Abstract
Cell fusion in yeast is the process by which two haploid cells fuse to form a diploid zygote. To dissect the pathway of cell fusion, we phenotypically and genetically characterized four cell fusion mutants, fus6/spa2, fus7/rvs161, fus1, and fus2. First, we examined the complete array of single and double mutants. In all cases but one, double mutants exhibited stronger cell fusion defects than single mutants. The exception was rvs161Δ fus2Δ, suggesting that Rvs161p and Fus2p act in concert. Dosage suppression analysis showed that Fus1p and Fus2p act downstream or parallel to Rvs161p and Spa2p. Second, electron microscopic analysis was used to define the mutant defects in cell fusion. In wild-type prezygotes vesicles were aligned and clustered across the cell fusion zone. The vesicles were associated with regions of cell wall thinning. Analysis of Fus− zygotes indicated that Fus1p was required for the normal localization of the vesicles to the zone of cell fusion, and Spa2p facilitated their clustering. In contrast, Fus2p and Rvs161p appeared to act after vesicle positioning. These findings lead us to propose that cell fusion is mediated in part by the localized release of vesicles containing components essential for cell fusion.
INTRODUCTION
The formation of one cell from the fusion of two progenitor cells is a perilous biological event. Cell fusion must be both complete and precise, allowing cytoplasmic continuity while avoiding cell lysis. Cell fusion is an essential process for the propagation of eukaryotic species that include a sexual mode of reproduction, as well as for developmental events such as myoblast fusion to form muscle cells (Doberstein et al., 1997).
Mating of the yeast Saccharomyces cerevisiae is an ideal system for studying the events and regulation of cell fusion. Conjugation in yeast is a relatively simple process in which two haploid cells of opposite mating type, a and α, fuse to form a diploid zygote (for review, see Sprague and Thorner, 1992; Marsh and Rose, 1997). The process begins when specific pheromones (a-factor and α-factor) produced by each cell type reciprocally stimulate a cascade of regulatory and morphological events essential to efficient mating. In response to pheromone binding, activation of an MAP kinase signal transduction system leads to the induction of a variety of mating specific genes (for reviews, see Errede et al., 1995; Herskowitz, 1995; Levin and Errede, 1995). The pheromone-stimulated cells agglutinate via surface glycoproteins (for review, see Lipke and Kurjan, 1992), arrest the cell cycle at G1 (for review, see Sprague and Thorner, 1992), and initiate directional cell growth toward a selected mating partner (for reviews, see Chenevert, 1994; Leberer et al., 1997). Once contact between partner cells is secured, the mating pair or prezygote undergoes cell fusion (for review, see Marsh and Rose, 1997). Finally, the two haploid nuclei fuse to form a single diploid nucleus in a process known as karyogamy (for reviews, see Rose, 1996; Marsh and Rose, 1997).
In S. cerevisiae mating is not essential for viability, and cells can be propagated asexually. Therefore, cell and nuclear fusion mutants can be generated without dire consequences to the organism. Characterization of such mutants has facilitated the identification of several genes involved in the efficiency and control of mating (Marsh and Rose, 1997). However, the genes required for cell fusion in yeast conjugation are only beginning to be understood. On a broad level, cell fusion requires the remodeling and removal of the intervening cell wall followed by the fusion of the two plasma membranes. In principle, failure to coordinate cell wall removal with cell fusion should be lethal, because the cells would become osmotically sensitive. Therefore, it seems likely that the regulation of cell fusion will be as critical to the cell as the mechanism. Pheromone levels appear to be at least one signal that is required for efficient initiation of cell fusion. Strains that produce reduced levels of pheromone accumulate prezygotes during mating (Brizzio et al., 1996). In addition, certain mutations in the a-factor pheromone transporter Ste6p also result in cell fusion defects (Elia and Marsh, 1996). The signal for cell fusion is likely to be propagated through the pheromone-stimulated protein kinase cascade, because the Fus3p kinase is required for efficient cell fusion (Elion et al., 1990, 1993; Fujimura, 1990a,b). Finally, cell fusion is influenced by protein kinase C, which is thought to be monitoring the osmotic state of the prezygote (Philips and Herskowitz, 1997).
Several genes have been identified that are likely to be involved in the actual mechanism of cell fusion. Upon pheromone addition, the cell fusion proteins expressed by FUS1 (McCaffrey et al., 1987; Trueheart et al., 1987; Trueheart and Fink, 1989), FUS2 (Trueheart et al., 1987; Elion et al., 1995), and RVS161/FUS7 (Brizzio et al., 1998) are induced and localized to the zone of cell fusion. Although their specific functions are not yet known, the regulation and localization of these proteins suggest that they might play a direct role in the process.
Other proteins have been identified that are likely to play a more indirect role in cell fusion. For example, mutations in certain genes required for membrane trafficking such as TPM1 (Liu and Bretscher, 1992) and CHS5 (discussed by Santos et al., 1997) cause an accumulation of prezygotes. In addition, genes implicated in cell polarization, including BNI1, SPA2, and PEA2, have been reported to have roles in cell fusion (Dorer et al., 1997). It is likely that these proteins are required for the localization or delivery of cell fusion specific components to the zone of cell fusion.
We have taken a genetic and ultrastructural approach to characterizing cell fusion in yeast. Our findings suggest that cell fusion is likely to occur through at least two parallel pathways, with Rvs161p and Fus2p defining one branch. From our ultrastructural analysis we concluded that cell fusion is likely to be mediated by vesicles that become clustered and aligned across the zone of cell fusion. We found that Fus1p appears to be important in localizing the vesicles to the zone of cell fusion and that Spa2p facilitates their clustering. The findings presented in this article suggest that both membrane trafficking and polarization are essential to the efficiency of the process and therefore play important roles in the mechanism of cell fusion.
MATERIALS AND METHODS
DNA Manipulations and Strain Construction
Yeast strains and plasmids used in the study are listed in Table 1. All standard plasmid manipulation and PCR methods were conducted essentially as described elsewhere (Sambrook et al., 1989). Yeast techniques were conducted according to published procedures (Rose et al., 1990). Lithium acetate transformations of yeast cells were conducted as described previously (Ito et al., 1983).
Table 1.
Strains and plasmids used in this study
| Strain | Genotype | Source |
|---|---|---|
| MY2787 | MATα ura3-52 leu2-Δ1 lys2-Δ202 | This laboratorya |
| MY2792 | MATα ura3-52 leu2-Δ1 his3-Δ200 | This laboratory |
| MY3371 | MATa ura3-52 leu2-Δ1 | This laboratory |
| MY3608 | MATa leu2 ura3-35 spa2-964 | This laboratory |
| MY3773 | MATα spa2-964 lys2-Δ202 ura3-52 | This laboratory |
| MY3776 | MATa leu2 ura3-52 spa2-Δ3::URA3 | This laboratory |
| MY3785 | MATα rvs161-1811 leu2 lys2-Δ202 ura3-52 | This laboratory |
| MY3905 | MATα rvs161-Δ1::LEU2 ura3-52 leu2-Δ1 his3-Δ200 | This laboratory |
| MY3909 | MATa rvs161-Δ1::LEU2 ura3-52 his3-Δ200 leu2-Δ1 trp1-Δ63 | This laboratory |
| MY4158 | MATa ura3-52 leu2 fus2-410 | This laboratory |
| MY4160 | MATa fus1-Δ1 fus2-Δ ura3-52 lys2-Δ202 | This laboratary |
| MY4161 | MATa trp1-Δ1 ura3-52 fus1-Δ1 | This laboratory |
| MY4164 | MATα trp1-Δ1 ura3-52 fus1-Δ1 | This laboratory |
| MY4178 | MATα ura3-52 leu2 lys2-Δ202 fus2-410 | This laboratory |
| MY4477 | MATa ura3-52 leu2 his3-Δ200 rvs161-1811 | This laboratory |
| MY4495 | MATα ura3-52 leu2 lys2-801 his3-Δ200 rvs161-Δ1::LEU2 | This laboratory |
| MY4742 | MATa fus2-Δ3 his4-Δ34 leu2-3 leu2-112 ura3-52 rvs161-Δ2::URA3 | This laboratory |
| MY4744 | MATα fus2-Δ3 his4-Δ34 trp1-Δ1 ura3-52 canr rvs161-Δ2::URA3 | This laboratory |
| MY4746 | MATa leu2 ura3-52 spa2-964 rvs161-Δ2::URA3 | This laboratory |
| MY4748 | MATα lys2-Δ202 ura3-52 spa2-964 rvs161-Δ2::URA3 | This laboratory |
| MY4801 | MATa fus2-Δ3 rvs161-Δ2::URA3 his4-Δ34 leu2-3 leu2-112 ura3-52 | This laboratory |
| MY4802 | MATα fus2-Δ3 rvs161-Δ2::URA3 his4-Δ34 trp1-Δ1 ura3-52 canr | This laboratory |
| MY4813 | MATa spa2-964 fus2-Δ::URA3 ura3-52 leu2 | This laboratory |
| MY4815 | MATα spa2-964 fus2-Δ::URA3 ura3-52 lys2-Δ202 | This laboratory |
| MY4817 | MATa spa2-964 fus1-Δ1 ura3-52 leu2 | This laboratory |
| MY4819 | MATα spa2-964 fus1-Δ1 ura3-52 lys2-Δ202 | This laboratory |
| MY4843 | MATα trp1-Δ1 ura3-52 fus1-Δ1 fus2-Δ3 p0 | This laboratory |
