Abstract
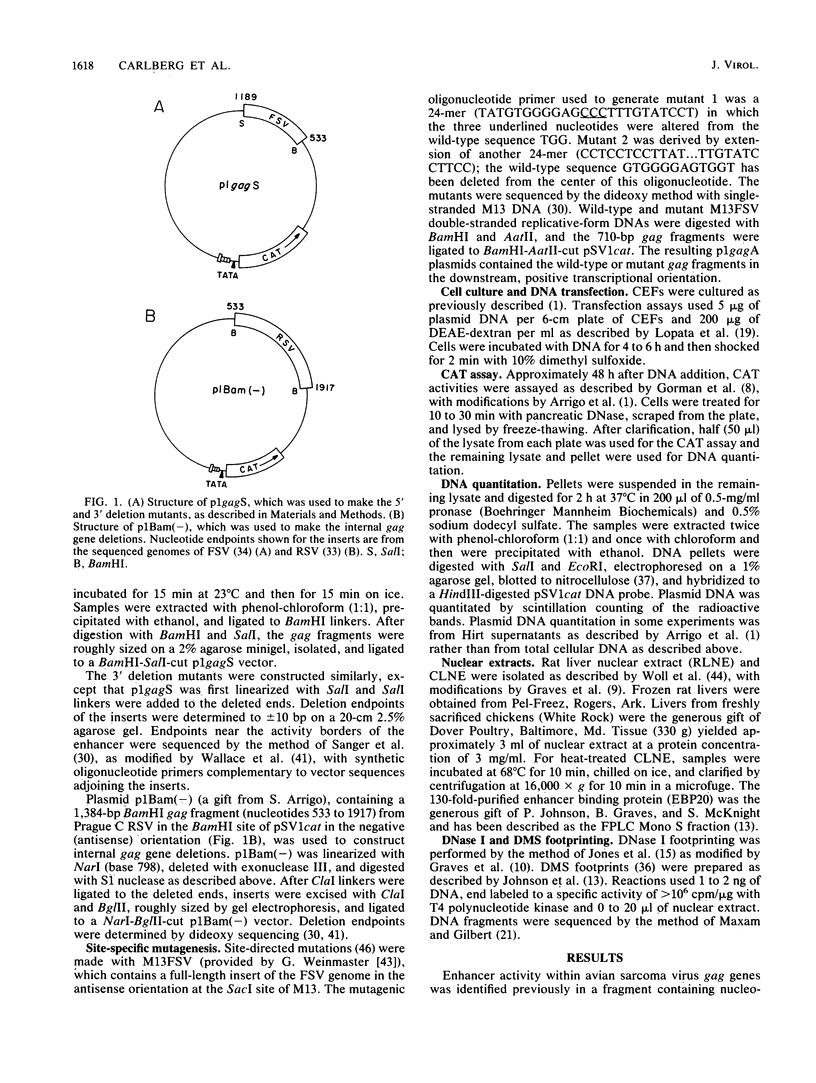
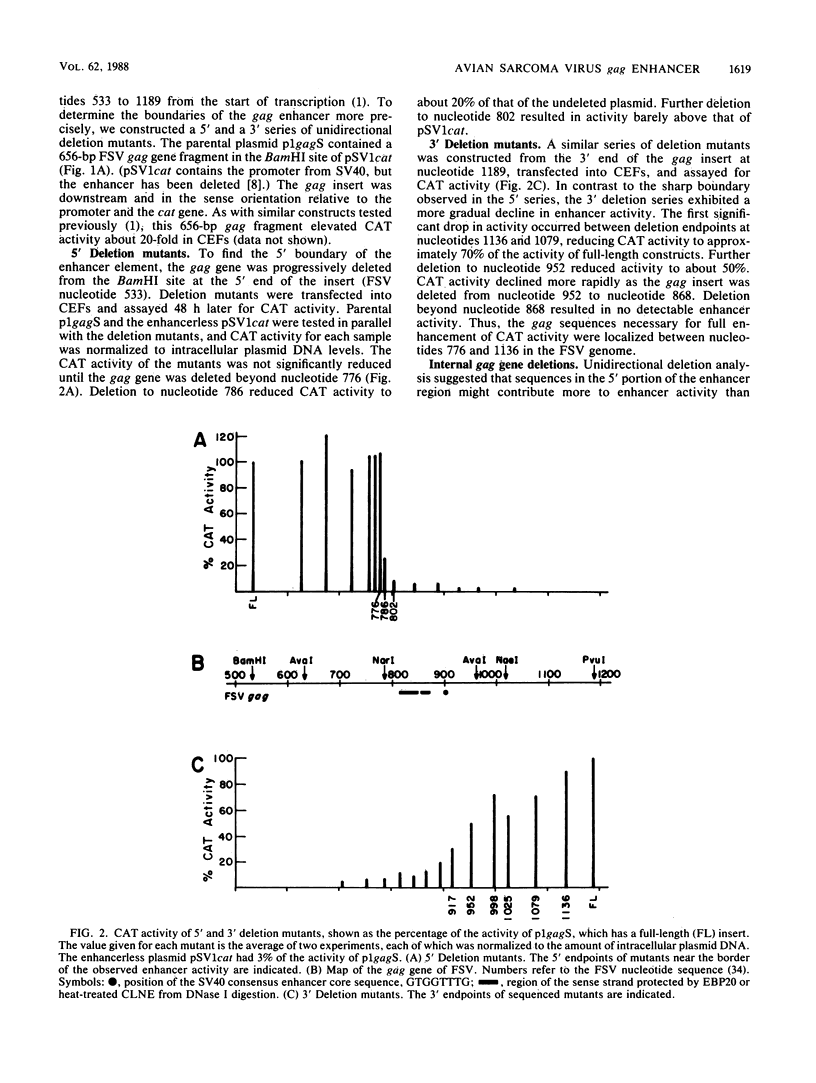
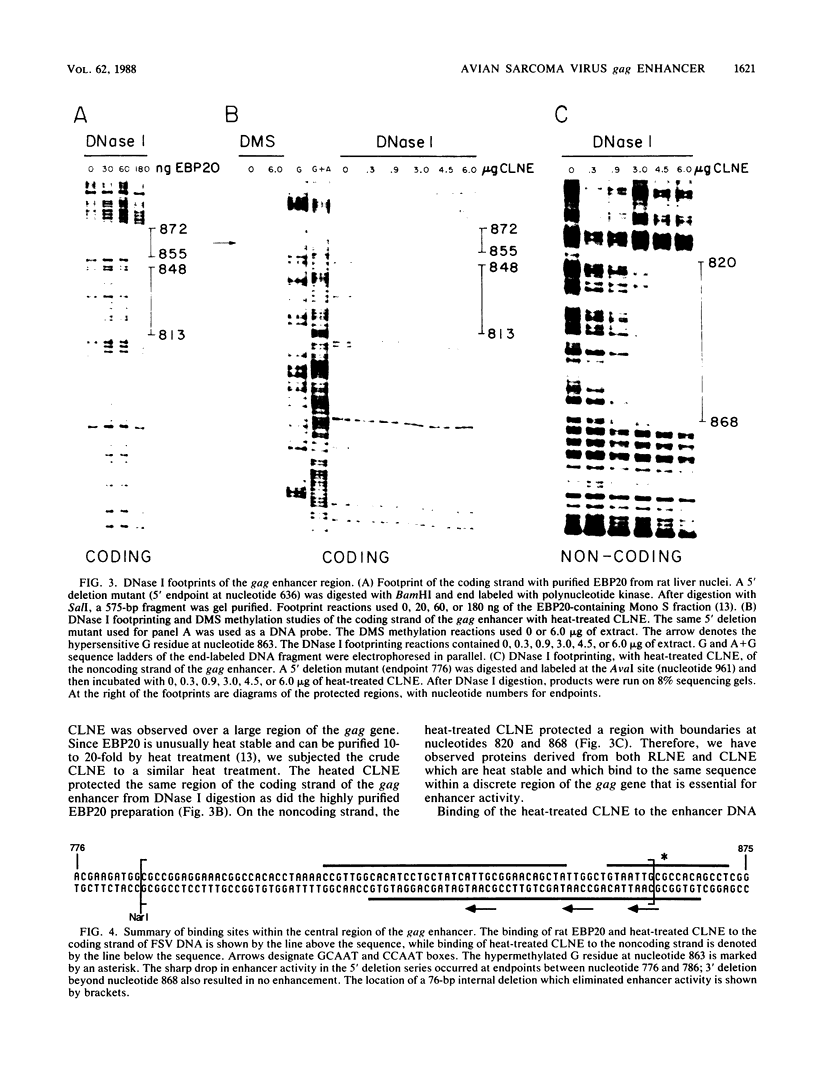
A cis-acting regulatory element within the gag gene of avian retroviruses has been localized by deletion analysis, and sites of protein interaction have been studied by DNase I footprinting. Unidirectional deletions were made from both the 5' and 3' ends of a 656-base-pair fragment of the gag gene of Fujinami sarcoma virus. These deletion mutants were tested for enhancer activity in a chloramphenicol acetyltransferase transient expression assay. A sharp 5' boundary for enhancer activity was observed between 776 and 786 nucleotides downstream from the transcription initiation site. In contrast, deletion from the 3' side resulted in a gradual loss of enhancer activity, reaching a near basal level of activity by nucleotide 868. Internal deletion of 76 nucleotides just downstream of the 5' boundary abolished enhancement. Mutagenesis of a consensus enhancer core sequence (GTGGTTTG) showed that this sequence was not necessary for enhancer activity in our transient assays. DNase I footprinting with both a highly purified enhancer-binding protein from rat liver (EBP20) and a partially purified chicken liver nuclear extract showed specific protection of nucleotides 813 to 872 within the localized enhancer region. Footprinting of unidirectional deletion mutants that had lost activity indicated that this binding was not sufficient to confer enhancement.
