Abstract
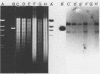
It was demonstrated for the first time that filamentous bacteriophage Cflt, which contains single-stranded DNA, can incorporate its genome into that of its host. Evidence in support of the incorporation was obtained from a Southern blot hybridization analysis of DNA isolated from Cflt-lysogenized cells. DNAs from different Cflt-lysogenized cells were purified, and the integration patterns were compared. Because all integration patterns were identical and only one fragment in Cflt replicative-form DNA was missing, it appears that the integration was site specific. Only one complement of viral DNA was integrated per host chromosome. To determine the attachment site on the viral DNA, the physical map of EcoRI, XhoI, SstII, and BglII on Cflt DNA was constructed. Based on this physical map and a Southern blot hybridization analysis of lysogen DNA with these restriction endonucleases, we demonstrated that DNA sequences from all regions of the Cflt genome were represented in the integrated viral sequences. The attachment site on the viral genome was located at 69.2 to 73.8 min on the Cflt DNA.
Full text
PDF
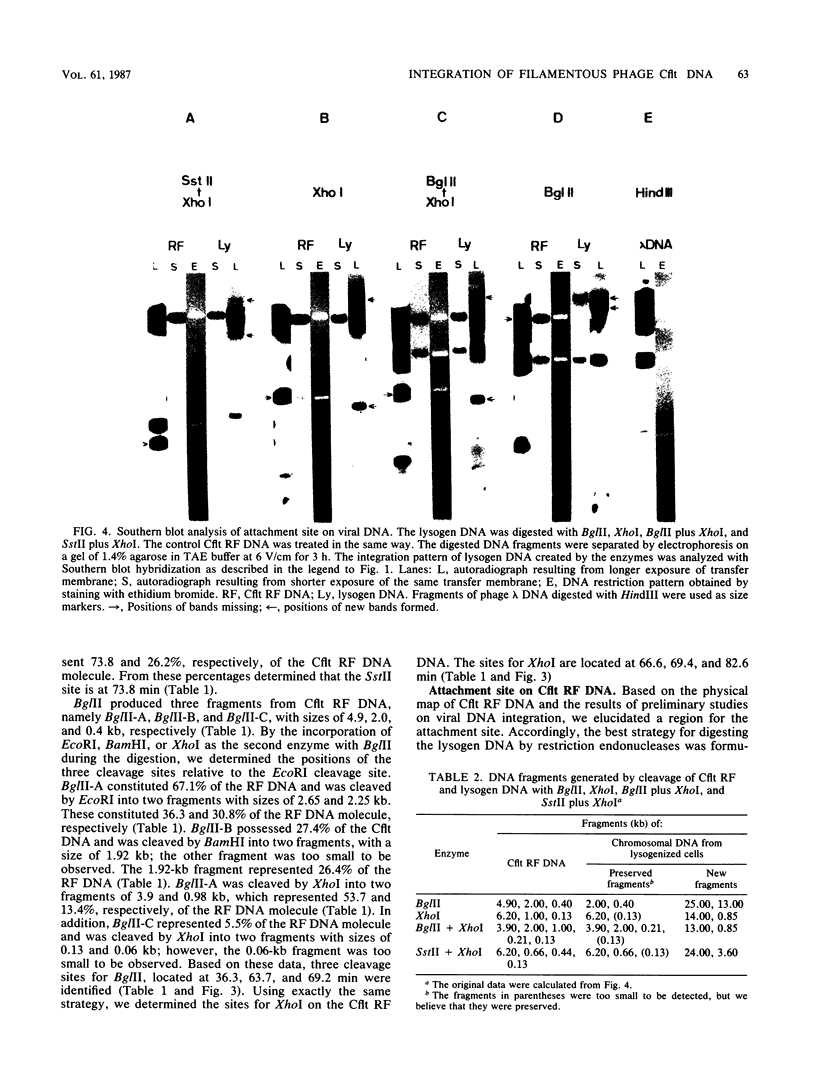





Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Baas P. D. DNA replication of single-stranded Escherichia coli DNA phages. Biochim Biophys Acta. 1985 Jun 24;825(2):111–139. doi: 10.1016/0167-4781(85)90096-x. [DOI] [PubMed] [Google Scholar]
- Barksdale L., Arden S. B. Persisting bacteriophage infections, lysogeny, and phage conversions. Annu Rev Microbiol. 1974;28(0):265–299. doi: 10.1146/annurev.mi.28.100174.001405. [DOI] [PubMed] [Google Scholar]
- Bukhari A. I., Metlay M. Genetic mapping of prophage Mu. Virology. 1973 Jul;54(1):109–116. doi: 10.1016/0042-6822(73)90120-7. [DOI] [PubMed] [Google Scholar]
- Howe M. M., Bade E. G. Molecular biology of bacteriophage mu. Science. 1975 Nov 14;190(4215):624–632. doi: 10.1126/science.1103291. [DOI] [PubMed] [Google Scholar]
- Ikeda H., Tomizawa J. I. Transducing fragments in generalized transduction by phage P1. I. Molecular origin of the fragments. J Mol Biol. 1965 Nov;14(1):85–109. doi: 10.1016/s0022-2836(65)80232-7. [DOI] [PubMed] [Google Scholar]
- Marvin D. A., Hohn B. Filamentous bacterial viruses. Bacteriol Rev. 1969 Jun;33(2):172–209. doi: 10.1128/br.33.2.172-209.1969. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Nash H. A. Integration and excision of bacteriophage lambda: the mechanism of conservation site specific recombination. Annu Rev Genet. 1981;15:143–167. doi: 10.1146/annurev.ge.15.120181.001043. [DOI] [PubMed] [Google Scholar]
- Rigby P. W., Dieckmann M., Rhodes C., Berg P. Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J Mol Biol. 1977 Jun 15;113(1):237–251. doi: 10.1016/0022-2836(77)90052-3. [DOI] [PubMed] [Google Scholar]
- Southern E. M. Detection of specific sequences among DNA fragments separated by gel electrophoresis. J Mol Biol. 1975 Nov 5;98(3):503–517. doi: 10.1016/s0022-2836(75)80083-0. [DOI] [PubMed] [Google Scholar]
- Sternberg N., Hoess R. The molecular genetics of bacteriophage P1. Annu Rev Genet. 1983;17:123–154. doi: 10.1146/annurev.ge.17.120183.001011. [DOI] [PubMed] [Google Scholar]
- Yang M. K., Kuo T. T. A physical map of the filamentous bacteriophage Cf genome. J Gen Virol. 1984 Jul;65(Pt 7):1173–1181. doi: 10.1099/0022-1317-65-7-1173. [DOI] [PubMed] [Google Scholar]