| MY4859 | MATa ura3-52 his3-Δ200 leu2-Δ1 trp1-Δ63 rvs161-Δ::LEU2 fus2-Δ::URA3 | This laboratory |
| MY4905 | MATa ura3-52 his3-Δ200 leu2-Δ1 trp1-Δ63 rvs1671::LEU2 fus1-Δ1 | This laboratory |
| MY4907 | MATα leu2 ura3-52 his3-Δ200 lys2-801 rvs161::LEU2 fus1-Δ1 | This laboratory |
| MY5040 | MATα leu2 ura3-52 his3-Δ200 lys2-801 rvs161::LEU2 fus2-Δ::URA3 | This laboratory |
| MY5062 | MATa spa2-964 rvs161-Δ2::ura3 leu2 ura3-52 | This laboratory |
| MY5067 | MATa spa2-964 fus2-Δ::ura3 leu2 ura3-52 | This laboratory |
| MY5224 | MATa ura3-52 his3-Δ200 leu2-Δ1 trp1-Δ63 rvs161-P203Q | This laboratory |
| MY5359 | MATα ura3-52 leu2-Δ1 rvs161-P203Q | This laboratory |
| MS2073 | MAT::LEU2 leu2-3 leu2-112 ura3-52 his3::TRP1 trp1-Δ1 | This laboratory |
| MS2104 | MAT::LEU2 leu2-3 leu2-112 ura3-52 his3::TRP1 trp1-Δ1 [MATα URA3] | This laboratory |
| MS2679 | MAT::LEU2 fus6-964 leu2-3 leu2-112 ura3-52 his3::TRP1 trp1-Δ1 [MATα URA3] | This laboratory |
| MS2680 | MAT::LEU2 fus6-964 leu2-3 leu2-112 ura3-52 his3::TRP1 trp1-Δ1 | This laboratory |
| MS2749 | MAT::LEU2 fus2-410 leu2-3 leu2-112 ura3-52 his3::TRP1 trp1-Δ1 [MATα URA3] | This laboratory |
| Y601 | MATα spa2-Δ3::URA3 ura3-52 his3-δ200 trp1-901 ade2-101 | M. Snyder, Yale University |
| JY424 | MATa fus2-Δ3 his4-Δ34 leu2-3 leu2-112 ura3-52 | G. Fink, Whitehead Institute |
| JY427 | MATa fus1-Δ1 leu2-3 leu2-112 ura3-52 | G. Fink, Whitehead Institute |
| JY428 | MATα fus2-Δ3 his4-Δ34 trp1-Δ1 ura3-52 canr | G. Fink, Whitehead Institute |
| JY429 | MATα trp1-Δ1 ura3-52 fus1-Δ1 fus2-Δ3 | G. Fink, Whitehead Institute |
| Plasmid | Yeast markers | Source |
| pMR3093 | SPA2 CEN ARS URA3 | This laboratory |
| pMR3234 | RVS161 CEN ARA URA3 | This laboratory |
| pMR3397 | RVS161 2μ URA3 | This laboratory |
| pMR3245 | rvs161-Δ1::LEU2 | This laboratory |
| pMR3261 | rvs161-Δ2::URA3 | This laboratory |
| pSB245 | FUS1 CEN ARS URA3 | G. Fink, Whitehead Institute |
| pSB257 | FUS2 2μ URA3 | G. Fink, Whitehead Institute |
| pSB265 | FUS2 CEN ARS URA3 | G. Fink, Whitehead Institute |
| pSB273 | FUS1 2μ URA3 | G. Fink, Whitehead Institute |
| pSB267 | fus2-Δ::URA3 | G. Fink, Whitehead Institute |
| pSB281 | fus1-Δ1 URA3 | G. Fink, Whitehead Institute |
| p182 | SPA2 2μ URA3 | M. Snyder, Yale University |
| pTS395 | PGAL-GFP CEN ARS URA3 | T. Stearns, Stanford University |
| pRS426 | 2μ URA3 | P. Hieter, Johns Hopkins |
| pRS424 | 2μ TRP1 | P. Hieter, Johns Hopkins |
The MS strains from the Rose Laboratory are congenic with S288C (hap2 mal kss1 gal2). The MY strains are also congenic with S288C, except that they are GAL2.
Generation of the rvs161Δ strains was done by the one-step gene replacement method (Rothstein, 1991) after linearizing pMR3245 or pMR3261 with HpaI (Brizzio et al., 1998). The following rvs161Δ strains, MY3905, MY4746, MY4742, MY4744, and MY4748, were constructed by this method from MY2792, MY3608, JY424, JY428, and MY3773, respectively.
The fus1Δ was done by the two-step gene replacement method (Scherer and Davis, 1979) after linearizing pSB281 (Fink laboratory, Whitehead Institute, Cambridge, MA) with KpnI. MY4817, MY4819, MY4905, and MY4907 were constructed by this method from MY3608, MY3773, MY3909, and MY4495, respectively. The fus2Δ::URA3 was constructed by the one-step gene replacement method after liberating the 3.2-kb ClaI-NruI fragment from pSB267 (Fink laboratory). MY4813, MY4815, MY4859, and MY5040 were constructed by this method from MY3608, MY3773, MY3909, and MY4495, respectively. We confirmed the fus1Δ and fus2Δ::URA3 deletions by PCR of potential positives. The primers to confirm the deletion of FUS1 were as follows: FUS1-1 (5′-CTCTGCAGGATGCCCT-3′), FUS1-2 (5′-CAGTTGTTGTCGTCTG-3′), FUS1-3 (5′-CACGGCAAGACCCCAT-3′), and FUS1-4 (5′-CAGTCGTATTCTTGGA-3′). For the detection of the deletion of FUS2, the following primers were used: FUS2-1 (5′-GTGATCCAAGATTCAA-3′), FUS2-2 (5′-TTTAATATCTCGCACA-3′), URA3-1 (5′-AAGCAGGCTGGGAAGC-3′), and URA3-2 (5′-TGTAGCTTTCGACATG-3′). The predicted bands (or lack of bands) clearly differentiated between the wild-type and the deletion loci.
In some experiments the URA3 marker was converted to ura3-52 by spontaneous gene conversion. Selection for this event was done by plating 200 μl of a saturated culture, grown in yeast extract with peptone and dextrose (YEPD),1 onto 5-fluoro-ortic acid plates. Resistant colonies were shown to still contain the desired cell fusion mutation either by plate matings or by PCR. These strains were then the recipients of URA3-containing plasmids. MY4801, MY4802, MY5067, and MY5062 were constructed by this method from MY4742, MY4744, MY4813, and MY4746, respectively. The ρo strain (MY4843) was constructed according to previously described methods (Rose et al., 1990).
Plate Matings and Microscopic Analysis of Cell Fusion Defects
For dosage suppression, MATa (MY4161, JY424, MY3608, and MY3909) and MATα cells (MY4164, JY428, MY3773, and MY4495) were each transformed with the following plasmids: pRS426, pMR3093, p182, pMR3234, pMR3397, pSB245, pSB273, pSB265, and pSB257. Three Ura+ colonies from each transformation were checked for their mating ability on plates. For almost all of the strains, mating to a fus1 fus2 (MY4843 or MY4160) lawn gave a good semiquantitative measure of the degree of suppression by the plasmid. For spa2, suppression was better visualized when the plasmid bearing strains were mated to a fus1 spa2 lawn (MY4817 or MY4819). For these plate matings, patches were grown on synthetic medium lacking uracil and mated to lawns for 3–5 h on YEPD and then replica printed to the appropriate medium to select for diploids. Occasionally, the partner strain was transformed with a vector plasmid bearing an auxotrophic marker to allow for selection of the diploid.
Quantitative analysis of the cell fusion phenotype was done microscopically using differential interference contrast (DIC) optics to assess the zygote morphology and 4′,6′-diamidino-2-phenylindole (DAPI) fluorescence to evaluate nuclear fusion. After determining the mating phenotype on plates, one representative transformant from each mating type was used in quantitative mating assays. The MATa and MATα strains were grown in synthetic complete medium lacking uracil to maintain the plasmids listed above. Filter matings were performed for 2.5 h on YEPD plates at 30°C as described previously (Brizzio et al., 1996).
The assay to evaluate cytoplasmic mixing in zygotes was such that one partner (MATa) expressed green fluorescent protein (GFP) as a soluble cytoplasmic marker. We transformed mutant and wild-type MATa ura3 strains with pTS395 (PGAL-GFP CEN ARS URA3; from T. Stearns, Stanford University, Stanford, CA). To induce the expression of GFP, the MATa cells were subcultured overnight in synthetic complete medium lacking uracil with 2% raffinose and 2% galactose at 23°C. The MATα partners were cultured in synthetic complete medium with 2% glucose. Filter matings were as described previously (Brizzio et al., 1996). For the 2.5-h matings, the filters were placed on YEPD (glucose-containing rich medium). Under these conditions, the glucose repression of PGAL-GFP did not significantly reduce the fluorescent GFP signal. The 8-h matings were conducted on plates with raffinose and galactose as carbon sources to maintain GFP expression in the cells. The mating mixtures were briefly fixed (5–15 min) in 4% formaldehyde and washed several times in PBS. Zygotes were identified using DIC optics, and the extent of mixing was scored by GFP fluorescence using a High Q FITC filter set (41001; Chroma Technology, Brattleboro, VT). We found that analysis of the cell fusion phenotype by cytoplasmic mixing of GFP was a more sensitive assay to quantify cell fusion defects than the DAPI–DIC analysis described above. The DAPI–DIC analysis of zygote and nuclear morphology led to an overestimation of the severity of the cell fusion phenotype for mutants such as fus2, which also has a nuclear fusion defects (Elion et al., 1995) (compare Figures 1 and 2B).
Figure 1.
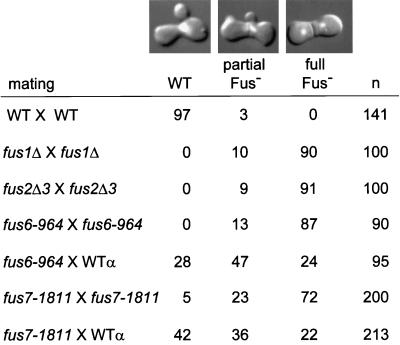
Defects of the cell fusion mutants. Filter matings were analyzed microscopically using DIC optics in combination with DAPI, a fluorescent DNA marker. The classes of zygotes scored were WT, which had no septum at the zone of fusion and a fused nucleus; partially defective (partial Fus−), which showed an obvious cell fusion septum and a fused nucleus; and completely defective (full Fus−), which had a pronounced septum and unfused nuclei. The percentages of each type of zygote are shown. The far right column shows the number of zygotes scored. The counting error is within 5%. The matings used in this analysis were WT × WT (MY3371 × MY2787), fus1Δ × fus1Δ (MY4161 × MY4164), fus2Δ × fus2Δ (JY424 × JY428), fus6-964 × fus6-964 (MS2680 × MS2679), fus6-964 × WT (MS2680 × MS2104), fus7-1811 × fus7-1811 (MY4477 × MY3785), and fus7-1811 × WT (MY4477 × MY2787). The photographs of the DIC–DAPI double images and the data from the mating between MY4477 × MY3785 have been reported elsewhere (Brizzio et al., 1998).
