Abstract
Staphylococcus aureus causes disease in humans and a wide array of animals. Of note, S. aureus mastitis of ruminants, including cows, sheep, and goats, results in major economic losses worldwide. Extensive variation in genome content exists among S. aureus pathogenic clones. However, the genomic variation among S. aureus strains infecting different animal species has not been well examined. To investigate variation in the genome content of human and ruminant S. aureus, we carried out whole-genome PCR scanning (WGPS), comparative genomic hybridizations (CGH), and the directed DNA sequence analysis of strains of human, bovine, ovine, and caprine origin. Extensive variation in genome content was discovered, including host- and ruminant-specific genetic loci. Ovine and caprine strains were genetically allied, whereas bovine strains were heterogeneous in gene content. As expected, mobile genetic elements such as pathogenicity islands and bacteriophages contributed to the variation in genome content between strains. However, differences specific for ruminant strains were restricted to regions of the conserved core genome, which contained allelic variation in genes encoding proteins of known and unknown function. Many of these proteins are predicted to be exported and could play a role in host-pathogen interactions. The genomic regions of difference identified by the whole-genome approaches adopted in the current study represent excellent targets for studies of the molecular basis of S. aureus host adaptation.
Staphylococcus aureus is responsible for a wide range of diseases in animals and humans. Mastitis of dairy cows, sheep, and goats typically is refractory to antibiotic treatment, and an effective vaccine has proved elusive to date.
Population genetic analyses have revealed extensive genetic variation in natural populations of S. aureus (27, 28, 46). Importantly, several previous studies have found that S. aureus clones are largely host specific and are rarely associated with cross-species transfer (22, 39, 47). In particular, the presence of ovine- and bovine-specific lineages of S. aureus was reported (30), and these data suggest that the majority of cases of bovine and ovine mastitis are caused by a small number of host-specialized clones (11).
The genome sequencing of numerous S. aureus strains of human origin has revealed a great deal of variation in gene content, particularly in the complement of mobile genetic elements (MGEs), which could reflect differences in virulence or tissue or disease tropism (32, 33). Several studies have shown that certain virulence genes are overrepresented in some clonal lineages and that some combinations correlate with pathogenic potential (35, 46).
Very recently, Herron-Olson et al. (18) sequenced the genome of a bovine strain (RF122) isolated from mastitis and identified many unique genes that were not found among the human sequenced strains. It is possible that these novel genes could contribute to the host specificity observed among bovine clones. Previously, we constructed a whole-genome microarray specific for the human methicillin-resistant S. aureus (MRSA) strain COL and carried out comparative genomic hybridizations (CGH) to explore the gene content of a small number of bovine and ovine S. aureus strains (12). We found that some bovine strains were genetically allied with the common ovine S. aureus lineage, suggesting that a similar gene complement may be required for bovine and ovine mastitis. However, the CGH analysis was limited by the microarray, which represented the genome of a single human strain and did not allow the identification of genes specific for bovine or ovine strains. Further, CGH does not allow the identification of the genomic location of components of the accessory genome. In contrast, the previously described whole-genome PCR scanning (WGPS) has been successfully used to investigate genome structure diversity in closely related strains of enterohemorrhagic Escherichia coli (34). WGPS is based on the long-range PCR (LR-PCR) amplification of bacterial chromosomes by using a set of primer pairs designed on a reference genome and gives an overall view of genome structure. We developed bioinformatic tools dedicated to WGPS to extend this approach to gram-positive species such as S. aureus (4, 5).
In order to investigate the genetic basis of the host adaptation of ruminant strains of S. aureus, we analyzed their genome structure and content using WGPS and CGH. The combination of these two complementary techniques allowed the identification and genome localization of genetic loci that were divergent in the ruminant strains examined. Unexpectedly, the majority of the regions of difference (RDs) specific for ruminant strains was located in the core genome and was not associated with MGEs. A majority of these determinants encode proteins predicted to be exported, suggesting a role in S. aureus-ruminant host interactions.
MATERIALS AND METHODS
Bacterial strains.
The WGPS and CGH studies were carried out on 12 strains (Table 1) isolated from human, bovine, ovine, and caprine hosts. Strain N315, whose genome sequence is publicly available, was used as a reference strain in the CGH experiments and in primer design for WGPS, and strain Mu50, another sequenced strain, was used to validate the experimental data obtained in WGPS. The bovine strain RF122 was isolated from clinical mastitis and corresponds to a predominant and widespread clone associated with bovine mastitis that has been completely sequenced. The three other bovine strains studied were isolated from subclinical mastitis and were chosen because they reproducibly induced subclinical mastitis in experimental udder infections in cows (3, 17, 36, and F. Gilbert, IASP-INRA Tours, personal communication). Ovine and caprine strains were isolated from clinical mastitis, the predominant form of the disease in those animals. Twenty-eight additional strains from human (n = 4), bovine (n = 12), and ovine-caprine (n = 12) origin were screened for host biotype-specific variations by diagnostic PCR tests.
TABLE 1.
Staphylococcus aureus strains used in this study
| Strain | Host | Source and/or type of infectiona | Origin | Year isolated | STb | CPc | Reference or source |
|---|---|---|---|---|---|---|---|
| N315 | Human | Pharynx infection MRSA | Japan | 1982 | 5 | 5 | 25 |
| Mu50 | Human | Wound infection MRSA, GISA | Japan | 1997 | 5 | 5 | 25 |
| 1178 | Human | Rachis infection | France | 2004 | 5 | ND | CHU Rennesd |
| 1183 | Human | Cystic fibrosis hypermutator | France | 2003 | 15 | ND | CHU Caene |
| RF122 | Bovine | Mastitis | Ireland | 1993 | 151 | 8 | 18; TCD, Dublin, Irelandf |
| 1166 | Bovine | Mastitis | France | 1964 | 97 | 5 | INRA Toursg |
| 1167 | Bovine | Mastitis | France | 1966 | 126 | 5 | INRA Tours |
| Newbould N305 | Bovine | Mastitis | United States | 1974 | 115 | 5 | INRA Tours |
| 1170 | Caprine | Mastitis | France | 1987 | 711 | NT | INRA Tours |
| 1171 | Caprine | Mastitis | France | 1966 | 712 | 8 | INRA Tours |
| 1173 | Ovine | Mastitis | Australia | 133 | NT | INRA Tours | |
| 1174 | Ovine | Mastitis | France | 1997 | 133 | 8 | INRA Tours |
| 1169 | Goat | Mastitis | France (central) | 1974 | ND | NT | INRA Tours |
| 1536 | Ovine | Clinical mastitis | France (southwest) | NA | ND | 8 | ENV-Toulouseh |
| 1367 | Ovine | Clinical mastitis | France (southeast) | 2001 | ND | 8 | AFSSA Sophia Antipolisi |
| 1281 | Ovine | Gangrenous mastitis | France (southeast) | 2001 | ND | 8 | AFSSA Sophia Antipolis |
| 1535 | Ovine | Subclinical mastitis | France (southwest) | NA | ND | 8 | ENV-Toulouse |
| 1366 | Ovine | Subclinical mastitis | France (Corsica) | 2001 | ND | 5 | AFSSA Sophia Antipolis |
| 1284 | Ovine | Udder abscess | France (southeast) | 2001 | ND | 8 | AFSSA Sophia Antipolis |
| 1533 | Goat | Udder skin | France (central) | NA | ND | 8 | INRA Aurillacj |
| 1524 | Goat | Udder skin | France (central) | NA | ND | 8 | INRA Aurillac |
| 1517 | Ovine | Nasal carriage | France (Bretagne) | 2006 | ND | 8 | INRA Rennesk |
| 1261 | Ovine | Nasal carriage | France (southeast) | 2003 | ND | NT | AFSSA Sophia Antipolis |
| 1213 | Ovine | Ovine biotype | NA | NA | ND | 8 | University of Gentl |
| 1259 | Bovine | Mastitis | Brazil (Rio de Janeiro) | 2005 | ND | NT | UFRJ, Rio de Janeiro, Brazilm |
| 1250 | Bovine | Mastitis | Belgium | 1974 | ND | 8 | University of Gent |
| 1245 | Bovine | Mastitis | France (Normandy) | 1967 | ND | NT | INRA Tours |
| 1241 | Bovine | Mastitis | France (central) | 2003 | ND | 5 | INRA Tours |
| 1242 | Bovine | Mastitis | France (west) | 2000 | ND | 8 | INRA Tours |
| 1231 | Bovine | Mastitis | France (central) | 1975 | ND | 5 | INRA Tours |
| 1247 | Bovine | Mastitis | Belgium | 1974 | ND | 5 | University of Gent |
| 1239 | Bovine | Mastitis | France (central) | 1996 | ND | 8 | INRA Tours |
| 1244 | Bovine | Mastitis | France (East) | 2000 | ND | 5 | INRA Tours |
| 1296 | Bovine | Cow milk | Brazil (Minas Gerais) | 2003 | ND | 8 | UFMG, Belo Horizonte, Braziln |
| 1465 | Bovine | Nasal carriage | France (Bretagne) | 2006 | ND | NT | INRA Rennes |
| 1479 | Bovine | Nasal carriage | France (Bretagne) | 2006 | ND | 5 | INRA Rennes |
| 1294 (MW2) | Human | MRSA, hospital pediatric infection | North Dakota | 1999 | ND | ND | 2 |
| 1177 | Human | Hospital-acquired gut infection | France (Bretagne) | 2003 | ND | ND | CHU Rennes |
| 1180 (MSSA476) | Human | Invasive community-acquired MSSA | United Kingdom | 1998 | ND | ND | 19 |
| 1181 (MRSA252) | Human | Epidemic MRSA | United Kingdom | 1997 | ND | ND | 19 |
GISA, glycopeptide-intermediate S. aureus; MSSA, methicillin-susceptible S. aureus.
STs were determined as described in Material and Methods. ND, not determined.
CP, capsular polysaccharide serotype, determined as described in Goerke et al. (15). 5, cap5 serotype; 8 cap8 serotype; NT, nontypeable (i.e., non-cap5, non-cap8).
Kindly provided by O. Gaillot, CHU Pontchaillou, Faculté de Médecine Rennes 1, France.
Kindly provided by R. Leclercq, CHU Côte de Nacre, Faculté de Médecine de Caen, France.
From the laboratory collection of J. R. Fitzgerald, University of Edinburgh, United Kingdom.
Kindly provided by F. Gilbert and P. Rainard, INRA de Tours, France.
Kindly provided by D. Bergonnier, Ecole Nationale Vétérinaire de Toulouse, France.
Kindly provided by E. Vautor, AFSSA Sophia Antipolis, France.
Kindly provided by M. C. Montel, INRA Aurillac, France.
From the laboratory collection of INRA Rennes, France.
Kindly provided by D. Vancraeynest, University of Ghent, Belgium.
Kindly provided by W. Lilenbaum, Federal University of Rio de Janeiro, Brazil.
Kindly provided by H. F. T. Cardoso, Federal University of Minas Gerais, Brazil.
Genotyping of the strains.
Strains were subjected to SmaI macrorestriction pulsed-field gel electrophoresis (PFGE). Using the unweighted-pair group method using average linkages and a Dice coefficient (with a tolerance limit of 1.1%), a phylogenetic tree was constructed as described previously (20). In parallel, allelic profiles of seven housekeeping genes were determined for the 12 strains initially examined, as described previously (10). They were compared to all of the other S. aureus STs found on the multilocus sequence typing (MLST) database (http://saureus.mlst.net) using eBURST software (http://www.mlst.net/BURST/burst.htm). A phylogenetic tree was constructed using MEGA 4.0 software and the neighbor-joining method with a bootstrap value of 1,000 (44).
