intensity of E6AP was quantitated using ImageJ software. (D) ChIP analysis was performed using anti-acH3, anti-acH4, anti-dimeH3K4, anti-c-Myc, and anti-NFE2 antibodies in E6-expressing cells with shE6AP no. 1 or shScr and quantitated by real-time PCR. NFE2 was used as a negative control. The enrichment of hTERT promoter DNA is relative to the β-globin gene. Error bars represent standard deviations of multiple PCRs from two independent experiments. (E) Map of E6AP (NM_130838) with indication of the target sites of shE6AP no. 1 and siE6AP no. 2 and no. 3. shE6AP no. 1 is targeted on site a (nucleotides 25 to 43). siE6AP no. 2 contains a combination of four different siE6APs targeting on sites b (nucleotides 633 to 651), c (nucleotides 662 to 680), d (nucleotides 1003 to 1020), and e (nucleotides 1589 to 1607). siE6AP no. 3 is targeted on site f (nucleotides 1809 to 1827). (F) Western blot analysis of E6AP in E6-expressing cells transfected with E6AP-specific siRNAs or the control siRNA (siE6AP no. 2, no. 3, and siCont). GAPDH was used as a loading control. (G) Levels of E6AP and hTERT mRNAs in E6-expressing cells with E6AP siRNA or the control siRNA were monitored by real-time PCR analysis of cDNAs using primers specific for E6AP, hTERT, and 36B4 (as a control).
NFX1-91 binds to the hTERT promoter and represses histone acetylation.
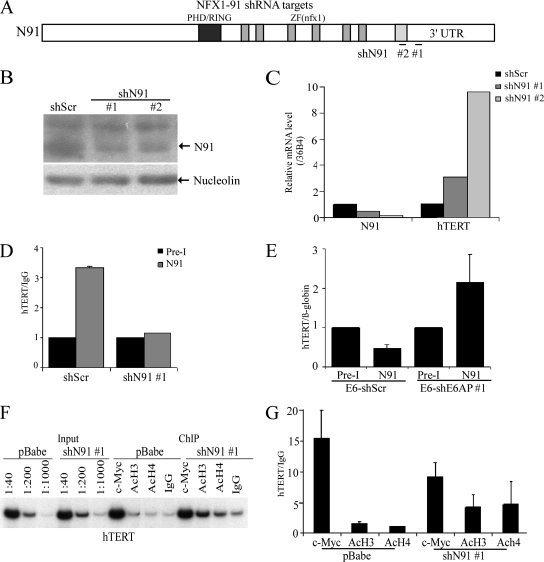
To manipulate gene expression of NFX1-91, two shRNAs were used to knock down NFX1-91. One shRNA is specific to the 3′ untranslated region of NFX1-91 (shN91 no. 1), and the other one is specific to the C terminus of NFX1-91 (shN91 no. 2). The target sites of these shRNAs are shown in Fig. 4A. The NFX1-91 protein level was knocked down by treatment with both shN91s compared to the scrambled shRNA, as determined by Western blotting (Fig. 4B). As expected, knockdown of NFX1-91 by each shN91 in HFK cells increased the hTERT mRNA level (Fig. 4C).
FIG. 4.
NFX1-91 binds to the hTERT promoter and represses acetylation of H3 and H4 in HFKs. (A) Map of NFX1-91 with indication of the target sites of N91 shRNA no. 1 and no. 2. shN91 no. 1 is targeted on the site (nucleotides 25 to 55) within the 3′ untranslated region of N91, and shN91 no. 2 is targeted on the site (nucleotides 2477 to 2495) within the unique C terminus of N91. (B)Western blot analysis of N91 in HFK cells with treatment of shN91 or shScr. Nucleolin was used as a loading control. (C) Levels of N91 and hTERT mRNA in HFK cells with shN91 or shScr were monitored by real-time PCR analysis of cDNAs using primers specific for N91, hTERT, and 36B4 (as a control). (D) ChIP analysis with rabbit anti-NFX1-91 (N91) and pre-I serum from HFKs transduced with shN91 no. 1 or shScr and quantitated by real-time PCR. Relative enrichment was calculated by the ratio of the signal obtained with the specific antibody relative to rabbit IgG as a control. (E) ChIP analysis with rabbit anti-N91 and pre-I serum from E6-expressing cells transduced with shE6AP no. 1 or shScr and quantitated by real-time PCR. The enrichment of hTERT promoter DNA is relative to the β-globin gene. (F and G) ChIP analysis with anti-c-Myc, anti-acH3, and anti-acH4 antibodies from HFKs transduced with pBABE-shN91 no. 1 or empty vector pBABE. Relative enrichment was calculated by the ratio of the signal obtained with the specific antibody relative to rabbit IgG as a control. Error bars represent standard deviations of multiple PCRs from two independent experiments.