Figure 2.
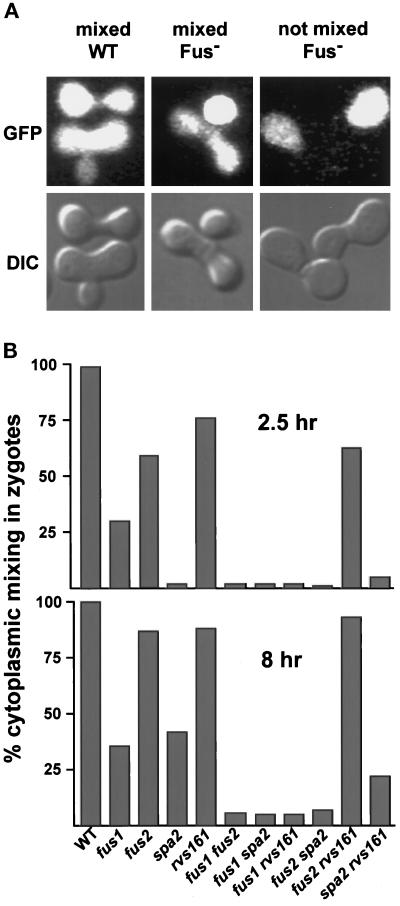
Cytoplasmic mixing in zygotes using a soluble fluorescent marker, GFP. (A) Three representative types of zygotes seen in the cytoplasmic mixing assay. The zygotes with mixed cytoplasms consisted of those with no obvious zone of cell fusion remaining (mixed WT) and zygotes with a cell fusion defect (mixed Fus−). Prezygotes with no removal of the intervening cell wall (not mixed Fus−) were also scored. The percentages of mixed zygotes (includes the mixed WT and the mixed Fus−) are shown in B. For each mating, a minimum 100 zygotes were counted. The counting error is within 5%. The matings were as follows: WT (MY3371 × MY2787), fus1 (JY427 × MY4164), fus2 (JY424 × JY428), spa2 (MY3608 × MY3773), rvs161 (MY3909 × MY4495), fus1 fus2 (MY4160 × JY429), fus1 spa2 (MY4817 × MY4819), fus1 rvs161 (MY4905 × MY4907), fus2 spa2 (MY5067 × MY4815), fus2 rvs161 (MY4801 × MY4802), and spa2 rvs161 (MY5062 × MY4748).
Cloning and Linkage of FUS6
MS2679, the original fus6-964 isolate, was back-crossed to MY3371 to obtain MY3608, a MATa fus6-964 strain for use in cloning the gene. The B1, C1, and C4 pools of a centromere-based genomic library (Rose et al., 1990) were transformed into MY3608 such that a total of ∼13 genome equivalents were introduced. Ura+ colonies were mated to lawns of MS2679 for 2.5–3.0 h. Diploids were selected on synthetic complete medium lacking leucine and histidine. Plasmids containing the LEU2 gene were discarded. The remaining candidates that mated well were retested and checked for the linkage of the suppression activity to the plasmid. The plasmids that passed these tests were isolated and transformed into a bacterial strain (XL1-Blue). Twelve independent plasmids contained a distinct 3.2-kb EcoRI restriction fragment. The fragment was isolated, labeled, and used to probe Prime λ-clone grid filters (obtained from Linda Riles, Washington University School of Medicine, St. Louis, MO) according to the protocol provided. The region identified on the grid filters contained SPA2, a gene known to have a bilateral mating defect. A plasmid containing only the SPA2 gene (obtained from M. Snyder, Yale University, New Haven, CT) was then shown to complement the fus6-964 defect. Linkage analysis between spa2::URA3 (M. Snyder, Yale University) and fus6-964 showed that the genes were tightly linked (for 19 tetrads, all were parental ditypes). Finally, we showed by microscopic analysis that an spa2Δ × spa2Δ mating gave a cell fusion defect identical to fus6-964. Western blotting analysis with α-Spa2p (M. Snyder, Yale University) confirmed that fus6-964 is a null allele.
Linkage of Mutation 410 to FUS2
Suppression of the cell fusion defect of mutant 410 by the FUS2 gene on a plasmid (Kurihara et al., 1994) suggested that this mutant might be an allele of FUS2. To confirm that the mutation 410 is allelic to FUS2, linkage analysis was performed. The 410 strain (MS2749) was crossed to JY424 (MATa fus2Δ). After sporulation, the tetrads were patched onto 5-fluoro-ortic acid to lose the MATα plasmid; thus all spore colonies mated as MATa cells. The spore colonies were mated to an MATα fus1 fus2 lawn (JY429) and selected on the appropriate synthetic medium. Tetrad analysis of the mating defect showed that for 33 tetrads, all of the spores were parental ditypes. The tight linkage between 410 and fus2Δ confirmed that the mutation is in FUS2 and therefore called fus2-410.
Electron Microscopy of Zygotes
Many published techniques for electron microscopy of yeast cells tend to destroy the ultrastructure of the cell wall. Therefore observations of the mechanism of cell fusion must be made where cell wall integrity has been preserved. To produce sufficient numbers of cells for electron microscopy, the limited filter mating protocol was scaled up 20-fold. Liquid cultures of a and α cells were grown in YEPD to equivalent cell densities in early exponential phase. Twenty milliliters of each culture were mixed and centrifuged to pellet the cells. All but 5 ml of the YEPD was decanted, and the cells were resuspended in the residual YEPD. The mating mixtures were filtered onto five 0.45-μm nitrocellulose filter disks (Millipore, Milford, MA). The filters were placed cell side up on a prewarmed YEPD plate for 2.5–3 h at 30°C. The cells were then rinsed off the filters with 1.5 ml of FIX (40 mM potassium phosphate, pH 7.4, 1 mM CaCl2, 1 mM MgCl2, 0.2 M sorbitol, 2% fresh gluteraldehyde). The cells were pelleted and resuspended in 1 ml of fresh FIX. The total incubation time in FIX was 30 min at room temperature. After washing three times in 50 mM potassium phosphate (pH 7.4), the samples were incubated in 4% potassium permanganate at 4°C for 4–6 h. The cells were washed four times with dH20 and then resuspended in 1 ml of 0.5–1% sodium periodate (Sigma, St. Louis, MO) for 15 min at room temperature. Treatment of the cells with 0.5–1.0% periodate did not alter cell wall appearance but significantly improved infiltration of the resin. The samples were washed once with 50 mM potassium phosphate and then resuspended in 50 mM ammonium phosphate for 15 min at room temperature. After two washes with dH20, the cells were resuspended in 2% filtered uranyl acetate and incubated at 4°C overnight with mixing. Cell dehydration was done by a series of washes in ethanol (50% ethanol and 70% ethanol for 5 min two times, 95% ethanol for 5 min, and 100% ethanol for 5 min three times). The samples were embedded in LR White resin (Polysciences, Warrington, PA). The sections were cut to 70–90 nm and stained with Reynold’s lead citrate (Reynolds, 1963).
RESULTS
SPA2 and RVS161/END6 Are Mating Type–nonspecific Cell Fusion Mutants
A bilateral mating screen conducted in our laboratory identified several mutations that resulted in blocks in cell fusion (Kurihara et al., 1994). The cell fusion mutants fell into two classes. The first class was a specific, in that the mutation in the MATa parent led to a Fus− defect, whereas the mutation had no detectable phenotype in MATα cells (Brizzio et al., 1996). The second class showed no cell type specificity for the cell fusion phenotypes. Two genes in this class, FUS1 (McCaffrey et al., 1987; Trueheart et al., 1987; Trueheart and Fink, 1989) and FUS2 (Trueheart et al., 1987; Elion et al., 1995), have been described previously. In addition to isolating an allele of FUS2 (see MATERIALS AND METHODS), we identified two new genes, FUS6 and FUS7, in the screen (Kurihara et al., 1994).
To characterize FUS6 and FUS7, we examined the phenotypes of two representative mutants, fus6-964 and fus7-1811, in comparison with wild-type, fus1 and fus2 deletion mutants (Figure 1). Mating mixtures were analyzed microscopically using DIC optics to assess the zygote morphology in combination with DAPI, a fluorescent DNA stain, to evaluate nuclear fusion. Three classes of zygotes were observed (Figure 1): wild type (WT), which had no septum and a fused nucleus; partially defective (partial Fus−), which contained an obvious septum but also showed a fused nucleus; and completely defective (full Fus−), which had a complete septum and unfused nuclei.
In contrast to the wild-type matings, fus1, fus2, fus6, and fus7 matings all contained zygotes and prezygotes with pronounced cell fusion defects (Figure 1). The quantitative data showed that the fus7 defect was less severe than the fus1, fus2, and spa2 defects. The fus6-964 and fus7-1811 mutations conferred bilateral mating defects, because the cell fusion phenotypes of the mutant by mutant matings were significantly stronger than the mutant by wild-type (unilateral) matings (Figure 1). It should be noted that the mutants had no observable defects in unilateral plate matings. However, when scored microscopically, there were obvious cell fusion defects when each mutant was mated to wild type (Figure 1). Furthermore, the unilateral defects were equally strong for both fus6-964 and fus7-1811 whether the mutation was in the a or α cell, confirming that fus6-964 and fus7-1811 were mating type–nonspecific mutants. Thus, fus6 and fus7 represented two new mutants displaying cell fusion phenotypes similar to the fus1 and fus2 mutants.
We cloned the FUS6 and FUS7 genes by complementation of the plate-mating defects (see MATERIALS AND METHODS and Brizzio et al., 1998). Molecular and genetic mapping allowed us to conclude that FUS6 and FUS7 were both previously characterized genes. FUS6 was determined to be SPA2, a gene implicated in cell polarity and morphogenesis in vegetative and mating cells (Snyder, 1989; Gehrung and Snyder, 1990; Snyder et al., 1991; Flescher et al., 1993; Chenevert et al., 1994; Zahner et al., 1996; Arkowitz and Lowe, 1997; Buehrer and Errede, 1997; Dorer et al., 1997; Xu and Kurjan, 1997). At the time of the cloning, SPA2 was reported to have a severe bilateral mating defect (Gehrung and Snyder, 1990); however, the role in cell fusion had not been determined. FUS7 was found to be allelic to RVS161 (Brizzio et al., 1998). Mutations in RVS161 are highly pleiotropic, conferring phenotypes that include reduced viability upon starvation (Crouzet et al., 1991; Desfarges et al., 1993), alterations in actin cytoskeleton organization (Sivadon et al., 1995), abnormalities in budding patterns (Durrens et al., 1995), and defects in endocytosis (Munn et al., 1995). A recent report also identified SPA2 as a gene involved in cell fusion (Dorer et al., 1997), although the nature of the defect was not investigated further.
