Abstract
Quaternary carbolinium salts have been reported to show improved antimalarial activity and reduced cytotoxicity as compared to electronically neutral β-carbolines. In this study mono- and di-methylated quaternary carbolinium cations of manzamine A were synthesized and evaluated for their in vitro antimalarial and antimicrobial activity, cytotoxicity and also their potential for glycogen synthase kinase (GSK-3β) inhibition using molecular docking studies. Among the analogs, 2-N-methylmanzamine A (2) exhibited antimalarial activity (IC50 0.7 – 1.0 µM) but was less potent than manzamine A. However the compound was significantly less cytotoxic to mammalian kidney fibroblasts and the selectivity index was in the same range as for manzamine A.
Keywords: SAR, Manzamine A, Antimalarial agent, Methylation
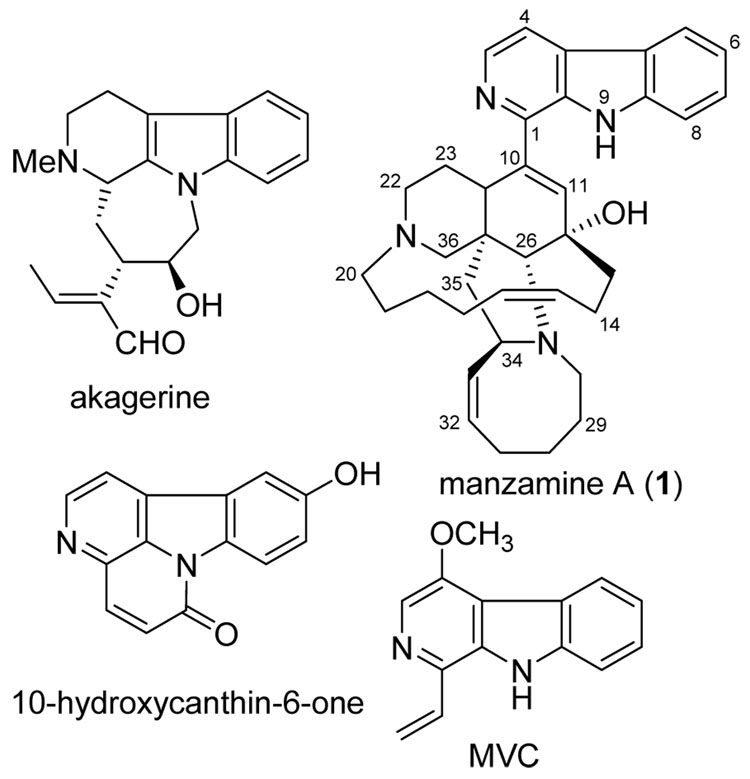
Malaria, a major tropical infectious disease caused primarily by the protozoan parasite Plasmodium falciparum, is one of the most serious health problems worldwide and is responsible for the death of over 1.12 million individuals every year with more than 40 percent of the global population at risk.[1][GS1] β-Carboline alkaloids are widely distributed in a number of plant and animal species, as well as marine invertebrates. Some of these alkaloids such as manzamine A (1), akagerine, 10-hydroxycanthin-6-one and 4-methoxy-1-vinyl-β-carboline (MVC) (Figure 1), display a diverse array of biological activities including antiplasmodial efficacy.[2–4] Manzamine A, as a first representative of the manzamine alkaloids, bearing a unique multi-heterocyclic ring system coupled to a β-carboline moiety, was isolated from the sponge Haliclona sp collected near Manzamo Island by Higa and coworkers in 1986.[5] Despite its potent in vitro activity as an antimalarial agent against P. falciparum the moderate toxicity of manzamine A hampered its development as a drug candidate.[4], [6–8]
Figure 1.
β-Carboline alkaloids with potential biological activities.
Recently, Ihara and coworkers have reported that the rhodacyanine family of compounds and the β-carboline system of MVC, owing in both cases to a π-delocalized lipophilic cationic (DLC) structure, display reasonable antimalarial and antileishmanial activities against P. falciparum and Leishmania major, respectively, with low cytotoxicity against mammalian cells.[9–13] The conceptual term, DLC, was originally proposed by Chen et al. in their studies related to anticancer agents.[14] It has been reported that several DLC compounds exhibit selective antitumor activity by accumulating in the mitochondria of carcinoma cells.[15–17] Vaidya et al. found that mitochondrial membrane potential collapse is an antimalarial mechanism of some drugs,[18] and many researchers concluded that DLCs could therefore represent a new antiplasmodial class of drugs.