Our previous data showed that NFX1-91 binds to the hTERT proximal X box in vitro (10). To determine whether NFX1-91 can bind to the hTERT promoter in vivo, a ChIP assay was performed using an antibody specific for NFX1-91 and was quantitated with real-time PCR using primers to the hTERT promoter. As shown in Fig. 4D, NFX1-91 binding to the hTERT promoter was enriched threefold relative to the pre-I serum, and knockdown of NFX1-91 expression by shRNA reduced the binding level of NFX1-91 to the hTERT promoter to background levels. Also, as anticipated from our previous data showing that the reduced level of NFX1-91 in E6 cells was dependent on the ability of E6 to bind E6AP (10), we also observed that there was only a background level of NFX1-91 binding to the hTERT promoter in E6-expressing cells, and knockdown of E6AP in E6-HFKs restored the level of NFX1-91 at the hTERT promoter (Fig. 4E). A small decrease (∼30%) of c-Myc binding to the hTERT promoter was detected in NFX1-91 knockdown cells (Fig. 4F and G). However, as anticipated, AcH3 and AcH4 levels increased ∼3- to 5-fold in HFK-shN91 cells versus control HFKs (Fig. 4F and G). Taken together, these data suggest that NFX1-91 may interact with a corepressor complex containing HDAC activity at the hTERT promoter.
NFX1-91 interacts with the corepressor mSin3A in vitro and in vivo.
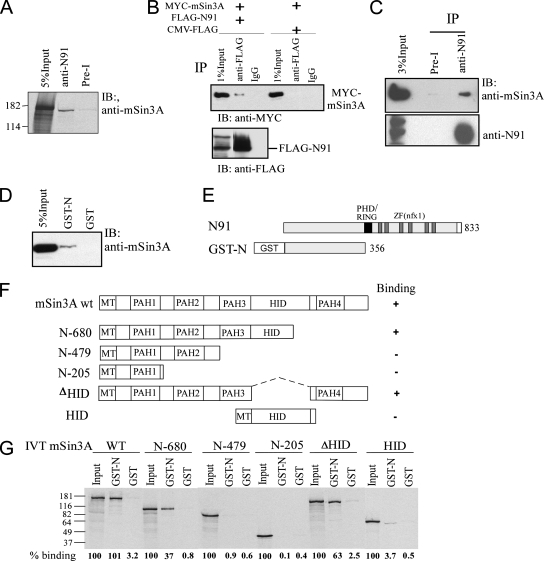
To establish whether NFX1-91 interacts with a corepressor complex containing HDAC activity at the hTERT promoter, the possible interaction of NFX1-91 and mSin3A was explored. First, in vitro coimmunoprecipitation experiments were performed from cell lysates containing endogenous NFX1-91 and in vitro-translated mSin3A. As shown in Fig. 5A, NFX1-91 formed a complex with mSin3A. An in vivo coimmunoprecipitation experiment showed that FLAG-tagged NFX1-91 bound to Myc-tagged mSin3A when both were transiently expressed in 293T cells (Fig. 5B). In addition, an association between endogenous NFX1-91 and mSin3A in 293T cells was detected by immunoprecipitation with anti-NFX1-91 antibody (Fig. 5C).
FIG. 5.