DNA-DNA microarray hybridization.
DNA-DNA microarray hybridization was used to assess variation in gene content among the 12 S. aureus strains. Hybridizations were performed as described previously (12, 48). Briefly, genomic DNA extracted from strains 1178, 1183, RF122, 1166, 1167, Newbould N305, 1170, 1171, 1173, and 1174 (hereafter designated test strains) were hybridized on the microarray with strain N315 (hereafter designated the control strain) genomic DNA as the reference. Three independent hybridizations were performed for each test strain with the probe set (gene) detection, as determined by using the one-sided Wilcoxon's signed rank test and the one-step Tukey's biweight estimate for signal detection (Affymetrix, Inc., Santa Clara, CA). Coding sequences (CDSs) that gave an equivalent hybridization signal for both the control and test strains were designated common and considered present in the two genomes (test and control strains). Common CDSs are indicated in Fig. 2 and in Table S1 in the supplemental material. CDSs that gave a low signal compared to that of the control N315 were designated poorly detected (see Table 4). They likely had enough sequence divergence that strong hybridization did not occur in one or two of the three independent hybridizations (allelic variation was suspected). They are indicated in Fig. 2 and in Table S1 in the supplemental material. Genes that gave no signal at all in each of the hybridizations compared to that of the control N315 were designated not detected and either were absent from the genome (gene loss was suspected) or had enough sequence divergence that hybridization did not occur under the conditions of stringency used (allelic variation was suspected). They are indicated in Fig. 2 and in Table S1 in the supplemental material. Genes that gave a signal in the test strain but not in the control N315 were designated additional (gene gain was suspected).
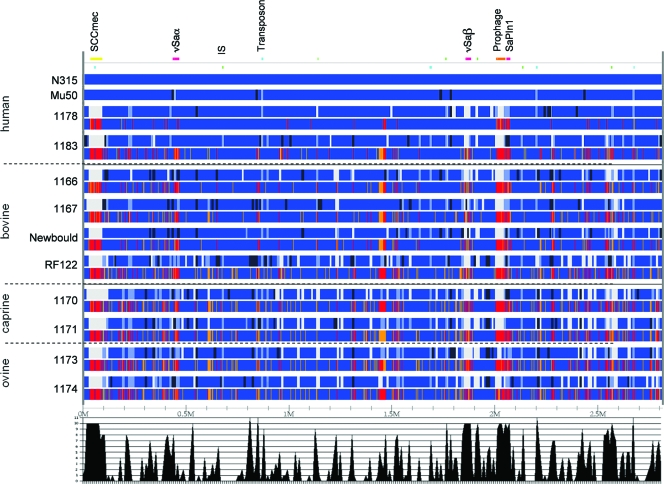
FIG. 2.
Summary and data comparison of the WGPS and the microarray analyses of the set of 12 S. aureus strains. Results of the WGPS of the 12 strains are shown in the upper lines. WGPS scanning analysis is presented as follows. Segments that yielded PCR products of identical, smaller, and bigger sizes compared to those in N315 are depicted by blue, dark blue, and light blue rectangles, respectively. Fragments that were not amplified are depicted by white rectangles. Fragments that were demonstrated as being absent by framing PCR experiments are depicted in gray. The positions of the MGEs, as determined in the N315 annotation, are given in the row above WGPS results. Results from the gene content analysis using the whole-genome microarray are shown in the lower lines. CGH analyses were carried out using N315 as the reference (Mu50, whose gene content is known, was not analyzed here). Genes that gave a signal in both N315 and the test strain were considered present and are indicated in blue. Genes that gave a low signal or no signal at all compared to that of N315 were judged as variable and are indicated in orange and red, respectively. The lower part of the figure summarizes the success rates of the WGPS. For each segment, the number of strains (1 to 12) that yielded an LR-PCR product is given.
TABLE 4.
Specific loci identified by both CGH and WGPS
| COG gene | PSORT determinationa | Description | Classification by functionb | Result by:
|
Suspected event | |
|---|---|---|---|---|---|---|
| CGHc | WGPSd | |||||
| Bovine specific | ||||||
| SA1091 | Membrane | FmhC protein; fmhC | CECP; cell wall | ND | 128-129 (NA) | Allelic variation or gene loss |
| SA1430 | Secreted/SP | Hypothetical protein; similar to enterotoxin A precursor | OF; pathogenic factors (toxins and colonization factors) | ND | 171 (<) | Allelic variation or gene loss |
| SA2003 | Secreted/SP | Hyaluronate lyase precursor; hysA | OF; pathogenic factors (toxins and colonization factors) | A | Gene gain | |
| SA2004 | Cytoplasm | Conserved hypothetical protein; possible secreted peptidase | Similar to unknown proteins | ND | 238-239 (NA) | Allelic variation or gene loss |
| SAR0436 | Secreted/SP | Hypothetical protein | No similarity | A | Allelic variation | |
| Ovine-caprine specific | ||||||
| SA0523 | Cytoplasm | Hypothetical protein; similar to poly(glycerol-phosphate) alpha-glucosyltransferase (teichoic acid biosynthesis) | CECP; cell wall | PD | 62 (var) | Allelic variation |
| SAS0861 | Secreted/PS | ABC-type dipeptide transport system | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SA1848 | Membrane | Probable ammonium transporter; nrgA | CECP; transport/binding proteins and lipoproteins | PD | 219 (NA) | Allelic variation |
| SA2475 | Membrane | Conserved hypothetical aminoacyl-tRNA synthetases class II signatures | CECP; transport/binding proteins and lipoproteins | ND | 292 (var) | Allelic variation |
| SA2476 | Membrane | Hypothetical protein; similar to cation ABC transporter; ATP-binding protein | CECP; transport/binding proteins and lipoproteins | ND | 292 (var) | Allelic variation |
| SA0579 | Membrane/CWA | Hypothetical protein; similar to Na+/H+ antiporter | CECP; membrane bioenergetics (electron transport chain and ATP synthase) | ND | 68 (NA) | Allelic variation or gene loss |
| SA0581 | Membrane | MnhD homologue; similar to Na+/H+ antiporter subunit | CECP; membrane bioenergetics (electron transport chain and ATP synthase) | PD | 68-69 (NA, <) | Gene loss |
| SA2184 | Cytoplasm | Nitrate reductase beta chain narH energy metabolism; anaerobic | CECP; membrane bioenergetics (electron transport chain and ATP synthase) | PD | 256-257 (=, NA) | Allelic variation |
| SA0171 | Secreted/SP | NAD-dependent formate dehydrogenase; fdh | IM; metabolism of carbohydrates and related molecules | ND | 19 (<) | Gene loss |
| SA1200 | Outside/no SP | Anthranilate synthase component II; trpG | IM; metabolism of amino acids and related molecules | PD | 143 (=) | Allelic variation |
| SA1201 | Membrane/no SP | Anthranilate phosphoribosyltransferase; trpD | IM; metabolism of amino acids and related molecules | PD | 143-144 (=, =) | Allelic variation |
| SA2120 | Cytoplasm | Hypothetical protein; similar to amino acid amidohydrolase | IM; metabolism of amino acids and related molecules | PD | 249-250 (=, =) | Allelic variation |
| SA2189 | Cytoplasm | Hypothetical protein; nirR nitrogen metabolism | IM; metabolism of amino acids and related molecules | PD | 257 (NA) | Allelic variation or gene loss |
| SA2469 | Cytoplasm | Hypothetical protein; similar to histidinol-phosphate transaminase | IM; metabolism of amino acids and related molecules | PD | 291 (=) | Allelic variation |
| SA1324 | Membrane | Ribosomal large subunit pseudouridine synthase b; rluB | IM; metabolism of nucleotides and nucleic acids | PD | 160 (NA) | Allelic variation or gene loss |
| SA0328 | Membrane | Hypothetical protein; similar to NADH-dependent fmn reductase | IM; metabolism of coenzymes and prosthetic groups | PD | 38 (=) | Allelic variation |
| SA1538 | Cytoplasm | Hypothetical protein; similar to iron-sulfur cofactor synthesis protein; nifZ | IM; metabolism of coenzymes and prosthetic groups | A | Gene gain | |
| SA1525 | Cytoplasm | DNA polymerase III, alpha chain; dnaE | IP; DNA replication | PD | 182-183 (=, =) | Allelic variation |
| SAR1898 | Cytoplasm | Putative type I restriction modification DNA specificity protein | IP; DNA modification and repair | A | Gene gain | |
| SA1145 | Cytoplasm | Hypothetical protein; similar to host factor 1 | OF; phage-related functions | PD | 136 (=) | Allelic variation |
| SACOL0350 | Cytoplasm | Hypothetical protein | OF; phage-related functions | A | Gene gain | |
| SACOL0336 | Cytoplasm | Hypothetical protein | OF; phage-related functions | A | Gene gain | |
| SA0909 | Membrane/SP | FmtA, autolysis and methicillin resistant-related protein; fmtA | OF; pathogenic factors (toxins and colonization factors) | PD | 107 (=) | Allelic variation |
| SA1941 | Cytoplasm | General stress protein 20U; dps | OF; adaptation to atypical conditions | PD | 230 (=) | Allelic variation |
| SA1193 | Membrane | Oxacillin resistance-related FmtC protein; fmtC | OF; miscellaneous | PD | 142-143 (=, =) | Allelic variation |
| SA0170 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 19 (<) | Gene loss |
| SAR0358 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | A | Allelic variation | |
| SA0355 | Cytoplasm | Hypothetical protein; similar to hypothetical protein virulence plasmid pXO1-38 | Similar to unknown proteins | ND | 41 (=) | Allelic variation |
| SAR0602 | Membrane | Putative membrane protein | Similar to unknown proteins | A | Allelic variation | |
| SA0783 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | PD | 91 (=) | Allelic variation |
| SA0976 | Membrane/SP | Conserved hypothetical protein; possesses a CWA | Similar to unknown proteins | PD | 114 (=) | Allelic variation |
| SA1578 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 190 (NA) | Allelic variation or gene loss |
| SA1957 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 232 (>) | Allelic variation |
| SA2477 | Membrane/SP | Conserved hypothetical protein; prokaryotic membrane lipoprotein lipid attachment site | Similar to unknown proteins | ND | 292 (var) | Allelic variation |
| SA2478 | Cytoplasm | Conserved hypothetical protein; similar to protein of unknown function in Helicobacter pylori | Similar to unknown proteins | ND | 292 (var) | Allelic variation |
| SA0372 | Cytoplasm | Hypothetical protein | No similarity | PD | 43 (>) | Allelic variation |
| SAR0627 | Cytoplasm | Hypothetical protein | No similarity | A | Allelic variation | |
| SA1284 | Cytoplasm | Hypothetical protein | No similarity | PD | 155 (=) | Allelic variation |
| SACOL0373 | Cytoplasm | Hypothetical protein (bacteriophage) | No similarity | A | Gene gain | |
| SAV1989 | Cytoplasm | Hypothetical protein (bacteriophage ΦMu50) | No similarity | A | Allelic variation | |
| SA2055 | Membrane | Hypothetical protein | No similarity | PD | 243 (=) | Allelic variation |
| SA2292 | Cytoplasm | Hypothetical protein | No similarity | PD | 269 (NA) | Allelic variation or gene loss |
| SAS2540 | Outside/CWA | Putative CWA protein | No similarity | A | Allelic variation | |
| SA2450 | Cytoplasm | Hypothetical protein | No similarity | PD | 290 (<) | Gene loss |
| SAR2739 | Cytoplasm | Hypothetical protein | No similarity | A | Allelic variation | |
| SA0129 | Secreted/SP | Hypothetical protein | No similarity | ND | 14 (=) | Allelic variation |
| SAS019 | Membrane | Hypothetical protein | No similarity | PD | 88 (=) | Allelic variation |
| SAS041 | Membrane/SP | Hypothetical protein | No similarity | PD | 138 (=) | Allelic variation |
| SAS057 | Membrane | Hypothetical protein | No similarity | PD | 210 (=) | Allelic variation |
| SACOL0329 | Cytoplasm | Conserved hypothetical protein (bacteriophage) | No similarity | A | Allelic variation | |