Genetic Characterization of Single and Double Cell Fusion Mutants
To gain a better understanding of the roles that SPA2, RVS161, FUS1, and FUS2 play in cell fusion, we examined how the genes interact genetically. Using a cytoplasmic mixing assay, we assessed the severity of the cell fusion defects for the spectrum of single and double mutants. In this assay the MATa cells expressed soluble GFP, whereas the partners contained no fluorescent marker. The mating progressed for either 2.5 or 8 h. The data (Figure 2) are represented as mixed (fluorescence seen throughout the zygote) or not mixed (fluorescence seen in one-half of the zygote). Using this scoring method, the class that we previously scored as partial Fus− (see above) would be included in the mixed category. The overall conclusions of the single and double mutant analyses were the same whether the matings were analyzed on plates (Brizzio et al., 1998; our unpublished observations) or microscopically (Figure 2). However, we found that the measurement of cytoplasmic mixing in zygotes gave the best resolution for distinguishing among the cell fusion mutants (see MATERIALS AND METHODS).
Figure 2 shows that after 2.5 h of mating, the single mutants rvs161 and fus2 showed the least severe defects (76 and 56% mixing, respectively). The spa2 mutant prezygotes showed the strongest defect (only 2% mixing), whereas fus1 exhibited an intermediate phenotype (30% mixing). After 8 h of mating, the single mutants fell into two classes; spa2 and fus1 showed comparable defects (42 and 36% mixing, respectively), as did fus2 and rvs161 (87 and 88% mixing, respectively). Analysis of the double mutants indicated that in all cases except one, the double mutant exhibited a more severe phenotype than either single mutant. The sole exception was the fus2Δ rvs161Δ double mutant (Figure 2). In this case, the double mutant defect was the same as the more severe single mutant phenotype (fus2 alone).
It is important to note that for our genetic analysis the mutants were constructed with deletions (fus1, fus2, and rvs161) or null alleles (spa2-964). When constructing and analyzing the double mutants, if the cell fusion phenotype was no worse when a second deletion was introduced, then we concluded that the two proteins act in concert or in the same pathway. In particular, we concluded that for Fus2p and Rvs161p, the proteins either act together or that one facilitates the function of the other in the process of cell fusion. Finally, because a strong synthetic phenotype was found for all of the other double mutant combinations (Figure 2), we concluded that cell fusion occurs through more than one partially redundant pathway, consistent with previous proposals (Trueheart et al., 1987). In support of this view, none of the deletion mutants was completely blocked in cell fusion (Figures 1 and 2).
Fus1p and Fus2p in High Copy Suppress All of the Cell Fusion Mutants
We next exploited high-copy suppression of mutant defects as another method of analyzing genetic interactions. High-copy suppression is often taken as an indication of a gene’s ability to act downstream or in a parallel pathway. Previous studies found that, in high copy, FUS1 partially suppressed fus2 and FUS2 partially suppressed fus1 (Trueheart et al., 1987). We tested the ability of SPA2, RVS161, FUS1, and FUS2 to suppress the various cell fusion mutants either in single copy (CEN plasmid mediated) or high copy (2μ plasmid mediated). We first performed quantitative filter matings in which both partners contained the plasmid (Figure 3A). This bilateral test for suppression was particularly important for spa2 matings. To show the unilateral suppression of mutant defects on plates, we next mated the various strains to severely compromised fus1 fus2 or fus1 spa2 double mutant partners (Figure 3B).
Figure 3.
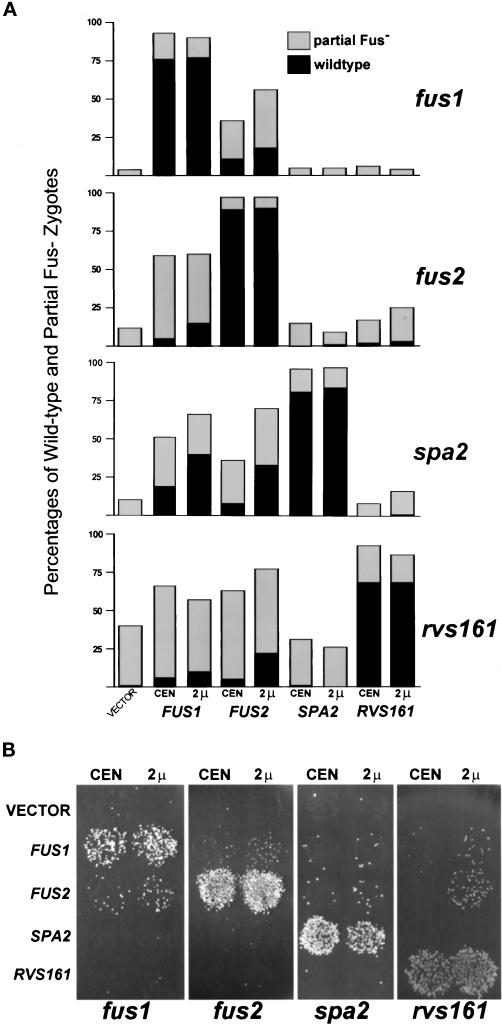
Dosage suppression of cell fusion defects. (A) Quantitation of the bilateral suppression of the cell fusion mating defects. The limited filter matings were fus1 (MY4161 × MY4164), fus2 (JY424 × JY428), spa2 (MY3608 × MY3773), and rvs161 (MY3909 × MY4495). In this analysis the MATa and MATα cells were each transformed with the following plasmids: vector, 2μ URA3 (pRS426), FUS1 CEN (pSB245), FUS1 2μ (pSB273), FUS2 CEN (pSB265), FUS2 2μ (pSB257), SPA2 CEN (pMR3093), SPA2 2μ (p182), RVS161 CEN (pMR3234)) and RVS161 2μ (pMR3397). The mating defects were scored as described for Figure 1. Black bars represent percentage of wild-type zygotes, and gray bars are for percentage of partial Fus− zygotes. A minimum of 200 zygotes were scored, and the error is within 5%. (B) Suppression of plate mating defects. The suppression of fus1, fus2, and rvs161 was tested by mating MATa cells fus1 (MY4161), fus2 (JY424), and rvs161 (MY3909) containing the plasmids listed above to a lawn of fus1 fus2 ρ0 (MY4843). The suppression of spa2 (MY3608) was determined by mating the spa2 cells containing the plasmids to a fus1 spa2 (MY4819) lawn. For the fus1 analysis MY4843 was transformed with pRS424 (2μ TRP1) to allow for selection of the diploid.
The results from both experiments showed that only FUS1 and FUS2 suppressed the cell fusion defect in all the mutants. In contrast, SPA2 and RVS161 only suppressed the cognate mutations. In all cases the suppression was dosage dependent (2μ suppressed better than CEN). However, the suppression was never complete. The fact that elevated levels of FUS1 and FUS2 can suppress all of the mutations we examined is consistent with the hypothesis that there are partially overlapping pathways for cell fusion. Furthermore, given that RVS161 and FUS2 appear to act in the same pathway, it is of interest that FUS2 suppresses rvs161Δ, but RVS161 does not suppress fus2Δ. Although our hypothesis is that FUS1 and FUS2 have the ability to act downstream, it should be noted that the failure of RVS161 and SPA2 to suppress in high copy might simply reflect the fact that these proteins are not rate limiting in the process of cell fusion.
Ultrastructural Characterization of the Diameter of the Zone of Cell Fusion in Wild-Type Mating Mixtures
To further characterize the pathway of cell fusion, we used electron microscopy methods to analyze wild-type and mutant zygotes. Because many electron microscopy fixation techniques remove the cell wall and therefore disrupt cell–cell contact during mating (Byers and Goetsch, 1975), we used a technique that preserved the cell wall and allowed for the observation of prezygotes (see MATERIALS AND METHODS). Furthermore, we used a method that is particularly useful for the visualization of membranes.
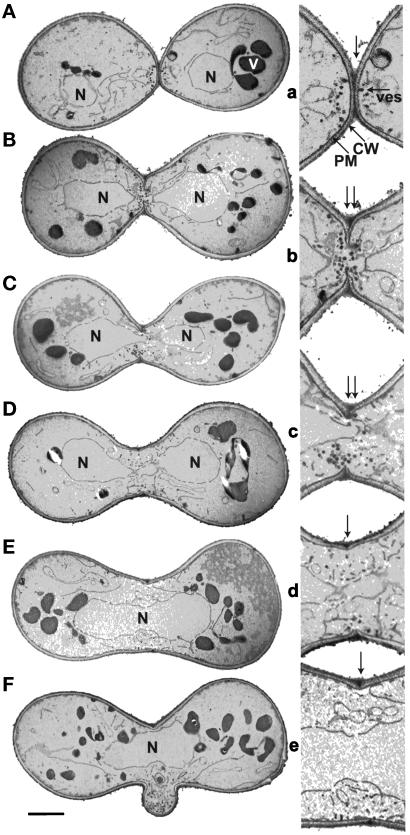
A detailed analysis of wild-type mating cells showed that prezygotes and zygotes in a mating mixture could be ordered according to their relative age by the presence of diagnostic landmarks. The first class was composed of prezygotes. The prezygotes were defined as two adhered cells with a degree of polarized growth toward each other, such that the cells became somewhat pear shaped (Figure 4A). Prezygotes also exhibited cell wall remodeling on the outer periphery of the zone of cell fusion. In addition, prezygotes often contained a region of close cell wall contact, cell wall thinning, and polarized vesicle clusters, discussed in detail below. By these criteria, a prezygote could be readily distinguished from large budded cells or from two closely apposed cells. In addition, the mother and daughter of large budded cells are more rounded and separated by cell wall material that has a distinctive chitin layer (not seen in prezygotes). The next class was the prekaryogamy zygotes (Figure 4, B and C), which had undergone cell fusion but not karyogamy. The prekaryogamy zygotes contained numerous vesicles and cell wall remnants at the cell fusion zone. The next class of zygotes was the early postkaryogamy zygotes (Figure 4D), which were defined by the presence of a fused nucleus containing an obvious constriction in the nuclear membrane. The nuclear constriction was found within the zone of cell fusion and is presumed to be the site of nuclear fusion. The postkaryogamy zygotes were also conspicuous for the absence of clustered vesicles at the cell fusion zone. When vesicles were observed (Figure 4D), they tended to be localized near the cell fusion scar. These early postkaryogamy zygotes also rarely had obvious cell wall remnants. The final class of zygotes was designated as mature zygotes and was defined either by the presence of a rounded nucleus with no constriction (Figure 4E) or by the presence of a bud (Figure 4F).
