Abstract
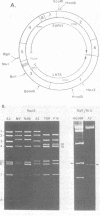
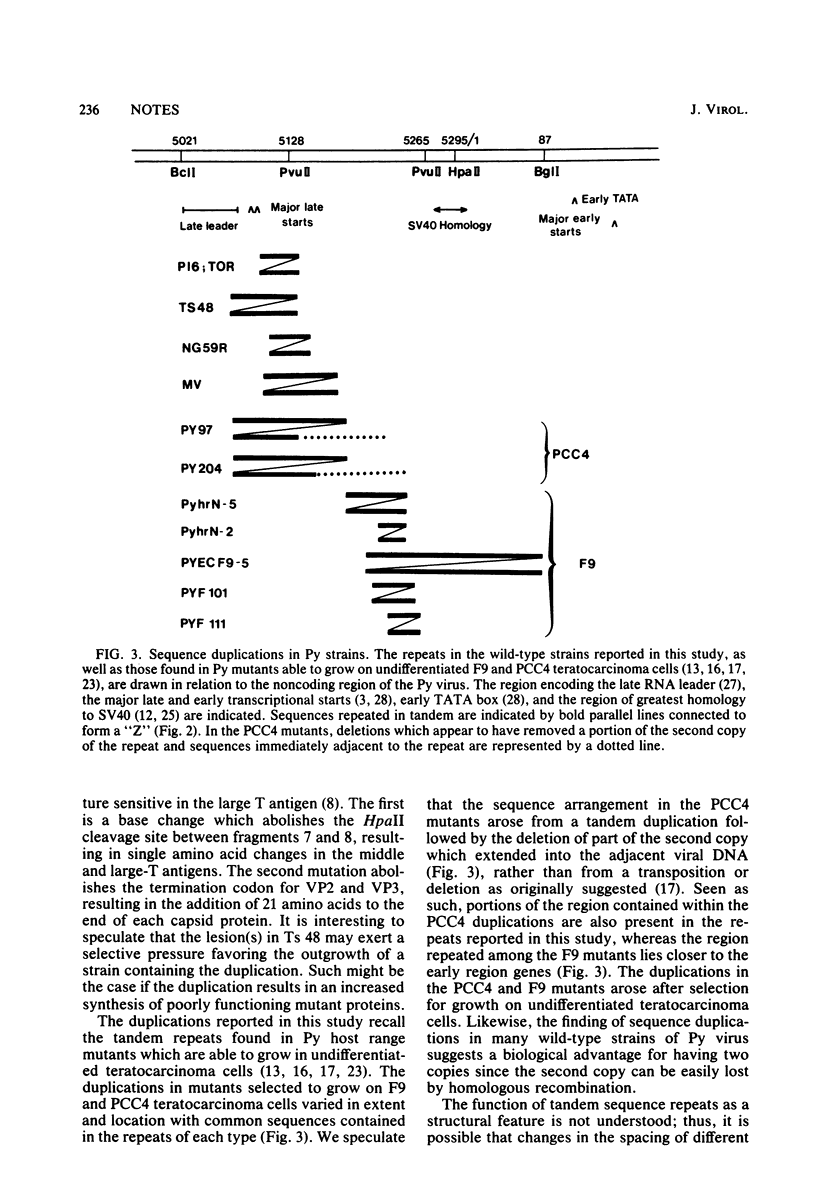
The sequenced prototype strains (A2 and A3) of polyoma virus lack sequence duplications characteristic of other papovaviruses. However, we found that five polyoma virus strains (P16, Toronto large plaque, MV, Ts 48, and NG59R) contain tandemly duplicated sequences in a region near the late RNA leader. Although the duplications vary in size (31 to 84 base pairs) and location (between nucleotide [nt] 5068 and nt 5185), the sequence between nt 5114 and nt 5137 is contained within all five duplicated segments. This region is known to be important in polyoma virus early gene expression, and it contains sequences capable of enhancing the expression of nonviral genes. Inspection of the sequences at and around the ends of the repeats indicated that the duplications do not arise by homologous recombination, and there was no indication that a sequence-specific mechanism results in their formation. However, the variation in the structure of the repeats among different polyoma virus strains suggests that these sequence duplications are a recent evolutionary occurrence. The potential biological significance of this variation is discussed.
Full text
PDF



Images in this article
Selected References
These references are in PubMed. This may not be the complete list of references from this article.