| CAC1895 | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
| CAC1934 | Cytoplasm | Conserved hypothetical protein | A | Gene gain | ||
| L103086 | Membrane | Unique predicted open reading frames | A | Gene gain | ||
| L28615 | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
| L42302 | Cytoplasm | Conserved hypothetical protein | A | Gene gain | ||
| lin0924 | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
| lin0925 | Membrane | Unique predicted open reading frames | A | Gene gain | ||
| MA2121 | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
| NMA2192 | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
| XF2526 | Membrane | Unique predicted open reading frames | A | Gene gain | ||
| yi5B | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
| Ruminant specific | ||||||
| SA1090 | Membrane/SP | lytN; LytN protein | CECP; cell wall | ND | 128-129 (var, NA) | Allelic variation or gene loss |
| SA2008 | Cytoplasm | Alpha-acetolactate synthase; alsS | IM; metabolism of carbohydrates and related molecules | PD | 239 (NA) | Allelic variation |
| SA0734 | Cytoplasm | Carboxyesterase precursor homologue; found in S. epidermidis and S. haemolyticus | IM; metabolism of lipids | PD | 86 (var) | Allelic variation |
| SA0386 | Secreted/SP | Exotoxin 10 SSL10 (pathogenicity island SaPIn2) | OF; pathogenic factors (toxins and colonization factors) | PD | 44 (var) | Allelic variation or gene loss |
| SA0742 | Membrane/SP | clfA; clumping factor A; CWA | OF; pathogenic factors (toxins and colonization factors) | ND | 86-87 (var, =) | Allelic variation |
| SAR2709 | Membrane/CWA | clfB; clumping factor B | OF; pathogenic factors (toxins and colonization factors) | A | Allelic variation | |
| SA0096 | Membrane/SP | Hypothetical protein; prokaryotic membrane lipoprotein lipid attachment site | No similarity | PD | 10 (NA) | Allelic variation |
| SAV1489 | Cytoplasm | Hypothetical protein | No similarity | ND | 159-160 (var, NA) | Allelic variation or gene loss |
| SA1753 | Membrane | Hypothetical protein (bacteriophage ΦN315) | No similarity | ND | 211 (NA) | Allelic variation or gene loss |
| SA1754 | Outside/SP | Hypothetical protein (bacteriophage ΦN315) | No similarity | ND | 211 (NA) | Allelic variation or gene loss |
| RF122 + ovine-caprine | ||||||
| SAR2501 | Outside/no SP | FmhA protein; fmhA | CECP; cell wall | A | Allelic variation | |
| FemAB family protein | ||||||
| SAR2649 | Membrane | Hypothetical protein; similar to acyltransferase; oatA | CECP; cell wall | PD | 276-277 (=, NA) | Allelic variation |
| SA0166 | Cytoplasm | Hypothetical protein; similar to nitrate transporter | CECP; transport/binding proteins and lipoproteins | PD | 18-19 (=, var) | Allelic variation or gene loss |
| SA0172 | Membrane | Hypothetical protein; similar to integral membrane protein LmrP; lipoprotein lipid attachment site | CECP; transport/binding proteins and lipoproteins | ND | 19 (var) | Allelic variation or gene loss |
| SA0294 | Membrane/SP | Hypothetical protein; similar to branched-chain amino acid uptake carrier; lipoprotein | CECP; transport/binding proteins and lipoproteins | PD | 34-35 (var, <) | Gene loss |
| SA0655 | Membrane | Fructose specific permease fruA | CECP; transport/binding proteins and lipoproteins | PD | 76 (=) | Allelic variation |
| SA1156 | Membrane | ABC transporter (ATP-binding protein) homolog | CECP; transport/binding proteins and lipoproteins | ND | 138 (var) | Allelic variation or gene loss |
| SA0848 | Cytoplasm | Oligopeptide transport system; ATP-binding protein; oppF homologue | CECP; transport/binding proteins and lipoproteins | A | Gene gain | |
| SAR1761 | Membrane/SP | Lysine-specific permease; lysP | CECP; transport/binding proteins and lipoproteins | PD | 179-180 (=, var) | Allelic variation or gene loss |
| SA1592 | Membrane | Arsenical pump membrane protein homolog | CECP; transport/binding proteins and lipoproteins | PD | 192 (=) | Allelic variation |
| SA1958 | Cytoplasm | Hypothetical protein; similar to ABC transporter | CECP; transport/binding proteins and lipoproteins | PD | 232 (var) | Allelic variation |
| SA1960 | Membrane | Phosphotransferase system; mannitol-specific IIBC component; mtlF | CECP; transport/binding proteins and lipoproteins | ND | 232 (var) | Allelic variation |
| SAR2268 | Secreted/SP | Putative transport system binding lipoprotein | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SAR2371 | Membrane | Putative membrane protein | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SA2142 | Membrane | Hypothetical protein; similar to multidrug resistance protein | CECP; transport/binding proteins and lipoproteins | ND | 252 (var) | Allelic variation |
| SA2191 | Membrane | Hypothetical protein; similar to nirC | CECP; transport/binding proteins and lipoproteins | ND | 257-258 (NA, NA) | Allelic variation |
| SA2242 | Membrane | Conserved hypothetical protein | CECP; transport/binding proteins and lipoproteins | ND | 263 (=) | Allelic variation |
| SA2243 | Cytoplasm | Hypothetical protein; similar to ABC transporter (ATP-binding protein) | CECP; transport/binding proteins and lipoproteins | ND | 263 (=) | Allelic variation |
| SAR2632 | Membrane/SP | Putative transport protein | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SAR2700 | Membrane/CWA | Putative membrane protein | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SAR2778 | Membrane | Putative nickel transport protein | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SAR2782 | Membrane | ABC transporter permease protein | CECP; transport/binding proteins and lipoproteins | A | Allelic variation | |
| SA1653 | Outside/no SP | Signal transduction protein TRAP | CECP; sensors (signal transduction) | ND | 198 (=) | Allelic variation |
| SA1158 | Membrane | Hypothetical protein; similar to two-component sensor histidine kinase | CECP; sensors (signal transduction) | ND | 138 (var) | Allelic variation or gene loss |
| SA0810 | Membrane | NA+/H+ antiporter subunit; mnhD | CECP; membrane bioenergetics (electron transport chain and ATP synthase) | PD | 94 (=) | Allelic variation |
| SA2446 | Membrane | Hypothetical protein; similar to preprotein translocase; secY | CECP; protein secretion | PD | 288-289 (=, var) | Allelic variation |
| SA2102 | Cytoplasm | Formate dehydrogenase homolog | IM; metabolism of carbohydrates and related molecules | PD | 247-248 (NA, var) | Allelic variation or gene loss |
| SA2304 | Outside/no SP | Fructose-bisphosphatase; fbp | IM; metabolism of carbohydrates and related molecules | PD | 271 (var) | Allelic variation or gene loss |
| SAR2635 | Cytoplasm | Putative acetyltransferase | IM; metabolism of carbohydrates and related molecules | A | Allelic variation | |
| SA2490 | Cytoplasm | Hypothetical protein; similar to n-hydroxyarylamine o-acetyltransferase | IM; metabolism of carbohydrates and related molecules | PD | 293-294 (var, =) | Allelic variation or gene loss |
| SAV1464 | Membrane | 3-Phosphoshikimate 1-carboxyvinyltransferase(5-enolpyruvylshikimate-3-phosphate synthase) (EPSP synthase) | IM; metabolism of amino acids and related molecules | A | Allelic variation | |
| SA1299 | Outside/no SP | Chorismate synthase; aroC | IM; metabolism of amino acids and related molecules | A | Gene gain | |
| SA1866 | Cytoplasm | Threonine dehydratase; ilvA | IM; metabolism of amino acids and related molecules | ND | 221-222 (=, =) | Allelic variation |
| SA0646 | Cytoplasm | Hypothetical protein; similar to deoxyribodipyrimidine photolyase | IM; metabolism of nucleotides and nucleic acids | PD | 75 (=) | Allelic variation |
| SAR1040 | Cytoplasm | Phosphoribosylaminoimidazole-succinocarboxamide synthetase; purC | IM; metabolism of nucleotides and nucleic acids | A | Allelic variation | |
| SA0533 | Membrane | Hypothetical protein, similar to long-chain fatty acid coenzyme A ligase; vraA | IM; metabolism of lipids | PD | 64 (=) | Allelic variation |
| SA1398 | Membrane | Hypothetical protein; similar to diacylglycerol kinase | IM; metabolism of lipids | PD | 168 (=) | Allelic variation |
| SA0317 | Cytoplasm | Hypothetical protein; similar to dihydroflavonol-4-reductase | IM; metabolism of coenzymes and prosthetic groups | PD | 37 (<) | Gene loss |
| SA0472 | Membrane | Dihydropteroate synthase chain A synthetase; folP | IM; metabolism of coenzymes and prosthetic groups | PD | 55 (NA) | Allelic variation or gene loss |
| SA1537 | Cytoplasm | Hypothetical protein; similar to thiamine biosynthesis protein; thiI | IM; metabolism of coenzymes and prosthetic groups | PD | 184 (=) | Allelic variation |
| SA0476 | Cytoplasm | Hypothetical protein; similar to transcription regulator; gntR family | IP; RNA synthesis | PD | 56-57 (>, =) | Allelic variation |
| SA0614 | Cytoplasm | Hypothetical protein; similar to two-component response regulator | IP; RNA synthesis | PD | 72 (=) | Allelic variation |
| SA0627 | Cytoplasm | Hypothetical protein; similar to LysR family transcriptional regulator | IP; RNA synthesis | PD | 73 (=) | Allelic variation |
| SA1159 | Not clear | Hypothetical protein; similar to two-component response regulator | IP; RNA synthesis | PD | 138 (var) | Allelic variation or gene loss |
| SA1591 | Cytoplasm | Arsenical resistance operon repressor homolog | IP; RNA synthesis | PD | 192 (=) | Allelic variation |
| SA1748 | Cytoplasm | Hypothetical protein; similar to transcription regulator; gntR family | IP; RNA synthesis | PD | 210 (=) | Allelic variation |
| SA1930 | Cytoplasm | Probable DNA-directed RNA polymerase delta subunit; rpoE | IP; RNA synthesis | PD | 229 (=) | Allelic variation |
| SA2340 | Cytoplasm | Hypothetical protein; similar to transcriptional regulator; tetR family | IP; RNA synthesis | PD | 275 (NA) | Allelic variation or gene loss |
| SA2364 | Cytoplasm | Hypothetical protein; similar to transcription regulator; acrR | IP; RNA synthesis | PD | 278 (NA) | Allelic variation or gene loss |
| SAR0836 | Cytoplasm | Putative RNase R; rnr | IP; RNA modification | A | Allelic variation | |
| SA1114 | Cytoplasm | tRNA pseudouridine 5S synthase; truB | IP; RNA modification | PD | 131-132 (NA, =) | Allelic variation or gene loss |
| SAR1278 | Cytoplasm | Putative tRNA delta(2)-isopentenylpyrophosphate transferase; miaA | IP; RNA modification | A | Allelic variation | |
| SA0330 | Cytoplasm | Hypothetical protein; similar to ribosomal-protein-serine N-acetyltransferase | IP; protein modification | PD | 38 (=) | Allelic variation |
| SA2482 | Cytoplasm | Pyrrolidone-carboxylate peptidase; pcp | IP; protein modification | PD | 292-293 (var, var) | Allelic variation or gene loss |
| SA0145 | Cytoplasm | Capsular polysaccharide synthesis enzyme Cap5B; capB | OF; adaptation to atypical conditions | PD | 16 (=) | Allelic variation |
| SA0356 | Cytoplasm | Truncated integrase | OF; phage-related functions | PD | 41 (=) | Allelic variation |
| SACOL0339 | Outside/no SP | Prophage L54a; single-stranded DNA binding protein (bacteriophage) | OF; phage-related functions | A | Gene gain | |
