Abstract
The use of antisense oligodeoxynucleotides (ODNs) to inhibit the expression of specific mRNA targets represents a powerful technology for control of gene expression. Cationic lipids and polymers are frequently used to improve the delivery of ODNs to cells, but the resulting complexes often aggregate, bind to serum components, and are trafficked poorly within cells. We show that the addition of a synthetic, pH-sensitive, membrane-disrupting polyanion, poly(propylacrylic acid) (PPAA), improves the in vitro efficiency of the cationic lipid, DOTAP, with regard to oligonucleotide delivery and antisense activity. In characterization studies, ODN complexation with DOTAP/ODN was maintained even when substantial amounts of PPAA were added. The formulation also exhibited partial protection of phosphodiester oligonucleotides against enzymatic digestion. In Chinese hamster ovary (CHO) cells, incorporation of PPAA in DOTAP/ODN complexes improved two- to threefold the cellular uptake of fluorescently tagged oligonucleotides. DOTAP/ODN complexes containing PPAA also maintained high levels of uptake into cells upon exposure to serum. Addition of PPAA to DOTAP/ODN complexes enhanced the antisense activity (using GFP as the target) over a range of PPAA concentrations in both serum-free, and to a lesser extent, serum-containing media. Thus, PPAA is a useful adjunct that improves the lipid-mediated delivery of oligonucleotides.
Introduction
Since its discovery nearly 30 years ago1, antisense technology has been used extensively as a research tool for sequence-specific gene silencing in elucidation of gene function and in drug target validation2, and it is currently being developed towards clinical application as well3,4. In theory, the concept of antisense technology is elegant. Short synthetic oligodeoxynucleotides (ODNs) bind selectively to the complementary mRNA transcript of a targeted gene via Watson-Crick hybridization, thereby inhibiting mRNA translation into the corresponding protein through any of several mechanisms, such as translational arrest or mRNA cleavage by ribonuclease H5. However, the practical application of antisense oligonucleotides has proven challenging, largely owing to their susceptibility to degradation by cellular nucleases and the difficulty in delivering efficacious and safe doses. To enhance their stability against nucleolytic degradation, as well as to increase their affinity for the target and decrease their toxicity, antisense oligonucleotides with various chemical modifications have been synthesized. These include among others phosphorothioate ODNs, methylphosphonates and phosphoramidate ODNs, 2′-O-modified RNA, locked nucleic acids and peptide nucleic acids3,6,7. Another longstanding problem exists in the delivery of antisense oligonucleotides to cells and tissues. Various nonviral oligonucleotide delivery systems have been developed to address several criteria such as improved stability under physiological conditions, efficient internalization by targeted cells, resistance to intracellular nucleases, and reduced nonspecific interactions with other macromolecules. Despite considerable research in this area, efficiency and specificity of delivery are not yet satisfactory in many cases8.
The core component of most oligonucleotide delivery systems is a synthetic polymer or a lipid (or lipid mixture). Commonly used for delivery of plasmids carrying therapeutic genes, cationic lipids are also promising reagents for oligonucleotide delivery.9,10 Cationic lipid/DNA complexes are usually formulated using an excess of positive charge (contributed by the lipid component), thus conferring upon the resulting particle more efficient interaction with the negatively charged cell membrane. Following cellular internalization, endosomes become acidified and progress to lysosomes, where their contents are degraded. Crucial, then, to the success of antisense downregulation is the efficient release of oligonucleotides from this endosomal-lysosomal pathway into the cytoplasm11. In view of this, some synthetic vectors are formulated with endosomal escape moieties (pH-sensitive chemical groups) that promote membrane disruption and release of DNA from the endosome into the cytoplasm. One approach is the incorporation of a neutral phospholipid, such as dioleylphosphatidylethanolamine (DOPE), which undergoes bilayer destabilization at the lower pH (5 to 6) of the endocytic compartment, fuses with the endosomal membrane and destabilizes the endosome, thereby releasing the DNA cargo into the cytoplasm.12 Synthetic, pH-responsive, amphipathic peptides such as GALA and JTS-1 have been designed as alternatives to the helper lipid approach13,14.
Prior studies have shown that the binding of negatively charged serum components to positively charged DNA complexes can significantly decrease transfection efficiency15,16. In order to decrease or reverse the overall particle charge and hence reduce serum opsonization, anionic polymers have been employed as an additional component of the delivery vector17-19. Previously, it was shown that a synthetic anionic polymer, poly(propylacrylic acid) (PPAA), was able to enhance the intracellular delivery of plasmid DNA by the commercially available cationic liposome, DOTAP, even in the presence of up to 50% serum. The presence of PPAA also increased the transfection of plasmids encoding antisense RNA in a mouse model20. While improved delivery of plasmids has been demonstrated by combining PPAA with DOTAP, this system has not been studied previously with respect to oligonucleotide delivery. Despite the considerable disparity in the molecular weights and topologies of plasmid DNA and oligonucleotides, each type of DNA interacts electrostatically with DOTAP14,15. We hypothesized that incorporating PPAA in the formulation would provide a means to improve the cationic liposome-mediated delivery of oligonucleotides. To address this question, we report the physicochemical characteristics of DOTAP/ODN complexes incorporating graded PPAA loadings (i.e. giving rise to various charge ratios), their cellular uptake by CHO cells, and their antisense inhibition in a model system consisting of CHO cells stably expressing pd1EGFP.