Our goal was to synthesize manzamine A analogs with decreased toxicity and improved therapeutic index for antimalarial efficacy and continuing structure-activity relationship studies. The rationale for the modification of the β-carboline moiety through quarternization of the pyridine nitrogen to form β-carbolinium salts is based on previous work that showed reduced toxicity and increased antimalarial activity of DLC’s formed upon transformation with either alkyl tosylates or alkyl halides as a correlation to the π–delocalization of the cationic species.[10, 11, 19]
Recently, we found that manzamine A inhibits GSK-3β with an IC50 of 10.2 µM so inhibition of this multifunctional serine/threonine kinase may in part be the reason for various therapeutic activities of manzamine A analogs.[20] Since, the crystal structure of GSK-3β is available, we initiated structure-based drug design paradigm of manzamine A analogs using molecular docking coupled with molecular dynamics. Quaternization of the neutral pyridine nitrogen to the desired quaternary ammonium cation was challenging but could finally be accomplished by methylation of manzamine A in the presence of methyl trifluoromethanesulfonate to afford three methylated products (2–4) in varying yields depending upon reaction conditions. The structures of 2, 3 and 4 were confirmed using high resolution ESI-MS, as well as 1H, 13C NMR, HMQC and HMBC. The in vitro antimalarial and antimicrobial activity in addition to cytotoxicity vs. mammalian cells of analogs 2–4 have also been evaluated.
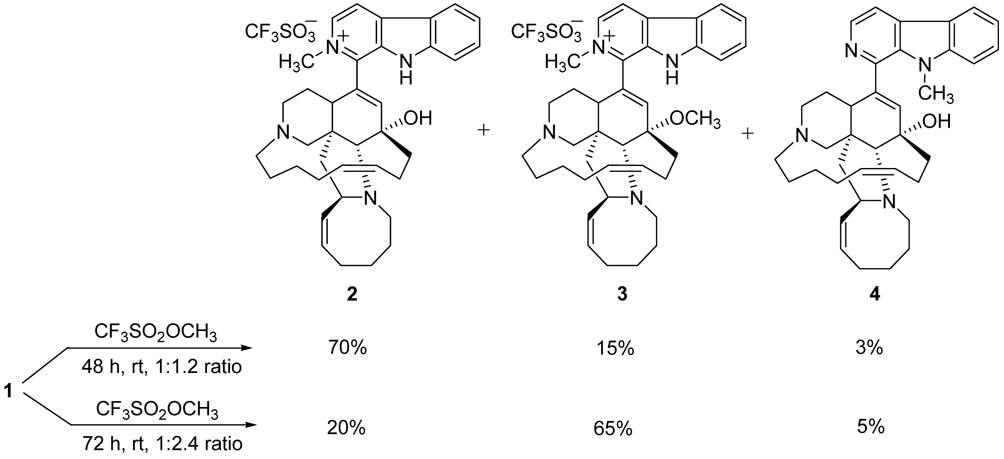
According to our previous unpublished work showing that N-methylation of the secondary amine of the β-carboline moiety drastically decreases antimalarial potency[GS3], along with the low yield of 4, we were not interested to pursue further evaluation of this compound. Using methyl iodide as the alkylating 2-N-Methylmanzamine A (2), 2-N,12-O-dimethyl manzamine A (3) sulfonate and 9-methyl manzamine A (4) (Scheme 1) are generated from the treatment of the β-carboline moiety with methyl trifluoromethanesulfonate to afford the mono- and dimethylated products.[11, 19][GS2] agent furnished the non-preferential analog 4 in higher yield.
Scheme 1.
The structures of analogs 2 and 3 were confirmed with spectroscopic and MS techniques. The high-resolution ESIMS of analog 2 showed the [M+H]+ ion peak at m/z 563.36747 amu in positive mode, in accordance with its molecular formula (C37H47N4O). The 1H NMR spectrum of 3 in MeOD in 400 MHz clearly indicated the presence of a new singlet methyl group at 4.57 (s, 3H, NCH3) ppm. This finding was confirmed by a new methyl singlet at 45.3 ppm in the 13C NMR spectrum. The HMBC spectrum of 2 supported the methylation of the pyridinium nitrogen as shown by a correlation between the CH3 singlet at 4.57 ppm and both adjacent aromatic carbon signals at 134.6 ppm (C-3) and 138.8 ppm (C-1) (Figure 2).
Figure 2.
Selected HMBC correlations of analogs 2 and 3.