| SA1222 | Cytoplasm | Truncated transposase | OF; transposon and IS | PD | 146 (NA) | Allelic variation or gene loss |
| SAR0566 | Membrane/SP | Serine-aspartate repeat-containing protein C precursor; sdrC | OF; pathogenic factors (toxins and colonization factors) | A | Allelic variation | |
| SA1751 | SP | Truncated map-w protein | OF; pathogenic factors (toxins and colonization factors) | PD | 211 (NA) | Allelic variation or gene loss |
| SA1973 | Membrane | Hypothetical protein; similar to hemolysin III | OF; pathogenic factors (toxins and colonization factors) | PD | 235 (=) | Allelic variation |
| SA2447 | Membrane/CWA | Hypothetical protein; similar to streptococcal hemagglutinin protein | OF; pathogenic factors (toxins and colonization factors) | PD | 289 (var) | Allelic variation or gene loss |
| SA0089 | Membrane | Hypothetical protein; similar to DNA helicase | Similar to unknown proteins | PD | 9 (NA) | Allelic variation or gene loss |
| SA0290 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | A | Allelic variation | |
| SA0333 | Membrane/SP | Conserved hypothetical protein | Similar to unknown proteins | PD | 38-39 (=, =) | Allelic variation |
| SA0477 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 56-57 (>, =) | Allelic variation |
| SAR0590 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | A | Allelic variation | |
| SA0584 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | PD | 69 (<) | gene loss |
| SA0648 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | ND | 75 (=) | Allelic variation |
| SA0649 | Cytoplasm | Conserved hypothetical protein; lipoprotein lipid attachment site | Similar to unknown proteins | PD | 75 (=) | Allelic variation |
| SAR1027 | Cytoplasm | GCN5-related N-α-acetyltransferase family protein | Similar to unknown proteins | A | Allelic variation | |
| SA1279 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 154-155 (NA, =) | Allelic variation |
| SA1601 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | PD | 192 (=) | Allelic variation |
| SA2101 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 247 (NA) | Allelic variation or gene loss |
| SA2133 | Membrane | Conserved hypothetical protein | Similar to unknown proteins | PD | 251 (=) | Allelic variation |
| SA2143 | Secreted/SP | Conserved hypothetical protein | Similar to unknown proteins | A | 252 (var) | Allelic variation |
| SA2190 | Cytoplasm | Conserved hypothetical protein | Similar to unknown proteins | PD | 257 (NA) | Allelic variation or gene loss |
| SA0203 | Membrane | Hypothetical protein | No similarity | ND | 24 (var) | Allelic variation or gene loss |
| SA0536 | Cytoplasm | Hypothetical protein | No similarity | PD | 64 (=) | Allelic variation |
| SA0611 | Cytoplasm | Hypothetical protein | No similarity | PD | 72 (=) | Allelic variation |
| SA0647 | Membrane | Hypothetical protein | No similarity | PD | 75 (=) | Allelic variation |
| SA1318 | Secreted/SP | Hypothetical protein; lipoprotein lipid attachment site | No similarity | ND | 159 (<) | Gene loss |
| SA1552 | Exported/SP | Hypothetical protein; CWA | No similarity | PD | 186 (=) | Allelic variation |
| SA1594 | Cytoplasm | Hypothetical protein | No similarity | PD | 192 (=) | Allelic variation |
| SA1610 | Cytoplasm | Hypothetical protein | No similarity | PD | 193 (<) | Gene loss |
| SA1619 | Exported/SP | Hypothetical protein; lipoprotein lipid attachment site | No similarity | A | Gene gain | |
| SA1635 | Cytoplasm | Hypothetical protein (pathogenicity island SaPIn3) | No similarity | ND | 196 (NA) | Allelic variation or gene loss |
| SA2059 | Cytoplasm | Hypothetical protein | No similarity | PD | 244 (NA) | Allelic variation or gene loss |
| SA2118 | Membrane | Hypothetical protein | No similarity | A | 249 (=) | Allelic variation |
| SA2267 | Cytoplasm | Hypothetical protein | No similarity | PD | 266 (NA) | Allelic variation or gene loss |
| SAS008 | Membrane | Hypothetical protein | No similarity | PD | 28 (=) | Allelic variation |
| SAS036 | Membrane | Hypothetical protein | No similarity | ND | 118-119 (NA, var) | Gene loss |
| SAS086 | Cytoplasm | Hypothetical protein | No similarity | PD | 251 (=) | Allelic variation |
| BH0452 | Membrane | Epidermin biosynthesis protein; epiC | A | Gene gain | ||
| BH1491 | Secreted | Epidermin leader peptide processing serine protease | A | Gene gain | ||
| BS_dnaC | Cytoplasm | Prophage l54a; replicative DNA helicase | A | Gene gain | ||
| MJ0348 | Cytoplasm | Hypothetical protein | A | Gene gain | ||
| MYPU_3080 | Membrane | Epidermin biosynthesis protein; epiB | A | Gene gain | ||
| SP0165 | Not clear | Epidermin biosynthesis protein D (epiD) | A | Gene gain | ||
| SPy1085 | Cytoplasm | Epidermin immunity protein f (epiF) | A | Gene gain | ||
| YML091c | Cytoplasm | Unique predicted open reading frames | A | Gene gain | ||
CWA, cell wall anchor; SP, signal peptide. Results were determined using PSORT or SignalP software.
The classification is as described in the N315 updated annotation available on the DOGAN website (http://www.bio.nite.go.jp/dogan/). CECP, cell envelope and cellular processes; OF, other functions; IP, information pathways; IM, intermediary metabolism.
CGH analysis (see Material and Methods for details). A, additional; PD, poorly detected; ND, not detected. Note that additional genes (A) cannot be localized on WGPS fragments.
WGPS results. The fragment number and size variation in the strains considered are shown: (=), same size; (<), smaller fragment; (>), bigger fragment; (NA), not amplified; (var), size variations within the strains considered.
WGPS.
Whole-genome PCR scanning (WGPS) recently was described as a method to identify previously undetected genome diversity in serotype O157 strains of Escherichia coli, which cause enterohemorrhagic infections (34). Genomic DNA used for WGPS experiments was extracted as previously described (5). WGPS was used to assess variation in gene content and genome structure among the 12 strains as described previously for S. aureus (5).
Primer design for WGPS.
WGPS first was employed in a study of Escherichia coli O157 isolates (34). Here, we intended to use it to gather information on the chromosomal structural diversity of highly divergent S. aureus strains. Accordingly, the design of primers was critical. Using GenoFrag, a software program dedicated to the design of primers optimized for WGPS that we previously developed (5), a set of 295 pairs of 25-mer primers was designed based on the N315 sequence. Primer specificity then was assessed by comparing primers to each of six publicly available S. aureus sequences (Mu50, MW2, COL, NCTC8325, MSSA476, and MRSA252) (2, 19, 25) using local software based on BLASTn (1). Primers that presented absolutely no annealing with any of the six genome sequences tested were kept, since they may be included in N315-specific regions. Using this set of primer pairs, 295 segments of ∼10 kb covering the whole N315 chromosome, with overlaps of ∼1 kb at every segment end, were predicted. Primer sequences are available upon request from the authors.
PCR analysis for WGPS.
PCRs were performed using genomic DNA as the template and LR-PCR kits (GeneAmpXL, Applera, France) using PCR conditions previously described (4, 5), and PCR products were routinely analyzed by 0.5% agarose gel electrophoresis. A fragment was considered significantly different when its size differed by more than 1-kb from the expected size in N315. This threshold was determined to take account of the resolution level of size estimation in routine agarose gel electrophoresis. When a fragment was not amplified after two attempts, additional PCR studies were performed using various combinations of primers, considering three types of genomic variation: (i) the lack of the target sequence due to deletion, (ii) nucleotide divergence at the hybridization site, and (iii) the expansion of the distance between the primers by a large insertion. By such additional experiments (named overlapping PCR and framing PCR), most variations of the first two types could be discriminated from the latter type. A total of 3,540 LR-PCRs were performed on the 12 tested strains. Results were graphically represented using GeneWiz software (21).
PCR of selected host-specific determinants.
PCR tests were carried out on the 30 strains studied (Table 1), using genomic DNA as the template and primers specific for the RDs that were identified by WGPS and CGH analyses (Table 2). Amplification reactions were carried out in a Bio-Rad iCycler (Bio-Rad, Marnes la Coquette, France) using the following program: 4 min at 95°C, followed by 30 cycles of 30 s at 95°C, 30 s at 55°C, and 30 s at 72°C. The program finished with an additional 7-min extension step at 72°C. PCR products were analyzed by agarose gel electrophoresis.
TABLE 2.
Primers used to detected ovine-caprine-specific regions of difference
| Region(s) of difference | No. of WGPS fragments | Forward primer | Reverse primer | Size (bp) |
|---|---|---|---|---|
| SA0170-SA0171 | 19 | TCAATATTTAGAGTATGATGTTGAAGC | TCAATAATTGCGTAAGTGAACACA | 454 |
| SA0317 | 37 | TGTTGAACAAAATCGTATTGCAG | TGTCGGTATCGTGTTCGGTA | 463 |
| SA0647-SA0648 | 75 | CCAAAACAAATAGTGGATCATCA | TTGAAACACATCAACTCAAACG | 1221 |
| SA1090 | 128 | AATTATTTTACCTCCTTCAATAGCTT | TGCTTTTTGGTGTAGTTGGTAAA | 400 |
| SA2475-SA2478 | 292 | CAAATTGATCAAATGAACCTTTCA | TCATTTTGCTTGGAACAGCT | 586 |
Microarray accession number.
The microarray data were deposited in the public repository database Gene Expression Omnibus (9) under the accession number GSE10187.
RESULTS
PFGE and MLST analysis of S. aureus ruminant strains.
The 12 strains analyzed by CGH and WGPS first were genotyped using MLST and PFGE. Allelic profiles determined by MLST were compared to all of the S. aureus STs in the MLST database, as described in Materials and Methods (Fig. 1A). Of the strains examined by CGH and WGPS in the current study, the limited number of human strains examined belonged to one of the most successful human clones, ST5, except strain 1183, which was of ST15. Bovine strains were represented by four different STs. Of note, strains 1166 and Newbould were closely related (a single-locus variant difference), and they clustered together with strain 1167 in a group that includes two other bovine strains and strains from human or nonspecified origin. Ovine strains shared an identical ST (ST133), and caprine strains were closely related and represented by novel single-locus variants (ST711 and ST712). Both were clustered in a group that included four ovine strains already deposited in the database (Fig. 1A). Bovine strain RF122 was more closely allied to the ovine and caprine strains than it was to the three other bovine strains. However, there was a considerable genetic distance between RF122 and the ovine and caprine strains, suggesting that they have not shared a very recent ancestor.
