Abstract
Covalent modification of histones by ubiquitylation is a prominent epigenetic mark that features in a variety of chromatin-based events such as histone methylation, gene silencing, and repair of DNA damage. The prototypical example of histone ubiquitylation is that of histone H2B in Saccharomyces cerevisiae. In this case, attachment of ubiquitin to lysine 123 (K123) of H2B is important for regulation of both active and transcriptionally silent genes and participates in trans to signal methylation of histone H3. It is generally assumed that H2B is monoubiquitylated at K123 and that it is this single ubiquitin moiety that influences H2B function. To determine whether this assumption is correct, we have re-examined the ubiquitylation status of endogenous H2B in yeast. We find that, contrary to expectations, H2B is extensively polyubiquitylated. Polyubiquitylation of H2B appears to occur within the context of chromatin and is not associated with H2B destruction. There are at least two distinct modes of H2B polyubiquitylation: one that occurs at K123 and depends on the Rad6–Bre1 ubiquitylation machinery and another that occurs on multiple lysine residues and is catalyzed by an uncharacterized ubiquitin ligase(s). Interestingly, these ubiquitylation events are under the influence of different combinations of ubiquitin-specific proteases, suggesting that they have distinct biological functions. These results raise the possibility that some of the biological effects of ubiquitylation of H2B are exerted via ubiquitin chains, rather than a single ubiquitin group.
INTRODUCTION
Over the last decade, it has become apparent that covalent modification of histones plays a prominent role in establishing epigenetic patterns of gene control in eukaryotic cells (Ruthenburg et al., 2007). A variety of histone modifications have been described, including phosphorylation, methylation, and ubiquitylation. These modifications frequently occur in distinct, interrelated patterns, giving rise to the notion that there is a histone code that is read by the cellular machinery to set the transcriptional status of a particular piece of chromatin (Jenuwein and Allis, 2001).
Interestingly, one of the first covalent histone modifications to be discovered was ubiquitylation. In their characterization of the nuclear protein “A24,” Busch and colleagues (Goldknopf et al., 1975; Goldknopf and Busch, 1977) described an isopeptide linkage between lysine 119 (K119) of histone H2A and the protein that is now known as ubiquitin (Ub). Early studies (e.g., Levinger and Varshavsky, 1982) demonstrated a connection between histone ubiquitylation and the transcriptional status of chromatin, and it is now clear that histone ubiquitylation, which has been reported for H2A, H2B, H3, and H4 (Muratani and Tansey, 2003), is an important regulatory modification involved in both gene silencing (e.g., de Napoles et al., 2004; Fang et al., 2004) and activation (e.g., Kao et al., 2004). Because of the genetics available in yeast, most of what is known about the functional significance of histone ubiquitylation comes from studies of Saccharomyces cerevisiae and histone H2B.
Yeast histone H2B is monoubiquitylated at lysine residue 123 (K123) by the Rad6 Ub-conjugating enzyme (Robzyk et al., 2000) and the Bre1 Ub-protein ligase (Hwang et al., 2003; Wood et al., 2003a). Ubiquitylation of H2B (H2B–Ub) at this site participates in a variety of processes, including establishment of cell size (Hwang et al., 2003), meiosis (Robzyk et al., 2000; Yamashita et al., 2004), gene activation (Kao et al., 2004), gene silencing (Dover et al., 2002; Sun and Allis, 2002), and histone H3 methylation (Dover et al., 2002; Sun and Allis, 2002). H2B ubiquitylation is reportedly higher at active genes than at the silenced telomeres (Emre et al., 2005), and both the recruitment and activity of the H2B-ubiquitylation machinery are regulated by interactions with the transcriptional apparatus, including pol II (Xiao et al., 2005), the Paf complex (Ng et al., 2003; Wood et al., 2003b; Xiao et al., 2005), and Bur kinases (Wood et al., 2005). At active genes, H2B ubiquitylation is essential for the processive methylation of histone H3 at lysine residues K4 and K79 (Dover et al., 2002; Sun and Allis, 2002; Dehe et al., 2005; Shahbazian et al., 2005), but not K36 (Li et al., 2003; Wood et al., 2003b), and for recruitment of proteasomal ATPases to chromatin (Ezhkova and Tansey, 2004). Although only ∼10% of steady-state H2B is monoubiquitylated (Robzyk et al., 2000), it is likely that a much larger percentage of H2B is actually ubiquitylated and that deubiquitylation is an active and crucial process (reviewed in Emre and Berger, 2004). In support of this notion, the H2B–Ub-specific protease Ubp8, which is part of the SAGA complex (Henry et al., 2003; Daniel et al., 2004), is important for gene activation (Henry et al., 2003), whereas Ubp10, which is thought to preferentially remove Ub from H2B at the telomeres (Emre et al., 2005; Gardner et al., 2005), is important for gene silencing (Kahana and Gottschling, 1999). These findings have led to a model in which cycles of H2B ubiquitylation and deubiquitylation provide directionality to different stages in the transcription process (e.g., Wyce et al., 2007).
Current understanding of the role of H2B–Ub in transcription, and the search for downstream effector molecules, is predicated on the notion that H2B is modified by a single Ub moiety at K123. This notion is based largely on the fact that total H2B isolated from yeast cells resolves into two predominant species: the unmodified form (∼90%; Robzyk et al., 2000) and the form that is monoubiquitylated at K123. It should be noted, however, that the existence of a substantial pool of a ubiquitylated protein such as this is unusual, and in most cases ubiquitylated proteins constitute only a small fraction of the total population of that protein (Kaiser and Tagwerker, 2005). For this reason, detection of Ub conjugates usually requires methods to specifically enrich for and stabilize ubiquitylated proteins (e.g., Yaglom et al., 1995). Interestingly, we have been unable to find an example in the literature where such directed methods were applied to detect ubiquitylated forms of H2B. We therefore performed such an analysis to reexamine the ubiquitylation status of endogenous H2B in yeast cells. We find that, contrary to expectations, H2B is polyubiquitylated, both at K123 and at multiple other lysine residues in the protein. There are at least two distinct pathways that contribute to H2B polyubiquitylation, and these are under the control of distinct deubiquitylating enzymes. These data reveal that previous analyses of the role of ubiquitylation of H2B at K123 have been performed within a backdrop of other H2B-ubiquitylation events and raise the possibility that poly-Ub chains mediate the transcriptional effects of H2B–Ub.
MATERIALS AND METHODS
Yeast Strains and Plasmids
Yeast strains used in this study are listed in Table 1.
Table 1.
Yeast strains used in this study
| Strain | Genotype | Source |
|---|---|---|
| JD47-13C | MATα his3Δ200 leu2-3,112 lys2-801 trp1Δ63 ura3-52 | Dohmen et al. (1995) |
| FGY1 | =JD47-13C, except HTB1-myc::LEU2 | This study |
| FGY2 | =JD47-13C, except HTB1-HA::LEU2 | This study |
| FG4742 | MATα hisΔ1 leu2Δ0 lys2Δ0 ura3Δ0 | Openbiosystem |
| FGY3 | =BY4742, except rad6::KanMX | Openbiosystem |
| FGY4 | =BY4742, except bre1::KanMX | Openbiosystem |
| FGY5 | =BY4742, except ubp8::LEU2 | This study |
| FGY6 | =BY4742, except ubp10::KanMX | This study |
| FGY7 | =BY4742, except ubp8::LEU2 ubp10::KanMX | This study |
| JR5-2A | MATa htb-1-1 htb2-1 leu2-3,-112 his3-11,-15 trp1-1 ura3-1 ade2-1 can1-100 〈Ycp50-HTB1〉 | Recht and Osley (1999) |
| FGY8 | =JR5-2A, except hta2-htb2::KanMX | This study |
| FGY9 | =FGY8, except bre1::LEU2 | This study |
| FGY10 | =FGY8, except ubp8::LEU2 | This study |
| FGY11 | =FGY8, except ubp10::TRP1 | This study |
| FGY12 | =FGY8, except ubp8::LEU2 ubp10::TRP1 | This study |
| FGY13 | =BY4742, except HTB1-HA::LEU2 | This study |
| FGY14 | =BY4742, except HTB1-His8::LEU2 | This study |
| FGY15 | =BY4742, except HTB1-His8-HA::LEU2 | This study |
| FGY16 | =BY4742, except HTB1-His8-HA::LEU2 Flag-SMT3 | This study |
| FGY17 | =BY4742, except Flag-SMT3 | This study |
The integration plasmid pRS305-HTB1-myc was created by inserting partial HTB1 sequence (51, +300) into the plasmid pRS305; in this case, the HTB1 coding sequence was modified to include a single, carboxy-terminal, myc-epitope tag. Plasmid pRS305-HTB1-HA is identical to pRS305-HTB1-myc except that HTB1 carries a single copy of the hemagglutinin (HA)-epitope tag. The plasmids were used to tag endogenous H2B by cutting with HindIII, transforming into the yeast strain JD37-13C (kindly provided by R. Deshaies [California Institute of Technology, Pasadena, CA]; Dohmen et al., 1995), and selecting for the LEU2+ phenotype, which creates strains FGY1 (HTB1-myc) and FGY2 (HTB1-HA). Correct integration was confirmed by PCR and Western blotting (WB).
