Abstract
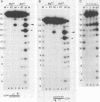
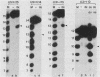
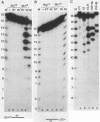
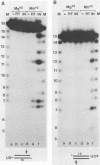
The purified integration protein (IN) of avian myeloblastosis virus is shown to nick double-stranded oligodeoxynucleotide substrates that mimic the ends of the linear form of viral DNA. In the presence of Mg2+, nicks are created 2 nucleotides from the 3' OH ends of both the U5 plus strand and the U3 minus strand. Similar cleavage is observed in the presence of Mn2+ but only when the extent of the reaction is limited. Neither the complementary strands nor sequences representing the termini of human immunodeficiency virus type 1 DNA were cleaved at analogous positions. Analysis of a series of substrates containing U5 base substitutions has defined the sequence requirements for site-selective nicking; nucleotides near the cleavage site are most critical for activity. The minimum substrate size required to demonstrate significant activity corresponds to the nearly perfect 15-base terminal inverted repeat. This in vitro activity of IN thus produces viral DNA ends that are joined to host DNA in vivo and corresponds to an expected early step in the integrative recombination reaction. These results provide the first enzymatic support using purified retroviral proteins for a linear DNA precursor to the integrated provirus.
Full text
PDF


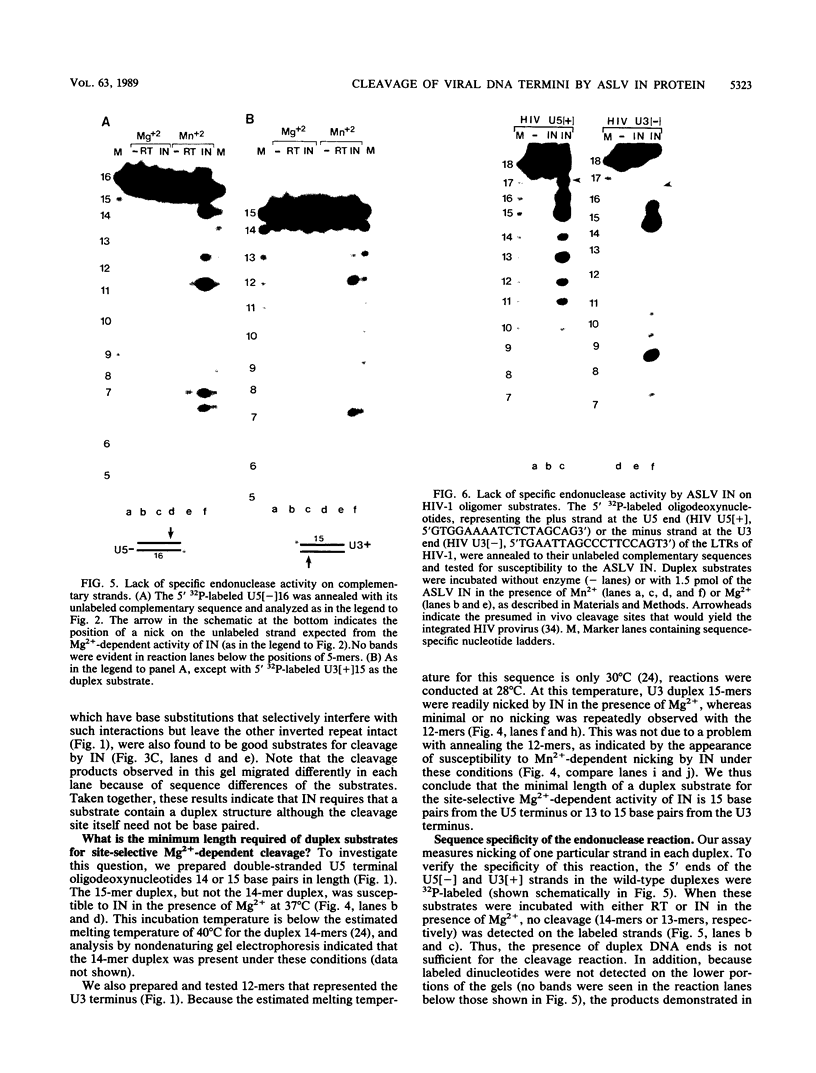
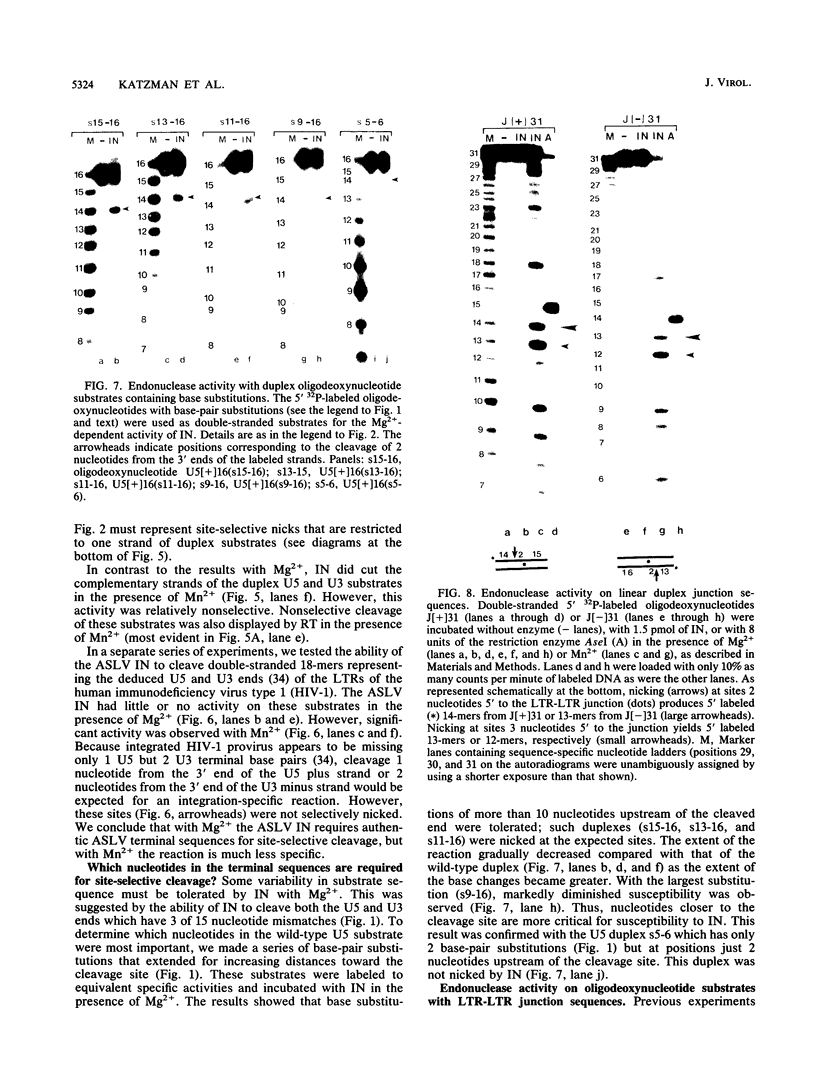



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Alexander F., Leis J., Soltis D. A., Crowl R. M., Danho W., Poonian M. S., Pan Y. C., Skalka A. M. Proteolytic processing of avian sarcoma and leukosis viruses pol-endo recombinant proteins reveals another pol gene domain. J Virol. 1987 Feb;61(2):534–542. doi: 10.1128/jvi.61.2.534-542.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brown P. O., Bowerman B., Varmus H. E., Bishop J. M. Correct integration of retroviral DNA in vitro. Cell. 1987 May 8;49(3):347–356. doi: 10.1016/0092-8674(87)90287-x. [DOI] [PubMed] [Google Scholar]
- Cobrinik D., Katz R., Terry R., Skalka A. M., Leis J. Avian sarcoma and leukosis virus pol-endonuclease recognition of the tandem long terminal repeat junction: minimum site required for cleavage is also required for viral growth. J Virol. 1987 Jun;61(6):1999–2008. doi: 10.1128/jvi.61.6.1999-2008.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donehower L. A. Analysis of mutant Moloney murine leukemia viruses containing linker insertion mutations in the 3' region of pol. J Virol. 1988 Nov;62(11):3958–3964. doi: 10.1128/jvi.62.11.3958-3964.