Experimental section
Materials
A phosphodiester oligonucleotide (5′-ATT GCA AGT AAA CAC AG-3′, RBG33)21 was used for preliminary physicochemical and stability studies. An oligonucleotide of identical sequence but fluorescently tagged with carboxyfluorescein (F-RBG33) was used for the cellular uptake studies. The 20mer phosphorothioate oligonucleotide (5′-TTG TGG CCG TTT ACG TCG CC-3′, EGFP157), previously selected for downregulation of EGFP (Dunham et al., manuscript in preparation), was used for the antisense effect studies. All ODNs were obtained from Integrated DNA Technologies (Coralville, IA) and delivered as HPLC grade. Before use, the lyophilized ODNs were resuspended to 1 mg/ml (phosphodiester ODNs) or 100 μM (phosphorothioate ODNs) concentration in TE buffer (10 mM Tris-HCl, 1 mM EDTA, pH 8.0). The cationic liposomal formulation N-[1-(2,3-dioleyloxy)propyl]-N,N,N-trimethylammonium methyl sulfate (DOTAP) was purchased from Roche Applied Science (Indianapolis, IN). Poly(α-propylacrylic acid) (PPAA) with 27,000 number average MW and polydispersity (Mw/Mn) of 1.8 was obtained from Polymer Source Inc. (Dorval, Quebec, Canada). The polymer was received as a dried powder and solubilized in a small quantity of tetrahydrofuran before further dilution (approx. 10-fold) to a concentration of 10 mg/ml in HEPES buffered saline (HBS; 20 mM HEPES, 150 mM NaCl, pH 7.4). All other reagents and solvents were purchased from Sigma-Aldrich (St. Louis, MO) and were used as received, unless noted otherwise. All buffers were prepared in MilliQ ultrapure water and filtered (0.22 μm) prior to use.
Vector preparation
The complexes were formed in HBS at a final DNA concentration of 1 μg/ 120μl for physicochemical assays and cellular uptake studies. Antisense experiments using CHO-pd1EGFP cells were performed at an ODN concentration of 300 nM. The components of the ternary complexes — DOTAP (D), PPAA (P) and ODN (O) — were prepared at a DOTAP:ODN (D:O) weight ratio of 10:1, corresponding to D:O charge ratios (+/-) of approximately 4. All DOTAP/PPAA/ODN complexes were formed by first pipette mixing together equal volumes of DOTAP and ODN followed by incubating for 30 minutes. PPAA was then added to the DOTAP/ODN solution, at concentrations producing the desired charge ratios. DOTAP and DNA were assumed to be completely ionized, and the share of negative charge due to the PPAA addition was calculated assuming 33% PPAA ionization at pH 7.420,22. DOTAP/ODN complexes were formed by mixing equal volumes of DOTAP and ODN, the former at appropriate concentrations to produce the desired charge ratios. All complexes were prepared in 1.5-ml Eppendorf tubes immediately before analysis or transfection.
OliGreen assay for ODN in complexes
Complexes (1 μg ODN per 120 μl) were formulated in HBS as previously described. The single-stranded DNA-binding reagent OliGreen (Invitrogen-Molecular Probes, Carlsbad, CA) was diluted 1:50 with HBS. The OliGreen working solution (60 μl) was added to 60 μl of D/P/O or D/O complex solution in a polystyrene 96-well black plate with clear bottoms (Corning, Corning, NY). The OliGreen fluorescence intensity was measured at excitation and emission wavelengths of 485 and 530 nm, respectively, with a Cytofluor 4000 Fluorescence Multi-well Plate Reader (Applied Biosystems, Foster City, CA). After background subtraction, each data point was normalized to that from uncomplexed ODN.
Zeta potential
The electrophoretic mobility of the DOTAP/PPAA/ODN and DOTAP/ODN complexes was determined by phase analysis light scattering using a Brookhaven ZetaPALS analyzer (Brookhaven Instruments Corp., Holtsville, NY). The phase shift is measured by comparing the phase of the light scattered by the particles with the phase of a reference beam. The software then computes the zeta potential (ζ) from the measured electrophoretic mobility using the Smoluchowski equation23. Complexes (120 μl) were diluted with 1.5 ml HBS in a standard 1 cm light path polystyrene cuvette, and a parallel plate electrode was inserted. The cuvette was placed in a temperature-controlled holder, after which measurements were performed ≥ 5 times at 25°C within 30 minutes of complex preparation and averaged for each sample.
Cell culture
The CHO-K1 cell line (ATCC CCL-61) was maintained in F-12K medium (Kaighn’s modification of Ham’s F-12; ATCC, Manassas, VA) supplemented with 10% fetal bovine serum (FBS), 100 units/mL penicillin and 100 mg/mL streptomycin. A CHO-K1 cell line containing a stably integrated destabilized EGFP (d1EGFP) transgene was produced by transfecting CHO cells with the 4.9-kb pd1EGFP-N1 plasmid (BD Biosciences Clontech, Palo Alto, CA, USA), and maintained under constant selective pressure by G418 (500 μg/ml) (Dunham et al., manuscript in preparation). All cell lines were cultivated in a humidified atmosphere of 5% CO2 at 37°C.
Transfection
CHO-K1 or CHO-pd1EGFP epithelial cells were plated at a density of 5 × 104 cells/well in 24-well plates ∼18 h prior to transfection. Cells were transfected with 1 μg of F-ODN or 0.9 μg (∼300 nM final concentration) anti-GFP ODN per well for the cellular uptake or antisense experiments, respectively. Prior to transfection, 380 μl of Opti-MEM reduced serum medium (Invitrogen) or medium containing 39% FBS, for experiments performed in the absence or presence of 30% serum, respectively, was added to each well. The transfection mixtures, prepared in 120 μl of HBS, were then added to the wells, and the cells were incubated with the complexes for 4 h. Thereafter, the cells were assayed by flow cytometry for F-ODN uptake, or in the case of the antisense experiment, the transfection medium replaced with normal growth medium containing 10% FBS and the cells cultured for a further 20 h before being assayed for EGFP activity.