- Banerji J., Rusconi S., Schaffner W. Expression of a beta-globin gene is enhanced by remote SV40 DNA sequences. Cell. 1981 Dec;27(2 Pt 1):299–308. doi: 10.1016/0092-8674(81)90413-x. [DOI] [PubMed] [Google Scholar]
- Benoist C., Chambon P. In vivo sequence requirements of the SV40 early promotor region. Nature. 1981 Mar 26;290(5804):304–310. doi: 10.1038/290304a0. [DOI] [PubMed] [Google Scholar]
- Cowie A., Tyndall C., Kamen R. Sequences at the capped 5'-ends of polyoma virus late region mRNAs: an example of extreme terminal heterogeneity. Nucleic Acids Res. 1981 Dec 11;9(23):6305–6322. doi: 10.1093/nar/9.23.6305. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Crawford L. V., Robbins A. K., Nicklin P. M., Osborn K. Polyoma DNA replication: location of the origin in different virus strains. Cold Spring Harb Symp Quant Biol. 1975;39(Pt 1):219–225. doi: 10.1101/sqb.1974.039.01.028. [DOI] [PubMed] [Google Scholar]
- Deininger P., Esty A., LaPorte P., Friedmann T. Nucleotide sequence and genetic organization of the polyoma late region: features common to the polyoma early region and SV40. Cell. 1979 Nov;18(3):771–779. doi: 10.1016/0092-8674(79)90130-2. [DOI] [PubMed] [Google Scholar]
- Eckhart W., Delbrück S., Deininger P., Friedmann T., Hunter T. A mutation increasing the size of the polyoma virion proteins, VP2 and VP3. Virology. 1981 Feb;109(1):35–46. doi: 10.1016/0042-6822(81)90469-4. [DOI] [PubMed] [Google Scholar]
- Eckhart W. Properties of temperature-sensitive mutants of polyoma virus. Cold Spring Harb Symp Quant Biol. 1975;39(Pt 1):37–40. doi: 10.1101/sqb.1974.039.01.007. [DOI] [PubMed] [Google Scholar]
- Fiers W., Contreras R., Haegemann G., Rogiers R., Van de Voorde A., Van Heuverswyn H., Van Herreweghe J., Volckaert G., Ysebaert M. Complete nucleotide sequence of SV40 DNA. Nature. 1978 May 11;273(5658):113–120. doi: 10.1038/273113a0. [DOI] [PubMed] [Google Scholar]
- Fried M., Griffin B. E., Lund E., Robberson D. L. Polyoma virus--a study of wild-type, mutant and defective DNAs. Cold Spring Harb Symp Quant Biol. 1975;39(Pt 1):45–52. doi: 10.1101/sqb.1974.039.01.009. [DOI] [PubMed] [Google Scholar]
- Fried M., Griffin B. E. Organization of the genomes of polyoma virus and SV40. Adv Cancer Res. 1977;24:67–113. doi: 10.1016/s0065-230x(08)61013-1. [DOI] [PubMed] [Google Scholar]
- Friedmann T., Esty A., LaPorte P., Deininger P. The nucleotide sequence and genome organization of the polyoma early region: extensive nucleotide and amino acid homology with SV40. Cell. 1979 Jul;17(3):715–724. doi: 10.1016/0092-8674(79)90278-2. [DOI] [PubMed] [Google Scholar]
- Fujimura F. K., Deininger P. L., Friedmann T., Linney E. Mutation near the polyoma DNA replication origin permits productive infection of F9 embryonal carcinoma cells. Cell. 1981 Mar;23(3):809–814. doi: 10.1016/0092-8674(81)90445-1. [DOI] [PubMed] [Google Scholar]
- Gruss P., Dhar R., Khoury G. Simian virus 40 tandem repeated sequences as an element of the early promoter. Proc Natl Acad Sci U S A. 1981 Feb;78(2):943–947. doi: 10.1073/pnas.78.2.943. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hirt B. Selective extraction of polyoma DNA from infected mouse cell cultures. J Mol Biol. 1967 Jun 14;26(2):365–369. doi: 10.1016/0022-2836(67)90307-5. [DOI] [PubMed] [Google Scholar]
- Katinka M., Vasseur M., Montreau N., Yaniv M., Blangy D. Polyoma DNA sequences involved in control of viral gene expression in murine embryonal carcinoma cells. Nature. 1981 Apr 23;290(5808):720–722. doi: 10.1038/290720a0. [DOI] [PubMed] [Google Scholar]
- Katinka M., Yaniv M., Vasseur M., Blangy D. Expression of polyoma early functions in mouse embryonal carcinoma cells depends on sequence rearrangements in the beginning of the late region. Cell. 1980 Jun;20(2):393–399. doi: 10.1016/0092-8674(80)90625-x. [DOI] [PubMed] [Google Scholar]
- Magnusson G., Nilsson M. G. Viable polyoma virus variant with two origins of DNA replication. Virology. 1982 May;119(1):12–21. doi: 10.1016/0042-6822(82)90060-5. [DOI] [PubMed] [Google Scholar]