Figure 4.
Cell fusion and karyogamy in wild-type mating mixtures (MY3371 × MY2787). (A) Prezygote; (B) early zygote before karyogamy; (C) early zygote during karyogamy; (D) early zygote after karyogamy; (E) mature unbudded zygote; and (F) mature budding zygote. The bars represent 1 μm for A–F. (a–e) Twofold enlargements of the respective cell fusion zones in A–E. At least one example of each salient cellular structure is noted as follows: N, nucleus; V, vacuole; CW, cell wall; PM, plasma membrane; and ves, vesicle. The arrows in A accentuate the zone of cell fusion. The double arrows in B and C point to zones that contain cell wall remnants after fusion. The arrows in D and E emphasize the cell fusion scar.
While analyzing the zone of cell fusion in the various classes of zygotes, we found that the diameter of the zone of cell adhesion in a prezygote and the region of cell fusion in a zygote are correlated with the age of the cell. Prezygotes showed narrow zones of cell fusion, with an average diameter of 1.0 ± 0.3 μm (Figure 4a). Prekaryogamy zygotes exhibited cell fusion zones similar in size to prezygotes, with an average diameter of 1.1 ± 0.2 μm (Figure 4, b and c). In contrast, postkaryogamy zygotes showed an average diameter of 1.6 ± 0.2 μm (Figure 4d). Finally, mature zygotes contained zones of cell fusion of ∼2.5 ± 0.5 μm (Figure 4f). We conclude from this analysis that as the zygote matures, cell wall remodeling continues such that the diameter at the zone of cell fusion expands. Therefore, the diameter of the cell fusion zone is an excellent indicator of the relative age of the zygote.
Cell Fusion Mutant Prezygotes Block at the Cell Wall Removal Step
The first step in cell fusion occurs when the mating pair becomes closely associated to form a prezygote. Unlike wild type, all of the mutant prezygotes contained extended zones of cell contact (2–3 μm on average; Figures 4A and 5). The spa2 mutant was the most extreme case, with zones of cell fusion averaging three times the diameter of wild type (Figure 5). Because the diameter of the zone of cell fusion in wild type correlated with the relative age of the zygote, we concluded that the Fus− prezygotes with broad regions of cell contact (>2 μm) had been attempting cell fusion for extended periods, possibly for the entire 2.5–3 h during the limited mating. Hence, these expanded cell fusion zones observed in the mutant prezygotes are likely to be aberrant structures that arise as a result of the block in cell fusion.
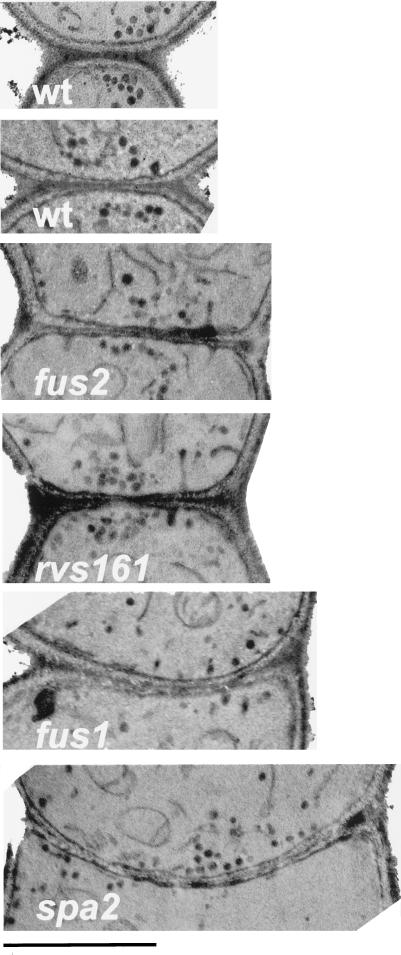
Figure 5.
Vesicle accumulation at the zone of cell fusion. The micrographs are from the following matings: WT (MY3371 × MY2787), fus2 (MY4158 × MY4178), rvs161 (MY5224 × MY5359), fus1 (MY4161 × MY4164), and spa2 (MY3608 × MY3773). The scale bar represents 1 μm.
After cell contact is secured, the intervening cell wall separating the partner cells must be removed. All of the mutants failed to efficiently remove the intervening cell wall. However, thinning of the cell wall at the zone of cell fusion was seen both in wild-type (Osumi et al., 1974) (Figure 5, second wt panel) and in Fus− zygotes (Figure 5, rvs161 panel). Our analysis of the wild-type matings suggested that the partner cells remove only a comparatively small region of cell wall at the zone of cell fusion. The “fusion pore” observed in early prekaryogamy zygotes was marked by a region 0.52 ± 0.19 μm in diameter that was initially degraded in the wild-type cell fusion event (Figure 4, B and b). For the Fus− prezygotes, the zone of cell fusion was much larger and therefore may require significantly more cell wall removal to give a phenotypically mature wild-type zygote with a smooth cell junction.
The formation of the fusion pore resulted in remnants of cell wall adjacent to the zone of cell fusion. The cell wall remnants projected into the cell, perpendicular to the long axis of the zygote. In wild-type zygotes the cell wall remnants existed only before or during karyogamy (Figure 4, b and c). In contrast, we found that all of the cell fusion mutant zygotes exhibited cell wall remnants at the zone of fusion (see Figure 7 and Brizzio et al., 1996). In wild type the remnants never projected more than 0.25 μm into the zygote (Figure 4b). Whereas in the cell fusion mutants the cell wall remnants could be as long as 3.0 μm, presumably as a result of the extended zones of contact. In mature wild-type zygotes, where the cell wall remnants were no longer visible, only a scar at the prior place of cell–cell contact remained (Figure 4, d and e).
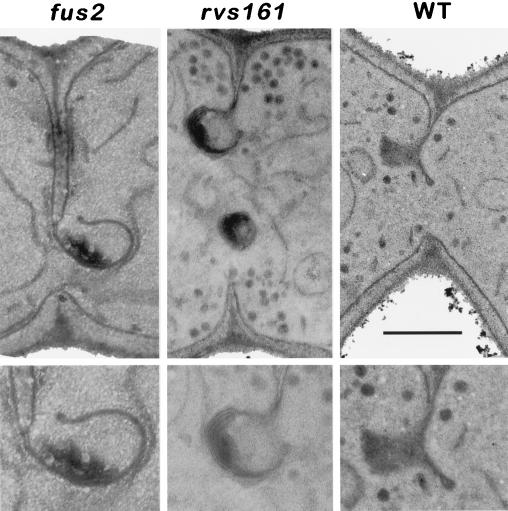
Figure 7.
Membrane structures after cell fusion. Micrographs are from matings of fus2 (MY4158 × MY4178), rvs161Δ (MY3909 × MY4495), and WT (MY3371 × MY2787). The bottom panels show a slight magnification that is underexposed to show the membranes in the electron-dense regions more clearly. Bar, 0.5 μm in the top panels.
Localization and Alignment of Vesicles at the Zone of Cell Fusion
One of the more striking features in wild-type prezygotes was the presence of vesicles clustered and aligned across the zone of cell fusion in mating pairs (see above). In wild-type prezygotes, the clustering was such that all the vesicles from both parents could be contained within a circle of ≤0.6 μm in diameter drawn over the cell fusion zone (Figures 4A and 5). Although the tight localization of the vesicles in wild-type prezygotes could be a consequence of the narrow zone of cell fusion, in many cases the region of clustering was smaller than the zone of cell fusion (Figure 5). That the vesicles are highly polarized directly across the cell fusion zone was more obvious when looking at the prezygotes of cell fusion mutants such as fus2 and rvs161. These prezygotes also localized the vesicles; however, they had an extended zone of cell fusion (average of 2 μm). In these cases, the vesicles were also found contained within a region of ∼0.6 μm in diameter. Interestingly, for both wild type and the mutants, the vesicles were often found directly across a region with localized cell wall thinning (Figure 5).
Analysis of cell fusion mutant prezygotes further supports the hypothesis that the vesicles play an important role in cell fusion. Whereas in wild type the vesicles were always clustered and aligned (Table 2, line 1), in two of the cell fusion mutants the vesicles were dispersed. Deletion of fus1 resulted in most prezygotes lacking vesicles at the zone (only 30% exhibited vesicles), and when present, the vesicles were often dispersed (53%) (Table 2, line 2). In spa2 prezygotes, the vesicles were present to the same extent as wild-type (60–70% of prezygotes had vesicles), but they were almost always dispersed (96%) (Table 2, line 3). In contrast, in both fus2 and rvs161 prezygotes, the vesicles were present, localized, and bundled to the same degree as wild-type prezygotes (Table 2, lines 4 and 5). In summary, Fus1p and Spa2p are required for normal vesicle positioning, whereas Fus2p and Rvs161p functions are not required for vesicle localization or clustering.
Table 2.
Vesicle localization and accumulation at the zone of cell fusion in prezygotes
| Matinga | % with vesiclesb | Vesicles per zonec | % aligned vesiclesd | % with plaquese | % with invaginationf |
|---|---|---|---|---|---|
| 1. WT | 71 | 13 | 100 | 0 | 0 |
| 2. fus1 | 30 | 6 | 47 | 0 | 0 |
| 3. spa2 | 60 | 19 | 4 | 0 | 0 |
| 4. fus2 | 33 | 15 | 100 | 20 | 6 |
| 5. rvs161Δ | 50 | 29 | 98 | 15 | 13 |
| 6. rvs161-P203Q | 80 | 12 | 100 | 10 | 25 |
| 7. fus1 fus2 | 12 | 7 | 40 | 0 | 0 |
| 8. fus1 rvs161 | 12 | 9 | 37 | 0 | 20 |
| 9. fus1 spa2 | 30 | 10 | 0 | 0 | 0 |
| 10. fus2 spa2 | 67 | 8 | 0 | 0 | 0 |
| 11. spa2 rvs161 | 80 | 9 | 0 | 0 | 7 |
| 12. fus2 rvs161 | 45 | 25 | 100 | 27 | 21 |
The data are from the following matings: 1. WT (MY3371 × MY2787); 2. fus1 (MY4161 × MY4164); 3. spa2 (MY3608 × MY3773); 4. fus2 (JY424 × JY428); 5. rvs161 (MY3909 × MY4495); 6. rvs161-P203Q (MY5224 × MY5359); 7. fus1 fus2 (MY4160 × JY429); 8. fus1 rvs161 (MY4905 × MY4907); 9. fus1 spa2 (MY4817 × MY4819); 10. fus2 spa2 (MY4813 × MY4815); 11. spa2 rvs161 (MY4746 × MY4748); and 12. fus2 rvs161 (MY4859 × MY4802).