High resolution TOF-ESI-MS of analog 3 provided a molecular mass of [M+H]+ at m/z 577.4245, corresponding to the addition of two methyl mass units, in comparison with parent molecule. The 1H and 13C NMR of compound 3 in DMSO-d6 contained two methyl proton signals at 4.41 (s, 3H, NCH3) and 3.13 ppm (s, 3H, OCH3) along with corresponding carbon resonances at 45.9 and 54.1 ppm, respectively, in agreement with the structure of analog 3 (Figure 2). In addition, HMBC experiment revealed an unambiguous correlation between the methoxy protons at 3.13 ppm and neighboring quaternary C-12 carbon at 62.1 ppm. These findings are consistent with the fact that we observed the NH group at position 9 of the β-carboline moiety still intact at 11.4 (s, 1H, NH), which in turn indicates that secondary amine group is not involved in the methylation.
Biological Evaluation
All synthetic analogs were evaluated for in vitro antimalarial and antimicrobial activity as well as cytotoxicity against mammalian cells.
Antimalarial activity was determined against chloroquine-sensitive (D6) and chloroquine-resistant (W2) strains of Plasmodium falciparum. As shown in Table 1, compound 2 showed antimalarial activity with an IC50 of 0.7 and 1.0 µM for D6 and W2 strains, respectively. Compounds 3 and 4 were inactive. None of the synthesized analogs (2–4) showed any cytotoxicity to mammalian kidney fibroblasts (Vero cells) while manzamine A was cytotoxic (IC50 0.5 µM). Although antimalarial activity of 2 was much less potent than manzamine A, the selectivity index of 2 remained comparable to manzamine A due to the loss of cytotoxicity. These results indicated that quarternization of the β-carboline moiety resulted in reduced toxicity but still retained antimalarial efficacy in case of 2. However, further methylation of 2 (to generate 3) resulted in loss of antimalarial activity. Methylation of the indoline nitrogen of manzamine A (4) resulted in the loss of both antimalarial activity and cytotoxicity.
Table 1.
In vitro Antimalarial Activity, Selectivity Index, Cytotoxicity to Vero Cells and Docking Scores.
| Compound | P. falciparum D6 | P. falciparum W2 | Cytotoxicity to Vero cells | Docking Score | ||
|---|---|---|---|---|---|---|
| IC50 (µM) | S.I. | IC50 (µM) | S.I. | IC50 (µM) | ChemScore† | |
| 1 | 0.017 | 29.4 | 0.020 | 25.23 | 0.501 | 24.9 (±1.0) |
| 2 | 0.736 | >24 | 1.011 | >17.5 | NC | 22.3 (±0.6) |
| 3 | NA | – | NA | – | NC | 20.6 (±0.8) |
| 4 | NA | – | NA | – | NC | – |
| Chloroquine | 0.013 | >1000 | 0.135 | >100 | NC | – |
| Artemisinin | 0.0063 | >2700 | 0.0045 | >3700 | NC | – |
NA= No antimalarial activity up to 8 µM; NC = No cytotoxicity up to 17 µM: Selectivity Index (S.I.) = IC50 (Vero Cells) / IC50 (P. falciparum).
GOLD ChemScore was an average over 3 runs. Standard deviation in parentheses.
Manzamine A and some of its analogs are known to inhibit Homo sapiens glycogen synthase kinase-3β (GSK-3β).[20] Based on the assumption that the antimalarial activity of these analogs may be due to kinase inhibition of P. falciparum, we used the crystal structure of HsGSK-3β (PDB: 1 gng)[21] for docking studies to predict the optimal binding positions and relative binding propensities of 1, 2 and 3. Based on the evidence that 1 was found to be an ATP-noncompetitive inhibitor of GSK-3β[20], we attempted to dock 1–3 into a pocket located in the vicinity of the activation pocket formed by three basic residues, Arg96, Arg180 and Lys205. We used the GOLD 3.1.1 docking program[22] with the scoring function ChemScore,[23] to compare the difference in binding affinity between the ligands and to compare with the experimental in vitro antimalarial activity. The co-crystallized ligands in 1gng, namely 2-amino-2-hydroxymethyl-propane-1,3-diol and sulfate ions, were removed before docking. The active site was defined as any atom that lay within a 15 Å radius of the δN of Arg96. The geometry of 1 was optimized using MMFF94 in Sybyl 7.2, starting from the published X-ray crystal structure of manzamine A.[5] 2 and 3 were built from the crystal structure of 1 followed by geometry optimization using the same force field. ChemScore is a dimensionless fitness function which is a measure of the free energy of a ligand binding to a protein. Since GOLD uses a genetic algorithm to dock ligands, consecutive docking runs do not give exactly the same fitness scores. To address this issue of inherent randomness of GOLD docking, we performed 3 consecutive docking runs to obtain average ChemScores for each ligand, which are given in Table 2 for 1–3.
Table 2.