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Donehower L. A., Varmus H. E. A mutant murine leukemia virus with a single missense codon in pol is defective in a function affecting integration. Proc Natl Acad Sci U S A. 1984 Oct;81(20):6461–6465. doi: 10.1073/pnas.81.20.6461. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duyk G., Leis J., Longiaru M., Skalka A. M. Selective cleavage in the avian retroviral long terminal repeat sequence by the endonuclease associated with the alpha beta form of avian reverse transcriptase. Proc Natl Acad Sci U S A. 1983 Nov;80(22):6745–6749. doi: 10.1073/pnas.80.22.6745. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Duyk G., Longiaru M., Cobrinik D., Kowal R., deHaseth P., Skalka A. M., Leis J. Circles with two tandem long terminal repeats are specifically cleaved by pol gene-associated endonuclease from avian sarcoma and leukosis viruses: nucleotide sequences required for site-specific cleavage. J Virol. 1985 Nov;56(2):589–599. doi: 10.1128/jvi.56.2.589-599.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujiwara T., Mizuuchi K. Retroviral DNA integration: structure of an integration intermediate. Cell. 1988 Aug 12;54(4):497–504. doi: 10.1016/0092-8674(88)90071-2. [DOI] [PubMed] [Google Scholar]
- Gibson W., Verma I. M. Studies on the reverse transcriptase of RNA tumor viruses. Structural relatedness of two subunits of avian RNA tumor viruses. Proc Natl Acad Sci U S A. 1974 Dec;71(12):4991–4994. doi: 10.1073/pnas.71.12.4991. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Golomb M., Grandgenett D. P. Endonuclease activity of purified RNA-directed DNA polymerase from avian myeloblastosis virus. J Biol Chem. 1979 Mar 10;254(5):1606–1613. [PubMed] [Google Scholar]
- Golomb M., Grandgenett D. P., Mason W. Virus-coded DNA endonuclease from avian retrovirus. J Virol. 1981 May;38(2):548–555. doi: 10.1128/jvi.38.2.548-555.1981. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandgenett D. P., Knaus R. J., Hippenmeyer P. J. Antibodies against a synthetic peptide of the avian retrovirus pp32 protein and the beta DNA polymerase subunit. Virology. 1983 Oct 15;130(1):257–262. doi: 10.1016/0042-6822(83)90137-x. [DOI] [PubMed] [Google Scholar]
- Grandgenett D. P., Vora A. C., Schiff R. D. A 32,000-dalton nucleic acid-binding protein from avian retravirus cores possesses DNA endonuclease activity. Virology. 1978 Aug;89(1):119–132. doi: 10.1016/0042-6822(78)90046-6. [DOI] [PubMed] [Google Scholar]