To express HTB1 from a single-copy plasmid under the control of the endogenous HTB1 promoter, the plasmid p413-HTB1-HA was constructed, which harbors the full HTB1 locus (−824, +300), modified to express H2B containing a single, carboxy-terminal, HA-epitope tag. Site-directed mutagenesis (Quickchange; Stratagene, La Jolla, CA) was used to create the following H2B mutants in p413-HTB1-HA: 1) K123R, lysine 123 changed to arginine; all other lysine residues intact; 2) K123+, lysine 123 intact and all other lysine residues changed to arginine; 3) K111+, lysine 111 intact and all other lysine residues changed to arginine; 4) K3+, lysine 3 intact and all other lysine residues changed to arginine; 5) K0, all lysine residues changed to arginine; 6) NK+, amino terminal lysine residues K3, 6, 7, 11, 16, 17, 21, 22, 30, and 31 intact and all other lysine residues changed to arginine; and 7) coreK+, core lysine residues 34, 37, 46, 49, 60, 82, 88, 89, and 111 intact and all other lysine residues changed to arginine. To express HA-tagged Htb1 proteins—WT and mutant—as the sole source of cellular H2B, a set of yeast strains were made by plasmid shuffle in strain FGY8 (Δhtb1Δhtb2; Table 1). The ubp8 and ubp10 deletion mutants were constructed by replacing the open reading frames (ORFs) with selection marker cassettes using a one-step PCR method (Knop et al., 1999). Deletion of ORFs was confirmed by PCR.
The plasmid pRS426-pCUP1-His7-UbiquitinG76A was constructed by inserting the CUP1 promoter and His7-Ubiquitin (mutant: G76A) coding sequences into pRS426. Plasmids pp389 (YepLac-TRP-pCUP-His6-ubiquitin;G76A), pp390 (YepLac-TRP-pCUP-His6-ubiquitin;G76A K48R), and pp391 (YepLac-TRP-pCUP-His6-ubiquitin;G76A K63R) were kindly provided by P. Kaiser (University of California, Irvine, CA; Flick et al., 2004). Plasmid pUB221 (pCUP1-his6-myc-Ubiquitin) was kindly provided by D. Finley (Harvard Medical School, Boston, MA; Yaglom et al., 1995).
To generate His8- and His8-HA-tagged H2B, pRS305-HTB1-HA was modified by site-directed mutagenesis to create pRS305-HTB1-His8 and pRS305-HTB1-His8-HA, respectively. The SUMO tagging plasmid pRS306-Flag-SMT3 was constructed by cloning into pRS306 the SMT3 gene with a Flag epitope sequence inserted after the second codon of SMT3. Endogenous HTB1 was then C-terminally HA-, his8-HA-, or his8-tagged by integrating the appropriate HindIII-digested plasmid and selecting for LEU2+. Endogenous SMT3 (encoding SUMO) was N-terminally Flag-tagged via a two-step gene replacement with PmlI-digested pRS306-Flag-SMT3. Correction integration was confirmed by PCR and Western blot.
Analysis of H2B Ubiquitylation and Stability
Ubiquitylation assays were carried out as described (Muratani et al., 2005) with some modification. Briefly, in the indicated strains, poly-histidine–tagged Ub was induced by addition of CuSO4 to a final concentration 0.2 mM. Cells were collected, disrupted by glass bead beating in buffer A2 (6 M guanidine-HCl, 100 mM Na2HPO4/NaH2PO4, pH 8.0, 10 mM imidazole, 250 mM NaCl, and 0.5% NP-40), and soluble material was recovered. At this point, 90% of the material was incubated with Ni-NTA agarose resin (Qiagen, Chatsworth, CA) for 2 h at 4°C. The resin was collected, washed twice with buffer A2, once with buffer A2/T2 (1 vol buffer A2 and 3 vol buffer T2: 50 mM Na2HPO4/NaH2PO4, pH 8.0, 250 mM NaCl, 20 mM imidazole, 0.5% NP-40) and twice with Buffer T2. Beads were collected and ubiquitylated proteins eluted with 2× SDS loading buffer containing 250 mM imidazole. Total proteins were recovered from the remaining 10% of the initial lysate (input) by ethanol precipitation, and resuspended in 1× SDS loading buffer. Proteins were then resolved by SDS-PAGE and analyzed by WB, using anti-HA HRP-conjugated antibody (Roche, Nutley, NJ). To assay the association of ubiquitylated H2B with chromatin, a similar approach was used, except that cellular material was first fractionated as described (Liang and Stillman, 1997). The Orc3 antibody was a gift from B. Stillman (Cold Spring Harbor Laboratory; Liang and Stillman, 1997).
Pulse-chase analysis was performed as described (Salghetti et al., 2000).
RNA Analyses
For GAL gene induction, yeast were grown overnight in rich media with 2% raffinose to an OD600 of ∼0.5 and were induced by addition of 2% galactose. Cells were collected after 1 h of induction and total RNA extracted with hot phenol. Random hexamer-primed reverse-transcription was performed with the TaqMan Reverse-Transcription Kit (Applied Biosystems, Foster City, CA). Real-time PCR was performed using an Opticon (Bio-Rad, Richmond, CA) thermocycler and the Cyber Green PCR Kit (Applied Biosystems). Gene-specific primers for GAL10 and 25s rRNA have been described (Muratani et al., 2005).
RESULTS
H2B Is Polyubiquitylated
It is commonly accepted that, in budding yeast, histone H2B is modified by covalent attachment of a single Ub moiety to lysine residue 123. This notion is based largely on the observation that ∼10% of H2B is present in the monoubiquitylated form in extracts from yeast cells (Robzyk et al., 2000). It should be noted, however, that this extent of ubiquitylation is unusual and that Ub-modified proteins are typically present in much lower amounts because of active deubiquitylation or proteasomal proteolysis. For the most part, therefore, ubiquitylated proteins can only efficiently be detected when they are selected for under conditions that block Ub removal and/or destruction by the proteasome. In our survey of the literature, we were unable to find an example of where this approach had been applied to histone H2B. Because models for how H2B–Ub functions are different when considering mono- versus polyubiquitylation or a single versus multiple sites of Ub conjugation, we used a directed method to characterize H2B ubiquitylation in exponentially growing yeast cells.