In Vitro Antimicrobial Activity (all values in µM).
| Compound | C. albicans | C. neoformans | MRSA | M. intracellulare | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| IC50 | MIC | MFC | IC50 | MIC | MFC | IC50 | MIC | MBC | IC50 | MIC | MBC | |
| 1 | 16.4 | - | - | 2.7 | 5.7 | 23 | 0.18 | - | - | 0.18 | 0.36 | 2.84 |
| 2 | - | - | - | 71.1 | - | - | 44.5 | 89 | 89 | 17.8 | 44.5 | 44.5 |
| 3 | - | - | - | 52 | 87 | 87 | - | - | - | - | - | - |
| 4 | - | - | - | - | - | - | - | - | - | - | - | - |
| Amphotericin B | 0.3 | 0.68 | 1.35 | 0.8 | 1.35 | 1.35 | NT | NT | NT | NT | NT | NT |
| Ciprofloxacin | NT | NT | NT | NT | NT | NT | 0.3 | 1.5 | - | 1.1 | 1.5 | 3 |
IC50 =The concentration that affords 50% inhibition of growth; MIC (Minimum Inhibitory Concentration) is the lowest test concentration that allows no detectable growth; Minimum Fungicidal/Bactericidal Concentration (MFC/MBC) is the lowest test concentration that kills the organism
“-“= Not Active at the highest test concentration of 91, 89, 87, 89, 5.4 and 3.0 for samples 1, 2, 3, 4, Amphotericin B and Ciprofloxacin, respectively; NT=Not Tested
Amphotericin B and ciprofloxacin are used as positive antifungal and antibacterial controls, respectively.
Docking scores agree with the in vitro antimalarial activity data and show that 1 > 2 > 3. This is consistent with the experimental findings that the first methylation on the tertiary nitrogen of the β-carboline ring (2) and the second methylation on the 12-hydroxy group (3) each decrease the antimalarial activity a step compared to having free tertiary nitrogen on the β-carboline ring and a 12-hydroxyl in manzamine A (1). The predicted binding positions of 1–3 within the ATP-noncompetitive binding pocket (as identified by the Lee and Richards solvent accessible molecular surface)[24] is shown in Figure 3.
Figure 3.
Binding positions of 1–3 within the ATP-noncompetitive binding pocket of GSK-3β (A:1, B:2, C:3). Ligands 1–3 are shown with green carbon; the protein is in ribbon format colored according to secondary structures; the binding pocket interior surface (as detected with the Lee/Richards molecular surface) is in white.
1 and 3 have the same binding mode whereas the binding position of 2 is very different. The polar and hydrophobic interactions of 1 are shown in Figure 4. Manzamine A (1) shows a strong hydrogen bonding of its 12-hydroxyl group with the backbone carbonyl of Arg92. Also, the hydrophobic β-carboline ring interacts with the side chains of Lys94, Asn95 and Arg96. The NH of the β-carboline ring is pointing towards Phe93 indicating a probable steric clash if any substitution is made at this position. In comparison to 1, 2 adopts a binding mode in a different region of the ATP-noncompetitive binding pocket, leading to less favorable hydrogen bonding and hydrophobic interactions. The crucial interactions for 2 are also shown in Figure 4. The 12-hydroxyl of 2 forms a hydrogen bond, but it is with Gln89, unlike that of 1 with Arg92. The hydrogen bonding between the NH2 of Gln89 and the 12-hydroxyl of 2 is weaker (more distant) than the interaction seen in 1. Also, 2 shows hydrophobic interactions with the side chain of Phe67. 3 adopts a binding position similar to 1, and 3 does not show any hydrogen bonding interaction, due to the second methylation at the 12-hydroxyl group. Thus the docking scores are in good agreement with the cytotoxicity results indicating their utility in the rational design of more active analogs from this class.
Figure 4.
Interactions of 1 (A) and 2 (B) with the GSK-3β ATP-noncompetitive binding pocket residues.
Antimicrobial activity was determined against a panel of human pathogenic bacteria and fungi (Table 2). Compounds 1 to 4 were inactive against the Gram negative bacteria E. coli and Pseudomonas aeruginosa, in addition to the opportunistic filamentous fungus Aspergillus fumigatus (data not shown). Moreover, compound 4 was inactive against all species tested, indicating that quaternarization of the secondary amine decreases bioactivity in general. Among compounds 1–3, manzamine A (1) remains the most potent with strong activity against the opportunistic yeast Cryptococcus neoformans, methicillin-resistant S. aureus (MRSA) and Mycobacterium intracellulare, with bioactivity against the latter microorganism rivaling the positive control Ciprofloxacin. Results indicated that the conversion of hydroxyl group to a methoxy in compound 3 also decreases in vitro antimicrobial activity. In summary, derivatization of either the pyridine or secondary amine of the β-carboline moiety, along with methylation of the C-12 hydroxyl group of manzamine A decrease in vitro activity against several pathogenic microorganisms.