NFX1-91 interacts with mSin3A in vitro and in vivo. (A) In vitro binding of full-length N91 and mSin3A. In vitro-translated 35S-labeled full-length mSin3A was incubated with 293T cell lysates (containing endogenous N91) and immunoprecipitated with rabbit anti-N91 antibody or pre-I serum as a negative control. (B) Top, in vivo interaction of full-length N91 and mSin3A. 293T cells were cotransfected with FLAG-N91 and Myc-mSin3A. CMV-FLAG is the empty vector control. Lysates were immunoprecipitated (IP) with rabbit anti-FLAG antibody and blotted (IB) with anti-Myc antibody. Bottom, a parallel experiment with probing with anti-FLAG to show the overexpression of FLAG-N91. (C) Coimmunoprecipitation assay was performed to determine the interaction of endogenous N91 and mSin3A. 293T cell lysates were immunoprecipitated with rabbit anti-N91 antibody or pre-I serum and blotted with anti-mSin3A antibody and anti-N91 antibody, respectively. The input lane contained 3% of the amount used in each IP assay. (D) In vitro binding of the GST-N91 N terminus with endogenous mSin3A. Purified GST-N protein was incubated with 293T cell nuclear extracts (containing endogenous mSin3A). Western blotting was performed with anti-mSin3A antibody. GST protein alone was used as a negative control. The input lane contained 5% of the amount used in the binding assay. (E) Schematic diagram of full-length N91 indicating the N-terminal domain used to construct the GST-N91 fusion. (F) Schematic summary of Myc-tagged mSin3A deletion constructs and N91 binding. wt, wild type. (G) GST pull-down assay using GST-N and in vitro-translated, 35S-labeled, Myc-tagged mSin3A deletion constructs. The input lane contained 5% of the amount used in the binding assay for each in vitro-translated protein. Percent binding was calculated relative to the 5% input for each mSin3A deletion construct.
To map the interaction between NFX1-91 and mSin3A, GST pull-down assays were performed. mSin3A bound to the N-terminal domain of the NFX1-91 encompassing amino acids 1 to 356 (Fig. 5D and E). The domain in mSin3A that interacted with GST-NFX1-91-N was mapped using in vitro-translated mSin3A constructs. The previously reported interaction between mSin3A and transcriptional repressors occurs mainly through four imperfect repeats of 70 to 90 amino acids, named PAH domains (Fig. 5F). The interaction between PAH2 and the Mad family of repressors, as well as other transcription factors, has been extensively characterized and involves a specific structural motif named the Sin interaction domain (5). The PAH3 domain has been shown to anchor DNA-binding proteins, such as the myeloid leukemia gene 1, AML1 (27), and ETS1. NFX1 lacks a recognizable Sin interaction domain, and Fig. 5G shows that all Myc-tagged mSin3A deletion constructs containing PAH3 bound the NFX1-91 N terminus, whereas Myc-tagged mSin3A deletion constructs without PAH3 were not able to bind the NFX1-91 N terminus, indicating that binding of NFX1-91 through its amino terminus is mediated at least in part by the PAH3 domain of mSin3A.
mSin3A is present at the hTERT promoter in HFKs.
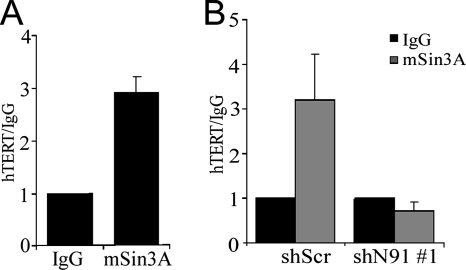
To determine whether mSin3A was present at the hTERT promoter, ChIP experiments were performed using anti-mSin3A antibody. As shown in Fig. 6A, mSin3A is present at the hTERT promoter in HFKs. Importantly, when NFX1-91 was knocked down by shN91 in HFKs, mSin3A levels at the hTERT promoter were reduced to background levels (Fig. 6B). This finding further supports our conclusion that mSin3A interacts with NFX1-91 at the hTERT promoter.