Full text
PDF





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Arrigo S., Yun M., Beemon K. cis-acting regulatory elements within gag genes of avian retroviruses. Mol Cell Biol. 1987 Jan;7(1):388–397. doi: 10.1128/mcb.7.1.388. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cullen B. R., Raymond K., Ju G. Transcriptional activity of avian retroviral long terminal repeats directly correlates with enhancer activity. J Virol. 1985 Feb;53(2):515–521. doi: 10.1128/jvi.53.2.515-521.1985. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dynan W. S., Tjian R. The promoter-specific transcription factor Sp1 binds to upstream sequences in the SV40 early promoter. Cell. 1983 Nov;35(1):79–87. doi: 10.1016/0092-8674(83)90210-6. [DOI] [PubMed] [Google Scholar]
- Ephrussi A., Church G. M., Tonegawa S., Gilbert W. B lineage--specific interactions of an immunoglobulin enhancer with cellular factors in vivo. Science. 1985 Jan 11;227(4683):134–140. doi: 10.1126/science.3917574. [DOI] [PubMed] [Google Scholar]
- Galas D. J., Schmitz A. DNAse footprinting: a simple method for the detection of protein-DNA binding specificity. Nucleic Acids Res. 1978 Sep;5(9):3157–3170. doi: 10.1093/nar/5.9.3157. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Goodenow M. M., Hayward W. S. 5' long terminal repeats of myc-associated proviruses appear structurally intact but are functionally impaired in tumors induced by avian leukosis viruses. J Virol. 1987 Aug;61(8):2489–2498. doi: 10.1128/jvi.61.8.2489-2498.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorman C. M., Merlino G. T., Willingham M. C., Pastan I., Howard B. H. The Rous sarcoma virus long terminal repeat is a strong promoter when introduced into a variety of eukaryotic cells by DNA-mediated transfection. Proc Natl Acad Sci U S A. 1982 Nov;79(22):6777–6781. doi: 10.1073/pnas.79.22.6777. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Gorman C. M., Moffat L. F., Howard B. H. Recombinant genomes which express chloramphenicol acetyltransferase in mammalian cells. Mol Cell Biol. 1982 Sep;2(9):1044–1051. doi: 10.1128/mcb.2.9.1044. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves B. J., Eisenman R. N., McKnight S. L. Delineation of transcriptional control signals within the Moloney murine sarcoma virus long terminal repeat. Mol Cell Biol. 1985 Aug;5(8):1948–1958. doi: 10.1128/mcb.5.8.1948. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Graves B. J., Johnson P. F., McKnight S. L. Homologous recognition of a promoter domain common to the MSV LTR and the HSV tk gene. Cell. 1986 Feb 28;44(4):565–576. doi: 10.1016/0092-8674(86)90266-7. [DOI] [PubMed] [Google Scholar]
- Herr W., Clarke J. The SV40 enhancer is composed of multiple functional elements that can compensate for one another. Cell. 1986 May 9;45(3):461–470. doi: 10.1016/0092-8674(86)90332-6. [DOI] [PubMed] [Google Scholar]
- Ishii S., Xu Y. H., Stratton R. H., Roe B. A., Merlino G. T., Pastan I. Characterization and sequence of the promoter region of the human epidermal growth factor receptor gene. Proc Natl Acad Sci U S A. 1985 Aug;82(15):4920–4924. doi: 10.1073/pnas.82.15.4920. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Johnson P. F., Landschulz W. H., Graves B. J., McKnight S. L. Identification of a rat liver nuclear protein that binds to the enhancer core element of three animal viruses. Genes Dev. 1987 Apr;1(2):133–146. doi: 10.1101/gad.1.2.133. [DOI] [PubMed] [Google Scholar]
- Jones K. A., Kadonaga J. T., Rosenfeld P. J., Kelly T. J., Tjian R. A cellular DNA-binding protein that activates eukaryotic transcription and DNA replication. Cell. 1987 Jan 16;48(1):79–89. doi: 10.1016/0092-8674(87)90358-8. [DOI] [PubMed] [Google Scholar]