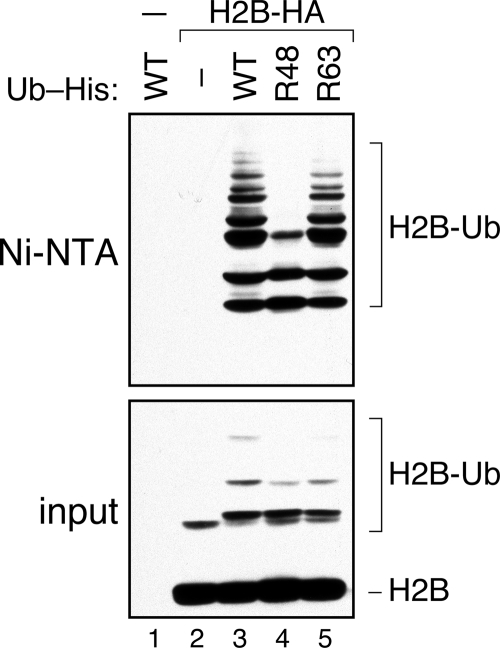
To detect H2B–Ub conjugates, we used an approach in which polyhistidine-tagged Ub was expressed in yeast (bearing HA epitope-tagged H2B) under the control of the copper-inducible CUP1 promoter (Yaglom et al., 1995). In this instance, the tagged Ub also carried a mutation (G76A) that prevents Ub removal by Ub-specific proteases (Hodgins et al., 1992). Total ubiquitylated proteins were purified under denaturing conditions by nickel-affinity (Ni-NTA) chromatography, and H2B–HA was detected by Western blotting (Figure 1). As expected (Robzyk et al., 2000), analysis of total H2B–HA (input) revealed the presence of two predominant H2B species, one corresponding to free H2B and the other corresponding to the monoubiquitylated form. Analysis of H2B–HA present in the Ni-NTA–purified material, in contrast, revealed a ladder of high-molecular-weight H2B conjugates (lane 3) that began at a size corresponding to monoubiquitylated H2B (∼25 kDa) and extended upward. At least nine distinct species could be resolved under these conditions, consistent with the notion that H2B is modified by addition of multiple Ub moieties. Although the relative levels of mono- versus polyubiquitylated H2B cannot be estimated from analysis of the Ni-NTA–purified material, we do note that a species corresponding in molecular weight to doubly ubiquitylated H2B can be detected in total extracts of cells expressing the noncleavable Ub (input; visible in the bracketed H2B–Ub region in lanes 3–5), suggesting that at a significant portion of H2B has at least two Ub groups attached. This conclusion was validated by purification of endogenous, polyhistidine-tagged H2B (Supplemental Figure S1); under these conditions, we could detect di- and triubiquitylated H2B in the absence of overexpressed, nonremovable Ub.
Figure 1.
Histone H2B is polyubiquitylated. Yeast cells were engineered to express either His-tagged WT or mutant (K48R or K63R) nonremovable (G76A) ubiquitin, together with either myc-tagged (FGY1; lane 1) or HA-epitope–tagged H2B (FGY2; lanes 2–5). Cells were lysed, and HA-tagged proteins present in the total lysate (input) or His-purified (Ni-NTA) material were detected by WB. The position of free H2B is indicated, as is the position of the H2B–Ub conjugates. Note that the monoubiquitylated form of H2B–HA appears as a doublet in the input; we presume that this corresponds to H2B–HA that is monoubiquitylated with either endogenous Ub (bottom panel) or the slightly larger His-Ub.
To determine whether the high-molecular-weight H2B–HA conjugates are modified by poly-Ub chains, we expressed mutant forms of polyhistidine-tagged Ub (G76A) in which one of the two prominent sites of Ub-chain formation—K48 or K63 (Pickart and Fushman, 2004)—was mutated to arginine. This analysis revealed that expression of the K48R mutant Ub reduced the number of H2B–Ub species to approximately three (lane 4), whereas expression of the K63 mutant (lane 5) had little if any effect on the extent of H2B modification. These data reveal that H2B is modified by at least one, and possibly more, K48-linked polyubiquitin chains and that K63-linked chains are unlikely to contribute significantly to the pool of ubiquitylated H2B.
There Are at Least Two Distinct Types of Polyubiquitylated H2B
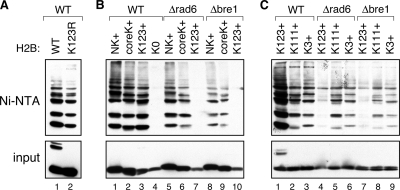
We next asked whether the poly-Ub chains associated with H2B were linked to the canonical lysine 123, other lysine residues in the protein, or both. We first used the His–Ub assay to examine the poly-Ub status of a mutant H2B in which K123 was changed to arginine (K123R; Figure 2A). As expected (Robzyk et al., 2000), the K123R mutation strongly reduced the levels of monoubiquitylated H2B present in total cell lysates (input; cf. lanes 1 and 2). Analysis of the Ni-NTA–purified material, in contrast, revealed that a significant pool of multiply ubiquitylated H2B remained in the K123R mutant, suggesting that residues in addition to K123 are sites for H2B ubiquitylation. Note, however, that total levels of multiply ubiquitylated H2B–HA are reduced in the K123R mutant, consistent with the notion that lysine 123 might also be polyubiquitylated.
Figure 2.
There are at least two distinct pathways of H2B polyubiquitylation. Ubiquitylation of the indicated HA-tagged H2B mutants was measured as described in the legend to Figure 1, except that H2B–HA was expressed from a centromeric plasmid in strain BY4742. (A) The K123R H2B mutant is polyubiquitylated. (B) Ubiquitylation at lysine 123 depends exclusively on Rad6–Bre1. The ubiquitylation status of four different H2B mutants—NK+, coreK+, K123+, and K0 (described in the text)—was determined in either WT cells (lanes 1–4), or congenic strains in which RAD6 (FGY3; lanes 5–7) or BRE1 (FGY4; lanes 8–10) were deleted. (C) H2B proteins carrying single lysine residues are polyubiquitylated. The ubiquitylation status of K123+, K111+, and K3+ H2B mutants was examined in WT (lanes 1–3), Δrad6 (lanes 4–6), and Δbre1 (lanes 7–9) cells. Note that a more sensitive form of detection was used in the experiment in C than in B, because of the lower absolute signal from the single lysine K111+ and K3+ mutants. For this reason, monoubiquitylated K123+ H2B is visible in the input in C (cf. lanes in B with lane 1 in C).
We further probed the nature of H2B–Ub by examining the ubiquitylation status of four additional H2B mutants in which groups of lysine residues were retained, whereas other lysine residues were changed to arginine: 1) NK+, which retained lysine residues in the amino-terminal portion of H2B (K3, 6, 7, 11, 16, 17, 21, 22, 30, 31); 2) coreK+, with lysine residues 34, 37, 46, 49, 60, 82, 88, 89, and 111 intact; 3) K123+, where K123 is the sole lysine in the protein; and 4) K0, a lysine-free mutant of H2B. This analysis revealed that, with the exception of the K0 protein (Figure 2B; lane 4), all three lysine-containing H2B mutants were multiply ubiquitylated (lanes 1–3). We conclude from this result that H2B is not subject to extensive lysine-free ubiquitylation (e.g., Bloom et al., 2003) and that multiple lysine residues within H2B can serve as sites of conjugation for Ub. Consistent with the results of the experiment presented in Figure 2A, this result also demonstrates that K123 can itself be subject to polyubiquitylation.
To examine the contribution of the previously described Rad6–Bre1 Ub-ligase pathway (Robzyk et al., 2000; Hwang et al., 2003; Wood et al., 2003a) to these ubiquitylation events, we also examined each mutant H2B in congenic Δrad6 and Δbre1 strains (lanes 5–10). We found that polyubiquitylation of the NK+ and coreK+ mutants was largely insensitive to disruption of either RAD6 or BRE1, revealing that this pathway is not responsible for ubiquitylation of these proteins. For the K123+ mutant, however, ubiquitylation was dependent on both RAD6 and BRE1: the K123+ protein in the Δrad6 and Δbre1 strains was largely indistinguishable from the K0 mutant. This finding reveals that both mono- and polyubiquitylation of H2B at K123 is dependent on Rad6–Bre1 and that there must be at least one other Ub-ligase system responsible for the ubiquitylation of the other H2B lysine residues.
