Abstract
It is hypothesized that the nanoscale organization of cell adhesion ligands in a synthetic ECM regulates non-viral gene delivery. This hypothesis was examined with pre-osteoblasts cultured on substrates which present varied density and spacing of synthetic adhesion ligands. The levels of gene transfer and expression were increased with the density of adhesion ligands, but decreased with the spacing of ligands, due to changes in the cell growth rate. This study provides a material-based control point at the nanometer scale for non-viral gene based therapies.
Keywords: RGD peptides, plasmid DNA, gel matrix, proliferation
Non-viral gene vectors [e.g., plasmid DNA (pDNA)] are being increasingly used in a variety of therapeutic applications1–3 because of safety concerns related to viral vectors.4 One of the major challenges with the use of non-viral gene vectors is the very limited range of transgene expression level, owing to the poor efficiency of gene transfer.5 Therefore, there have been extensive efforts to design novel gene carriers to improve the levels of gene transfer and expression using lipids, polycations, dendrimers, nanorods, and oligopeptides5–7 and efforts to spatially concentrate the pDNA.8 In contrast, the design of gene delivery systems has largely ignored the importance of the cellular microenvironment in gene uptake and expression, although target tissues for in vivo gene delivery present varying cellular microenvironments.
Gene transfer is related to the rate of cell proliferation,9 and the chemistry and mechanics of the ECM to which a cell adheres regulates many aspects of cell phenotype, including proliferation, apoptosis and differentiation.10–14 These findings suggest that alteration of the ECM may allow one to regulate a cell’s ability to take up and express exogeneous genes. Recently, we have demonstrated that the mechanical rigidity of synthetic ECM used as adhesion substrates plays a critical role in modulating gene transfer and expression.15 In this study, we demonstrate a new material-based control point to regulate non-viral delivery by engineering the nanoscale organization of cell adhesion molecules in the synthetic ECM. The underlying mechanism of this regulation appears to be related to the material control over cell proliferation. The spatial distribution of cell adhesion molecules at the nanometer scale was controlled using a gel matrix formed from cross-linked alginate molecules containing covalently bound synthetic oligopeptides containing the Arg-Gly-Asp (RGD) sequence (RGD peptides) which mediate cell adhesion.16 Because unmodified alginate molecules are highly inert to cell adhesion and protein adsorption,17 cellular adhesion to gels functionalized with RGD peptides is solely attributed to the bonding between cell receptors and RGD peptides.
The overall density of RGD peptides (NRGD) and the distance between islands of RGD peptides (dRGD), which are known to regulate cell proliferation11, were used as variables to control the spatial organization of RGD peptides. NRGD was varied from 3 × 109 to 60 × 109/mm2 by altering the degree of substitution, defined as the number of RGD peptides bound to a single polymer chain (Fig. 1a). The area occupied by a single polymer chain modified with RGD peptides was defined as an island of RGD peptides. Analysis with gel permeation chromatography showed that the degree of substitution had little influence on the radius of gyration of the polymer chains, inferring a minimal change in the size of islands of RGD peptides and the distance between them (dRGD) with varying degree of substitution. dRGD was varied from 36 to 120 nm, at a given NRGD of 6 × 109/mm2, by diluting RGD peptide-modified polymer chains with unmodified polymer chains in the gel matrix (Fig. 1d). dRGD was calculated with a Monte Carlo simulation based on a non-overlapping random percolation model.18 Increasing the dilution ratio leads to larger dRGD, and the number of RGD peptides in an one island was raised in parallel from 2 to 20 to maintain a constant NRGD. These variations in the spatial organization of RGD peptides had minimal influences on the physical properties of gel matrix (i.e., elastic modulus and swelling ratio), or the interaction between the gel and gene vectors.
Figure 1.