Flow cytometry
Replicate wells of cells were washed in phosphate buffered saline (PBS; Invitrogen), detached with trypsin-EDTA (Invitrogen), washed with serum-containing medium, pelleted by centrifugation for 5 min at 200 g, washed with PBS, resuspended in 500 μl of PBS containing 1 μg/ml 7-aminoactinomycin D (7AAD; Sigma-Aldrich) and maintained on ice before being subjected to flow cytometry analysis. The DNA-intercalating agent 7AAD was used for excluding nonviable cells from the analysis. Mean cellular fluorescence intensities for 10,000 cells were determined on a FACSCalibur three-laser flow cytometer (BD Biosciences). CellQuest software was used to acquire and analyze the results. Viable cells were gated according to their typical forward/side scatter characteristics and 7AAD staining. The flow cytometer was calibrated with fluorescent beads (CaliBRITE Beads, BD Biosciences) prior to each use to ensure comparable readings over the period of the study.
Confocal fluorescent microscopy
Complexes were formulated as previously described but using an anti-GFP ODN labeled on the 5′ end with the fluorophore Cy5 (Integrated DNA Technologies). For the uptake study, 250,000 CHO-pd1EGFP cells were seeded onto 35-mm tissue culture dishes (MatTek Corporation) in 2 ml complete growth medium (containing 500 μg/ml G418) and grown overnight. DOTAP/ODN (D:O weight ratio = 10:1; charge ratio = 4) and DOTAP/PPAA/ODN (charge ratio 0.025) containing Cy5 anti-GFP ODN as well as naked Cy5 anti-GFP ODN were placed on the cells in serum-free media (Opti-MEM) at 300 nM final concentration of the ODN. After a 4 h incubation the cells were washed twice with PBS and incubated with complete growth for another 20 h. GFP fluorescent cells were then visualized at 63X magnification, 488 nm excitation and 500-555 nm emission using a Leica TCS SP2 microscope (Leica Microsystems, Inc., Exton, PA). Cy5 fluorescence was excited at 633 nm and light emission collected at 650-750 nm.
Results
DNA accessibility in complexes
The interaction of DOTAP/ODN with PPAA was analyzed using OliGreen, a dye that fluoresces upon binding to single-stranded DNA (closed symbols in Figure 1)24. The molar charge ratio represents the ratio of positive charge contributed by the cationic lipid component (DOTAP) to the sum of the negative charges from the DNA and PPAA components; thus, increasing amounts of PPAA in the ternary complex correspond to decreasing charge ratios (from left to right, on the x-axis). The highest charge ratio, approximately 4, corresponds to DOTAP and ODN only without PPAA. At this ratio, roughly 25% of the fluorescence remained, indicative of ODN that is complexed by the DOTAP but still accessible to dye. We have observed comparable patterns of dye binding for ODN complexation with the cationic polymer, poly-l-lysine24. Addition of increasing concentrations of PPAA caused an increase in the fluorescence of the solution, indicating increased availability of the ODNs for dye binding. The effect of added PPAA was relatively minor until the charge ratio became less than approximately 0.1, at which point the fluorescence increased significantly above the baseline. Negligible fluorescence, comparable to background, appeared in samples containing only DOTAP or DOTAP/PPAA (data not shown).
Figure 1.
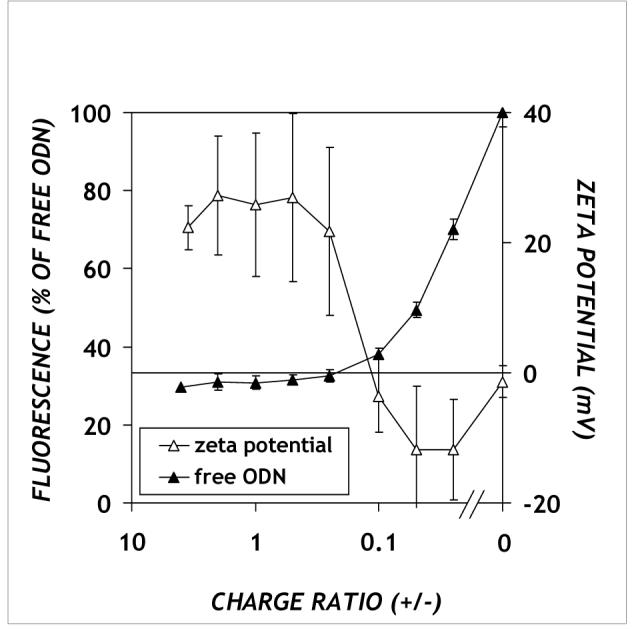
DNA accessibility (closed symbols) and zeta potentials (open symbols) of DOTAP/PPAA/ODN complexes. The complexes were prepared as described in Materials and Methods. Means of three (fluorescence) or two (zeta potential) separate experiments ± SEM are shown.
The electrokinetic properties of the DOTAP/PPAA/ODN complexes formed at varying ratios of constituents were examined using phase analysis light scattering (open symbols in Figure 1). The zeta potential of DOTAP alone, at a concentration corresponding to that in DOTAP/PPAA/ODN complexes, was found to be 28.3 ± 2.6 mV (data not shown). The zeta potential of DOTAP/PPAA/ODN complexes is positive for charge ratios greater than 2 (only that for a charge ratio of 4 is shown), which is roughly the point at which the DOTAP concentration exceeds its critical aggregation concentration (c.a.c.) of 70 μM25. Similar to the OliGreen assay, the effect of added PPAA was relatively minor until the charge ratio became less than approximately 0.1, at which point the zeta potential exhibited a transition from positive to negative values (open symbols in Figure 1).