We subsequently analyzed a panel of mutant H2B proteins, each of which contained a single lysine residue. The majority of these mutants were ubiquitylated (not shown), and we therefore selected two for further analysis: K111+ and K3+. We selected these two mutants because they carried lysine residues in different domains of the protein (K111+ in the core, K3+ in the amino terminus). This analysis is shown in Figure 2C. Both the K111+ and K3+ mutants were polyubiquitylated (lanes 2 and 3), although to a significantly less extent than K123+ (lane 1). In all both cases, loss of either Rad6 or Bre1 did not eliminate polyubiquitylation (cf. lanes 1–3 with 4–9), although some small differences were observed in the various mutants (e.g., the K111+ mutant in the Δbre1 strain; lane 8). This result emphasizes that ubiquitylation events at K123 are uniquely dependent on the Rad6–Bre1 proteins, whereas ubiquitylation at other sites within H2B must be directed by a distinct Ub-ligase system(s).
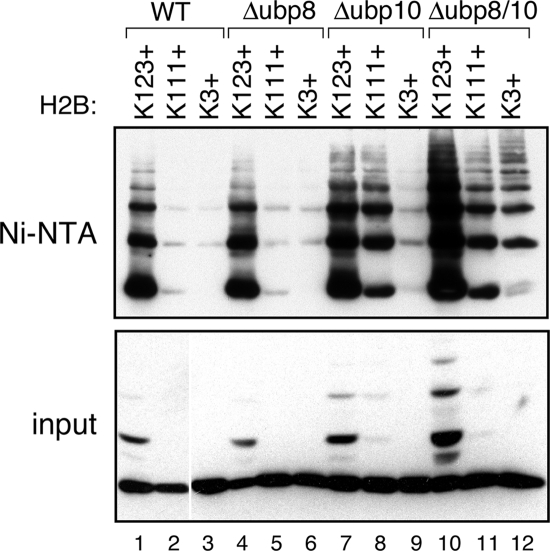
Histone H2B deubiquitylation plays an important role in mediating the transcriptional effects of H2B ubiquitylation (Emre and Berger, 2004). Two deubiquitylating enzymes—Ubp8 (Henry et al., 2003; Daniel et al., 2004) and Ubp10 (Emre et al., 2005; Gardner et al., 2005)—have been shown to deubiquitylate H2B at K123. The finding that other sites in H2B are ubiquitylated and that K123 can be polyubiquitylated raises the possibility that these noncanonical ubiquitylation events can also be regulated by Ubp8 and/or Ubp10. We therefore examined the ubiquitylation status of three single lysine H2B mutants (K123+, K111+, and K3+) in Δubp8, Δubp10, and Δubp8/Δubp10 yeast strains to compare how different sites of H2B ubiquitylation respond to loss of these deubiquitylating enzymes (Figure 3). In this analysis, the K123+ mutant responded little if any to loss of UBP8 (cf. lanes 1 and 4), whereas deletion of UBP10 increased the ubiquitylation levels of this protein considerably (lane 7). The failure of Δubp8 to affect K123+ ubiquitylation appears to be the result of redundancy between Ubp8 and Ubp10, because simultaneous disruption of both proteins increased the levels of mono- and polyubiquitylated K123+ considerably (lane 10). This result is consistent with previous observations showing that loss of Ubp8 and Ubp10 synergize to promote ubiquitylation of WT H2B (Gardner et al., 2005). Importantly, because the Δubp8/Δubp10 mutant accumulates significant levels of poly ubiquitylated K123+ H2B (visible in both the input and Ni-NTA–purified material), this result also reveals that these two deubiquitylating enzymes also play a role in mediating the extent of Ub-chain formation at lysine residue 123.
Figure 3.
Deubiquitylating enzymes differentially affect Ub-chain length at particular H2B lysine residues. Ubiquitylation of the indicated HA-tagged H2B mutants was measured as described in the legend to Figure 1, except that H2B–HA was expressed from a centromeric plasmid in BY4742. Note also that to detect the effects of deleting UBP8 and/or UBP10, we used a cleavable form of His-Ub in which residue G76 of Ub is intact (Yaglom et al., 1995). Four congenic yeast strains were analyzed: WT (BY4742; lanes 1–3), Δubp8 (FGY5; lanes 4–6), Δubp10 (FGY6; lanes 7–9), and Δubp8/Δubp10 (FGY7; lanes 10–12).
Analysis of the K3+ mutant revealed a similar pattern of behavior as K123+ (cf. lanes 3, 6, 9, and 12), suggesting that Ubp8 and Ubp10 both work to limit the extent of ubiquitylation at this site. Intriguingly, however, analysis of the K111+ mutant revealed a different pattern of response to deletion of UBP8 and UBP10. Like the K123+ mutant, deletion of UBP8 had little effect on the levels of ubiquitylated K111+ (cf. lanes 2 and 5), whereas deletion of UBP10 increased the extent of K111+ ubiquitylation considerably (lane 8). Unlike the K123+ mutant, however, there appeared to be little redundancy between Ubp8 and Ubp10, because the ubiquitylation status of the K111+ mutant was very similar in the Δubp8 and Δubp8/Δubp10 strains (cf. lanes 8 and 11). From this result we conclude that ubiquitylation at lysine 111 of H2B is predominantly under the control of Ubp10 and that Ubp8 does little to limit Ub-chain formation at this site. The differential behavior of the K111+ mutant demonstrates that deubiquitylating enzymes Ubp8 and Ubp10 do not work globally on ubiquitylated H2B, but display specificity for particular sites of Ub conjugation. This finding further supports the idea that multiple pathways regulate H2B ubiquitylation and suggests that ubiquitylation, and deubiquitylation, of different lysine residues within H2B may mediate distinct functional outcomes.
Polyubiquitylated H2B Is Chromatin-associated
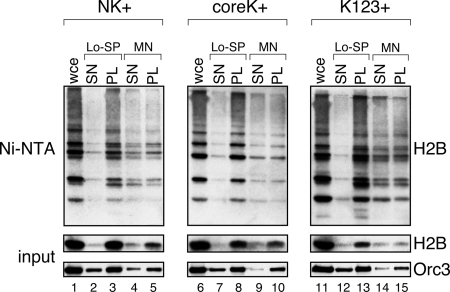
A central factor in models for how ubiquitylation of H2B at K123 controls gene expression is that H2B–Ub is incorporated into chromatin and acts to either recruit factors to sites of gene regulation or to disrupt local chromatin structure (Osley, 2004). It is conceivable, however, that the additional ubiquitylated forms of H2B that we detect do not exist within the context of chromatin, in which case a role for these modifications in the control of chromatin-mediated events would be more difficult to imagine. We therefore asked whether forms of H2B with noncanonical ubiquitin chains can associate with chromatin, as shown in Figure 4. We measured the ubiquitylation status of the NK+, coreK+, and K123+ H2B mutants after fractionation of chromatin using the method of Liang and Stillman (1997). In this assay, total cell extracts are first subject to low-speed centrifugation to separate soluble proteins from the insoluble matrix and chromatin. The insoluble material is then treated with micrococcal nuclease to release chromatin from this fraction, leading to the appearance of chromatin-associated proteins in the supernatant after a second round of centrifugation. The DNA replication factor Orc3 was our control in this assay (Liang and Stillman, 1997). As shown in the figure, ubiquitylated forms of all three H2B mutants were present in the insoluble material (pellet) after the first round of centrifugation (lanes 3, 8, and 13), and all three could be released from the pellet by treatment with micrococcal nuclease, to an extent similar to that observed for total levels of Orc3 (lanes 4, 9, and 14). From this result, we conclude that a significant portion of K123-ubiquitylated H2B, as well as H2B ubiquitylated at other sites, is tightly associated with chromatin. Interestingly, we also observed that ubiquitylated forms of H2B were generally released from chromatin more efficiently than the nonubiquitylated species (cf. Ni-NTA vs. input for lanes 4 and 5, 9 and 10, and 14 and 15), suggesting that chromatin containing ubiquitylated H2B may be more accessible than that containing the unmodified form.
Figure 4.