In conclusion, the preliminary structure-activity relationship study in regards to the first and second methylation showed that the introduction of the first methyl at the pyridine nitrogen in the β-carboline moiety of manzamine A significantly decreases the cytotoxicity but also reduced the antimalarial activity of manzamine A. Introduction of a second methyl group at C-12-OH position completely eliminated the antimalarial activity indicating that that OH group is an essential pharmacophore for the antimalarial activity of the manzamine class of alkaloids. The docking studies of these analogs on GSK-3β delineated the ability of manzamine A (1) to form a stronger hydrogen bond than 2 and the inability of 3 to interact favorably with the ATP-noncompetitive site amino acids of GSK-3β, because 3 lacks a complementary donor atom at its C-12-OH position.
Experimental Section
General Preparation of Compounds 3–4
Treatment of manzamine A (54.8 mg, 0.1 mmol) with methyl trifluoromethanesulfonate (135.5 µL, 1.2 mmol) in a molar ratio of 1 to 1.2 in dry methylene chloride while stirring for 48 hours at ambient temperature resulted in the formation of major product 2 in 70% yield in addition to two minor products 3 in 15% and 4 in 3% yield. Purification was completed via HPLC using a reverse-phase C8 column with a mobile phase gradient of water/acetonitrile and a flow rate of 10 mL/min. The major compound appears at a retention time of 40 minutes.
By increasing the molar ratio of methyl trifluoromethanesulfonate from 1.2 to 2.4 and stirring for 72 hours, compound 3 was generated in 65% yield alongside the minor products 2 in 20% and 4 in 5% yield. The resultant yellow solid was washed with diethyl ether and purified using the method above affording compound 3 at a retention time of 30 minutes.
2-N-Methylmanzamine A trifluoromethanesulfonate (2)
[α]25D + 57.5 (c = 0.08, MeOH); UV λmax (MeOH) 260, 310, 375 nm; IR νmax (CHCl3) 3207 (NH), 3073, 3011, 2928, 2855, 1671, 1628, 1577, 1520, 1415, 1296, 1198, 1030 cm−1; 1H NMR (MeOD) δ 8.65 (2H, m), 8.43 (1H, d), 7.87 (2H, m), 7.52 (1H, t), 6.27 (1H, s), 5.68 (2H, m), 5.53 (1H, t), 4.71 (1H, m), 4.57 (3H, s, NCH3), 4.43 (1H, m), 3.89 (1H, s), 3.65 (1H, m), 3.48 (1H, t) and 3.35-0.90 (complex); 13C NMR (MeOD) δ 144.8, 141.8, 138.8, 135.4, 135.2, 134.6, 132.6, 132.2, 132.0, 126.9, 122.7,122.1, 119.9, 116.3, 113.1, 74.8, 69.7, 68.7, 66.6, 56.9, 53.1, 48.7, 48.4, 46.8, 45.3, 43.2, 40.1, 39.7, 33.0, 28.4, 26.4, 25.4, 24.9, 24.3 and 20.8; HRESIMS m/z calcd for C37H47N4O [M]+ 563.3750, found 563.3674.
2-N,12-O-dimethyl manzamine A trifluoromethanesulfonate (3)
[α]25D + 25.7 (c = 0.14, MeOH); UV λmax (MeOH) 225, 260, 314, 380 nm; IR νmax (CHCl3) 3443 (NH), 3331, 3258, 2981, 2935, 1679, 1606, 1527, 1457, 1391, 1365, 1250, 1157, 1051 cm−1; 1H NMR (DMSO-d6) δ 11.40 (1H, s, NH), 8.84 (2H, m), 8.48 (1H, d), 8.23 (1H, d), 7.72 (1H, t), 7.42 (1H, t), 5.88 (1H, s), 5.52 (1H, m), 5.40 (2H, m), 5.22 (1H, m), 4.41 (3H, s, NCH3), 4.01 (2H, m), 3.81 (1H, d), 3.56 (2H, m), 3.32 (5H, m), 3.20 (2H, m), 3.13 (3H, s, OCH3), 2.89 (3H, m), 2.74 (1H, m) and 1.43-2.19 (complex); 13C NMR (DMSO-d6) δ 145.4, 145.1, 141.8, 138.8, 137.2, 136.1, 135.3, 133.4, 131.9, 129.6, 129.3, 125.8, 123.6, 121.9, 119.8, 117.2, 114.6, 74.1, 62.9, 62.1, 61.5, 59.1, 58.2, 54.1, 50.4, 45.9, 45.0, 43.8, 43.0, 36.7, 27.6, 26.4, 25.8, 25.4, 25.0, 24.2, 23.2 and 18.7; HRESIMS m/z calcd for C38H49N4O [M]+ 577.3906, found 577.4044.