- Jones K. A., Yamamoto K. R., Tjian R. Two distinct transcription factors bind to the HSV thymidine kinase promoter in vitro. Cell. 1985 Sep;42(2):559–572. doi: 10.1016/0092-8674(85)90113-8. [DOI] [PubMed] [Google Scholar]
- Laimins L. A., Khoury G., Gorman C., Howard B., Gruss P. Host-specific activation of transcription by tandem repeats from simian virus 40 and Moloney murine sarcoma virus. Proc Natl Acad Sci U S A. 1982 Nov;79(21):6453–6457. doi: 10.1073/pnas.79.21.6453. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Laimins L. A., Tsichlis P., Khoury G. Multiple enhancer domains in the 3' terminus of the Prague strain of Rous sarcoma virus. Nucleic Acids Res. 1984 Aug 24;12(16):6427–6442. doi: 10.1093/nar/12.16.6427. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Levantis P., Gillespie D. A., Hart K., Bissell M. J., Wyke J. A. Control of expression of an integrated Rous sarcoma provirus in rat cells: role of 5' genomic duplications reveals unexpected patterns of gene transcription and its regulation. J Virol. 1986 Mar;57(3):907–916. doi: 10.1128/jvi.57.3.907-916.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lopata M. A., Cleveland D. W., Sollner-Webb B. High level transient expression of a chloramphenicol acetyl transferase gene by DEAE-dextran mediated DNA transfection coupled with a dimethyl sulfoxide or glycerol shock treatment. Nucleic Acids Res. 1984 Jul 25;12(14):5707–5717. doi: 10.1093/nar/12.14.5707. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Luciw P. A., Bishop J. M., Varmus H. E., Capecchi M. R. Location and function of retroviral and SV40 sequences that enhance biochemical transformation after microinjection of DNA. Cell. 1983 Jul;33(3):705–716. doi: 10.1016/0092-8674(83)90013-2. [DOI] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- McKeon C., Schmidt A., de Crombrugghe B. A sequence conserved in both the chicken and mouse alpha 2(I) collagen promoter contains sites sensitive to S1 nuclease. J Biol Chem. 1984 May 25;259(10):6636–6640. [PubMed] [Google Scholar]
- McKnight S. L., Gavis E. R. Expression of the herpes thymidine kinase gene in Xenopus laevis oocytes: an assay for the study of deletion mutants constructed in vitro. Nucleic Acids Res. 1980 Dec 20;8(24):5931–5948. doi: 10.1093/nar/8.24.5931. [DOI] [PMC free article] [PubMed] [Google Scholar]
- McKnight S., Tjian R. Transcriptional selectivity of viral genes in mammalian cells. Cell. 1986 Sep 12;46(6):795–805. doi: 10.1016/0092-8674(86)90061-9. [DOI] [PubMed] [Google Scholar]
- Oskarsson M., McClements W. L., Blair D. G., Maizel J. V., Vande Woude G. F. Properties of a normal mouse cell DNA sequence (sarc) homologous to the src sequence of Moloney sarcoma virus. Science. 1980 Mar 14;207(4436):1222–1224. doi: 10.1126/science.6243788. [DOI] [PubMed] [Google Scholar]
- Payvar F., DeFranco D., Firestone G. L., Edgar B., Wrange O., Okret S., Gustafsson J. A., Yamamoto K. R. Sequence-specific binding of glucocorticoid receptor to MTV DNA at sites within and upstream of the transcribed region. Cell. 1983 Dec;35(2 Pt 1):381–392. doi: 10.1016/0092-8674(83)90171-x. [DOI] [PubMed] [Google Scholar]
- Queen C., Stafford J. Fine mapping of an immunoglobulin gene activator. Mol Cell Biol. 1984 Jun;4(6):1042–1049. doi: 10.1128/mcb.4.6.1042. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Robinson H. L., Gagnon G. C. Patterns of proviral insertion and deletion in avian leukosis virus-induced lymphomas. J Virol. 1986 Jan;57(1):28–36. doi: 10.1128/jvi.57.1.28-36.1986. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sakonju S., Bogenhagen D. F., Brown D. D. A control region in the center of the 5S RNA gene directs specific initiation of transcription: I. The 5' border of the region. Cell. 1980 Jan;19(1):13–25. doi: 10.1016/0092-8674(80)90384-0. [DOI] [PubMed] [Google Scholar]