The number of prezygotes with vesicles that could be scored divided by the total number of zygotes is represented in the % with vesicles column.
The number of vesicles at the cell fusion zone per 70 to 90-nm section is shown in the vesicles per zone column.
The prezygotes were scored as having aligned vesicles (% aligned) if the vesicles were clustered in ∼0.5-μm-diameter regions directly across the cell fusion zone.
The percentages of prezygotes with electron-dense regions on one or both sides of the cell fusion zone (Figure 6) were scored as containing plaques (% with plaques).
The percentage of prezygotes with one or more invagination (Figure 6) is indicated.
A second vesicle-associated phenotype was observed in rvs161Δ prezygotes. We observed that the vesicles at the cell fusion zone were present at two- to threefold higher numbers than wild-type (Table 2). However, we also noticed that vesicles were present at higher levels within buds of rvs161Δ cells, suggesting that the phenotype was not cell fusion specific. In support of this we found that cell fusion–defective point mutants of rvs161 (rvs161-P203Q) that exhibit no apparent endocytic abnormalities (Brizzio et al., 1998) did not accumulate vesicles above wild-type levels. Furthermore, the rvs161-P203Q prezygotes were indistinguishable from fus2 prezygotes (Table 2, line 6, and Figure 5). These results suggest that in rvs161Δ a block in endocytosis results in an accumulation of vesicles, but data from the point mutant suggest that the accumulation is not likely to be relevant to the defect in cell fusion.
Double mutant analysis allowed us to begin to define the progression of events during cell fusion, particularly with respect to vesicle clustering. In general, we found that double mutants containing fus1 exhibited prezygote morphologies similar to fus1 single mutants, and the double mutants containing spa2 displayed prezygote phenotypes similar to the spa2 single mutants (Table 2). Specifically, fus1 fus2 and fus1 rvs161 showed a decrease in the number of prezygotes with vesicles and a reduction in vesicle clustering similar to fus1 levels. The fus2 spa2 and rvs161 spa2 mutants showed an almost complete absence of vesicle clustering. The fus1 spa2 double mutant prezygotes tended to look like a superimposition of the mutant defects, in that fewer prezygotes contained observable vesicles at the zone of cell fusion (a characteristic of fus1), and when present, the vesicles always failed to cluster (a spa2 phenotype). Finally, as described in more detail below, the ultrastructural morphologies of the fus2Δ rvs161Δ prezygotes were very similar to the fus2Δ and the rvs161Δ single mutant prezygotes. These observations are consistent with the genetic findings discussed earlier that suggested that Fus2p and Rvs161p function in a similar part of the process of cell fusion. Taken together, the analysis of the double mutants suggested that the localization of the vesicles and the clustering across the cell fusion zone are early events in the cell fusion process. These events require the functions of Fus1p and Spa2p, whereas Rvs161p and Fus2p are needed for subsequent steps in the process.
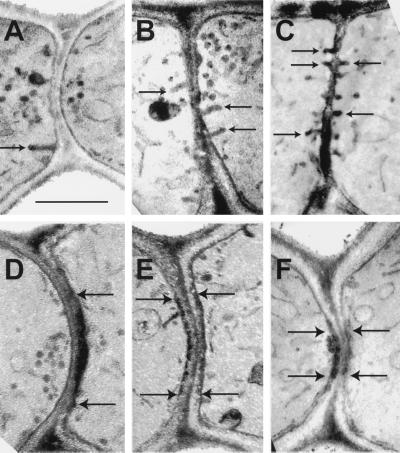
Electron-dense Plaques and Membrane Invaginations Are Observed at the Zone of Cell Fusion in fus2 and rvs161 Matings
The fus2 and rvs161 prezygotes exhibited unusual electron-dense plaques along the plasma membranes at the cell fusion zone (Figure 6, D–F) that were not seen in wild-type, fus1, and spa2 prezygotes (Table 2). Analysis of double mutants revealed that the presence of the plaques was dependent on both FUS1 and SPA2 function (Table 2), because the structures were not observed in double mutants containing a mutation in either fus1 or spa2. These results suggest that these structures are derived from the deposition of material from the vesicles, which first must be localized to the cell fusion zone.
Figure 6.
Electron-dense plaques and invaginations at the zones of cell fusion in fus2 and rvs161 prezygotes. The micrographs are from matings of rvs161-P203Q (MY5224 × MY5359) (A), rvs161▵ (MY3909 × MY4495) (B, C, and F), and fus2 (MY4158 × MY4178) (D and E). Examples of invaginations (A–C) are accentuated with arrows. The electron-dense plaques are along the plasma membrane between the arrows in D–F. Bar, 0.5 μm.
In addition, numerous membrane invaginations were also observed in fus2 and in rvs161 prezygotes (Figures 5 and 6 and Table 2). The mating-defective, endocytosis-competent mutant rvs161-P203Q (Brizzio et al., 1998) (Figure 6A) also showed the invaginations but at a lower density than seen with rvs161Δ (Figure 6, B and C). The invaginations in fus2 prezygotes were dependent on FUS1 and SPA2 function (Table 2), because they were not observed in fus1 fus2 or in spa2 fus2 prezygotes. However, at least a subset of the invaginations found in rvs161Δ were independent of FUS1 and SPA2 function (the genetic analysis was not done with the rvs161 point mutant rvs161-P203Q). The rvs161 deletion strains tended to show more invaginations per cell fusion zone (Figure 6, B and C) than did fus2 (Figure 5) or rvs161-P203Q (Figure 6A). Therefore, it is likely that most of the invaginations seen in double mutant strains (Table 2) are a consequence of the nonmating functions of Rvs161p. This suggests that the invaginations may be formed by different mechanisms, such that their presence in the rvs161Δ is a combination of cell fusion–associated and endocytosis-related defects.
One hypothesis for the origin of the invaginations is that membrane flux is perturbed in the fus2 and rvs161 mutants during cell fusion. It is likely that the polarized secretion at the zone of cell fusion must be balanced by endocytosis to maintain the plasma membrane at a constant surface area. Failure to initiate fusion in the fus2 and rvs161 prezygotes might lead to an imbalance between polarized secretion and membrane trafficking. Our observations of mutant prezygotes are consistent with this explanation. Moreover, the rvs161Δ, which has both cell fusion and endocytic defects, has even more invaginations than the fus2 mutant. On the other hand, mutants such as spa2 and fus1, which appear to have problems in polarization, contained no invaginations. Presumably the cell fusion zones in these mutants are less active sites for membrane flux.
Fused Plasma Membrane Structures at the Zone of Cell Fusion
Finally, we observed structures in wild-type, fus2, and rvs161 zygotes that might be intermediates in the fusion and removal of the plasma membrane. Typically, these structures looked like the plasma membranes from each partner in the mating pair had joined (fused), liberating a free end, which terminated with a loop structure (Figure 7). The double layer of membrane was generally curved. In fus2 and rvs161 zygotes, we also observed electron-dense regions associated with the converged plasma membranes. The electron-dense regions resemble the plaques observed in the prezygotes. The electron density of these structures was such that exposures of the negative required special care to see the double membrane (Figure 7, bottom panels). Interestingly, large (0.3 μm) rolled membrane structures were often found near the zone of cell fusion in many prezygotes and zygotes (our unpublished observations; Osumi et al., 1974). These rolled structures differed from the ones shown in Figure 7 in that they did not appear to be continuous with the existing plasma membranes of either partner. The origin of the rolls is not known, but it is possible that they are derived from the converged membrane structures (Figure 7). The rolled structures could also be derived from peripheral endoplasmic reticulum, which is prevalent at the zone of cell fusion.
DISCUSSION
Summary
To gain a better understanding of the events and regulation of cell fusion, we exploited the model system of conjugation in S. cerevisiae. Part of our approach involved a genetic analysis of the phenotypes of wild type and four cell fusion mutants: fus1 (Trueheart et al., 1987; Trueheart and Fink, 1989), fus2 (Elion et al., 1995), fus6/spa2, and fus7/rvs161 (Kurihara et al., 1994; Brizzio et al., 1998). Double mutant and dosage suppression analysis suggested that cell fusion occurs through at least two partially redundant or parallel pathways. Furthermore, our findings suggest that at least one branch of the process requires the concerted functions of Fus2p and Rvs161p. In support of this finding, we have concurrently determined that these two proteins physically interact, and that the stability of Fus2p depends on the mating function of Rvs161p (Brizzio et al., 1998).
Ultrastructural analysis confirmed the genetic data and led to a model for the functions of the cell fusion genes. We propose that the release of the contents of highly polarized vesicles into the cell fusion zone contributes to breakdown of the cell wall, a prerequisite for cell fusion in yeast. Analysis of the mutant prezygotes suggested that the vesicles are integral to the process of efficient cell fusion. Two of the cell fusion proteins, Fus1p and Spa2p, appeared to be important for the vesicle positioning. Fus1p seems to be required for the efficient localization of the vesicles to the cell fusion region. Spa2p, on the other hand, appears to be crucial for vesicle clustering across the zone. Fus2p and Rvs161p are likely to function after vesicle positioning and to contribute to cell fusion independently of vesicle clustering and alignment.