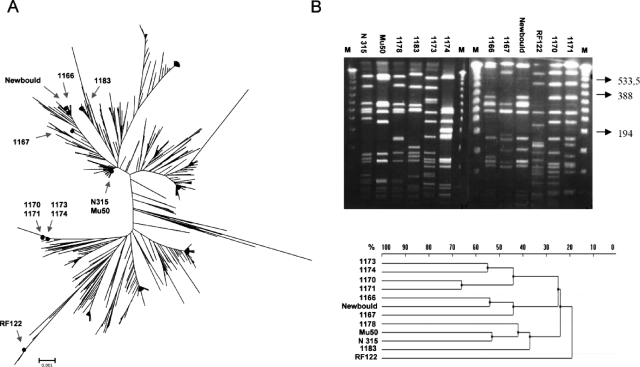
FIG. 1.
MLST and PFGE genotyping of the 12 S. aureus strains. (A) Radial tree depicting the phylogenetic relationships of the STs of the 12 strains used in this study and all of the S. aureus STs available in the MLST database (1,095 as of March 2008). MEGA 4.0 software and the neighbor-joining method with a bootstrap value of 1,000 were used to build the tree. (B) PFGE macrorestriction analysis of the chromosome of the 12 strains (upper). Dendrogram presenting the percentage of genetic similarity between the 12 strains (lower). The unweighted-pair group method using average linkages and a Dice coefficient (with a tolerance limit of 1.1%) were used to build the dendrogram. M, molecular size marker.
PFGE analysis revealed that all strains were genotypically different but grouped together according to host origin, except for bovine strain RF122, which appeared as a unique and quite distant profile from those of other strains (Fig. 1B). The highest profile similarity between two strains was not above 67% in the caprine biotype. In the human cluster, similarity was below 40%. This is mainly due to 1183, a hypermutator strain isolated from a cystic fibrosis patient. S. aureus strains isolated from cystic fibrosis reportedly are genotypically distinct among human isolates (16). SmaI restriction sites also may have been altered by the accumulation of point mutations because of the 1183 hypermutator phenotype.
Further, the additional isolates used to screen by PCR for the distribution of host-specific determinants were analyzed by PFGE and compared to each other (see the additional strains in Table 1, except 1465). All strains were genotypically distinct. Using a similarity cutoff of 50%, four groups were distinguished and correlated well with host origin. In the two ovine-caprine groups (two caprine, six ovine, one bovine, and three ovine, respectively), profile similarities ranged from 51 to 90%. In the bovine group (nine bovine and one caprine), profile similarities ranged from 54 to 88%. By PFGE analysis, a fourth group containing a bovine strain (LMA1259) appeared to be distantly related to the other strains (see Fig. S2 in the supplemental material).
Variation in genome content identified by CGH.
CGH experiments were performed using an oligonucleotide microarray that was representative of the genome sequences of the human strains COL, N315, Mu50, EMRSA16, MSSA476, and 8325 and the bovine strain RF122. Strain N315 was used as a reference in all hybridization experiments. Three independent hybridizations were performed for each strain examined. In Fig. 2, each CDS was ordered along the chromosome according to the genome structure of the reference strain N315. The genome location of CDSs that are designated variable (i.e., low or no hybridization signal) or present by CGH was predicted by their location in the genome of the reference strain.
Overall, CGH revealed genome diversity within the S. aureus species that was greater than that previously reported. Among the 3,894 S. aureus CDSs spotted on the microarray, 1,875 CDSs were shared by the 12 strains studied, which is equivalent to 70.5% of the average number of CDSs found in the strains (n = 2,656). Among the 697 genes that were strain dependent or variable, 204 (29.3%) genes were predicted to be associated with MGEs or related islands (see Table S1 in the supplemental material).
The classification of the variable genes according to their putative or known function revealed that some categories contained more variable genes than others (Table 3). For example, in the functional categories defense mechanisms, replication, recombination and repair, and unclassified genes (i.e., not in clusters of orthologous groups [COGs]), 47.9, 37.0, and 62.8%, respectively, of genes were variable. This is partly explained by the fact that many of the variable genes are encoded on MGEs, such as S. aureus pathogenicity islands (SaPIs) or prophages, which contain genes for virulence, antibiotic resistance, replication, repair, and phage-specific functions in addition to numerous genes of unknown function. Several other categories contain a relatively high proportion (>20%) of variable and additional genes. This is the case for cell cycle control, cell division, chromosome partitioning (30.4%), inorganic ion transport and metabolism (23.2%), signal transduction mechanisms (22.9%), cell wall/membrane/envelope biogenesis (22%), amino acid transport and metabolism (21.8%), and unknown function (20.6%). In contrast, categories such as carbohydrate transport and metabolism (9.5%), energy production and conversion (6.8%), and nucleotide transport and metabolism (7.9%) contain a low level of variable genes. These data suggest that some categories contain a larger proportion of genes involved in niche adaptation or contingency function than other categories.
TABLE 3.
Categorization and conservation of the genes in the S. aureus strains
| General and specific COG | Commona | Not or poorly detectedb | Additionalc | Total |
|---|---|---|---|---|
| Metabolism | ||||
| Amino acid transport and metabolism | 133 | 36 | 1 | 170 |
| Carbohydrate transport and metabolism | 95 | 9 | 1 | 105 |
| Coenzyme transport and metabolism | 69 | 8 | 77 | |
| Energy production and conversion | 82 | 6 | 88 | |
| Inorganic ion transport and metabolism | 96 | 28 | 1 | 125 |
| Lipid transport and metabolism | 43 | 8 | 51 | |
| Nucleotide transport and metabolism | 58 | 5 | 63 | |
| Secondary metabolites biosynthesis, transport, and catabolism | 18 | 3 | 21 | |
| Cellular processes and signaling | ||||
| Cell cycle control, cell division, and chromosome partitioning | 16 | 7 | 23 | |
| Cell wall/membrane/envelope biogenesis | 71 | 19 | 1 | 91 |
| Defense mechanisms | 25 | 18 | 5 | 48 |
| Intracellular trafficking, secretion, and vesicular transport | 15 | 3 | 18 | |
| Posttranslational modification, protein turnover, and chaperones | 50 | 9 | 59 | |
| Signal transduction mechanisms | 27 | 7 | 1 | 35 |
| Information storage and processing | ||||
| Replication, recombination, and repair | 80 | 44 | 3 | 127 |
| Transcription | 92 | 24 | 1 | 117 |
| Translation, ribosomal structure, and biogenesis | 128 | 17 | 145 | |
| Poorly characterized | ||||
| Function unknown | 173 | 44 | 1 | 218 |
| General function, prediction only | 178 | 40 | 218 | |
| Not in COGs | 198 | 315 | 19 | 532 |
| Other | 228 | 47 | 2 | 275 |
| Total | 1,875 | 697 | 36 | 2,608 |
Genes that were present in all strains tested in CGH.
Genes that were not detected at all (i.e., no signal in three independent hybridizations) or gave a low signal for at least one strain.
Genes that were present in at least one tested strain but not in N315.
Host-specific variation in gene content.
The gene content of the strains was analyzed with regard to their host origin (see Table S2 in the supplemental material). In each group (bovine strains and ovine-caprine strains), approximately 2,000 CDSs were common to the four strains (i.e., present in the four strains). The amount of variable CDSs (absent in at least one strain of a given group compared to the content of N315) and additional CDSs (i.e., present in at least one strain but absent in N315) were comparable. However, the distribution of these variable or additional CDSs within the bovine strains and the ovine-caprine strains indicated extensive variation. Specifically, strain RF122 contained 228 of the 569 variable genes and 253 of the 406 additional genes found in bovine strains, indicating that it has not shared a recent ancestor with the other bovine strains. In contrast, in the ovine and caprine group, 394 variable genes out of 508 and 185 additional genes out of 327 were common to all four strains, indicating the genomic homogeneity of strains isolated from ovine-caprine hosts.
Structural diversity of S. aureus chromosomes.
A limitation of CGH analysis is that it cannot identify genes present in the test strain but absent from the reference strains represented on the microarray. Also, genes that are present in the test strain but have undergone diversification such that hybridization with the array does not occur cannot be discriminated from those genes that are absent. The WGPS approach takes advantage of the sequence data available for one or more strains to localize and identify the regions of the genomic variation found in other strains (5, 34). A set of 295 primer pairs was designed from the N315 genome sequence as previously described (5) and used in the LR-PCR analysis of the 12 strains. As expected, the LR-PCR of strain N315 resulted in 100% amplification of 295 PCR products and a range of 80 to 99% amplification for the other strains examined (see Fig. S1 and Fig. S2 in the supplemental material). Interestingly, the hypermutator strain 1183 of human origin had an LR-PCR success rate of 95% in spite of the higher rate of nucleotide substitution expected for that strain, which was manifested in a very distinct PFGE type. Among bovine strains, RF122 had an amplification success rate of 86%, while that for the other bovine strains was 94%, consistently with the difference in gene content identified for strain RF122 in contrast to the other bovine strains. Among the successfully amplified WGPS fragments, a great majority (from ∼97% in Mu50 to ∼82% in ovine strains) were indistinguishable in length from those generated with the N315 genome.
Consistently with PFGE and MLST data, WGPS amplification profiles of the strains of human origin were very similar to but distinct from ovine and caprine strains that were closely related to each other (Fig. 2). The WGPS profiles of bovine strains 1166, 1167, and Newbould contained many genomic variations at the same sites that were largely distinct from those of bovine strain RF122.
Impact of MGEs on genome diversity.
The MGEs, such as staphylococcal cassette chromosome mec, SaPI, or prophages, found in N315 were considered present in a test strain when the corresponding PCR products were successfully amplified and were of a size identical to that of products generated with N315. Sometimes, WGPS fragments that included the left and right extremities of an MGE were obtained, but size variations were observed within the internal fragments, suggesting the presence of a variant MGE in the strain. In some cases, several internal fragments of a large MGE were amplified, but amplification across the predicted site of insertion failed, suggesting the possible existence of related MGEs localized elsewhere on the chromosome. The staphylococcal cassette chromosome mec was absent from all of the strains except Mu50. Considerable variation in the number and variety of MGEs was observed. Of note, CGH results showed that tst, the gene encoding TSST-1, was present in strains RF122, 1170, 1171, and 1174, whereas the corresponding WGPS fragment that contains the SaPI-encoding tst gene in N315 was not amplified. These data indicate a distinct location for the tst-encoded SaPIs in these strains, consistently with a previous study and recent genome sequence that defined the site of insertion of SaPIbov in RF122 (13, 18). Similarly, the presence of the 29.3-kb-long vSaα (previously annotated as SaPIn2), found in N315, which encodes the staphylococcal superantigen-like family of proteins (26), was predicted to be present by WGPS in the four human strains examined. This correlated well with the CGH results for human strain 1178, whereas for human strain 1183, 17 of the 31 CDSs that compose vSaα were variable (not detected or with a low signal), suggesting a high nucleotide divergence in this locus. In all of the animal strains, WGPS fragments were amplified at the vSaα locus but contained size variations that were reflected in CGH results: up to 15 out of the 31 CDSs of vSaα were not detected in the bovine strains. In ovine-caprine strains, CGH and WGPS data suggested the presence of variants of vSaα. The 26.2-kb vSaβ (previously annotated as SaPIn3), corresponding to the enterotoxin gene cluster (seg, sen, yent2, yent1, sei, sem, and seo) and lukDE, was not amplified in most animal strains. Only one LR-PCR product at the expected size was obtained in the bovine strains 1167 and Newbould, which correlated well with the CGH data. WGPS and CGH results suggest that the vSaβ locus is highly variable in animal strains. The lukDE genes are well conserved in the animal strains, but the enterotoxin gene cluster is absent from all bovine, ovine, and caprine strains except RF122. This observation was confirmed by RF122 sequence data (18). Taken together, these data demonstrate the considerable variety in gene content and chromosomal location that exists among SaPIs encoded by different strains. None of the SaPI variations observed were host specific. Prophage ΦN315 was absent in all other strains except Mu50. However, some internal fragments were amplified in strains 1178 (human), 1173 (ovine), and 1174 (ovine), indicating the presence of a related phage inserted elsewhere in the genome. Most of the insertion sequence (IS) elements found in N315 were absent in the corresponding fragments in the animal strains. Similarly, none of the four copies of Tn554 found in the genome of N315 were found among the other strains.