Ubiquitylated H2B is associated with chromatin. Wild-type yeast cells (JR5-2A) were engineered by plasmid shuffle to express the indicated H2B-HA mutants (from a centromeric plasmid) as well as His-tagged G76A ubiquitin. Chromatin fractionation was then performed as described (Liang and Stillman, 1997). Whole cell extracts (WCE; lanes 1, 6, and 11) were subject to a low-speed centrifugation (Lo-SP) and separated into soluble material (SN; lanes 2, 7, and 12) and an insoluble pellet containing chromatin (PL; lanes 3, 8, and 13). The insoluble material was then treated with micrococcal nuclease (MN) to release mono- and polynucleosomes into the soluble fraction. These extracts were then subject to a second round of centrifugation to separate remaining insoluble material (PL; lanes 5, 10, and 15) from the solubilized nucleosomes (SN; lanes 4, 9, and 14). Nickel-affinity chromatography was then used to isolate ubiquitylated proteins in each fraction. H2B–HA in the Ni-NTA and input material, as well as Orc3 (control) in the input, were detected by WB.
Ubiquitylation of Lysine 123 of H2B Can Be Essential
Rad6–Bre1-mediated ubiquitylation of histone H2B at K123 is a nonessential process in yeast. Yeast deleted for RAD6 or BRE1, or carrying the K123R H2B mutant, display reduced GAL gene activation (Kao et al., 2004), as well as defects in meiosis (Robzyk et al., 2000; Yamashita et al., 2004), gene silencing (Dover et al., 2002; Sun and Allis, 2002), and histone H3 methylation (Dover et al., 2002; Sun and Allis, 2002), but are nonetheless viable. Our finding that there are multiple ubiquitylation events on histone H2B raises the important point that previous analyses of the functional consequences of Rad6–Bre1-dependent H2B ubiquitylation (at K123) have been performed within a backdrop of other H2B-ubiquitylation events, which in turn may mask effects resulting from loss of Rad6–Bre1 ubiquitylation. Although we do not know the identity of the cellular machinery that ubiquitylates the noncanonical sites on H2B, our finding that ubiquitylation of the K123+ mutant entirely dependent on Rad6–Bre1 (Figure 2) gives us the opportunity to examine the contribution of this site of ubiquitylation in isolation.
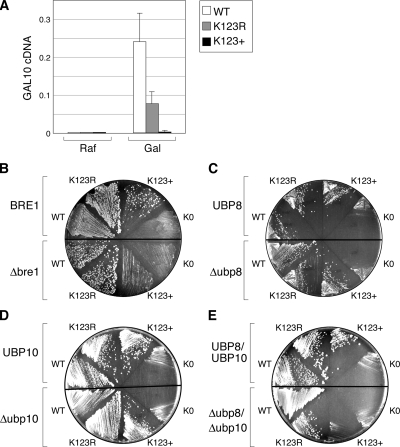
We therefore asked whether H2B with a single lysine at position 123 would support GAL gene activation and yeast cell viability, and—if so—whether this was dependent on Bre1. We examined galactose-mediated induction of the GAL10 gene (Figure 5A). Consistent with previous reports (Henry et al., 2003; Kao et al., 2004), we found that the K123R mutation reduced GAL10 induction about threefold. Under identical conditions, however, the K123+ mutant showed little if any induction of GAL10, arguing that other lysine residues within H2B (besides K123) are important for GAL activation. Although this result itself is not informative, it does contrast strongly with the effects of these mutations on cell viability (Figure 5B), as described below.
Figure 5.
Ubiquitylation at lysine 123 of H2B can be essential for viability. Plasmid shuffle was used to express WT H2B—and the K123R and K123+ mutants—as the sole source of histone H2B in strains FGY8, FGY9, and FGY10 (Table 1). (A) Lysine 123 of H2B is required for full GAL gene induction, but is not sufficient. Yeast were grown in raffinose and either uninduced (Raf) or induced with galactose (Gal) for 60 min. RNA was collected and reverse-transcribed into cDNA, and levels of GAL10 and 25S rRNA cDNAs were determined by quantitative PCR. “GAL10 cDNA” is expressed relative to the 25S rRNA control. (B) The K123+ and Δbre1 mutations are synthetically lethal. The indicated H2B proteins were expressed in either WT (BRE1; FGY8) or Δbre1 (FGY9) cells, together with a WT H2B protein expressed from a plasmid bearing the URA3 gene. Yeast were struck onto media containing 5-fluoroorotic acid (FOA) to eliminate cells expressing WT H2B. The K123+ H2B mutant supports viability in the BRE1, but not Δbre1, cells. (C) No genetic interaction between K123+ and Δubp8. Assay performed as in B but with UBP8 (FGY8) and Δubp8 (FGY10) cells. (D) No genetic interaction between K123+ and Δubp10. Assay performed as in B but with UBP10 (FGY8) and Δubp10 (FGY11) cells. (E) The K123+ and Δubp8/Δubp10 mutations are synthetically lethal. Assay performed as in B but with UBP8/10 (FGY8) and Δubp8/Δubp10 (FGY12) cells.
We used a plasmid shuffle assay to compare the ability of wild-type (WT), K123R, K123+, and K0 H2B proteins to support yeast growth, both in the presence and absence of BRE1 (Figure 5B). As expected, the WT and K123R H2B proteins were functional in this assay and were not affected by loss of BRE1. The lysine-free H2B protein (K0), in contrast, did not support viability in either strain background. Interestingly, however, H2B bearing only a single lysine at position 123 (K123+) did allow yeast growth in BRE1 cells, albeit at a reduced level compared with the WT protein. Moreover, the ability of the K123+ mutant to support cell growth was dependent on BRE1, as the K123+/Δbre1 combination was inviable. We conclude from this result that most of the lysine residues in H2B are dispensable for the essential functions of this histone and that in the absence of these lysine residues ubiquitylation at K123 is an essential process.
Because of the role for histone deubiquitylation in mediating the transcriptional effects of H2B–Ub (Emre and Berger, 2004), we repeated the above analysis, but this time in cells that either expressed or lacked Ubp8 (Figure 5C), Ubp10 (Figure 5D), or both (Figure 5E). Deletion of either UBP8 or UBP10 was not synthetic with the K123+ H2B mutation. Remarkably, however, the double ubp8/Δubp10 deletion was synthetically lethal with the K123+ mutation. This result reveals that deubiquitylation of H2B at K123, like ubiquitylation itself, can be an essential process.
DISCUSSION
Analysis of H2B ubiquitylation in the genetically tractable yeast S. cerevisiae has been instrumental in revealing the extent to which this process can regulate gene activity. Exactly how ubiquitylation regulates H2B function is, however, unknown. To understand this mechanism, it is essential to understand the nature of how H2B is modified by ubiquitin. We used a directed method to examine the ubiquitylation status of H2B, and various mutants, in yeast cells. We found that, contrary to expectations, H2B is subject to extensive polyubiquitylation, both at the canonical K123 residue (which is assumed to be monoubiquitylated) and at multiple other sites within the protein. Ubiquitylation at K123 depends on the characterized Rad6–Bre1 pathway, whereas ubiquitylation at other sites occurs independent of Rad6 or Bre1. The existence of these multiple ubiquitylation events challenges the notion that a single Ub moiety is responsible for the biological activities of H2B–Ub and suggests that poly-Ub chains, perhaps attached to multiple sites on H2B, may influence its function.