FIG. 6.
mSin3A binds to the hTERT promoter in HFKs. (A) ChIP analysis with rabbit anti-mSin3A and rabbit IgG as a control from HFKs, quantitated by real-time PCR. Relative enrichment was calculated by the ratio of the signal obtained with the specific antibody relative to rabbit IgG as control. Error bars represent standard deviations of multiple PCRs from two independent experiments. (B) ChIP analysis with rabbit anti-mSin3A and rabbit IgG from HFKs transduced with shN91 or shScr, quantitated by real-time PCR. Error bars represent standard deviations of multiple PCRs from two independent experiments.
NFX1-91 immunoprecipitates contain HDAC activity.
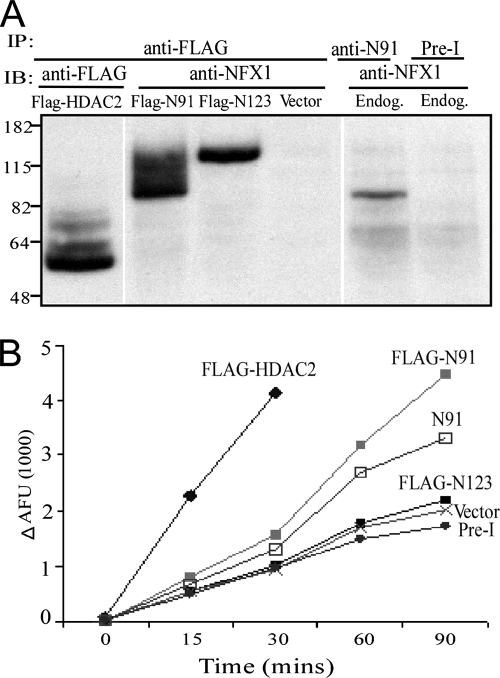
To test whether NFX1-91 interacts with a functional mSin3A/HDAC complex, the association of NFX1-91 with HDAC activity was examined. Both isoforms of NFX1 (NFX1-91 and NFX1-123) were expressed as FLAG-tagged proteins in 293T cells, and FLAG-HDAC2 was used as a positive control (Fig. 7A). After immunoprecipitation with anti-FLAG antibody, the immunocomplexes were assayed for HDAC activity. Indeed, the FLAG-NFX1-91 complex contained substantial HDAC activity, while FLAG-NFX1-123 showed HDAC activity comparable to that of pre-I serum (Fig. 7B). These results are consistent with the lack of repressor activity for NFX1-123 (10). To determine whether endogenous NFX1-91 could copurify with HDAC, immunoprecipitation of the untransfected 293T cell lysates with NFX1-91 antibody was assayed for HDAC activity. This showed that endogenous NFX1-91 contained HDAC activity (Fig. 7B). These data suggest that NFX1-91 may interact with mSin3A/HDAC complexes to target genes and thus maintain H3 and H4 deacetylation at the hTERT promoter.
FIG. 7.
HDAC activity is immunoprecipitated by NFX1-91. (A) Western analysis of immunoprecipitated proteins used in the HDAC assay. (B) HADC assay. FLAG-N91, FLAG-NFX1-123 (FLAG-N123), and FLAG-HDAC2 were overexpressed in 293T cells. Two hours prior to harvesting cells, 10 μM MG-132 was added to the culture. Whole-cell lysates (12.5 mg) were immunoprecipitated with rabbit anti-FLAG antibody. FLAG-HDAC2 was used as a positive control and CMV-based empty vector as a negative control. For endogenous N91, 293T cell lysate (12.5 mg) was immunoprecipitated with rabbit anti-N91 antibody or pre-I serum. HDAC activity was assayed at different time points during incubation.