Cell Fusion in Wild Type
Ultimately, our goal is to understand the mechanism of cell fusion in wild-type yeast. Therefore, we characterized the events of cell fusion at an ultrastructural level in wild-type mating mixtures. In a prezygote, the region of contact between partner cells averaged a diameter of 1.0 μm from the outer borders and contained a 0.5-μm region of very close association. Vesicles in the region were found to be clustered and aligned within a 0.6-μm diameter directly across the intervening cell wall. Cell wall thinning was often observed between the vesicle clusters, and the initial cell fusion pore in wild-type zygotes averaged a 0.5-μm diameter (similar to the size of the vesicle bundle). Before karyogamy, cell fusion–associated vesicles were still present either at the recently formed 0.5-μm pore or associated with the peripheral cell wall remnants. During or just after karyogamy, cell wall remodeling continued such that the remnants disappeared, and the zygotic zone of cell fusion expanded from 1.1 to 1.6 μm. As zygotes matured further, dilation of the region seemed to occur until the zygotes averaged 2.5 μm. Finally, only a slight scar at the prior place of cell–cell contact was observed in mature wild-type zygotes.
The Role of Polarization in Vesicle Positioning
Our observations of cell fusion in wild type provided a framework within which to build models for cell fusion in yeast. After the mating cells make initial contact, the next step appears to be clustering and alignment of the cell fusion–associated vesicles. The high degree of polarization of these vesicles has strong implications for cell-to-cell communication. For example, the alignment could be a consequence of structural components positioned by a specific type of extracellular contact. Alternatively, a highly sensitive polarization apparatus positioned in response to pheromone signaling could mediate the juxtaposition. An obvious candidate for the polarization apparatus is the MAP kinase complex, which has recently been shown to include a myriad of components, including Bem1p, Cdc24p, and Cdc42p, that are known to effect polarization (Leeuw et al., 1995). This latter hypothesis is in keeping with our earlier work that showed that elevated pheromone levels are needed for efficient cell fusion (Brizzio et al., 1996). Further support for this model comes from the fact that high-copy suppressors of the spa2 cell fusion defect (unclustered vesicles) are genes that have genetic interactions with CDC24 and CDC42 (our unpublished observations).
Models for Removal of the Intervening Cell Wall
After the partner cells have positioned the cell fusion–associated vesicles, the cell wall must be removed for fusion to progress. In agreement with previous work (Osumi et al., 1974), we noticed a thinning of the cell wall at the zone of cell fusion in prezygotes. The thinning is thought to ultimately lead to the juxtaposition of the partner plasma membranes and eventually to membrane fusion. It is likely that localized thinning of the intervening cell wall in prezygotes is achieved by the deposition of hydrolytic enzymes into the zone of cell fusion. Accordingly, intracellular secretory vesicles would be likely to serve as carriers of the cell fusion hydrolytic activity. Byers and Goetsch (1975) noted the presence of vesicles in wild-type zygotes after cell fusion but before karyogamy. Furthermore, Baba et al. (1989) observed a concentration of vesicles in mating projections. The presence of these vesicles found in early zygotes or in mating projections is thought to be the consequence of polarized secretion, an event known to occur during mating (see review by Welch et al., 1994).
Our observations of prezygotes lead us to incorporate a vesicle-mediated mechanism for cell wall thinning and breakdown. The model we favor assumes that the cell fusion–associated vesicles are largely secretory. Clearly, our ultrastructural analyses cannot determine whether the observed vesicles are endocytic or secretory in origin. However, we have concurrently found that certain mutants that have strong endocytic defects do not have cell fusion defects, suggesting that endocytosis does not contribute significantly to the efficiency of cell fusion (Brizzio et al., 1998).
The vesicle-mediated model that we favor suggests that the highly polarized release of vesicle contents causes the localized thinning and eventual breakdown of the cell wall material separating the mating pair. Presumably, the vesicles carry cell fusion–specific proteins, including cell wall–degrading enzymes. In support of this, Baba et al. (1989) carefully analyzed vesicles in mating projections and determined that the vesicle population in cells responding to pheromone were on average 65 nm in diameter, whereas vesicles in vegetative cells averaged 90 nm. Their findings suggest that there are mating-specific vesicles polarized to the mating projection. It is of interest that tpm1 mutants accumulate a distinct class of vesicles and also exhibit cell fusion defects (Liu and Bretscher, 1992).
The contents of the vesicles we observed at the cell fusion zone remain speculative, however certain cell wall–degrading enzymes are likely candidates. The mechanism of cell wall breakdown is not clear; however, the cell wall remnants found in wild-type and in mutant zygotes may begin to shed some light on the process. We observed that both in the mutants (see Figure 7, middle panel) and in early wild-type zygotes (Figure 4b) the remnants had a similar tapered appearance, and the cell wall maintained the characteristic layered structure (see review by Cid et al., 1995). The tapering suggested that cell wall thinning had occurred adjacent to the region. Because the remnants maintained the layered appearance, we suggest that cell wall removal during fusion is not the consequence of a profusion of hydrolytic components, which completely obliterates the cell wall structure at the cell fusion zone. Rather, we believe that the thinning process is such that a great deal of cell wall integrity is maintained during the process. Breakdown of the cell wall might simply reflect a shift in the balance between hydrolysis and synthesis of structural components. For example, increasing the expression of hydrolases, although decreasing (or maintaining) the activity of synthases, could result in such a shift.
At this stage of the analysis we cannot distinguish between models for regulated or constitutive release of the vesicles. One hypothesis is that the vesicles might be constitutively released from both partners at a defined location, which would lead to cell wall thinning and eventually to breakdown of the intervening material. Alternatively, by analogy to the events in nerve terminals, the vesicles could be positioned across the zone of cell fusion, poised to be released upon the appropriate triggering event. Finally, the mechanism might involve a combination of both mechanisms, such that some vesicles are constitutively released, whereas others are tethered intracellularly until the signal for cell fusion is propagated. Such a model would be consistent with the finding that very high levels of pheromone are required to signal the process of cell fusion in prezygotes (Brizzio et al., 1996).
Similarities to Other Cell Fusion Paradigms
Despite their differences, the events of cell fusion of myoblasts to form a muscle fiber and the fusion of haploid yeast cells to form a diploid share features worth noting. In each system the intervening cell material must be removed to permit cell-cell contact, after which the membranes must fuse to allow for cytoplasmic mixing without cell lysis. After initial formation of pores in fusing myoblasts, vesiculation is thought to partially mediate complete removal of plasma membranes (Doberstein et al., 1997). Although we cannot definitively rule out vesiculation in the mechanism of yeast cell fusion, we do not observe vesiculating plasma membrane intermediates during early zygote formation. We did observe a structure (Figure 7) in both wild-type and certain mutant zygotes that was more consistent with converged membranes with a single pore.
In contrast to the apparent differences, a recent ultrastructural analysis in Drosophila showed that paired vesicles aligned across the fusion zone are likely to be important for myoblast fusion (Doberstein et al., 1997). The alignment of vesicles is strikingly similar to what we have observed in yeast, suggesting that both systems have mechanisms for polarization across the cell fusion zone. Furthermore, Doberstein et al. (1997) observed electron-dense plaques across the zone of cell fusion in myoblasts. These structures found in Drosophila were remarkably similar to the plaques we observed in fus2 and rvs161 mutant zygotes, further suggesting that some aspects of the mechanisms of cell fusion in the two organisms are conserved.
ACKNOWLEDGMENTS
We are grateful to the laboratories of M. Snyder, G. Fink, T. Stearns, and P. Hieter for generously supplying plasmids and strains. We thank Joe Goodhouse for training in electron microscopic techniques and technical assistance. This work was supported by National Institutes of Health grant GM37739 awarded to M.R. A.G. was supported in part by The Jane Coffin Childs Memorial Fund for Medical Research. V.B. was supported by a National Institutes of Health institutional grant.
Footnotes
Abbreviations used: DAPI, 4′,6′-diamidino-2-phenylindole; GFP, green fluorescent protein; DIC, differential interference contrast; WT, wild-type; YEPD, yeast extract with peptone and dextrose.