Our studies have revealed two previously unrecognized types of H2B ubiquitylation: ubiquitylation at noncanonical lysine residues (i.e., not K123), and polyubiquitylation at lysine 123. Using “chain terminating” forms of Ub (Figure 1), we have shown that most of these polyubiquitylation events involve K48-linked Ub chains and that there are multiple sites of Ub-chain formation within H2B. Analysis of single lysine H2B mutants reveals that most of the lysine residues within the protein can be subject to some ubiquitylation (Figure 2 and data not shown), suggesting that there are either multiple pathways directing ubiquitylation at the various sites on H2B or that a smaller number of pathways can ubiquitylate H2B without targeting selected lysine residues. The latter scenario is particularly common in the Ub–proteasome system, where finding specific lysine residues required for Ub conjugation is the exception, rather than the rule (e.g., Crook et al., 1996). It is important to note, however, that loss of Rad6 or Bre1 has little effect on the ubiquitylation status of NK+, coreK+, K111+, and K3+ mutants (Figure 2), revealing that 1) Rad/Bre1 is fairly specific to K123 and 2) at least one other Ub-ligase must mediate ubiquitylation events at the other sites. It is also important to note that although the other Ub-ligase(s) may not have specificity toward particular lysine residues, the output of its activity could be very specific, because Ubp8 and Ubp10 appear to have different preferences for removing Ub from select lysine residues (Figure 3).
What is the function of these noncanonical K48-linked polyubiquitin chains? Although K48-linked chains are typically implicated in proteolysis (Pickart and Fushman, 2004), we think it is unlikely that these modifications direct H2B destruction. Histones, including H2B, are reported to be metabolically quite stable (Russev et al., 1975). We have confirmed this result with bulk H2B–HA in our system (Supplemental Figure S2). This finding also suggests that the function of H2B polyubiquitylation is not linked to the degradation of excess histones (Gunjan and Verreault, 2003). We favor the idea that K48-linked H2B polyubiquitylation plays a nonproteolytic, regulatory role in some aspect of chromatin function. This notion is supported by our finding that the ubiquitylated forms of the NK+ and coreK+ H2B mutants are associated with chromatin (Figure 4) and is analogous to what has been observed for the Met4 transcription factor, where limited K48-linked poly-Ub chains modulate Met4 function without altering its stability (Flick et al., 2004; Flick et al., 2006). It is possible that multiple Ub chains on H2B recruit Ub-dependent chaperones, such as Cdc48/p97 and the base of the 19S proteasome (Jentsch and Rumpf, 2007), which can in turn alter chromatin structure. Recently, it was found that Cdc48/p49 can extract polyubiquitylated Aurora B kinase from chromatin (Ramadan et al., 2007). By analogy, perhaps polyubiquitylation of H2B is important for mediating its eviction from chromatin as part of nonreplicative histone H2B exchange (Jamai et al., 2007).
In addition to revealing ubiquitylation at noncanonical sites in H2B, our data also demonstrate that lysine 123 itself is subject to polyubiquitylation (Figure 2). Intriguingly, both mono- and polyubiquitylation at K123 is dependent on RAD6/BRE1 (Figure 2), revealing that the Rad6–Bre1 complex has selectivity for this residue and is the only ubiquitylation pathway that can target lysine 123. Given that many lysine residues on H2B can potentially serve as sites for Ub conjugation, the specificity between Rad6–Bre1 and K123 is striking and raises the issue of why other Ub-ligases fail to ubiquitylate K123. One possibility would be if the other Ub-ligases accessed a different pool of H2B than Rad6–Bre1, one in which K123 is not exposed in a context that is appropriate for ubiquitylation.
Regardless of how specificity between K123 and Rad6–Bre1 is established, these findings raise the distinct possibility that the previously described functions of H2B–K123 ubiquitylation are mediated by poly-Ub chains. This is an important realization, because poly-Ub chains may be able to interact with protein factors that cannot bind tightly to a single Ub moiety (Jamai et al., 2007). As described above, poly-Ub chains could recruit Ub-dependent chaperones to chromatin, which in turn facilitate the known functions of K123 ubiquitylation. Indeed, we have previously found that H2B ubiquitylation is required for recruitment of 19S proteasomal ATPases to chromatin (Ezhkova and Tansey, 2004) and that mutations in these ATPases result in loss of histone H3 (K4 and K79) methylation, which is itself a Rad6–Bre1/K123-dependent process. This finding has led us to propose that 19S proteins come to chromatin in a Ub-dependent manner and promote H3 methylation by altering local chromatin structure. One weakness in this model has been the fact that proteasome subunits have low affinity for mono-Ub (Deveraux et al., 1994), making it difficult to imagine how monoubiquitylation of H2B at K123 could be linked to 19S recruitment. The demonstration that H2B is polyubiquitylated at this site makes a direct link between Rad6–Bre1-dependent ubiquitylation and 19S recruitment to chromatin more likely.
Our data also raise the possibility that transcriptional effects mediated by Ubp8 and Ubp10 could be a result of trimming poly-Ub chains, rather than deubiquitylating H2B entirely. It is difficult to dissect the relative contribution of mono- versus polyubiquitylation at lysine 123. Although the steady-state levels of polyubiquitylated H2B are considerably lower than the K123-monoubiquitylated form, this does not necessarily mean that H2B bearing a single Ub at K123 is the active species. Not only is ubiquitylation a highly dynamic process, but H2B polyubiquitylation may be spatially or temporally restricted, making it possible that H2B is extensively ubiquitylated at some sites in the genome where it the modification exerts its function. At this point, we do not know whether Rad6–Bre1 catalyzes K123-linked poly-Ub chains, or whether it primes H2B for polyubiquitylation by another ligase. We have found, however, that the E4, Ufd2, which is involved in Ub-chain elongation (Koegl et al., 1999), is not required for polyubiquitylation at this site (data not shown), and we note that Rad6 can polyubiquitylate H2B in vitro (Sung et al., 1988), suggesting that perhaps Rad6–Bre1 are sufficient for driving Ub-chain formation at K123.
Our genetic data (Figure 5) show that, in the presence of the K123+ form of H2B, disruption of either the Bre1 Ub-ligase or the Ubp8/Ubp10 deubiquitylating enzymes blocks yeast cell viability. This result strongly suggests that both H2B ubiquitylation and deubiquitylation can be essential, supporting the notion that cycling of H2B ubiquitylation is a critical part of histone function (Emre and Berger, 2004). Whether this phenomenon reflects cycling of total K123 ubiquitylation, or essential trimming of the poly-Ub chain on K123, however, remains to be determined.
Are histones polyubiquitylated in other contexts? Early studies identified polyubiquitylated forms of both H2A and H2B (Davie et al., 1987, 1991), and found that both mono- and polyubiquitylated H2B are preferentially localized to transcriptionally active chromatin domains (Nickel et al., 1989; Davie et al., 1991). It is thus possible that poly-Ub chains broadly regulate histone function in eukaryotic cells. The demonstration of H2B polyubiquitylation in a genetically tractable model organism should permit future analyses of the role these polyubiquitylation events play in control of chromatin dynamics.
Supplementary Material
ACKNOWLEDGMENTS
We thank G. Collins, A. Daulny, and A. Leung for helpful discussion and comments on the manuscript. For reagents we thank R. Deshaies, P. Kaiser, D. Finley, M.-A. Osley (University of New Mexico Health Sciences Center, Albuquerque, NM), and B. Stillman. W.P.T. was a Kimmel Foundation and Leukemia and Lymphoma Society Scholar. This work is supported by National Institutes of Health Grant GM067728.
Footnotes
This article was published online ahead of print in MBC in Press (http://www.molbiolcell.org/cgi/doi/10.1091/mbc.E08-01-0050) on June 18, 2008.