9N-Methylmanzamine A (4)
[α]25D +18.6 (c = 0.07, MeOH); UV λmax (MeOH) 260, 310, 375 nm; IR νmax (CHCl3) 3648 (NH), 3294, 3011, 2924, 2838, 1675, 1628, 1519, 1443, 1329, 1199, 1017 cm−1; 1H NMR (MeOD) δ 8.63, (1H, br s), 8.40 (1H, d), 7.78 (1H, d), 7.53 (1H, d), 7.40 (1H, dd), 6.40 (1H, s), 5.91 (1H, br s), 5.62 (1H, q), 5.52 (1H, td), 5.26 (1H, t), 4.31 (1H, br s), 3.92 (3H, s, NCH3), 3.50 (1H, d), 3.08 (2H, m), 2.78 (1H, m), 2.56 (2H, m), and 1.2-2.5 (complex); 13C NMR (MeOD) δ 154.2, 143.5, 139.9, 138.2, 137.4, 135.1, 134.5, 134.2, 132.3, 129.9, 129.2, 128.5, 122.2, 118.3, 117.2, 113.3, 112.5, 103.5, 75.1, 70.0, 68.7, 56.0, 55.0, 53.5, 50.9, 49.6, 47.1, 44.7, 40.9, 40.3, 32.7, 31.7, 28.1, 26.8, 26.0, 25.7, and 21.7; HRESIMS m/z calcd for C37H47N4O [M+H]+ 563.3750, found 563.3764.
Assay for in vitro Antimalarial Activity and Cytotoxicity
A 200 µL suspension of red blood cells infected with D6 or W2 strain of P. falciparum (2% parasitemia and 2% hematocrit in RPMI 1640 medium supplemented with 10% human serum and 60 µg/mL Amikacin) was added to the wells of a 96-well plate containing 10 µL of serially diluted samples. The plate was flushed with a gas mixture of 90% N2, 5% O2, and 5% CO2 and incubated at 37 °C, for 72 h in a modular incubation chamber (Billups-Rothenberg, CA). Parasitic LDH activity was determined by using MalstatTM reagent (Flow Inc., Portland, OR) according to the procedure of Makler and Hinrichs.[25] 20 µL of the incubation mixture was mixed with 100 µL of the MalstatTM reagent and incubated at room temperature for 30 min followed by addition of 20 µL of a 1:1 mixture of NBT/PES (Sigma, St. Louis, MO) and further incubation in the dark for 1 h. The reaction was stopped by the addition of 100 µL of 5% acetic acid. The plate was read at 650 nm. Artemisinin and chloroquine were included as the drug controls. IC50 values were computed from the dose response curves. To determine the selectivity index of antimalarial activity of compounds their in vitro cytotoxicity to mammalian cells was also determined. The assay was performed as described earlier.[26] Vero cells (monkey kidney fibroblasts) were seeded to the wells of 96-well plate at a density of 25,000 cells/well and incubated for 24 h. Samples at different concentrations were added and plates were again incubated for 48 h. The number of viable cells was determined by Neutral Red assay. IC50 values were obtained from dose response curves. Doxorubicin was used as a positive control.
Assay for Antimicrobial Activity
All organisms are obtained from the American Type Culture Collection (Manassas, VA) and include the fungi Candida albicans ATCC 90028, Cryptococcus neoformans ATCC 90113, and Aspergillus fumigatus ATCC 90906 and the bacteria methicillin-resistant Staphylococcus aureus ATCC 43300 (MRS), Escherichia coli ATCC 35218, Pseudomonas aeruginosa ATCC 27853, and Mycobacterium intracellulare ATCC 23068. Susceptibility testing is performed using a modified version of the CLSI (formerly NCCLS) methods 27–31. M. intracellulare is tested using a modified method of Franzblau, et al.31 Samples are serially-diluted in 20% DMSO/saline and transferred in duplicate to 96 well flat bottom microplates. Microbial inocula are prepared by correcting the OD630 of microbe suspensions in incubation broth to afford final target inocula. Drug controls [Ciprofloxacin (ICN Biomedicals, Ohio) for bacteria and Amphotericin B (ICN Biomedicals, Ohio) for fungi] are included in each assay. All organisms are read at either 630nm using the EL-340 Biokinetics Reader (Bio-Tek Instruments, Vermont) or 544ex/590em, (M. intracellulare, A. fumigatus) using the Polarstar Galaxy Plate Reader (BMG LabTechnologies, Germany) prior to and after incubation. Minimum fungicidal or bactericidal concentrations are determined by removing 5ul from each clear well, transferring to agar and incubating. The MFC/MBC is defined as the lowest test concentration that kills the organism (allows no growth on agar).