Combining WGPS and CGH results to identify host-specific chromosomal RDs.
CGH analysis provides information on the gene composition but not the chromosomal locations. It also fails to identify strain-specific genes that are not represented on the microarray. Conversely, WGPS detects variable loci in which genetic events have taken place, while the identity of the gene content cannot be determined. In order to explore the genomic diversity of S. aureus with regard to host adaptation, we combined the two datasets (Fig. 2). Based on CGH results, we grouped the variable CDSs into four categories (Table 4): (i) bovine-specific RDs (five CDSs) found to be variable (i.e., either not detected, poorly detected, or additional compared to the CDSs of the N315 reference strain) in the four bovine strains tested, whereas they were not variable in either of the other (human or ovine-caprine) strains; (ii) ovine-caprine RDs (61 CDSs) were found to be variable in ovine-caprine strains and not in the other strains; (iii) ruminant-specific RDs (13 CDSs) were found to be variable in all of the ruminant strains and not in the human ones; (iv) a fourth category comprising variable CDSs of the four ovine-caprine strains plus RF122 (101 CDSs) was identified. Once those categories had been defined, WGPS data enabled discrimination between CDSs that were likely to be absent from the genome tested (the shorter WGPS fragment) and, thus, did not hybridize in CGH (gene loss was suspected) (Table 4) and CDSs that were present in the strains (no significant size variation in the corresponding WGPS fragment compared to that of the N315 reference) but likely had undergone sufficient nucleotide divergence such that they did not hybridize in CGH (allelic variation was suspected) (Table 4).
Two of five bovine-specific CDSs belong to genes classified as pathogenic factors, two other genes were unclassified, and one gene, fmhC, was classified in the cell wall category, and its product shares identity with FemA and FemB (45). Most of the ovine-caprine-specific CDSs (26 out of 61) are represented by unclassified genes. Interestingly, 35 CDSs were suspected to have undergone allelic variation (Table 4), among which are two fmt genes that reportedly are involved in autolysis and methicillin resistance (24) and two trp genes that are predicted to exist in an operon (49). Homologs of these genes made by Mycobacterium tuberculosis are involved in virulence and colonization (42). All 13 ruminant-specific CDSs were variable (i.e., low or no hybridization signal) and were located in loci harboring size variations according to WGPS results. Interestingly, genes involved in virulence (SSL10 and clfA) and autolysis (lytN) were found among those variable genes and, thus, may have undergone considerable divergence or may be absent from ruminant strains. In the fourth category, allelic variation was suspected for 59 CDSs. These 59 CDSs included genes involved in transport (e.g., fruA, encoding a fructose permease; and mtlF, encoding a component of a mannitol-specific phosphotransferase system), in membrane bioenergetics (mnhD, encoding an Na+/H+ antiporter subunit), in protein secretion (secY, encoding a preprotein translocase), in RNA synthesis (one lysR and two gntR transcriptional regulators), and in virulence (CDSs similar to those of streptococcal hemagglutinin and hemolysin III). Among the other CDSs of the fourth category, 12 CDSs corresponded to gene gain and could not be located in WGPS fragments (Table 4). Five of the latter genes are involved in epidermin (an S. epidermidis bacteriocin) biosynthesis, suggesting that horizontal gene transfer occurred in these strains.
Compared to N315 gene content, 151 host-specific CDSs were suspected to be absent or to have undergone allelic variation, and 29 other genes were identified as additional genes. Interestingly, the analysis of the known or putative location of these genes using PSORT (31) or SignalP (6) revealed that 50.5% were predicted to be exported or secreted proteins, suggesting a possible role in bacterium-host or tissue interactions.
The categorization of these 180 genes according to their functions revealed that 65 genes were of unknown function or of no similarity. Among the 36 genes involved in cell envelope and cellular processes, all of the genes were found in subcategories such as transport/binding proteins and lipoproteins (24 genes), cell wall (5 genes), sensors (2 genes), membrane bioenergetics (4 genes), and protein secretion (1 gene) (Table 4). These subcategories group together genes that putatively are involved in interactions with the host or the environment. No genes were found in other subcategories, such as cell division, sporulation, germination, and transformation/competence. The 25 genes found in the intermediary metabolism category were evenly distributed among the subcategories. Among the 16 genes involved in information pathways, the majority were found in subcategories such as RNA synthesis (9 genes) and RNA modification (3 genes). Most of the 19 genes found in the category of other functions were classified in subcategories like pathogenic factors (10 genes), adaptation to atypical conditions, and miscellaneous (3 genes). Only six genes were found to be associated with the subcategories phage-related functions and transposon and ISs, confirming the low incidence of MGE in RDs predicted to be host specific.
Selected host-specific determinants are widely distributed among a panel of animal strains.
In order to test the hypothesis that the genomic variation identified by WGPS and CGH among a small number of isolates was more widely distributed among isolates of a specific host association, we screened an additional 28 strains of human (n = 4), bovine (n = 12), ovine (n = 9), and caprine hosts (n = 3) for the presence of five selected RDs by PCR (Table 1). RDs representing predicted allelic variation in the ovine and caprine strains were sequenced in the ovine strain 1174, and oligonucleotide primers specific for the RDs were designed. For regions of gene loss, the genome sequence of strain N315 was used to design oligonucleotides for PCR (Table 2).
PCR indicated the presence of each region of difference in a large proportion (62.5 to 100%) of the ovine-caprine strains examined and their absence (or weak occurrence) in strains of human and bovine origin, indicating ovine and caprine host specificity (Fig. 3, Table 5). Of note, strains 1524 (caprine origin) and 1231 (bovine origin) gave expected PCR results, consistently with that predicted by their host origin, even though they were more closely related to strains from other animal origins, as determined by PFGE analysis (see Fig. S2 in the supplemental material).
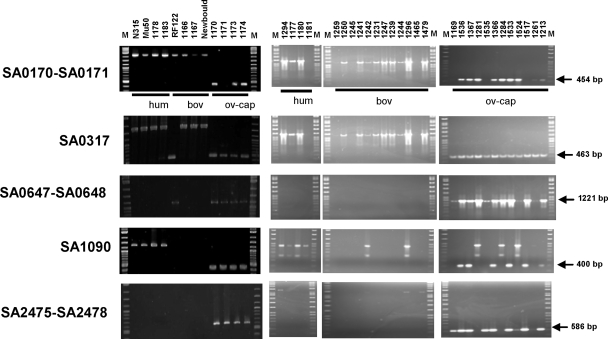
FIG. 3.
Demonstration of host specificity for six ovine-caprine traits. PCR tests were run on 40 different strains isolated from human (hum) (8), bovine (bov) (16), and ovine-caprine (ov-cap) (16) hosts. PCR products were analyzed by electrophoresis on agarose gels. Host origins are indicated at the top of each gel. In the gels on the right, M indicates the RaoulI linear DNA marker (Appligene; Qbiogene, Strasbourg, France). In the gels on the left, M indicates a 2-log ladder linear DNA marker (New England Biolabs, Ozyme, France).
TABLE 5.
Description of six ovine-caprine-specific regions
| Region | Gene and/or function | Location (by PSORT/SignalP) | CGH/WGPS determination | No. (%) of PCR products at the expected size
|
Specificity | ||
|---|---|---|---|---|---|---|---|
| Human | Bovine | Ovine-caprine | |||||
| SA0170 | Conserved hypothetical protein | Cytoplasm/no SPa | Gene lossb | 0 (0) | 2 (12.5) | 12 (75) | Ovine-caprine |
| SA0171 | fdh; NAD-dependent formate dehydrogenase | Secreted/SP | Gene lossb | ||||
| SA0317 | Hypothetical protein; similar to dihydroflavonol-4-reductase | Outside/SP | Gene lossc | 0 (0) | 1 (6.25) | 16 (100) | Ovine-caprine |
| SA0647 | Hypothetical protein | Membrane | Allelic variantd | 0 (0) | 1 (6.25) | 16 (100) | Ovine-caprine |
| SA0648 | Conserved hypothetical protein | Membrane/no SP | Allelic variant | ||||
| SA1090 | lytN; LytN protein | Secreted/SP | Control/NAe | 0 (0) | 0 (0) | 10 (62.5) | Ovine-caprine |
| SA2475-SA2478 | Conserved hypothetical protein | Membrane/no SP | Allelic variant | 0 (0) | 0 (0) | 14 (87.5) | Ovine-caprine |
| SA2476 | Conserved hypothetical protein | Membrane/no SP | Allelic variant | ||||
| SA2477 | Conserved hypothetical protein | Membrane/SP | Allelic variant | ||||
| SA2478 | Conserved hypothetical protein | Cytoplasm/no SP | Allelic variant | ||||
SP, signal peptide.
No signal or a weak signal in CGH and WGPS fragments that were shorter in ovine-caprine strains than in human strains.
No signal or a weak signal in CGH and WGPS fragments that were shorter in ovine-caprine plus RF122 strains than in human strains.
WGPS fragment harboring the same size as that in human strains.
NA, not amplified in WGPS.
DISCUSSION
The combination of CGH and WGPS reveals extensive genome diversity among S. aureus human and ruminant isolates.
Microarray-based CGH is a powerful means of exploring bacterial gene content that has made an important contribution to our understanding of bacterial diversity (12, 23, 28, 51). However, CGH analysis alone cannot discriminate between genes that are highly divergent in or completely absent from the test strain compared to the gene content of the index strain. Further, gene content analysis using CGH is limited by the set of targets spotted on the microarray. To overcome these limitations, a combination of CGH and WGPS was used to examine the genomic diversity of S. aureus isolates from human, bovine, ovine, and caprine hosts. WGPS previously has been used to explore the diversity of closely related Escherichia coli O157 (34) and group A streptococcus strains (7). Here, we demonstrate that WGPS can be used to examine the diversity of genetically divergent bacterial strains, provided that the primer design is robust (4, 5). Combining the CGH and WGPS approaches allowed (i) the determination of the gene content relative to that of the genes represented on the microarray, (ii) the identification of the chromosomal sites of inserted or deleted genetic elements, and (iii) the identification of genes in the core genome that have undergone considerable nucleotide divergence. Our data indicate greater levels of diversity within the S. aureus species than has been reported previously. Not surprisingly, many of the variable genes belonged to functional categories that reflect horizontal gene transfer. However, a high proportion of variable genes also was associated with genes involved in defense mechanisms, the cell envelope, and amino acid and inorganic ion metabolism, suggesting that such genes contribute to niche adaptation. The greater-than-expected levels of genome diversity could reflect the fact that the strains examined were obtained from different hosts and could require distinct gene complements for survival. Further, the fact that strains inhabit different ecological niches would reduce the opportunities for the lateral transfer of genes between strains, and a distinct gene pool is likely to exist in different niches.