REFERENCES
- Bloom J., Amador V., Bartolini F., DeMartino G., Pagano M. Proteasome-mediated degradation of p21 via N-terminal ubiquitinylation. Cell. 2003;115:71–82. doi: 10.1016/s0092-8674(03)00755-4. [DOI] [PubMed] [Google Scholar]
- Crook T., Ludwig R. L., Marston N. J., Willkomm D., Vousden K. H. Sensitivity of p53 lysine mutants to ubiquitin-directed degradation targeted by human papillomavirus E6. Virology. 1996;217:285–292. doi: 10.1006/viro.1996.0115. [DOI] [PubMed] [Google Scholar]
- Daniel J. A., Torok M. S., Sun Z. W., Schieltz D., Allis C. D., Yates J. R., 3rd, Grant P. A. Deubiquitination of histone H2B by a yeast acetyltransferase complex regulates transcription. J. Biol. Chem. 2004;279:1867–1871. doi: 10.1074/jbc.C300494200. [DOI] [PubMed] [Google Scholar]
- Davie J. R., Delcuve G. P., Nickel B. E., Moirier R., Bailey G. Reduced levels of histones H1o and H1b, and unaltered content of methylated DNA in rainbow trout hepatocellular carcinoma chromatin. Cancer Res. 1987;47:5407–5410. [PubMed] [Google Scholar]
- Davie J. R., Lin R., Allis C. D. Timing of the appearance of ubiquitinated histones in developing new macronuclei of Tetrahymena thermophila. Biochem. Cell Biol. 1991;69:66–71. doi: 10.1139/o91-009. [DOI] [PubMed] [Google Scholar]
- de Napoles M., et al. Polycomb group proteins Ring1A/B link ubiquitylation of histone H2A to heritable gene silencing and X inactivation. Dev. Cell. 2004;7:663–676. doi: 10.1016/j.devcel.2004.10.005. [DOI] [PubMed] [Google Scholar]
- Dehe P. M., Pamblanco M., Luciano P., Lebrun R., Moinier D., Sendra R., Verreault A., Tordera V., Geli V. Histone H3 lysine 4 mono-methylation does not require ubiquitination of histone H2B. J. Mol. Biol. 2005;353:477–484. doi: 10.1016/j.jmb.2005.08.059. [DOI] [PubMed] [Google Scholar]
- Deveraux Q., Ustrell V., Pickart C., Rechsteiner M. A 26 S protease subunit that binds ubiquitin conjugates. J. Biol. Chem. 1994;269:7059–7061. [PubMed] [Google Scholar]
- Dohmen R. J., Stappen R., McGrath J. P., Forrova H., Kolarov J., Goffeau A., Varshavsky A. An essential yeast gene encoding a homolog of ubiquitin-activating enzyme. J. Biol. Chem. 1995;270:18099–18109. doi: 10.1074/jbc.270.30.18099. [DOI] [PubMed] [Google Scholar]
- Dover J., Schneider J., Tawiah-Boateng M. A., Wood A., Dean K., Johnston M., Shilatifard A. Methylation of histone H3 by COMPASS requires ubiquitination of histone H2B by Rad6. J. Biol. Chem. 2002;277:28368–28371. doi: 10.1074/jbc.C200348200. [DOI] [PubMed] [Google Scholar]
- Emre N. C., Berger S. L. Histone H2B ubiquitylation and deubiquitylation in genomic regulation. Cold Spring Harb. Symp. Quant. Biol. 2004;69:289–299. doi: 10.1101/sqb.2004.69.289. [DOI] [PubMed] [Google Scholar]
- Emre N. C., et al. Maintenance of low histone ubiquitylation by Ubp10 correlates with telomere-proximal Sir2 association and gene silencing. Mol. Cell. 2005;17:585–594. doi: 10.1016/j.molcel.2005.01.007. [DOI] [PubMed] [Google Scholar]
- Ezhkova E., Tansey W. P. Proteasomal ATPases link ubiquitylation of histone H2B to methylation of histone H3. Mol. Cell. 2004;13:435–442. doi: 10.1016/s1097-2765(04)00026-7. [DOI] [PubMed] [Google Scholar]
- Fang J., Chen T., Chadwick B., Li E., Zhang Y. Ring1b-mediated H2A ubiquitination associates with inactive X chromosomes and is involved in initiation of X inactivation. J. Biol. Chem. 2004;279:52812–52815. doi: 10.1074/jbc.C400493200. [DOI] [PubMed] [Google Scholar]
- Flick K., Ouni I., Wohlschlegel J. A., Capati C., McDonald W. H., Yates J. R., Kaiser P. Proteolysis-independent regulation of the transcription factor Met4 by a single Lys 48-linked ubiquitin chain. Nat. Cell Biol. 2004;6:634–641. doi: 10.1038/ncb1143. [DOI] [PubMed] [Google Scholar]
- Flick K., Raasi S., Zhang H., Yen J. L., Kaiser P. A ubiquitin-interacting motif protects polyubiquitinated Met4 from degradation by the 26S proteasome. Nat. Cell Biol. 2006;8:509–515. doi: 10.1038/ncb1402. [DOI] [PubMed] [Google Scholar]
- Gardner R. G., Nelson Z. W., Gottschling D. E. Ubp10/Dot4p regulates the persistence of ubiquitinated histone H2B: distinct roles in telomeric silencing and general chromatin. Mol. Cell. Biol. 2005;25:6123–6139. doi: 10.1128/MCB.25.14.6123-6139.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldknopf I. L., Busch H. Isopeptide linkage between nonhistone and histone 2A polypeptides of chromosomal conjugate-protein A24. Proc. Natl. Acad. Sci. USA. 1977;74:864–868. doi: 10.1073/pnas.74.3.864. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goldknopf I. L., Taylor C. W., Baum R. M., Yeoman L. C., Olson M. O., Prestayko A. W., Busch H. Isolation and characterization of protein A24, a “histone-like” non-histone chromosomal protein. J. Biol. Chem. 1975;250:7182–7187. [PubMed] [Google Scholar]
- Gunjan A., Verreault A. A Rad53 kinase-dependent surveillance mechanism that regulates histone protein levels in S. cerevisiae. Cell. 2003;115:537–549. doi: 10.1016/s0092-8674(03)00896-1. [DOI] [PubMed] [Google Scholar]
- Henry K. W., Wyce A., Lo W. S., Duggan L. J., Emre N. C., Kao C. F., Pillus L., Shilatifard A., Osley M. A., Berger S. L. Transcriptional activation via sequential histone H2B ubiquitylation and deubiquitylation, mediated by SAGA-associated Ubp8. Genes Dev. 2003;17:2648–2663. doi: 10.1101/gad.1144003. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hodgins R. R., Ellison K. S., Ellison M. J. Expression of a ubiquitin derivative that conjugates to protein irreversibly produces phenotypes consistent with a ubiquitin deficiency. J. Biol. Chem. 1992;267:8807–8812. [PubMed] [Google Scholar]
- Hwang W. W., Venkatasubrahmanyam S., Ianculescu A. G., Tong A., Boone C., Madhani H. D. A conserved RING finger protein required for histone H2B monoubiquitination and cell size control. Mol. Cell. 2003;11:261–266. doi: 10.1016/s1097-2765(02)00826-2. [DOI] [PubMed] [Google Scholar]
- Jamai A., Imoberdorf R. M., Strubin M. Continuous histone H2B and transcription-dependent histone H3 exchange in yeast cells outside of replication. Mol. Cell. 2007;25:345–355. doi: 10.1016/j.molcel.2007.01.019. [DOI] [PubMed] [Google Scholar]
- Jentsch S., Rumpf S. Cdc48 (p97): a “molecular gearbox” in the ubiquitin pathway? Trends Biochem. Sci. 2007;32:6–11. doi: 10.1016/j.tibs.2006.11.005. [DOI] [PubMed] [Google Scholar]
- Jenuwein T., Allis C. D. Translating the histone code. Science. 2001;293:1074–1080. doi: 10.1126/science.1063127. [DOI] [PubMed] [Google Scholar]
- Kahana A., Gottschling D. E. DOT4 links silencing and cell growth in Saccharomyces cerevisiae. Mol. Cell. Biol. 1999;19:6608–6620. doi: 10.1128/mcb.19.10.6608. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kaiser P., Tagwerker C. Is this protein ubiquitinated? Methods Enzymol. 2005;399:243–248. doi: 10.1016/S0076-6879(05)99016-2. [DOI] [PubMed] [Google Scholar]