One role of vectors used for oligonucleotide delivery is to protect the DNA from digestion by nucleases. Hence, we evaluated the effect of PPAA on ODN recovery and integrity in serum-free and serum-containing media. Samples of ODN, DOTAP/ODN (charge ratio 4.0) and DOTAP/PPAA/ODN (charge ratio 0.025) were incubated at 37°C with media containing 0% or 10% fetal bovine serum, and the ODN recovery measured over 24 h. Under serum-containing conditions, the ODN was nearly completely digested, regardless of whether they were complexed by DOTAP or DOTAP/PPAA (results not shown). Since, under the conditions studied, neither DOTAP nor DOTAP/PPAA aids in protection against serum, any beneficial effects of these compounds are due to other effects. No degradation occurred and significant recovery of ODN was observed under serum-free conditions (results not shown).
Enhancement of DOTAP/F-ODN uptake into cells by PPAA
Previously, it was shown that PPAA enhances drug and gene delivery mediated by DOTAP20,26. To determine the effect of PPAA on the cellular uptake levels of fluorescently labeled oligonucleotides complexed with DOTAP, we followed the cell-associated fluorescence in Chinese hamster ovary (CHO) cells (Figure 2). The delivery experiments were performed both in the absence and in the presence of 30% serum, a level intermediate between standard cell culture (10%) and the work of Kyriakides et al.20, who investigated plasmid transfection in serum levels up to 50%. The cells were treated with DOTAP/PPAA/F-ODN complexes for 4 h, after which they were analyzed by flow cytometry. The negative control, comprising complexes containing the untagged ODN previously used in the physicochemical studies, exhibited negligible background fluorescence, similar to that of cells treated with vehicle buffer (data not shown). When F-ODNs were added alone in culture medium, no uptake was observed. In contrast, complexation of F-ODN to DOTAP led to increased cellular fluorescence intensity in both serum-free and serum-containing media (Figures 2A and B, respectively). We then decreased the charge ratio by adding increasing amounts of PPAA to a fixed amount of DOTAP/F-ODN (1 μg ODN and 10 μg DOTAP per well in 24-well plates, corresponding to an initial charge ratio of 4). Increasing PPAA concentrations resulted in an increase of cellular uptake. The trends were fairly similar under serum-free and serum-containing conditions. However, significantly greater uptake was observed in serum-free medium, as shown by the approximately 10-fold mean cell fluorescence compared to those in serum-containing medium (see raw mean fluorescence intensities).
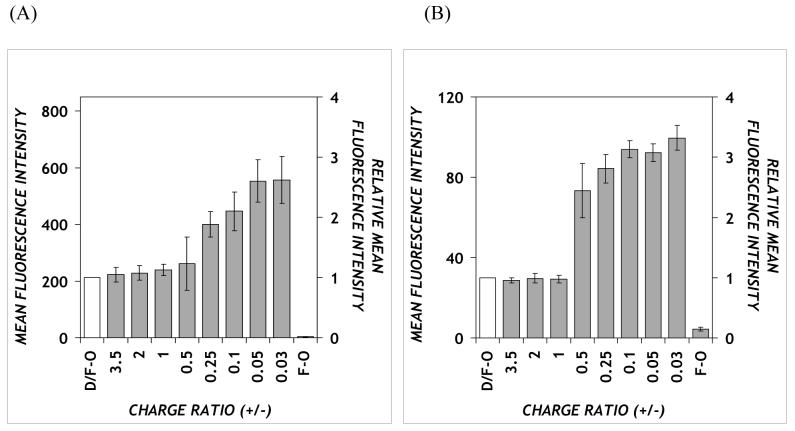
Figure 2.
Cellular uptake of DOTAP/PPAA/F-ODN in the (A) absence or (B) presence of 30% serum. DOTAP/PPAA/ODN complexes containing a FAM-labeled ODN were presented to CHO-K1 cells in: (A) Opti-MEM or (B) complete growth medium (supplemented with 30% serum and antibiotics). Secondary y-axis values represent the geometric mean fluorescence intensities normalized to DOTAP/ODN (denoted D/F-O in the axis labels). The negative control is fluorescent ODN without carrier (F-O). Error bars represent the SEM (n = 3) for each group.
Inhibition of d1EGFP expression by DOTAP/PPAA/ODN complexes
Recent studies on the mechanism by which cationic liposomes deliver antisense oligonucleotides into cells have suggested that the ability of complexes to destabilize the endosomal membrane and release their DNA cargo into the cytoplasm is critical for achieving antisense inhibition8,27. We hypothesized that, due to its pH-dependent membrane destabilization ability26,28, PPAA addition to DOTAP/ODN complexes would result in improved antisense activity. To evaluate the biological activity of antisense oligonucleotides, we employed a model cellular system consisting of a clonal CHO-K1 cell line stably transfected with destabilized EGFP. A synthetic phosphorothioate AS ODN, previously determined as effective in our laboratory (Dunham et al., manuscript in preparation), was directed against the pd1EGFP mRNA. The short (1 h) intracellular half-life of the protein encoded by pd1EGFP provides a close coupling between silencing at the mRNA level and downregulation of protein levels. Furthermore, this system allows the extent of target downregulation to be quantitatively assessed by flow cytometry.
The inhibition of d1EGFP following exposure of pd1EGFP-CHO cells to complexes of DOTAP/PPAA/ODN at various charge ratios for 4 h was investigated in the absence (Figure 3A) or presence (Figure 3B) of 10% serum. A non-complementary phosphorothioate ODN served as a sequence control. All data, shown as the geometric mean fluorescence intensity of the cells, were normalized to GFP levels of control cells (i.e., CHO-pd1EGFP treated with HBS vehicle). Under serum-free conditions, addition of PPAA to the AS ODN complexed with DOTAP enabled significant downregulation levels, in contrast with quite modest inhibition (∼30%) for the DOTAP/ODN formulation (in the absence and presence of serum, respectively). Large amounts of PPAA (corresponding to the lowest net charge ratios) led to strikingly high effectiveness, with approximately 90% downregulation exhibited by cells treated with DOTAP/PPAA/ODN complexes at the lowest tested charge ratio of 0.025 (Figure 3A). Control treatments of ternary complexes, where the DOTAP or PPAA component was replaced by vehicle buffer or buffer containing a small amount of tetrahydrofuran (equivalent to that in which the PPAA stock was formulated), did not differ from the blank or DOTAP/ODN equivalent, respectively (data not shown). Very few cells were excluded by the 7-AAD analysis, indicating that the relatively short treatment time (4h) prevented excessive toxicity effects from compromising the results.