The majority of RDs among isolates from host-specific lineages are located in the core genome.
In the present study, we investigated the genetic basis of S. aureus host specificity by comparing the genome content of strains causing mastitis in ruminants to the genome content of representative human isolates. A total of 180 CDSs were specific for lineages associated with ruminant hosts, including 61 CDSs that were found to be specific for strains infecting small ruminants (goat and sheep). Unexpectedly, our analyses indicated that the majority of host-specific differences were located outside of predicted MGEs that together make up the accessory genome and instead belong to the core genome. Lindsay et al. recently reported the existence of such variable core genes in community-acquired invasive isolates and nasal carriage isolates of human origins. For each dominant lineage, they found several unique combinations of core variable genes, suggesting a common ancestor followed by evolutionary divergence (28). Only 23 of the ruminant lineage-specific CDSs identified in our study (e.g., lytN, oppF, fmhC, and mtlF) were found in the list of 728 core variable genes determined by Lindsay et al., suggesting that distinct sets of genes have undergone diversification in different host-specific lineages.
Combining WGPS and CGH results, we discovered that some genes are absent from ovine-caprine strains but present in isolates from other hosts. For example, we found that SA0170, encoding a hypothetical protein, and SA0171 (fdh, encoding an NAD-dependent formate dehydrogenase) are absent from ovine-caprine isolates but are encoded together in an operon in human and bovine isolates (49). SA0170 previously was shown to be up-regulated upon internalization in human epithelial cells (14), and both genes were more upregulated in biofilm than planktonic culture conditions (38). These results suggest that these genes are involved in interactions with and persistence in the human host. Further functional analyses are required to check whether the operon is dispensable for the infection of ovine or caprine hosts and whether its loss is a true host-specific adaptation. We also identified seven genes that were present in all S. aureus strains tested but contained allelic variation in ovine and caprine strains. Of these, six genes encoded proteins of unknown function, including five that are predicted to be membrane associated or secreted. Of note, SA0647 is found in several other staphylococcal species, including Staphylococcus haemolyticus, Staphylococcus saprophyticus, and Staphylococcus epidermidis, and is upregulated by mgr, a global regulator of gene expression in S. aureus (29). Also, gene SA2478, part of a four-gene operon, also is induced upon internalization in human epithelial cells (14). The RDs specific for ovine and caprine strains included lytN, a gene encoding a 372-amino-acid protein with a putative muramidase activity that may be involved in autolysis (37, 43) and that is downregulated by mgr (29) and upregulated by rot (40).
Importance of predicted secreted and exported proteins in host-specific determinants.
Remarkably, we discovered that 91 (50.5%) of the putative host-specific determinants were predicted to be extracytoplasmic, indicating a possible role in host-pathogen interactions. These proteins represent a very high proportion of the 130 to 145 proteins (i.e., ∼5% of the entire predicted proteome of S. aureus) predicted to be exported or secreted by S. aureus in a secretomic study (41). Similarly, in human isolates, Lindsay et al. found that many core variable genes were known or predicted to be expressed on the bacterial surface and to interact with the host during nasal colonization and infection (28). Recently, Herron-Olson et al. reported that bovine strain RF122 genes encoding surface proteins predicted to be involved in host interactions show higher-than-average rates of nonsynonymous substitution and gene decay relative to those of their homologs in the sequenced human isolates (18). They also showed that some virulence-associated genes, like clfA, encoding clumping factor, exist as pseudo-genes in the RF122 genome (18). We found that clfA represented an allelic variant in RF122 and ovine-caprine strains. Taken together, these data are consistent with an important role for the conserved secretome in host adaptation.
Identification of a clonal lineage of S. aureus that is specific for ovine and caprine hosts.
On the basis of PFGE and MLST analysis, strains isolated from sheep and goats were closely related, which is consistent with previous findings (47). CGH and WGPS also indicated the close relatedness of ovine-caprine strains, whereas bovine isolates contained much greater variation in gene content, consistently with previous studies of the virulence gene content of bovine mastitis-associated isolates (47, 50). In contrast to bovine mastitis, which usually is subclinical, S. aureus ovine mastitis typically is clinical in nature, whereas coagulase-negative staphylococci are associated with subclinical mastitis (8). It is possible that variation in S. aureus genome content specific for the ovine-caprine clone contributes to the increased severity of infection associated with sheep and goat mastitis.
Three of the four strains isolated from cows were closely related to each other, whereas strain RF122 (a strain that reproducibly induced clinical mastitis in experimental infections) was more closely related to the ovine-caprine strains. RF122 was found to share more variable CDSs with the ovine-caprine clone than with the other bovine strains, as indicated in a previous CGH study (14). In fact, we found that 117 genes varied similarly in RF122 and the ovine and caprine strains but not in all other strains. In agreement with MLST and PFGE results, these data suggest that some bovine strains are more closely related to the ovine-caprine-specific clone than to the other bovine strains. Therefore, it is possible that the determinants found in common for RF122 and ovine-caprine strains likely are due to their similar clonal origin. However, this does not rule out the possibility that some of these differences represent adaptations to the ruminant mammary gland, even though they are not shared with the other bovine strains of more distant relatedness.
All of the strains of goat and caprine origin were found to be closely related. It is apparent from our work and that of others that there is a single major lineage of S. aureus associated with mastitis infections of sheep and goat (30). At this point, it is not possible to identify which of the RDs represent true host-specific genetic adaptations and which are only lineage specific. However, host-specific determinants selected in this study gave a PCR profile consistent with their host origin in the vast majority of strains tested, notably for a caprine strain that grouped in a bovine-associated clonal lineage (as determined by PFGE analysis) and a bovine strain that grouped in an ovine-caprine cluster.
This study highlights the extensive variation that exists among human and animal isolates of S. aureus. We found that the core genome contains the majority of RDs that were specific for ruminant strains, including many that encoded surface-associated or extracellular proteins. These loci represent excellent candidates for studies of the molecular basis of S. aureus host specificity.
Supplementary Material
Acknowledgments
We are grateful to S. D. Ehrlich and A. Sorokin (UGM, INRA Jouy en Josas) and to Pascal Rainard (IASP, INRA Tours) for helpful and constructive discussion at the beginning of this work.
N.B.Z. was the recipient of a Ph.D. grant from the French Ministry of Research and Education. P.D.A. is the recipient of a CAPES grant from the Brazilian government (CAPES-COFECUB project 539/06). J.R.F. is supported by the Biotechnology and Biological Sciences Research Council, United Kingdom, research grant BB/D521222/1.
Footnotes
Published ahead of print on 20 June 2008.
Supplemental material for this article may be found at http://jb.asm.org/.
REFERENCES
- 1.Altschul, S. F., W. Gish, W. Miller, E. W. Myers, and D. J. Lipman. 1990. Basic local alignment search tool. J. Mol. Biol. 215403-410. [DOI] [PubMed] [Google Scholar]
- 2.Baba, T., F. Takeuchi, M. Kuroda, H. Yuzawa, K. Aoki, A. Oguchi, Y. Nagai, N. Iwama, K. Asano, T. Naimi, H. Kuroda, L. Cui, K. Yamamoto, and K. Hiramatsu. 2002. Genome and virulence determinants of high virulence community-acquired MRSA. Lancet 3591819-1827. [DOI] [PubMed] [Google Scholar]
- 3.Bannerman, D. D., M. J. Paape, J. W. Lee, X. Zhao, J. C. Hope, and P. Rainard. 2004. Escherichia coli and Staphylococcus aureus elicit differential innate immune responses following intramammary infection. Clin. Diagn. Lab Immunol. 11463-472. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 4.Ben Zakour, N., C. Grimaldi, M. Gautier, P. Langella, V. Azevedo, E. Maguin, and Y. Le Loir. 2006. Testing of a whole genome PCR scanning approach to identify genomic variability in four different species of lactic acid bacteria. Res. Microbiol. 157386-394. [DOI] [PubMed] [Google Scholar]
- 5.Ben Zakour, N., M. Gautier, R. Andonov, D. Lavenier, M. F. Cochet, P. Veber, A. Sorokin, and Y. Le Loir. 2004. GenoFrag: software to design primers optimized for whole genome scanning by long-range PCR amplification. Nucleic Acids Res. 3217-24. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 6.Bendtsen, J. D., H. Nielsen, G. von Heijne, and S. Brunak. 2004. Improved prediction of signal peptides: SignalP 3.0. J. Mol. Biol. 340783-795. [DOI] [PubMed] [Google Scholar]
- 7.Beres, S. B., G. L. Sylva, D. E. Sturdevant, C. N. Granville, M. Liu, S. M. Ricklefs, A. R. Whitney, L. D. Parkins, N. P. Hoe, G. J. Adams, D. E. Low, F. R. DeLeo, A. McGeer, and J. M. Musser. 2004. Genome-wide molecular dissection of serotype M3 group A Streptococcus strains causing two epidemics of invasive infections. Proc. Natl. Acad. Sci. USA 10111833-11838. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 8.Bergonier, D., R. de Cremoux, R. Rupp, G. Lagriffoul, and X. Berthelot. 2003. Mastitis of dairy small ruminants. Vet. Res. 34689-716. [DOI] [PubMed] [Google Scholar]
- 9.Edgar, R., M. Domrachev, and A. E. Lash. 2002. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 30207-210. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Enright, M. C., N. P. J. Day, C. E. Davies, S. J. Peacock, and B. G. Spratt. 2000. Multilocus sequence typing for characterization of methicillin-resistant and methicillin-susceptible clones of Staphylococcus aureus. J. Clin. Microbiol. 381008-1015. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 11.Fitzgerald, J. R., W. J. Meaney, P. J. Hartigan, C. J. Smyth, and V. Kapur. 1997. Fine-structure molecular epidemiological analysis of Staphylococcus aureus recovered from cows. Epidemiol. Infect. 119261-269. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Fitzgerald, J. R., D. E. Sturdevant, S. M. Mackie, S. R. Gill, and J. M. Musser. 2001. Evolutionary genomics of Staphylococcus aureus: insights into the origin of methicillin-resistant strains and the toxic shock syndrome epidemic. Proc. Natl. Acad. Sci. USA 988821-8826. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Fitzgerald, J. R., S. R. Monday, T. J. Foster, G. A. Bohach, P. J. Hartigan, W. J. Meaney, and C. J. Smyth. 2001. Characterization of a putative pathogenicity island from bovine Staphylococcus aureus encoding multiple superantigens. J. Bacteriol. 18363-70. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 14.Garzoni, C., P. Francois, A. Huyghe, S. Couzinet, C. Tapparel, Y. Charbonnier, A. Renzoni, S. Lucchini, D. P. Lew, P. Vaudaux, W. L. Kelley, and J. Schrenzel. 2007. A global view of Staphylococcus aureus whole genome expression upon internalization in human epithelial cells. BMC. Genomics 8171. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Goerke, C., S. Esser, M. Kummel, and C. Wolz. 2005. Staphylococcus aureus strain designation by agr and cap polymorphism typing and delineation of agr diversification by sequence analysis. Int. J. Med. Microbiol. 29567-75. [DOI] [PubMed] [Google Scholar]