Table 3.
Anti-HCV Activity in the Huh-7 Replicon Assay.
| Compds | ΔCt HCV | ΔCt rRNA | % Inhibition |
|
|---|---|---|---|---|
| HCV | rRNA | |||
| 2 | 9.56 | 15.74 | 99.87 | 100.00 |
| RS-446* | 7.48 | −0.25 | 99.43 | −19.22 |
RS-446 (2-Me-C) was used as the control. The compound and the control are tested at 10 µM in triplicate immediately after seeding.
Acknowledgements
We are very grateful to Drs. Subagus Wahyuono and Jiangnan Peng for manzamine A purification; Ms. Marsha Wright and Mr. John Trott for biological testing. Raymond Schinazi’s group is gratefully acknowledged for HCV assays. This work was supported by the NIH, NIAID, Division of AIDS, Grant No. AI 27094 and[GS4] the Public Health Service, grant number R-01 AI 36596-12 from the National Institute of Allergy and Infectious Diseases. Fellowship support for M.A.I was provided through the Ministry of Higher Education and Scientific Research – Egypt.
Footnotes
Publisher's Disclaimer: This is a PDF file of an unedited manuscript that has been accepted for publication. As a service to our customers we are providing this early version of the manuscript. The manuscript will undergo copyediting, typesetting, and review of the resulting proof before it is published in its final citable form. Please note that during the production process errors may be discovered which could affect the content, and all legal disclaimers that apply to the journal pertain.
References and Notes
- 1.Toure YT, Oduola A. Nature Reviews Microbiology. 2004;2(4):276–277. doi: 10.1038/nrmicro870. [DOI] [PubMed] [Google Scholar]
- 2.Ang KKH, Holmes MJ, Higa T, Hamann MT, Kara UAK. Antimicrobial Agents and Chemotherapy. 2000;44(6):1645–1649. doi: 10.1128/aac.44.6.1645-1649.2000. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 3.Chan KL, Oneill MJ, Phillipson JD, Warhurst DC. Planta Medica. 1986;2:105–107. doi: 10.1055/s-2007-969091. [DOI] [PubMed] [Google Scholar]
- 4.Wright CW, Bray DH, Oneill MJ, Warhurst DC, Phillipson JD, Quetinleclercq J, Angenot L. Planta Medica. 1991;57(4):337–340. doi: 10.1055/s-2006-960112. [DOI] [PubMed] [Google Scholar]
- 5.Sakai R, Higa T. Journal of the American Chemical Society. 1986;108(20):6404–6405. [Google Scholar]
- 6.El Sayed KA, Kelly M, Kara UAK, Ang KKH, Katsuyama I, Dunbar DC, Khan AA, Hamann MT. Journal of the American Chemical Society. 2001;123(9):1804–1808. doi: 10.1021/ja002073o. [DOI] [PubMed] [Google Scholar]
- 7.Hu J-F, Hamann MT, Hill R, Kelly M. Alkaloids Chem Biol. 2003;60:207–285. doi: 10.1016/s0099-9598(03)60004-0. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Rao KV, Santarsiero BD, Mesecar AD, Schinazi RF, Tekwani BL, Hamann MT. Journal of Natural Products. 2003;66(6):823–828. doi: 10.1021/np020592u. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 9.Takasu K, Inoue H, Kim HS, Suzuki M, Shishido T, Wataya Y, Ihara M. Journal of Medicinal Chemistry. 2002;45(5):995–998. doi: 10.1021/jm0155704. [DOI] [PubMed] [Google Scholar]
- 10.Takasu K, Shimogama T, Saiin C, Kim HS, Wataya Y, Brun R, Ihara M. Chemical & Pharmaceutical Bulletin. 2005;53(6):653–661. doi: 10.1248/cpb.53.653. [DOI] [PubMed] [Google Scholar]