- Grandgenett D. P., Vora A. C. Site-specific nicking at the avian retrovirus LTR circle junction by the viral pp32 DNA endonuclease. Nucleic Acids Res. 1985 Sep 11;13(17):6205–6221. doi: 10.1093/nar/13.17.6205. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Grandgenett D. P., Vora A. C., Swanstrom R., Olsen J. C. Nuclease mechanism of the avian retrovirus pp32 endonuclease. J Virol. 1986 Jun;58(3):970–974. doi: 10.1128/jvi.58.3.970-974.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hishinuma F., DeBona P. J., Astrin S., Skalka A. M. Nucleotide sequence of acceptor site and termini of integrated avian endogenous provirus ev1: integration creates a 6 bp repeat of host DNA. Cell. 1981 Jan;23(1):155–164. doi: 10.1016/0092-8674(81)90280-4. [DOI] [PubMed] [Google Scholar]
- Hsu T. W., Sabran J. L., Mark G. E., Guntaka R. V., Taylor J. M. Analysis of unintegrated avian RNA tumor virus double-stranded DNA intermediates. J Virol. 1978 Dec;28(3):810–818. doi: 10.1128/jvi.28.3.810-818.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hughes S. H., Shank P. R., Spector D. H., Kung H. J., Bishop J. M., Varmus H. E., Vogt P. K., Breitman M. L. Proviruses of avian sarcoma virus are terminally redundant, co-extensive with unintegrated linear DNA and integrated at many sites. Cell. 1978 Dec;15(4):1397–1410. doi: 10.1016/0092-8674(78)90064-8. [DOI] [PubMed] [Google Scholar]
- Hurwitz J., Leis J. P. RNA-dependent DNA polymerase activity of RNA tumor viruses. I. Directing influence of DNA in the reaction. J Virol. 1972 Jan;9(1):116–129. doi: 10.1128/jvi.9.1.116-129.1972. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leis J. P. RNA-dependent DNA polymerase activity of RNA tumor virus. VI. Processive mode of action of avian myeloblastosis virus polymerase. J Virol. 1976 Sep;19(3):932–939. doi: 10.1128/jvi.19.3.932-939.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leis J., Baltimore D., Bishop J. M., Coffin J., Fleissner E., Goff S. P., Oroszlan S., Robinson H., Skalka A. M., Temin H. M. Standardized and simplified nomenclature for proteins common to all retroviruses. J Virol. 1988 May;62(5):1808–1809. doi: 10.1128/jvi.62.5.1808-1809.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leis J., Duyk G., Johnson S., Longiaru M., Skalka A. Mechanism of action of the endonuclease associated with the alpha beta and beta beta forms of avian RNA tumor virus reverse transcriptase. J Virol. 1983 Feb;45(2):727–739. doi: 10.1128/jvi.45.2.727-739.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Meinkoth J., Wahl G. Hybridization of nucleic acids immobilized on solid supports. Anal Biochem. 1984 May 1;138(2):267–284. doi: 10.1016/0003-2697(84)90808-x. [DOI] [PubMed] [Google Scholar]