- 16.Goerke, C., and C. Wolz. 2004. Regulatory and genomic plasticity of Staphylococcus aureus during persistent colonization and infection. Int. J. Med. Microbiol. 294195-202. [DOI] [PubMed] [Google Scholar]
- 17.Hensen, S. M., M. J. Pavicic, J. A. Lohuis, J. A. de Hoog, and B. Poutrel. 2000. Location of Staphylococcus aureus within the experimentally infected bovine udder and the expression of capsular polysaccharide type 5 in situ. J. Dairy Sci. 831966-1975. [DOI] [PubMed] [Google Scholar]
- 18.Herron-Olson, L., J. R. Fitzgerald, J. M. Musser, and V. Kapur. 2007. Molecular correlates of host specialization in Staphylococcus aureus. PLoS ONE. 2e1120. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 19.Holden, M. T. G., E. J. Feil, J. A. Lindsay, S. J. Peacock, N. P. J. Day, M. C. Enright, T. J. Foster, C. E. Moore, L. Hurst, R. Atkin, A. Barron, N. Bason, S. D. Bentley, C. Chillingworth, T. Chillingworth, C. Churcher, L. Clark, C. Corton, A. Cronin, J. Doggett, L. Dowd, T. Feltwell, Z. Hance, B. Harris, H. Hauser, S. Holroyd, K. Jagels, K. D. James, N. Lennard, A. Line, R. Mayes, S. Moule, K. Mungall, D. Ormond, M. A. Quail, E. Rabbinowitsch, K. Rutherford, M. Sanders, S. Sharp, M. Simmonds, K. Stevens, S. Whitehead, B. G. Barrell, B. G. Spratt, and J. Parkhill. 2004. Complete genomes of two clinical Staphylococcus aureus strains: evidence for the rapid evolution of virulence and drug resistance. Proc. Natl. Acad. Sci. USA 1019786-9791. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 20.Ichiyama, S., M. Ohta, K. Shimokata, N. Kato, and J. Takeuchi. 1991. Genomic DNA fingerprinting by pulsed-field gel electrophoresis as an epidemiological marker for study of nosocomial infections caused by methicillin-resistant Staphylococcus aureus. J. Clin. Microbiol. 292690-2695. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Jensen, L. J., C. Friis, and D. W. Ussery. 1999. Three views of microbial genomes. Res. Microbiol. 150773-777. [DOI] [PubMed] [Google Scholar]
- 22.Kapur, V., W. M. Sischo, R. S. Greer, T. S. Whittam, and J. M. Musser. 1995. Molecular population genetic analysis of Staphylococcus aureus recovered from cows. J. Clin. Microbiol. 33376-380. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 23.Koessler, T., P. Francois, Y. Charbonnier, A. Huyghe, M. Bento, S. Dharan, G. Renzi, D. Lew, S. Harbarth, D. Pittet, and J. Schrenzel. 2006. Use of oligoarrays for characterization of community-onset methicillin-resistant Staphylococcus aureus. J. Clin. Microbiol. 441040-1048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Komatsuzawa, H., M. Sugai, K. Ohta, T. Fujiwara, S. Nakashima, J. Suzuki, C. Y. Lee, and H. Suginaka. 1997. Cloning and characterization of the fmt gene, which affects the methicillin resistance level and autolysis in the presence of triton X-100 in methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 412355-2361. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Kuroda, M., T. Ohta, I. Uchiyama, T. Baba, H. Yuzawa, I. Kobayashi, L. Cui, A. Oguchi, K. Aoki, Y. Nagai, J. Lian, T. Ito, M. Kanamori, H. Matsumaru, A. Maruyama, H. Murakami, A. Hosoyama, Y. Mizutani-Ui, N. K. Takahashi, T. Sawano, R. Inoue, C. Kaito, K. Sekimizu, H. Hirakawa, S. Kuhara, S. Goto, J. Yabuzaki, M. Kanehisa, A. Yamashita, K. Oshima, K. Furuya, C. Yoshino, T. Shiba, M. Hattori, N. Ogasawara, H. Hayashi, and K. Hiramatsu. 2001. Whole genome sequencing of methicillin-resistant Staphylococcus aureus. Lancet 3571225-1240. [DOI] [PubMed] [Google Scholar]
- 26.Lina, G., G. A. Bohach, S. P. Nair, K. Hiramatsu, E. Jouvin-Marche, and R. Mariuzza. 2004. Standard nomenclature for the superantigens expressed by Staphylococcus. J. Infect. Dis. 1892334-2336. [DOI] [PubMed] [Google Scholar]
- 27.Lindsay, J. A., and M. T. Holden. 2006. Understanding the rise of the superbug: investigation of the evolution and genomic variation of Staphylococcus aureus. Funct. Integr. Genomics. 6186-201. [DOI] [PubMed] [Google Scholar]
- 28.Lindsay, J. A., C. E. Moore, N. P. Day, S. J. Peacock, A. A. Witney, R. A. Stabler, S. E. Husain, P. D. Butcher, and J. Hinds. 2006. Microarrays reveal that each of the ten dominant lineages of Staphylococcus aureus has a unique combination of surface-associated and regulatory genes. J. Bacteriol. 188669-676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 29.Luong, T. T., P. M. Dunman, E. Murphy, S. J. Projan, and C. Y. Lee. 2006. Transcription profiling of the mgrA regulon in Staphylococcus aureus. J. Bacteriol. 1881899-1910. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 30.Musser, J. M., and R. K. Selander. 1990. Genetic analysis of natural populations of Staphylococcus aureus, p. 59-68. In R. P. Novick (ed.), Molecular biology of the staphylococci. VCH Publishers, New York, NY.
- 31.Nakai, K., and P. Horton. 1999. PSORT: a program for detecting sorting signals in proteins and predicting their subcellular localization. Trends Biochem. Sci. 2434-36. [DOI] [PubMed] [Google Scholar]
- 32.Novick, R. P., and A. Subedi. 2007. The SaPIs: mobile pathogenicity islands of Staphylococcus. Chem. Immunol. Allergy. 9342-57. [DOI] [PubMed] [Google Scholar]
- 33.Novick, R. P. 2003. Mobile genetic elements and bacterial toxinoses: the superantigen-encoding pathogenicity islands of Staphylococcus aureus. Plasmid 4993-105. [DOI] [PubMed] [Google Scholar]
- 34.Ohnishi, M., J. Terajima, K. Kurokawa, K. Nakayama, T. Murata, K. Tamura, Y. Ogura, H. Watanabe, and T. Hayashi. 2002. Genomic diversity of enterohemorrhagic Escherichia coli O157 revealed by whole genome PCR scanning. Proc. Natl. Acad. Sci. USA 9917043-17048. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 35.Peacock, S. J., C. E. Moore, A. Justice, M. Kantzanou, L. Story, K. Mackie, G. O'Neill, and N. P. Day. 2002. Virulent combinations of adhesin and toxin genes in natural populations of Staphylococcus aureus. Infect. Immun. 704987-4996. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 36.Poutrel, B., and C. Lerondelle. 1978. Induced staphylococcal infections in the bovine mammary gland. Influence of the month of lactation and other factors related to the cow. Ann. Rech. Vet. 9119-128. [PubMed] [Google Scholar]
- 37.Renzoni, A., C. Barras, P. Francois, Y. Charbonnier, E. Huggler, C. Garzoni, W. L. Kelley, P. Majcherczyk, J. Schrenzel, D. P. Lew, and P. Vaudaux. 2006. Transcriptomic and functional analysis of an autolysis-deficient, teicoplanin-resistant derivative of methicillin-resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 503048-3061. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 38.Resch, A., R. Rosenstein, C. Nerz, and F. Gotz. 2005. Differential gene expression profiling of Staphylococcus aureus cultivated under biofilm and planktonic conditions. Appl. Environ. Microbiol. 712663-2676. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 39.Rodgers, J. D., J. J. McCullagh, P. T. McNamee, J. A. Smyth, and H. J. Ball. 1999. Comparison of Staphylococcus aureus recovered from personnel in a poultry hatchery and in broiler parent farms with those isolated from skeletal disease in broilers. Vet. Microbiol. 69189-198. [DOI] [PubMed] [Google Scholar]
- 40.Saïd-Salim, B., P. M. Dunman, F. M. McAleese, D. Macapagal, E. Murphy, P. J. McNamara, S. Arvidson, T. J. Foster, S. J. Projan, and B. N. Kreiswirth. 2003. Global regulation of Staphylococcus aureus genes by Rot. J. Bacteriol. 185610-619. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 41.Sibbald, M. J., A. K. Ziebandt, S. Engelmann, M. Hecker, J. A. de, H. J. Harmsen, G. C. Raangs, I. Stokroos, J. P. Arends, J. Y. Dubois, and J. M. van Dijl. 2006. Mapping the pathways to staphylococcal pathogenesis by comparative secretomics. Microbiol. Mol. Biol. Rev. 70755-788. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Smith, D. A., T. Parish, N. G. Stoker, and G. J. Bancroft. 2001. Characterization of auxotrophic mutants of Mycobacterium tuberculosis and their potential as vaccine candidates. Infect. Immun. 691142-1150. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 43.Sugai, M., T. Fujiwara, H. Komatsuzawa, and H. Suginaka. 1998. Identification and molecular characterization of a gene homologous to epr (endopeptidase resistance gene) in Staphylococcus aureus. Gene 22467-75. [DOI] [PubMed] [Google Scholar]
- 44.Tamura, K., J. Dudley, M. Nei, and S. Kumar. 2007. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) software version 4.0. Mol. Biol. Evol. 241596-1599. [DOI] [PubMed] [Google Scholar]
- 45.Tschierske, M., C. Mori, S. Rohrer, K. Ehlert, K. J. Shaw, and B. Berger-Bachi. 1999. Identification of three additional femAB-like open reading frames in Staphylococcus aureus. FEMS Microbiol. Lett. 17197-102. [DOI] [PubMed] [Google Scholar]
- 46.van Belkum, A., D. C. Melles, S. V. Snijders, W. B. van Leeuwen, H. F. Wertheim, J. L. Nouwen, H. A. Verbrugh, and J. Etienne. 2006. Clonal distribution and differential occurrence of the enterotoxin gene cluster, egc, in carriage- versus bacteremia-associated isolates of Staphylococcus aureus. J. Clin. Microbiol. 441555-1557. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 47.van Leeuwen, W. B., D. C. Melles, A. Alaidan, M. Al-Ahdal, H. A. Boelens, S. V. Snijders, H. Wertheim, E. van Duijkeren, J. K. Peeters, P. J. van der Spek, R. Gorkink, G. Simons, H. A. Verbrugh, and A. van Belkum. 2005. Host- and tissue-specific pathogenic traits of Staphylococcus aureus. J. Bacteriol. 1874584-4591. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 48.Voyich, J. M., K. R. Braughton, D. E. Sturdevant, A. R. Whitney, B. Said-Salim, S. F. Porcella, R. D. Long, D. W. Dorward, D. J. Gardner, B. N. Kreiswirth, J. M. Musser, and F. R. DeLeo. 2005. Insights into mechanisms used by Staphylococcus aureus to avoid destruction by human neutrophils. J. Immunol. 1753907-3919. [DOI] [PubMed] [Google Scholar]
- 49.Wang, L., J. D. Trawick, R. Yamamoto, and C. Zamudio. 2004. Genome-wide operon prediction in Staphylococcus aureus. Nucleic Acids Res. 323689-3702. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 50.Zecconi, A., L. Cesaris, E. Liandris, V. Dapra, and R. Piccinini. 2006. Role of several Staphylococcus aureus virulence factors on the inflammatory response in bovine mammary gland. Microb. Pathog. 40177-183. [DOI] [PubMed] [Google Scholar]
- 51.Zhang, R., and C. T. Zhang. 2006. The impact of comparative genomics on infectious disease research. Microbes. Infect. 81613-1622. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.