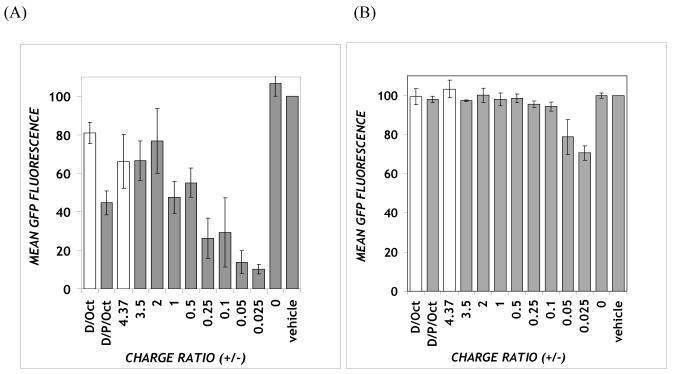
Figure 3.
Stably pd1EGFP-expressing CHO cells were treated with DOTAP/PPAA/ODN complexes containing 300 nM ODN under: (A) serum-free or (B) 10% serum conditions. The RBG33 ODN was used as a sequence control (ct) and delivered using DOTAP/PPAA/ODN and DOTAP/ODN at the lowest and highest charge ratios, respectively. GFP fluorescence values were normalized to untreated control cells (100%) and are expressed as geometric mean ± SEM (n = 3 for A; n = 4 for B).
When serum was present in the medium, the effectiveness of the ODNs was greatly reduced under all conditions. Without PPAA, DOTAP/ODN was completely ineffective at inhibiting pd1EGFP expression. Large amounts of PPAA (net charge ratios ≤ 0.05) were required to produce downregulation levels of up to 30%. Taken together, the cellular uptake and antisense activity assays suggest that PPAA improves the efficiency of DOTAP/ODN delivery.
A common complication in the use of phosphorothioate oligonucleotides for antisense experiments is the presence of nonspecific effects on irrelevant genes29. Indeed, in our studies a nonspecific AS ODN sequence (control: D/P/O ct in Figure 3A) was able to mediate partial GFP downregulation, which also suggests the possibility that nonspecific inhibition of other cellular proteins could also take place. It is generally believed that more efficient delivery systems will result in lower ODN doses and thus fewer nonspecific effects30,31. Thus, further optimization of this system to enhance efficiency may also enhance specificity.
Endosomal escape and nuclear accumulation of DOTAP/PPAA/ODN complexes
To determine the effect of PPAA on the intracellular fate of DOTAP/ODN complexes, fluorescently labeled ODNs and pd1EGFP were monitored by confocal microscopy. The stably expressing CHO-pd1EGFP cells were treated with complexes containing anti-GFP ODNs that were 5′-end labeled with a Cy5 (blue) fluorescent tag and incorporated into DOTAP/PPAA/ODN complexes at a charge ratio of 0.025. Compared to cells treated with vehicle buffer only (Figure 4A), those treated with ODN in the absence of vehicle exhibited a low level of punctuate ODN fluorescence and no decrease in GFP fluorescence (Figure 4B). Delivery of ODN using DOTAP without PPAA resulted in higher levels of ODN fluorescence, but the intracellular distribution remained punctate and only modest inhibition of GFP fluorescence was observed (Figure 4C). In stark contrast, visibly significant silencing of GFP expression was detected in cells treated with DOTAP/PPAA/ODN complexes (Figure 4D). Furthermore, the ODN fluorescence distribution in these cells is considerably more uniform, encompassing the cytoplasm and the nucleus, indicative of PPAA-mediated release of the Cy5-ODN from DOTAP and endosomes. These results corroborate and provide a mechanistic basis for the flow cytometric analyses showing the effect of PPAA on DOTAP/ODN complexes.
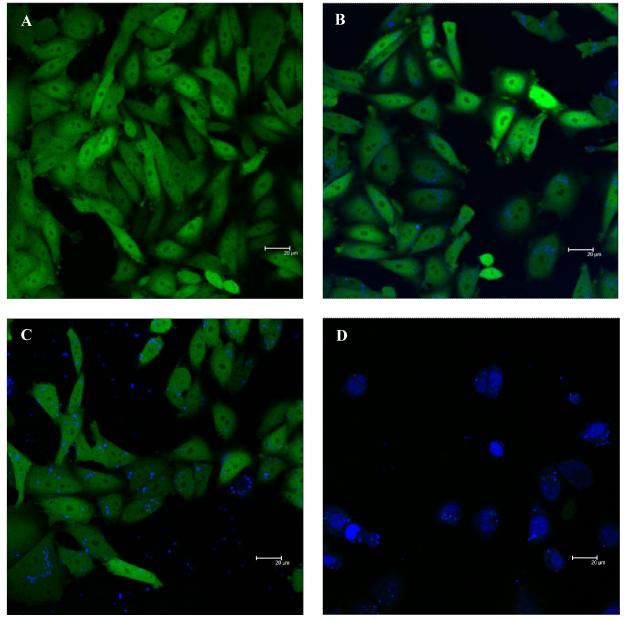
Figure 4.