REFERENCES
- Arkowitz R A, Lowe. N. A small conserved domain in the yeast Spa2p is necessary and sufficient for its polarized localization. J Cell Biol. 1997;138:17–36. doi: 10.1083/jcb.138.1.17. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Baba M, Baba N, Ohsumi Y, Kanaya K, Osumi M. Three-dimensional analysis of morphogenesis induced by mating pheromone alpha factor in Saccharomyces cerevisiae. J Cell Sci. 1989;94:207–216. doi: 10.1242/jcs.94.2.207. [DOI] [PubMed] [Google Scholar]
- Brizzio V, Gammie AE, Nijbroek G, Michaelis S, Rose MD. Cell fusion during yeast mating requires high levels of a-factor mating pheromone. J Cell Biol, 1996;135:1727–1739. doi: 10.1083/jcb.135.6.1727. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Brizzio, V., Gammie, A.E., and Rose, M.D. (1998). Rvs161p interacts with Fus2p to promote cell fusion in Saccharomyces cerevisiae. J. Cell Biol. (in press). [DOI] [PMC free article] [PubMed]
- Buehrer BM, Errede B. Coordination of the mating and cell integrity mitogen-activated protein kinase pathways in Saccharomyces cerevisiae. Mol Cell Biol. 1997;17:6517–6525. doi: 10.1128/mcb.17.11.6517. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Byers B, Goetsch L. Behavior of spindles and spindle plaques in the cell cycle and conjugation of Saccharomyces cerevisiae. J Bacteriol. 1975;124:511–523. doi: 10.1128/jb.124.1.511-523.1975. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chenevert J. Cell polarization directed by extracellular cues in yeast. Mol Biol Cell. 1994;5:1169–1175. doi: 10.1091/mbc.5.11.1169. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Chenevert J, Valtz N, Herskowitz I. Identification of genes required for normal pheromone-induced cell polarization in Saccharomyces cerevisiae. Genetics. 1994;136:1287–1296. doi: 10.1093/genetics/136.4.1287. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Cid VJ, Duran A, del Rey F, Snyder MP, Nombela C, Sanchez M. Molecular basis of cell integrity and morphogenesis in Saccharomyces cerevisiae. Microbiol Rev. 1995;59:345–386. doi: 10.1128/mr.59.3.345-386.1995. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crouzet M, Urdaci M, Dulau L, Aigle M. Yeast mutant affected for viability upon nutrient starvation: characterization and cloning of the RVS161 gene. Yeast. 1991;7:727–743. doi: 10.1002/yea.320070708. [DOI] [PubMed] [Google Scholar]
- Desfarges L, Durrens P, Juguelin H, Cassagne C, Bonneu M, Aigle M. Yeast mutants affected in viability upon starvation have a modified phospholipid composition. Yeast. 1993;9:267–277. doi: 10.1002/yea.320090306. [DOI] [PubMed] [Google Scholar]
- Doberstein SK, Fetter RD, Mehta AY, Goodman CS. Genetic analysis of myoblast fusion: blown fuse is required for progression beyond the prefusion complex. J Cell Biol. 1997;136:1249–1261. doi: 10.1083/jcb.136.6.1249. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Dorer R, Boone C, Kimbrough T, Kim J, Hartwell LH. Genetic analysis of default mating behavior in Saccharomyces cerevisiae. Genetics. 1997;146:39–55. doi: 10.1093/genetics/146.1.39. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Durrens P, Revardel E, Bonneu M, Aigle M. Evidence for a branched pathway in the polarized cell division of Saccharomyces cerevisiae. Curr Genet. 1995;27:213–216. doi: 10.1007/BF00326151. [DOI] [PubMed] [Google Scholar]
- Elia L, Marsh L. Role of the ABC transporter Ste6 in cell fusion during yeast conjugation. J Cell Biol. 1996;135:741–751. doi: 10.1083/jcb.135.3.741. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elion EA, Grisafi PL, Fink GR. FUS3 encodes a cdc2/CDC28-related kinase required for the transition from mitosis into conjugation. Cell. 1990;60:649–664. doi: 10.1016/0092-8674(90)90668-5. [DOI] [PubMed] [Google Scholar]
- Elion EA, Satterberg B, Kranz JE. FUS3 phosphorylates multiple components of the mating signal transduction cascade: evidence for STE12 and FAR1. Mol Biol Cell. 1993;4:495–510. doi: 10.1091/mbc.4.5.495. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Elion EA, Trueheart J, Fink GR. Fus2 localizes near the site of cell fusion and is required for both cell fusion and nuclear alignment during zygote formation. J Cell Biol. 1995;130:1283–1296. doi: 10.1083/jcb.130.6.1283. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Errede B, Cade RM, Yashar BM, Kamada Y, Levin DE, Irie K, Matsumoto K. Dynamics and organization of MAP kinase signal pathways. Mol Reprod Dev. 1995;42:477–485. doi: 10.1002/mrd.1080420416. [DOI] [PubMed] [Google Scholar]
- Flescher EG, Madden K, Snyder M. Components required for cytokinesis are important for bud site selection in yeast. J Cell Biol. 1993;122:373–386. doi: 10.1083/jcb.122.2.373. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujimura H. Identification and characterization of a mutation affecting the division arrest signaling of the pheromone response pathway in Saccharomyces cerevisiae. Genetics. 1990a;124:275–282. doi: 10.1093/genetics/124.2.275. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Fujimura H. Molecular cloning of the DAC2/FUS3 gene essential for pheromone-induced G1-arrest of the cell cycle in Saccharomyces cerevisiae. Curr Genet. 1990b;18:395–400. doi: 10.1007/BF00309907. [DOI] [PubMed] [Google Scholar]
- Gehrung S, Snyder M. The SPA2 gene of Saccharomyces cerevisiae is important for pheromone- induced morphogenesis and efficient mating. J Cell Biol, 1990;111:1451–1464. doi: 10.1083/jcb.111.4.1451. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Herskowitz I. MAP kinase pathways in yeast: for mating and more. Cell. 1995;80:187–197. doi: 10.1016/0092-8674(95)90402-6. [DOI] [PubMed] [Google Scholar]
- Ito H, Fukuda Y, Murata K, Kimura A. Transformation of intact yeast cells treated with alkali cations. J Bacteriol. 1983;153:163–168. doi: 10.1128/jb.153.1.163-168.1983. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kurihara LJ, Beh CT, Latterich M, Schekman R, Rose MD. Nuclear congression and membrane fusion: two distinct events in the yeast karyogamy pathway. J Cell Biol. 1994;126:911–923. doi: 10.1083/jcb.126.4.911. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Leberer E, Thomas DY, Whiteway M. Pheromone signalling and polarized morphogenesis in yeast. Curr Opin Genet Dev. 1997;7:59–66. doi: 10.1016/s0959-437x(97)80110-4. [DOI] [PubMed] [Google Scholar]
- Leeuw T, Fourest-Lieuvin A, Wu C, Chenevert J, Clark K, Whiteway M, Thomas DY, Leberer E. Pheromone response in yeast: association of Bem1p with proteins of the MAP kinase cascade and actin. Science. 1995;270:1210–1213. doi: 10.1126/science.270.5239.1210. [DOI] [PubMed] [Google Scholar]
- Levin DE, Errede B. The proliferation of MAP kinase signaling pathways in yeast. Curr Opin Cell Biol. 1995;7:197–202. doi: 10.1016/0955-0674(95)80028-x. [DOI] [PubMed] [Google Scholar]
- Lipke PN, Kurjan J. Sexual agglutination in budding yeasts: structure, function, and regulation of adhesion glycoproteins. Microbiol Rev. 1992;56:180–194. doi: 10.1128/mr.56.1.180-194.1992. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Liu H, Bretscher A. Characterization of TPM1 disrupted yeast cells indicates an involvement of tropomyosin in directed vesicular transport. J Cell Biol. 1992;118:285–299. doi: 10.1083/jcb.118.2.285. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Marsh L, Rose MD. In: The pathway of cell and nuclear fusion during mating in S. cerevisiae. In: The Molecular and Cellular Biology of the Yeast Saccharomyces: Cell Cycle and Cell Biology. Pringle JR, Broach JR, Jones EW, editors. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1997. pp. 827–888. [Google Scholar]
- McCaffrey G, Clay FJ, Kelsay K, Sprague GF., Jr Identification and regulation of a gene required for cell fusion during mating of the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1987;7:2680–2690. doi: 10.1128/mcb.7.8.2680. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Munn AL, Stevenson BJ, Geli MI, Riezman H. end5, end6, and end7: mutations that cause actin delocalization and block the internalization step of endocytosis in Saccharomyces cerevisiae. Mol Biol Cell. 1995;6:1721–1742. doi: 10.1091/mbc.6.12.1721. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Osumi M, Shimoda C, Yanagishima N. Mating reaction in Saccharomyces cerevisiae. V. Changes in the fine structure during the mating reaction. Arch Mikrobiol. 1974;97:27–38. [PubMed] [Google Scholar]
- Philips J, Herskowitz I. Osmotic balance regulates cell fusion during mating in Saccharomyces cerevisiae. J Cell Biol. 1997;138:961–974. doi: 10.1083/jcb.138.5.961. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reynolds ES. The use of lead citrate at high pH as an electron opaque stain in electron microscopy. J Cell Biol. 1963;17:208–212. doi: 10.1083/jcb.17.1.208. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Rose MD. Nuclear fusion in the yeast, Saccharomyces cerevisiae. Annu Rev Cell Dev Biol. 1996;12:663–695. doi: 10.1146/annurev.cellbio.12.1.663. [DOI] [PubMed] [Google Scholar]
- Rose MD, Winston F, Hieter P. Methods of Yeast Genetics. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1990. , 198. [Google Scholar]
- Rothstein R. Targeting, disruption, replacement, and allele rescue: integrative DNA transformation in yeast. Methods Enzymol. 1991;194:281–301. doi: 10.1016/0076-6879(91)94022-5. [DOI] [PubMed] [Google Scholar]
- Sambrook J, Fritsch EF, Maniatis T. Molecular Cloning: A Laboratory Manual. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1989. [Google Scholar]
- Santos B, Duran A, Valdivieso MH. CHS5, a gene involved in chitin synthesis and mating in Saccharomyces cerevisiae. Mol Cell Biol. 1997;17:2485–2496. doi: 10.1128/mcb.17.5.2485. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Scherer S, Davis RW. Replacement of chromosome segments with altered DNA sequences constructed in vitro. Proc Natl Acad Sci USA. 1979;76:4951–4955. doi: 10.1073/pnas.76.10.4951. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sivadon P, Bauer F, Aigle M, Crouzet M. Actin cytoskeleton and budding pattern are altered in the yeast rvs161 mutant: the Rvs161 protein shares common domains with the brain protein amphiphysin. Mol Gen Genet. 1995;246:485–495. doi: 10.1007/BF00290452. [DOI] [PubMed] [Google Scholar]
- Snyder M. The SPA2 protein of yeast localizes to sites of cell growth. J Cell Biol. 1989;108:1419–1429. doi: 10.1083/jcb.108.4.1419. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Snyder M, Gehrung S, Page BD. Studies concerning the temporal and genetic control of cell polarity in Saccharomyces cerevisiae. J Cell Biol. 1991;114:515–532. doi: 10.1083/jcb.114.3.515. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Sprague GF, Thorner JW. In: Pheromone response and signal transduction during the mating process of Saccharomyces cerevisiae. In: The Molecular and Cellular Biology of the Yeast Saccharomyces: Gene Expression. Jones EW, Pringle JR, Broach JR, editors. II. Cold Spring Harbor, NY: Cold Spring Harbor Laboratory Press; 1992. pp. 657–744. [Google Scholar]
- Trueheart J, Boeke JD, Fink GR. Two genes required for cell fusion during yeast conjugation: evidence for a pheromone-induced surface protein. Mol Cell Biol. 1987;7:2316–2328. doi: 10.1128/mcb.7.7.2316. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Trueheart J, Fink GR. The yeast cell fusion protein FUS1 is O-glycosylated and spans the plasma membrane. Proc Natl Acad Sci USA. 1989;86:9916–9920. doi: 10.1073/pnas.86.24.9916. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Welch MD, Holtzman DA, Drubin DG. The yeast actin cytoskeleton. Curr Opin Cell Biol. 1994;6:110–119. doi: 10.1016/0955-0674(94)90124-4. [DOI] [PubMed] [Google Scholar]
- Xu BE, Kurjan J. Evidence that mating by the Saccharomyces cerevisiae gpa1Val50 mutant occurs through the default mating pathway and a suggestion of a role for ubiquitin-mediated proteolysis. Mol Biol Cell. 1997;8:1649–1664. doi: 10.1091/mbc.8.9.1649. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Zahner JE, Harkins HA, Pringle JR. Genetic analysis of the bipolar pattern of bud site selection in the yeast Saccharomyces cerevisiae. Mol Cell Biol. 1996;16:1857–1870. doi: 10.1128/mcb.16.4.1857. [DOI] [PMC free article] [PubMed] [Google Scholar]