- 11.Takasu K, Shimogama T, Saiin C, Kim HS, Wataya Y, Ihara M. Bioorganic & Medicinal Chemistry Letters. 2004;14(7):1689–1692. doi: 10.1016/j.bmcl.2004.01.055. [DOI] [PubMed] [Google Scholar]
- 12.Takasu K, Terauchi H, Inoue H, Kim HS, Wataya Y, Ihara M. Journal of Combinatorial Chemistry. 2003;5(3):211–214. doi: 10.1021/cc020119z. [DOI] [PubMed] [Google Scholar]
- 13.Takasu K, Terauchi H, Inoue H, Takahashi M, Sekita S, Ihara M. Heterocycles. 2004;64:215–221. [Google Scholar]
- 14.Chen LB. Annual Review of Cell Biology. 1988;4:155–181. doi: 10.1146/annurev.cb.04.110188.001103. [DOI] [PubMed] [Google Scholar]
- 15.Kawakami M, Koya K, Ukai T, Tatsuta N, Ikegawa A, Ogawa K, Shishido T, Chen LB. Journal of Medicinal Chemistry. 1997;40(20):3151–3160. doi: 10.1021/jm9702692. [DOI] [PubMed] [Google Scholar]
- 16.Modica-Napolitano JS, Aprille JR. Advanced Drug Delivery Reviews. 2001;49(1–2):63–70. doi: 10.1016/s0169-409x(01)00125-9. [DOI] [PubMed] [Google Scholar]
- 17.Weissig V, Torchilin VP. Advanced Drug Delivery Reviews. 2001;49(1–2):127–149. doi: 10.1016/s0169-409x(01)00131-4. [DOI] [PubMed] [Google Scholar]
- 18.Srivastava IK, Rottenberg H, Vaidya AB. Journal of Biological Chemistry. 1997;272(7):3961–3966. doi: 10.1074/jbc.272.7.3961. [DOI] [PubMed] [Google Scholar]
- 19.Blom JF, Brutsch T, Barbaras D, Bethuel Y, Locher HH, Hubschwerlen C, Gademann K. Organic Letters. 2006;8(4):737–740. doi: 10.1021/ol052968b. [DOI] [PubMed] [Google Scholar]
- 20.Hamann MT, Alonso D, Martin-Aparicio E, Fuertes A, Perez-Puerto MJ, Castro A, Morales S, Navarro ML, del Monte-Millan M, Medina M, Pennaka H, Balaiah A, Peng JN, Cook J, Wahyuono S, Martinez A. Journal of Natural Products. 2007;70(9):1397–1405. doi: 10.1021/np060092r. [DOI] [PubMed] [Google Scholar]
- 21.Bax B, Carter PS, Lewis C, Guy AR, Bridges A, Tanner R, Pettman G, Mannix C, Culbert AA, Brown MJB, Smith DG, Reith AD. Structure. 2001;9(12):1143–1152. doi: 10.1016/s0969-2126(01)00679-7. [DOI] [PubMed] [Google Scholar]
- 22.Jones G, Willett P, Glen RC, Leach AR, Taylor R. Journal of Molecular Biology. 1997;267(3):727–748. doi: 10.1006/jmbi.1996.0897. [DOI] [PubMed] [Google Scholar]
- 23.Eldridge MD, Murray CW, Auton TR, Paolini GV, Mee RP. Journal of Computer-Aided Molecular Design. 1997;11(5):425–445. doi: 10.1023/a:1007996124545. [DOI] [PubMed] [Google Scholar]
- 24.Lee B, Richards FM. Journal of Molecular Biology. 1971;55:379–400. doi: 10.1016/0022-2836(71)90324-x. [DOI] [PubMed] [Google Scholar]
- 25.Makler MT, Hinrichs DJ. American Journal of Tropical Medicine and Hygiene. 1993;48(2):205–210. doi: 10.4269/ajtmh.1993.48.205. [DOI] [PubMed] [Google Scholar]
- 26.Mustafa J, Khan SI, Ma G, Walker LA, Khan IA. Lipids. 2004;39(2):167–172. doi: 10.1007/s11745-004-1215-5. [DOI] [PubMed] [Google Scholar]
- 27.NCCLS. NCCLS Document M27-A2. 2002;22(15):1–51. [Google Scholar]
- 28.NCCLS. NCCLS Document M7-A7. 2006;26(2):1–64. [Google Scholar]
- 29.NCCLS. NCCLS Document M38-A. 2002;22(16):1–51. [Google Scholar]
- 30.NCCLS. NCCLS Document M24-A. 2003;23(18):1–84. [Google Scholar]
- 31.Franzblau SG, Witzig RS, McLaughlin JC, Torres P, Madico G, Hernandez A, Degnan MT, Cook MB, Quenzer VK, Ferguson RM, Gilman RH. Journal of Clinical Microbiology. 1998;36(2):362–366. doi: 10.1128/jcm.36.2.362-366.1998. [DOI] [PMC free article] [PubMed] [Google Scholar]