Confocal fluorescence image of CHO-pd1EGFP cells transfected with: (A) vehicle buffer, (B) ODN only, (C) DOTAP/ODN and (D) DOTAP/PPAA/ODN at a charge ratio of 0.025. Cells were transfected with 300 nM final concentration of Cy5 (blue) labeled ODNs for 4 h, after which they were washed and used for microscopy.
Discussion
Most gene and oligonucleotide delivery systems utilize endocytotic pathways for entry11. Thus, for efficient antisense activity to occur in the cytosol or nucleus, oligonucleotides must escape from endolysosomal compartments before lysosomal degradation occurs. Poly(propylacrylic acid) (PPAA), a synthetic anionic polymer that exhibits membrane disruption over a narrow pH range, is known to enhance transfection for both cationic lipid- and polymer-based gene delivery20,32-34,35. In view of this, we have employed PPAA for antisense oligonucleotide delivery. The transfection reagent DOTAP, by itself and in combination with other lipids or polymers, has been employed previously for antisense ODN delivery14,15,36. Our study represents the first attempt to apply the DOTAP/PPAA/DNA formulation to antisense oligonucleotides. Possible ways in which PPAA might enhance ODN delivery are altering the stability of the complexes, protection of the ODNs from extracellular and intracellular nucleases, and/or destabilization of endosomal membranes.
A possible explanation for the increase in ODN uptake and antisense activity is stabilization of DOTAP/ODN complexes with respect to serum and self-aggregation. The DOTAP/PPAA/ODN formulation entailed pre-complexation of ODN with DOTAP, followed by the addition of PPAA polymer. The extent of ODN complexation by DOTAP, prior to PPAA addition, was found to be a function of DOTAP concentration, with maximal complexation requiring a DOTAP concentration exceeding its c.a.c. of 70 μM25. No significant change was observed in the particle size, as determined by light scattering, when PPAA was present (results not shown), suggesting that PPAA did not exert its effects on ODN delivery by aggregating the DOTAP/ODN complexes. Interestingly, PPAA also enhanced intracellular release and antisense effectiveness when DOTAP/ODN/PPAA complexes were formulated at DOTAP concentration below the c.a.c. (results not shown); however, the overall levels of uptake and effectiveness were less than when DOTAP was used at levels above the c.a.c.
Previous work has shown that serum does not dissociate ODNs from DOTAP lipids but rather inhibits cellular uptake15. Reducing the cationic charge of DOTAP/ODN complexes by inclusion of neutral lipids, such as DOPE or cholesterol, enhances cellular uptake in the presence of serum15. Moreover, an anionic polymer, poly(acrylic acid), which does not possess any membrane disruptive activity, enhanced the gene transfection efficiency of DOTAP/DNA complexes in a manner that included avoidance of serum inhibition18. Thus, it is likely that the addition of PPAA in our system exerted at least some of its effect by lowering vector interactions with serum and/or extracellular matrix components via the change in electrostatic properties of the DOTAP/ODN/PPAA complexes. It may be possible to exploit this mechanism further and improve the efficiency of our formulation by the incorporation of altered or additional amphipathic moieties.
Another potential mechanism of PPAA enhancement is protection of ODNs from rapid degradation in the intra- and extracellular milieu, which is a major obstacle limiting the success of antisense therapeutics, especially in vivo. The addition of PPAA at levels leading to enhanced intracellular release (Figure 3) also causes increased accessibility of ODNs to OliGreen. The exact mechanism for this behavior is not clear, but the fact that particles are still detected by light scattering suggests that the lipoplexes are not completely dissociated, but rather may be dynamically altered due to adsorption of the PPAA to the cationic lipid. Although DOTAP has been reported to enhance nuclease resistance of ODNs 37, neither DOTAP/ODN nor DOTAP/PPAA/ODN complexes exhibited appreciable differences in degradation rates in serum-containing medium in vitro (results not shown); i.e., in all cases substantial degradation of ODN occurred after 2 hours exposure to serum-containing medium. However, this is sufficient time for cellular entry and thus not likely to be rate-limiting in our experiments38.
ODNs delivered with DOTAP typically reside in vesicular structures (endosomes) with a relatively low rate of escape39. The unique characteristic of PPAA is its ability to disrupt membranes in a pH-dependent fashion. As the pH drops from the extracellular value of 7.4, protonation of the carboxylate anion form of PPAA increases, neutralizing the charge on the polymer. In the uncharged state, the hydrophobicity of PPAA is enhanced, lending it the ability to disrupt biological membranes. In hemolysis experiments, PPAA exhibited maximal membrane disruption activity at pH 6, which corresponds to the pH of early endosomes.26 In our work, complexes that did not contain PPAA demonstrated little or no antisense activity against pd1EGFP (Figures 3 and 4). This result is consistent with the failure of the ODN to move into the cytoplasm, a view that is supported by confocal microscopy images of the distribution of ODN throughout the CHO cells when PPAA was included, in contrast to those localized in punctuate, possibly endolysosomal, structures when it was not (Figure 4). This effect on distribution is similar to that observed for chitosan/plasmid/PPAA complexes delivered to HeLa cells 34.
If cytosolic delivery of antisense ODNs is indeed the limiting step in activity, we would expect a correlation between the antisense activity and cellular uptake. We find that these quantities are positively correlated for DOTAP/PPAA/ODN complexes, with a characteristic saturating dose-response relationship (Figure 5). As with OliGreen®, the fluorescence of labeled ODNs is masked by complexation, and thus the fluorescence uptake measured in our assays should be interpreted as intracellular ODN that is released from complexes. Since lipids including DOTAP are known to be left behind when ODNs are released from endosomes40, the measured uptake likely consists primarily of cytoplasmic ODN, although contributions from uncomplexed ODN in other compartments cannot be ruled out.