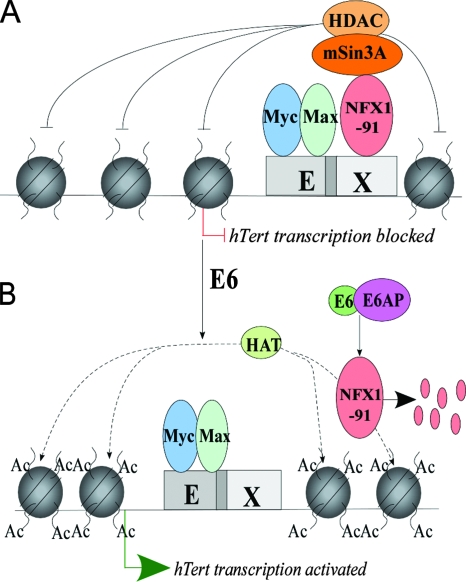
Based on the results presented here and our previous data (9, 10), we propose a model for how HPV16 E6 activates hTERT transcription in HFKs (Fig. 8). In normal HFKs, c-Myc binds to E boxes at the hTERT promoter. NFX1-91 binds to the X box overlapping the proximal E box at the hTERT promoter and binds to mSin3A/HDAC complexes, maintaining chromatin at the hTERT promoter in a deacetylated state. We do not know whether NFX1-91 occupancy at the hTERT promoter blocks the ability of c-Myc to recruit HATs such as TRRAP or whether NFX1-91 blocks HAT activity. Upon expression of HPV16 E6, the E6/E6AP complex induces NFX1-91 ubiquitination and degradation and removes the mSin3A/HDAC inhibitory complex. Acetylation and methylation of histones are seen at the hTERT promoter in E6-expressing cells. It is not known whether c-Myc or other factors recruit the HATs to the hTERT promoter, nor are the identities of the HATs known. Our attempt to immunoprecipitate TRRAP at the hTERT promoter was not successful (data not shown). In conclusion, our data support a mechanism in which E6/E6AP removes an NFX1-91/mSin3A/HDAC repressor complex to promote transcription of hTERT in human epithelial cells.
FIG. 8.
Model of HPV16 E6 activation of hTERT in HFKs. (A) In normal HFKs, c-Myc binds E boxes at the hTERT promoter. NFX1-91 interacts with the mSin3A/HDAC repressor complex and binds to the X box overlapping the proximal E box at the hTERT promoter. The presence of the NFX1-91/mSin3A/HDAC repressor complex maintains the deacetylated status of histones. (B) E6 expression induces degradation of NFX1-91 and relieves the mSin3A/HDAC inhibitory complex, which allows the acetylation of histone mediated by either c-Myc or other, unknown transcriptional factors.
Acknowledgments
We thank Lindy Gewin for reagents and advice, Kristin Robinson for excellent technical assistance, the Eisenman laboratory for mSin3A reagents, the Vasioukhin laboratory for the FUGW lentiviral vector, and Rachel Katzenellenbogen for critical reading of the manuscript.
This work was supported by NIH grant R01CA064795 to D.A.G.
Footnotes
Published ahead of print on 27 May 2008.
REFERENCES
- 1.Ayer, D. E., Q. A. Lawrence, and R. N. Eisenman. 1995. Mad-Max transcriptional repression is mediated by ternary complex formation with mammalian homologs of yeast repressor Sin3. Cell 80767-776. [DOI] [PubMed] [Google Scholar]
- 2.Bartz, S. R., and M. A. Vodicka. 1997. Production of high-titer human immunodeficiency virus type 1 pseudotyped with vesicular stomatitis virus glycoprotein. Methods 12337-342. [DOI] [PubMed] [Google Scholar]
- 3.Bayne, S., and J. P. Liu. 2005. Hormones and growth factors regulate telomerase activity in ageing and cancer. Mol. Cell Endocrinol. 24011-22. [DOI] [PubMed] [Google Scholar]
- 4.Bernstein, B. E., M. Kamal, K. Lindblad-Toh, S. Bekiranov, D. K. Bailey, D. J. Huebert, S. McMahon, E. K. Karlsson, E. J. Kulbokas III, T. R. Gingeras, S. L. Schreiber, and E. S. Lander. 2005. Genomic maps and comparative analysis of histone modifications in human and mouse. Cell 120169-181. [DOI] [PubMed] [Google Scholar]
- 5.Brubaker, K., S. M. Cowley, K. Huang, L. Loo, G. S. Yochum, D. E. Ayer, R. N. Eisenman, and I. Radhakrishnan. 2000. Solution structure of the interacting domains of the Mad-Sin3 complex: implications for recruitment of a chromatin-modifying complex. Cell 103655-665. [DOI] [PubMed] [Google Scholar]