- Kao C. F., Hillyer C., Tsukuda T., Henry K., Berger S., Osley M. A. Rad6 plays a role in transcriptional activation through ubiquitylation of histone H2B. Genes Dev. 2004;18:184–195. doi: 10.1101/gad.1149604. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Knop M., Siegers K., Pereira G., Zachariae W., Winsor B., Nasmyth K., Schiebel E. Epitope tagging of yeast genes using a PCR-based strategy: more tags and improved practical routines. Yeast. 1999;15:963–972. doi: 10.1002/(SICI)1097-0061(199907)15:10B<963::AID-YEA399>3.0.CO;2-W. [DOI] [PubMed] [Google Scholar]
- Koegl M., Hoppe T., Schlenker S., Ulrich H. D., Mayer T. U., Jentsch S. A novel ubiquitination factor, E4, is involved in multiubiquitin chain assembly. Cell. 1999;96:635–644. doi: 10.1016/s0092-8674(00)80574-7. [DOI] [PubMed] [Google Scholar]
- Levinger L., Varshavsky A. Selective arrangement of ubiquitinated and D1 protein-containing nucleosomes within the Drosophila genome. Cell. 1982;28:375–385. doi: 10.1016/0092-8674(82)90355-5. [DOI] [PubMed] [Google Scholar]
- Li B., Howe L., Anderson S., Yates J. R., 3rd, Workman J. L. The Set2 histone methyltransferase functions through the phosphorylated CTD of RNA polymerase II. J. Biol. Chem. 2003;278:8897–8903. doi: 10.1074/jbc.M212134200. [DOI] [PubMed] [Google Scholar]
- Liang C., Stillman B. Persistent initiation of DNA replication and chromatin-bound MCM proteins during the cell cycle in cdc6 mutants. Genes Dev. 1997;11:3375–3386. doi: 10.1101/gad.11.24.3375. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Muratani M., Kung C., Shokat K. M., Tansey W. P. The F box protein Dsg1/Mdm30 is a transcriptional coactivator that stimulates Gal4 turnover and cotranscriptional mRNA processing. Cell. 2005;120:887–899. doi: 10.1016/j.cell.2004.12.025. [DOI] [PubMed] [Google Scholar]
- Muratani M., Tansey W. P. How the ubiquitin-proteasome system controls transcription. Nat. Rev. Mol. Cell Biol. 2003;4:192–201. doi: 10.1038/nrm1049. [DOI] [PubMed] [Google Scholar]
- Ng H. H., Dole S., Struhl K. The Rtf1 component of the Paf1 transcriptional elongation complex is required for ubiquitination of histone H2B. J. Biol. Chem. 2003;278:33625–33628. doi: 10.1074/jbc.C300270200. [DOI] [PubMed] [Google Scholar]
- Nickel B. E., Allis C. D., Davie J. R. Ubiquitinated histone H2B is preferentially located in transcriptionally active chromatin. Biochemistry. 1989;28:958–963. doi: 10.1021/bi00429a006. [DOI] [PubMed] [Google Scholar]
- Osley M. A. H2B ubiquitylation: the end is in sight. Biochim. Biophys. Acta. 2004;1677:74–78. doi: 10.1016/j.bbaexp.2003.10.013. [DOI] [PubMed] [Google Scholar]
- Pickart C. M., Fushman D. Polyubiquitin chains: polymeric protein signals. Curr. Opin. Chem. Biol. 2004;8:610–616. doi: 10.1016/j.cbpa.2004.09.009. [DOI] [PubMed] [Google Scholar]
- Ramadan K., Bruderer R., Spiga F. M., Popp O., Baur T., Gotta M., Meyer H. H. Cdc48/p97 promotes reformation of the nucleus by extracting the kinase Aurora B from chromatin. Nature. 2007;450:1258–1262. doi: 10.1038/nature06388. [DOI] [PubMed] [Google Scholar]
- Recht J., Osley M. A. Mutations in both the structured domain and N-terminus of histone H2B bypass the requirement for Swi-Snf in yeast. EMBO J. 1999;18:229–240. doi: 10.1093/emboj/18.1.229. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robzyk K., Recht J., Osley M. A. Rad6-dependent ubiquitination of histone H2B in yeast. Science. 2000;287:501–504. doi: 10.1126/science.287.5452.501. [DOI] [PubMed] [Google Scholar]
- Russev G., Anachkova B., Tsanev R. Fractionation of rat-liver-chromatin nonhistone proteins into two groups with different metabolic rates. Eur. J. Biochem. 1975;58:253–257. doi: 10.1111/j.1432-1033.1975.tb02370.x. [DOI] [PubMed] [Google Scholar]
- Ruthenburg A. J., Li H., Patel D. J., Allis C. D. Multivalent engagement of chromatin modifications by linked binding modules. Nat. Rev. Mol. Cell Biol. 2007;8:983–994. doi: 10.1038/nrm2298. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Salghetti S. E., Muratani M., Wijnen H., Futcher B., Tansey W. P. Functional overlap of sequences that activate transcription and signal ubiquitin-mediated proteolysis. Proc. Natl. Acad. Sci. USA. 2000;97:3118–3123. doi: 10.1073/pnas.050007597. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Shahbazian M. D., Zhang K., Grunstein M. Histone H2B ubiquitylation controls processive methylation but not monomethylation by Dot1 and Set1. Mol. Cell. 2005;19:271–277. doi: 10.1016/j.molcel.2005.06.010. [DOI] [PubMed] [Google Scholar]
- Sun Z. W., Allis C. D. Ubiquitination of histone H2B regulates H3 methylation and gene silencing in yeast. Nature. 2002;418:104–108. doi: 10.1038/nature00883. [DOI] [PubMed] [Google Scholar]
- Sung P., Prakash S., Prakash L. The RAD6 protein of Saccharomyces cerevisiae polyubiquitinates histones, and its acidic domain mediates this activity. Genes Dev. 1988;2:1476–1485. doi: 10.1101/gad.2.11.1476. [DOI] [PubMed] [Google Scholar]
- Wood A., et al. Bre1, an e3 ubiquitin ligase required for recruitment and substrate selection of rad6 at a promoter. Mol. Cell. 2003a;11:267–274. doi: 10.1016/s1097-2765(02)00802-x. [DOI] [PubMed] [Google Scholar]
- Wood A., Schneider J., Dover J., Johnston M., Shilatifard A. The Paf1 complex is essential for histone monoubiquitination by the Rad6/Bre1 complex, which signals for histone methylation by COMPASS and Dot1p. J. Biol. Chem. 2003b;278:34739–34742. doi: 10.1074/jbc.C300269200. [DOI] [PubMed] [Google Scholar]
- Wood A., Schneider J., Dover J., Johnston M., Shilatifard A. The Bur1/Bur2 complex is required for histone H2B monoubiquitination by Rad6/Bre1 and histone methylation by COMPASS. Mol. Cell. 2005;20:589–599. doi: 10.1016/j.molcel.2005.09.010. [DOI] [PubMed] [Google Scholar]
- Wyce A., Xiao T., Whelan K. A., Kosman C., Walter W., Eick D., Hughes T. R., Krogan N. J., Strahl B. D., Berger S. L. H2B ubiquitylation acts as a barrier to Ctk1 nucleosomal recruitment prior to removal by Ubp8 within a SAGA-related complex. Mol. Cell. 2007;27:275–288. doi: 10.1016/j.molcel.2007.01.035. [DOI] [PubMed] [Google Scholar]
- Xiao T., Kao C. F., Krogan N. J., Sun Z. W., Greenblatt J. F., Osley M. A., Strahl B. D. Histone H2B ubiquitylation is associated with elongating RNA polymerase II. Mol. Cell Biol. 2005;25:637–651. doi: 10.1128/MCB.25.2.637-651.2005. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yaglom J., Linskens M. H., Sadis S., Rubin D. M., Futcher B., Finley D. p34Cdc28-mediated control of Cln3 cyclin degradation. Mol. Cell. Biol. 1995;15:731–741. doi: 10.1128/mcb.15.2.731. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yamashita K., Shinohara M., Shinohara A. Rad6-Bre1-mediated histone H2B ubiquitylation modulates the formation of double-strand breaks during meiosis. Proc. Natl. Acad. Sci. USA. 2004;101:11380–11385. doi: 10.1073/pnas.0400078101. [DOI] [PMC free article] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.