- Marinus M. G. Influence of uvrD3, uvrE502, and recL152 mutations on the phenotypes of Escherichia coli K-12 dam mutants. J Bacteriol. 1980 Jan;141(1):223–226. doi: 10.1128/jb.141.1.223-226.1980. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Maxam A. M., Gilbert W. Sequencing end-labeled DNA with base-specific chemical cleavages. Methods Enzymol. 1980;65(1):499–560. doi: 10.1016/s0076-6879(80)65059-9. [DOI] [PubMed] [Google Scholar]
- Moreau P., Hen R., Wasylyk B., Everett R., Gaub M. P., Chambon P. The SV40 72 base repair repeat has a striking effect on gene expression both in SV40 and other chimeric recombinants. Nucleic Acids Res. 1981 Nov 25;9(22):6047–6068. doi: 10.1093/nar/9.22.6047. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Reddy V. B., Thimmappaya B., Dhar R., Subramanian K. N., Zain B. S., Pan J., Ghosh P. K., Celma M. L., Weissman S. M. The genome of simian virus 40. Science. 1978 May 5;200(4341):494–502. doi: 10.1126/science.205947. [DOI] [PubMed] [Google Scholar]
- Seif I., Khoury G., Dhar R. The genome of human papovavirus BKV. Cell. 1979 Dec;18(4):963–977. doi: 10.1016/0092-8674(79)90209-5. [DOI] [PubMed] [Google Scholar]
- Sekikawa K., Levine A. J. Isolation and characterization of polyoma host range mutants that replicate in nullipotential embryonal carcinoma cells. Proc Natl Acad Sci U S A. 1981 Feb;78(2):1100–1104. doi: 10.1073/pnas.78.2.1100. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Soeda E., Arrand J. R., Smolar N., Walsh J. E., Griffin B. E. Coding potential and regulatory signals of the polyoma virus genome. Nature. 1980 Jan 31;283(5746):445–453. doi: 10.1038/283445a0. [DOI] [PubMed] [Google Scholar]
- Staneloni R. J., Fluck M. M., Benjamin T. L. Host range selection of transformation-defective hr-t mutants of polyoma virus. Virology. 1977 Apr;77(2):598–609. doi: 10.1016/0042-6822(77)90485-8. [DOI] [PubMed] [Google Scholar]
- Treisman R. Characterisation of polyoma late mRNA leader sequences by molecular cloning and DNA sequence analysis. Nucleic Acids Res. 1980 Nov 11;8(21):4867–4888. doi: 10.1093/nar/8.21.4867. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Treisman R., Cowie A., Favaloro J., Jat P., Kamen R. The structures of the spliced mRNAs encoding polyoma virus early region proteins. J Mol Appl Genet. 1981;1(2):83–92. [PubMed] [Google Scholar]
- Tyndall C., La Mantia G., Thacker C. M., Favaloro J., Kamen R. A region of the polyoma virus genome between the replication origin and late protein coding sequences is required in cis for both early gene expression and viral DNA replication. Nucleic Acids Res. 1981 Dec 11;9(23):6231–6250. doi: 10.1093/nar/9.23.6231. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Van Heuverswyn H., Fiers W. Nucleotide sequence of the Hind-C fragment of simian virus 40 DNA. Comparison of the 5'-untranslated region of wild-type virus and of some deletion Mutants. Eur J Biochem. 1979 Oct;100(1):51–60. doi: 10.1111/j.1432-1033.1979.tb02032.x. [DOI] [PubMed] [Google Scholar]
- Vogt M., Bacheler L. T., Boice L. Proposed structure of two defective viral DNA oligomers produced in 3T3 cells transformed by the ts-a mutant of polyoma virus. J Virol. 1976 Mar;17(3):1009–1026. doi: 10.1128/jvi.17.3.1009-1026.1976. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Weaver R. F., Weissmann C. Mapping of RNA by a modification of the Berk-Sharp procedure: the 5' termini of 15 S beta-globin mRNA precursor and mature 10 s beta-globin mRNA have identical map coordinates. Nucleic Acids Res. 1979 Nov 10;7(5):1175–1193. doi: 10.1093/nar/7.5.1175. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Yang R. C., Wu R. BK virus DNA: complete nucleotide sequence of a human tumor virus. Science. 1979 Oct 26;206(4417):456–462. doi: 10.1126/science.228391. [DOI] [PubMed] [Google Scholar]
- de Villiers J., Schaffner W. A small segment of polyoma virus DNA enhances the expression of a cloned beta-globin gene over a distance of 1400 base pairs. Nucleic Acids Res. 1981 Dec 11;9(23):6251–6264. doi: 10.1093/nar/9.23.6251. [DOI] [PMC free article] [PubMed] [Google Scholar]