Figure 5.
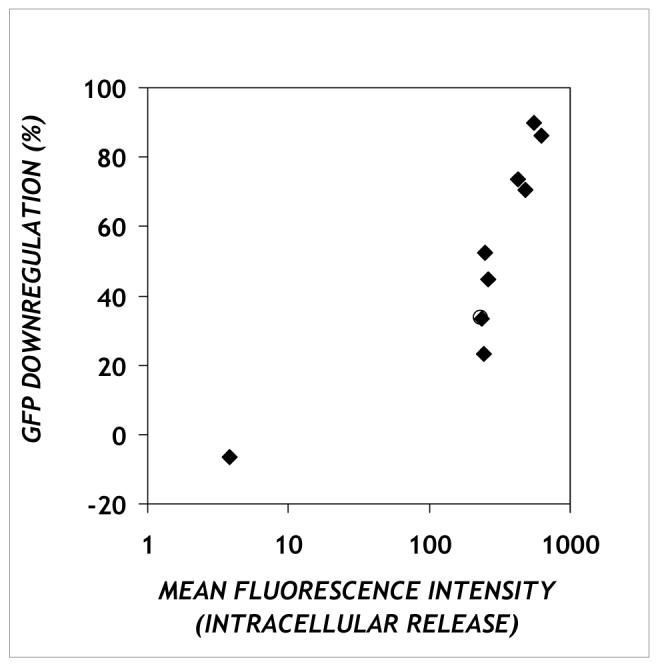
The relationship between GFP downregulation and cellular uptake. Each point represents one charge ratio at which the percentage downregulation of GFP by complexes containing an antisense oligonucleotide against pd1EGFP (y-axis) and geometric mean fluorescence value of complexes containing a fluorescent oligonucleotide (x-axis) were measured. The data refer to experiments performed under serum-free conditions.
In summary, we have investigated a formulation which has the capability of overcoming one of the major challenges associated with antisense oligonucleotide delivery, namely the release of endocytosed DNA complexes into the cytoplasm of the cell. As with previous studies investigating the use of PPAA in complexes of DOTAP and plasmid DNA20, we found enhanced intracellular delivery of oligonucleotides in the absence and presence of serum. Although the cellular antisense activity in the presence of serum was rather low, the favorable intracellular delivery and activity of the DOTAP/PPAA/ODN formulation make it a promising basic formulation for antisense oligonucleotide delivery. The type and amount of lipid are variables that may be optimized to improve the level of activity in the system employed here or in transferring this concept to other systems. Furthermore, PPAA may prove useful for enhancing delivery and facilitating endosomal escape of other ODN delivery systems.
Acknowledgments
We thank Dr. Patrick Johnson and Prof. Prabhas Moghe for access to the CytoFluor and confocal microscopy collaborative resources under the auspices of NIH Grant P41 EB001046 for the biomedical technology resource RESBIO; and Sumati Sundaram, Dr. Nicole Harris and Dr. Jaap Schut for helpful discussions. Support for this work was provided by NIH Grant R01 GM65913 and by The Petroleum Research Fund, administered by the American Chemical Society.
References
- (1).Zamecnik PC, Stephenson ML. Proceedings of the National Academy of Science USA. 1978;75:280–284. doi: 10.1073/pnas.75.1.280. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (2).Taylor MF, Wiederholt K, Sverdrup F. Drug Discov Today. 1999;4:562–567. doi: 10.1016/s1359-6446(99)01392-6. [DOI] [PubMed] [Google Scholar]
- (3).Crooke ST. Annu Rev Med. 2004;55:61–95. doi: 10.1146/annurev.med.55.091902.104408. [DOI] [PubMed] [Google Scholar]
- (4).Lee LK, Roth CM. Curr Opin Biotechnol. 2003;14:505–511. doi: 10.1016/s0958-1669(03)00115-0. [DOI] [PubMed] [Google Scholar]
- (5).Opalinska JB, Gewirtz AM. Nat Rev Drug Discov. 2002;1:503–514. doi: 10.1038/nrd837. [DOI] [PubMed] [Google Scholar]
- (6).Crooke ST. Curr Mol Med. 2004;4:465–487. doi: 10.2174/1566524043360375. [DOI] [PubMed] [Google Scholar]
- (7).Grunweller A, Wyszko E, Bieber B, Jahnel R, Erdmann VA, Kurreck J. Nucleic Acids Res. 2003;31:3185–3193. doi: 10.1093/nar/gkg409. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (8).Shi F, Hoekstra D. J Control Release. 2004;97:189–209. doi: 10.1016/j.jconrel.2004.03.016. [DOI] [PubMed] [Google Scholar]
- (9).Dass CR. J Pharm Pharmacol. 2002;54:3–27. doi: 10.1211/0022357021771887. [DOI] [PubMed] [Google Scholar]
- (10).Fenske DB, Cullis PR. Methods Enzymol. 2005;391:7–40. doi: 10.1016/S0076-6879(05)91001-X. [DOI] [PubMed] [Google Scholar]
- (11).Roth CM, Sundaram S. Annu Rev Biomed Eng. 2004;6:397–426. doi: 10.1146/annurev.bioeng.6.040803.140203. [DOI] [PubMed] [Google Scholar]
- (12).Farhood H, Serbina N, Huang L. Biochim Biophys Acta. 1995;1235:289–295. doi: 10.1016/0005-2736(95)80016-9. [DOI] [PubMed] [Google Scholar]
- (13).Hughes JA, Aronsohn AI, Avrutskaya AV, Juliano RL. Pharm Res. 1996;13:404–410. doi: 10.1023/a:1016044609972. [DOI] [PubMed] [Google Scholar]
- (14).Jaaskelainen I, Peltola S, Honkakoski P, Monkkonen J, Urtti A. Eur J Pharm Sci. 2000;10:187–193. doi: 10.1016/s0928-0987(00)00068-3. [DOI] [PubMed] [Google Scholar]