- Panganiban A. T., Temin H. M. Circles with two tandem LTRs are precursors to integrated retrovirus DNA. Cell. 1984 Mar;36(3):673–679. doi: 10.1016/0092-8674(84)90347-7. [DOI] [PubMed] [Google Scholar]
- Panganiban A. T., Temin H. M. The retrovirus pol gene encodes a product required for DNA integration: identification of a retrovirus int locus. Proc Natl Acad Sci U S A. 1984 Dec;81(24):7885–7889. doi: 10.1073/pnas.81.24.7885. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Quinn T. P., Grandgenett D. P. Genetic evidence that the avian retrovirus DNA endonuclease domain of pol is necessary for viral integration. J Virol. 1988 Jul;62(7):2307–2312. doi: 10.1128/jvi.62.7.2307-2312.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rho H. M., Grandgenett D. P., Green M. Sequence relatedness between the subunits of avian myeloblastosis virus reverse transcriptase. J Biol Chem. 1975 Jul 10;250(13):5278–5280. [PubMed] [Google Scholar]
- Sabran J. L., Hsu T. W., Yeater C., Kaji A., Mason W. S., Taylor J. M. Analysis of integrated avian RNA tumor virus DNA in transformed chicken, duck and quail fibroblasts. J Virol. 1979 Jan;29(1):170–178. doi: 10.1128/jvi.29.1.170-178.1979. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiff R. D., Grandgenett D. P. Partial phosphorylation in vivo of the avian retrovirus pp32 DNA endonuclease. J Virol. 1980 Dec;36(3):889–893. doi: 10.1128/jvi.36.3.889-893.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schiff R. D., Grandgenett D. P. Virus-coded origin of a 32,000-dalton protein from avian retrovirus cores: structural relatedness of p32 and the beta polypeptide of the avian retrovirus DNA polymerase. J Virol. 1978 Oct;28(1):279–291. doi: 10.1128/jvi.28.1.279-291.1978. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartzberg P., Colicelli J., Goff S. P. Construction and analysis of deletion mutations in the pol gene of Moloney murine leukemia virus: a new viral function required for productive infection. Cell. 1984 Jul;37(3):1043–1052. doi: 10.1016/0092-8674(84)90439-2. [DOI] [PubMed] [Google Scholar]
- Shank P. R., Hughes S. H., Kung H. J., Majors J. E., Quintrell N., Guntaka R. V., Bishop J. M., Varmus H. E. Mapping unintegrated avian sarcoma virus DNA: termini of linear DNA bear 300 nucleotides present once or twice in two species of circular DNA. Cell. 1978 Dec;15(4):1383–1395. doi: 10.1016/0092-8674(78)90063-6. [DOI] [PubMed] [Google Scholar]
- Terry R., Soltis D. A., Katzman M., Cobrinik D., Leis J., Skalka A. M. Properties of avian sarcoma-leukosis virus pp32-related pol-endonucleases produced in Escherichia coli. J Virol. 1988 Jul;62(7):2358–2365. doi: 10.1128/jvi.62.7.2358-2365.1988. [DOI] [PMC free article] [PubMed] [Google Scholar]