- Sanger F., Nicklen S., Coulson A. R. DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A. 1977 Dec;74(12):5463–5467. doi: 10.1073/pnas.74.12.5463. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schwartz D. E., Tizard R., Gilbert W. Nucleotide sequence of Rous sarcoma virus. Cell. 1983 Mar;32(3):853–869. doi: 10.1016/0092-8674(83)90071-5. [DOI] [PubMed] [Google Scholar]
- Schöler H. R., Gruss P. Specific interaction between enhancer-containing molecules and cellular components. Cell. 1984 Feb;36(2):403–411. doi: 10.1016/0092-8674(84)90233-2. [DOI] [PubMed] [Google Scholar]
- Shibuya M., Hanafusa H. Nucleotide sequence of Fujinami sarcoma virus: evolutionary relationship of its transforming gene with transforming genes of other sarcoma viruses. Cell. 1982 Oct;30(3):787–795. doi: 10.1016/0092-8674(82)90283-5. [DOI] [PubMed] [Google Scholar]
- Siebenlist U., Simpson R. B., Gilbert W. E. coli RNA polymerase interacts homologously with two different promoters. Cell. 1980 Jun;20(2):269–281. doi: 10.1016/0092-8674(80)90613-3. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Stoltzfus C. M., Chang L. J., Cripe T. P., Turek L. P. Efficient transformation by Prague A Rous sarcoma virus plasmid DNA requires the presence of cis-acting regions within the gag gene. J Virol. 1987 Nov;61(11):3401–3409. doi: 10.1128/jvi.61.11.3401-3409.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Swift R. A., Boerkoel C., Ridgway A., Fujita D. J., Dodgson J. B., Kung H. J. B-lymphoma induction by reticuloendotheliosis virus: characterization of a mutated chicken syncytial virus provirus involved in c-myc activation. J Virol. 1987 Jul;61(7):2084–2090. doi: 10.1128/jvi.61.7.2084-2090.1987. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Wallace R. B., Johnson M. J., Suggs S. V., Miyoshi K., Bhatt R., Itakura K. A set of synthetic oligodeoxyribonucleotide primers for DNA sequencing in the plasmid vector pBR322. Gene. 1981 Dec;16(1-3):21–26. doi: 10.1016/0378-1119(81)90057-3. [DOI] [PubMed] [Google Scholar]
- Weiher H., König M., Gruss P. Multiple point mutations affecting the simian virus 40 enhancer. Science. 1983 Feb 11;219(4585):626–631. doi: 10.1126/science.6297005. [DOI] [PubMed] [Google Scholar]
- Weinmaster G., Zoller M. J., Smith M., Hinze E., Pawson T. Mutagenesis of Fujinami sarcoma virus: evidence that tyrosine phosphorylation of P130gag-fps modulates its biological activity. Cell. 1984 Jun;37(2):559–568. doi: 10.1016/0092-8674(84)90386-6. [DOI] [PubMed] [Google Scholar]
- Woll W. W., Duffy J. J., Giese N. A., Lindell T. J. Nuclear isolation by a modified method of Hewish and Burgoyne: implications for the study of nuclear enzymology. Life Sci. 1981 Dec 28;29(26):2709–2719. doi: 10.1016/0024-3205(81)90530-0. [DOI] [PubMed] [Google Scholar]
- Zenke M., Grundström T., Matthes H., Wintzerith M., Schatz C., Wildeman A., Chambon P. Multiple sequence motifs are involved in SV40 enhancer function. EMBO J. 1986 Feb;5(2):387–397. doi: 10.1002/j.1460-2075.1986.tb04224.x. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zoller M. J., Smith M. Oligonucleotide-directed mutagenesis: a simple method using two oligonucleotide primers and a single-stranded DNA template. DNA. 1984 Dec;3(6):479–488. doi: 10.1089/dna.1.1984.3.479. [DOI] [PubMed] [Google Scholar]