- (15).Zelphati O, Uyechi LS, Barron LG, Szoka FC., Jr. Biochim Biophys Acta. 1998;1390:119–133. doi: 10.1016/s0005-2760(97)00169-0. [DOI] [PubMed] [Google Scholar]
- (16).Zhang Y, Anchordoquy TJ. Biochim Biophys Acta. 2004;1663:143–157. doi: 10.1016/j.bbamem.2004.03.004. [DOI] [PubMed] [Google Scholar]
- (17).Trubetskoy VS, Loomis A, Hagstrom JE, Budker VG, Wolff JA. Nucleic Acids Res. 1999;27:3090–3095. doi: 10.1093/nar/27.15.3090. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (18).Trubetskoy VS, Wong SC, Subbotin V, Budker VG, Loomis A, Hagstrom JE, Wolff JA. Gene Ther. 2003;10:261–271. doi: 10.1038/sj.gt.3301888. [DOI] [PubMed] [Google Scholar]
- (19).Finsinger D, Remy JS, Erbacher P, Koch C, Plank C. Gene Ther. 2000;7:1183–1192. doi: 10.1038/sj.gt.3301227. [DOI] [PubMed] [Google Scholar]
- (20).Kyriakides TR, Cheung CY, Murthy N, Bornstein P, Stayton PS, Hoffman AS. J Control Release. 2002;78:295–303. doi: 10.1016/s0168-3659(01)00504-1. [DOI] [PubMed] [Google Scholar]
- (21).Walton SP, Stephanopoulos GN, Yarmush ML, Roth CM. Biophysical Journal. 2002;82:366–377. doi: 10.1016/S0006-3495(02)75401-5. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (22).Sennato S, Bordi F, Cametti C, Di Biasio A, Diociaiuti M. Colloids and Surfaces A: Physicochemical and Engineering Aspects. 2005;270-271:138–147. [Google Scholar]
- (23).Evans DF, Wennerstrom H. The colloidal domain: where physics, chemistry, biology, and technology meet. 2nd ed. Wiley; New York: 1999. [Google Scholar]
- (24).Sundaram S, Viriyayuthakorn S, Roth CM. Biomacromolecules. 2005;6:2961–2968. doi: 10.1021/bm0502314. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (25).Zuidam NJ, Barenholz Y. Biochim Biophys Acta. 1997;1329:211–222. doi: 10.1016/s0005-2736(97)00110-7. [DOI] [PubMed] [Google Scholar]
- (26).Jones RA, Cheung CY, Black FE, Zia JK, Stayton PS, Hoffman AS, Wilson MR. Biochem J. 2003;372:65–75. doi: 10.1042/BJ20021945. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (27).Marcusson EG, Bhat B, Manoharan M, Bennett CF, Dean NM. Nucleic Acids Res. 1998;26:2016–2023. doi: 10.1093/nar/26.8.2016. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (28).Kusonwiriyawong C, van de Wetering P, Hubbell JA, Merkle HP, Walter E. Eur J Pharm Biopharm. 2003;56:237–246. doi: 10.1016/s0939-6411(03)00093-6. [DOI] [PubMed] [Google Scholar]
- (29).Sohail M. Drug Discov Today. 2001;6:1260–1261. doi: 10.1016/s1359-6446(01)02115-8. [DOI] [PubMed] [Google Scholar]
- (30).Gewirtz AM, Sokol DL, Ratajczak MZ. Blood. 1998;92:712–736. [PubMed] [Google Scholar]
- (31).Cho YS, Kim MK, Cheadle C, Neary C, Becker KG, Cho-Chung YS. Proc Natl Acad Sci U S A. 2001;98:9819–9823. doi: 10.1073/pnas.171314398. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (32).Cheung CY, Murthy N, Stayton PS, Hoffman AS. Bioconjug Chem. 2001;12:906–910. doi: 10.1021/bc0100408. [DOI] [PubMed] [Google Scholar]
- (33).Cheung CY, Stayton PS, Hoffman AS. J Biomater Sci Polym Ed. 2005;16:163–179. doi: 10.1163/1568562053115390. [DOI] [PubMed] [Google Scholar]
- (34).Kiang T, Bright C, Cheung CY, Stayton PS, Hoffman AS, Leong KW. J Biomater Sci Polym Ed. 2004;15:1405–1421. doi: 10.1163/1568562042368112. [DOI] [PubMed] [Google Scholar]
- (35).Targeted Genetics Corporation. 2003 http://www.targen.com/investor/pr-detail.php/id=130.
- (36).Aronsohn AI, Hughes JA. J Drug Target. 1998;5:163–169. doi: 10.3109/10611869808995871. [DOI] [PubMed] [Google Scholar]
- (37).Capaccioli S, Di Pasquale G, Mini E, Mazzei T, Quattrone A. Biochem Biophys Res Commun. 1993;197:818–825. doi: 10.1006/bbrc.1993.2552. [DOI] [PubMed] [Google Scholar]
- (38).Roth CM. Biophys J. 2005;89:2286–2295. doi: 10.1529/biophysj.104.054080. [DOI] [PMC free article] [PubMed] [Google Scholar]
- (39).Weyermann J, Lochmann D, Zimmer A. J Control Release. 2004;100:411–423. doi: 10.1016/j.jconrel.2004.08.027. [DOI] [PubMed] [Google Scholar]
- (40).Zelphati O, Szoka FC. Proc. Natl. Acad. Sci. 1996;93:11493–11498. doi: 10.1073/pnas.93.21.11493. [DOI] [PMC free article] [PubMed] [Google Scholar]