- 6.Cong, Y. S., and S. Bacchetti. 2000. Histone deacetylation is involved in the transcriptional repression of hTERT in normal human cells. J. Biol. Chem. 27535665-35668. [DOI] [PubMed] [Google Scholar]
- 7.Counter, C. M., A. A. Avilion, C. E. LeFeuvre, N. G. Stewart, C. W. Greider, C. B. Harley, and S. Bacchetti. 1992. Telomere shortening associated with chromosome instability is arrested in immortal cells which express telomerase activity. EMBO J. 111921-1929. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Dannenberg, J. H., G. David, S. Zhong, J. van der Torre, W. H. Wong, and R. A. Depinho. 2005. mSin3A corepressor regulates diverse transcriptional networks governing normal and neoplastic growth and survival. Genes Dev. 191581-1595. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Gewin, L., and D. A. Galloway. 2001. E box-dependent activation of telomerase by human papillomavirus type 16 E6 does not require induction of c-myc. J. Virol. 757198-7201. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Gewin, L., H. Myers, T. Kiyono, and D. A. Galloway. 2004. Identification of a novel telomerase repressor that interacts with the human papillomavirus type-16 E6/E6-AP complex. Genes Dev. 182269-2282. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Gomez-Roman, N., C. Grandori, R. N. Eisenman, and R. J. White. 2003. Direct activation of RNA polymerase III transcription by c-Myc. Nature 421290-294. [DOI] [PubMed] [Google Scholar]
- 12.Goueli, B. S., and R. Janknecht. 2003. Regulation of telomerase reverse transcriptase gene activity by upstream stimulatory factor. Oncogene. 228042-8047. [DOI] [PubMed] [Google Scholar]
- 13.Goueli, B. S., and R. Janknecht. 2004. Upregulation of the catalytic telomerase subunit by the transcription factor ER81 and oncogenic HER2/Neu, Ras, or Raf. Mol. Cell. Biol. 2425-35. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Grandori, C., S. M. Cowley, L. P. James, and R. N. Eisenman. 2000. The Myc/Max/Mad network and the transcriptional control of cell behavior. Annu. Rev. Cell Dev. Biol. 16653-699. [DOI] [PubMed] [Google Scholar]
- 15.Greenberg, R. A., R. C. O'Hagan, H. Deng, Q. Xiao, S. R. Hann, R. R. Adams, S. Lichtsteiner, L. Chin, G. B. Morin, and R. A. DePinho. 1999. Telomerase reverse transcriptase gene is a direct target of c-Myc but is not functionally equivalent in cellular transformation. Oncogene 181219-1226. [DOI] [PubMed] [Google Scholar]
- 16.Ho, J. S., W. Ma, D. Y. Mao, and S. Benchimol. 2005. p53-dependent transcriptional repression of c-myc is required for G1 cell cycle arrest. Mol. Cell. Biol. 257423-7431. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 17.Horikawa, I., and J. C. Barrett. 2003. Transcriptional regulation of the telomerase hTERT gene as a target for cellular and viral oncogenic mechanisms. Carcinogenesis 241167-1176. [DOI] [PubMed] [Google Scholar]
- 18.Horikawa, I., P. L. Cable, C. Afshari, and J. C. Barrett. 1999. Cloning and characterization of the promoter region of human telomerase reverse transcriptase gene. Cancer Res. 59826-830. [PubMed] [Google Scholar]
- 19.Hou, M., X. Wang, N. Popov, A. Zhang, X. Zhao, R. Zhou, A. Zetterberg, M. Bjorkholm, M. Henriksson, A. Gruber, and D. Xu. 2002. The histone deacetylase inhibitor trichostatin A derepresses the telomerase reverse transcriptase (hTERT) gene in human cells. Exp. Cell Res. 27425-34. [DOI] [PubMed] [Google Scholar]
- 20.James, M. A., J. H. Lee, and A. J. Klingelhutz. 2006. HPV16-E6 associated hTERT promoter acetylation is E6AP dependent, increased in later passage cells and enhanced by loss of p300. Int. J. Cancer 1191878-1885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Kyo, S., M. Takakura, T. Taira, T. Kanaya, H. Itoh, M. Yutsudo, H. Ariga, and M. Inoue. 2000. Sp1 cooperates with c-Myc to activate transcription of the human telomerase reverse transcriptase gene (hTERT). Nucleic Acids Res. 28669-677. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 22.Laherty, C. D., W. M. Yang, J. M. Sun, J. R. Davie, E. Seto, and R. N. Eisenman. 1997. Histone deacetylases associated with the mSin3 corepressor mediate mad transcriptional repression. Cell 89349-356. [DOI] [PubMed] [Google Scholar]
- 23.Li, H., T. H. Lee, and H. Avraham. 2002. A novel tricomplex of BRCA1, Nmi, and c-Myc inhibits c-Myc-induced human telomerase reverse transcriptase gene (hTERT) promoter activity in breast cancer. J. Biol. Chem. 27720965-20973. [DOI] [PubMed] [Google Scholar]
- 24.Lin, S. Y., and S. J. Elledge. 2003. Multiple tumor suppressor pathways negatively regulate telomerase. Cell 113881-889. [DOI] [PubMed] [Google Scholar]
- 25.Liu, X., H. Yuan, B. Fu, G. L. Disbrow, T. Apolinario, V. Tomaic, M. L. Kelley, C. C. Baker, J. Huibregtse, and R. Schlegel. 2005. The E6AP ubiquitin ligase is required for transactivation of the hTERT promoter by the human papillomavirus E6 oncoprotein. J. Biol. Chem. 28010807-10816. [DOI] [PubMed] [Google Scholar]
- 26.Lois, C., E. J. Hong, S. Pease, E. J. Brown, and D. Baltimore. 2002. Germline transmission and tissue-specific expression of transgenes delivered by lentiviral vectors. Science 295868-872. [DOI] [PubMed] [Google Scholar]
- 27.Lutterbach, B., J. J. Westendorf, B. Linggi, S. Isaac, E. Seto, and S. W. Hiebert. 2000. A mechanism of repression by acute myeloid leukemia-1, the target of multiple chromosomal translocations in acute leukemia. J. Biol. Chem. 275651-656. [DOI] [PubMed] [Google Scholar]
- 28.Meyerson, M., C. M. Counter, E. N. Eaton, L. W. Ellisen, P. Steiner, S. D. Caddle, L. Ziaugra, R. L. Beijersbergen, M. J. Davidoff, Q. Liu, S. Bacchetti, D. A. Haber, and R. A. Weinberg. 1997. hEST2, the putative human telomerase catalytic subunit gene, is up-regulated in tumor cells and during immortalization. Cell 90785-795. [DOI] [PubMed] [Google Scholar]
- 29.Misiti, S., S. Nanni, G. Fontemaggi, Y. S. Cong, J. Wen, H. W. Hirte, G. Piaggio, A. Sacchi, A. Pontecorvi, S. Bacchetti, and A. Farsetti. 2000. Induction of hTERT expression and telomerase activity by estrogens in human ovary epithelium cells. Mol. Cell. Biol. 203764-3771. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Nakamura, T. M., G. B. Morin, K. B. Chapman, S. L. Weinrich, W. H. Andrews, J. Lingner, C. B. Harley, and T. R. Cech. 1997. Telomerase catalytic subunit homologs from fission yeast and human. Science 277955-959. [DOI] [PubMed] [Google Scholar]
- 31.Ng, H. H., F. Robert, R. A. Young, and K. Struhl. 2003. Targeted recruitment of Set1 histone methylase by elongating Pol II provides a localized mark and memory of recent transcriptional activity. Mol. Cell 11709-719. [DOI] [PubMed] [Google Scholar]
- 32.Nugent, C. I., and V. Lundblad. 1998. The telomerase reverse transcriptase: components and regulation. Genes Dev. 121073-1085. [DOI] [PubMed] [Google Scholar]
- 33.Oh, S. T., S. Kyo, and L. A. Laimins. 2001. Telomerase activation by human papillomavirus type 16 E6 protein: induction of human telomerase reverse transcriptase expression through Myc and GC-rich Sp1 binding sites. J. Virol. 755559-5566. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 34.Orlando, V., H. Strutt, and R. Paro. 1997. Analysis of chromatin structure by in vivo formaldehyde cross-linking. Methods 11205-214. [DOI] [PubMed] [Google Scholar]
- 35.Pray-Grant, M. G., J. A. Daniel, D. Schieltz, J. R. Yates, 3rd, and P. A. Grant. 2005. Chd1 chromodomain links histone H3 methylation with SAGA- and SLIK-dependent acetylation. Nature 433434-438. [DOI] [PubMed] [Google Scholar]
- 36.Santos-Rosa, H., R. Schneider, A. J. Bannister, J. Sherriff, B. E. Bernstein, N. C. Emre, S. L. Schreiber, J. Mellor, and T. Kouzarides. 2002. Active genes are tri-methylated at K4 of histone H3. Nature 419407-411. [DOI] [PubMed] [Google Scholar]
- 37.Takakura, M., S. Kyo, T. Kanaya, H. Hirano, J. Takeda, M. Yutsudo, and M. Inoue. 1999. Cloning of human telomerase catalytic subunit (hTERT) gene promoter and identification of proximal core promoter sequences essential for transcriptional activation in immortalized and cancer cells. Cancer Res. 59551-557. [PubMed] [Google Scholar]
- 38.Takakura, M., S. Kyo, Y. Sowa, Z. Wang, N. Yatabe, Y. Maida, M. Tanaka, and M. Inoue. 2001. Telomerase activation by histone deacetylase inhibitor in normal cells. Nucleic Acids Res. 293006-3011. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Veldman, T., I. Horikawa, J. C. Barrett, and R. Schlegel. 2001. Transcriptional activation of the telomerase hTERT gene by human papillomavirus type 16 E6 oncoprotein. J. Virol. 754467-4472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 40.Veldman, T., X. Liu, H. Yuan, and R. Schlegel. 2003. Human papillomavirus E6 and Myc proteins associate in vivo and bind to and cooperatively activate the telomerase reverse transcriptase promoter. Proc. Natl. Acad. Sci. USA 1008211-8216. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Wang, H., I. Clark, P. R. Nicholson, I. Herskowitz, and D. J. Stillman. 1990. The Saccharomyces cerevisiae SIN3 gene, a negative regulator of HO, contains four paired amphipathic helix motifs. Mol. Cell. Biol. 105927-5936. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Wang, J., L. Y. Xie, S. Allan, D. Beach, and G. J. Hannon. 1998. Myc activates telomerase. Genes Dev. 121769-1774. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Wick, M., D. Zubov, and G. Hagen. 1999. Genomic organization and promoter characterization of the gene encoding the human telomerase reverse transcriptase (hTERT). Gene 23297-106. [DOI] [PubMed] [Google Scholar]
- 44.Won, J., J. Yim, and T. K. Kim. 2002. Opposing regulatory roles of E2F in human telomerase reverse transcriptase (hTERT) gene expression in human tumor and normal somatic cells. FASEB J. 161943-1945. [DOI] [PubMed] [Google Scholar]
- 45.Wu, K. J., C. Grandori, M. Amacker, N. Simon-Vermot, A. Polack, J. Lingner, and R. Dalla-Favera. 1999. Direct activation of TERT transcription by c-MYC. Nat. Genet. 21220-224. [DOI] [PubMed] [Google Scholar]
- 46.Xu, D., N. Popov, M. Hou, Q. Wang, M. Bjorkholm, A. Gruber, A. R. Menkel, and M. Henriksson. 2001. Switch from Myc/Max to Mad1/Max binding and decrease in histone acetylation at the telomerase reverse transcriptase promoter during differentiation of HL60 cells. Proc. Natl. Acad. Sci. USA 983826-3831. [DOI] [PMC free article] [PubMed] [Google Scholar]