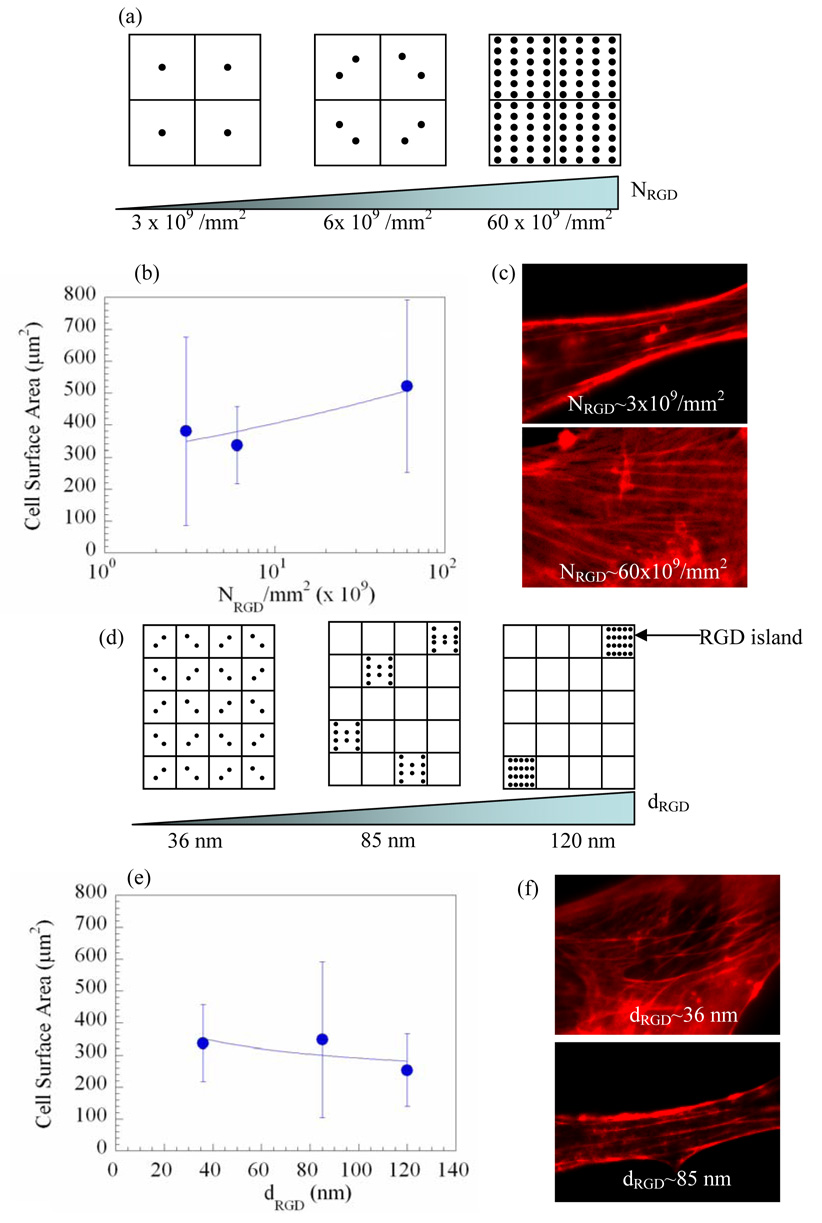
Schematic illustration of nanoscale spatial organization of RGD peptides in the gel matrix to which a cell adheres, and its influence on cell adhesion. Total density of RGD peptides (NRGD) was varied from 3 × 109, to 6 × 109 and 60 × 109/mm2 (a) using polymer molecules with different degree of substitution of RGD peptides. NRGD did not significantly alter the surface area of cells adhered to the gel matrix (b), but enhanced the formation of intracellular actin stress fibers (labeled with rhodamine phalloidin) (c). The distance between RGD islands (dRGD) was varied from 36 to 85 and 120 nm (d) by diluting RGD peptide-modified polymer molecules with unmodified polymer molecules. The increase in dRGD at a given NRGD of 6 × 109/mm2 did not significantly change the surface area of cells (e), but limited formation of stress fibers (f). Each grid in the illustration represents an island of RGD peptides with a scale of 30 nm by 30 nm. • in grid represents RGD peptides bound to polymer chain. Data points and error bars in (b) and (e) represent the mean and standard deviation from more than 10 cells at each condition.
We first examined whether these material variables modulate the adhesion of preosteoblasts to the gel matrix, as this can significantly influence cell fate.11 The dependency of the extent of cell spreading, quantified with the surface area of cells on NRGD, was statistically insignificant, although the average values were increased with NRGD (Fig. 1b). In contrast, the number of stress fibers formed in a single cell was increased with NRGD (Fig. 1c). The variation in dRGD at a given NRGD had a statistically insignificant influence on the cell morphology, although the average values were decreased with increasing dRGD (Fig. 1e). However, increases in dRGD significantly limited the formation of stress fibers (Fig. 1f). These results suggest that both NRGD and dRGD mediate cytoskeletal rearrangements, which have been related to signaling pathways promoting cell proliferation.19
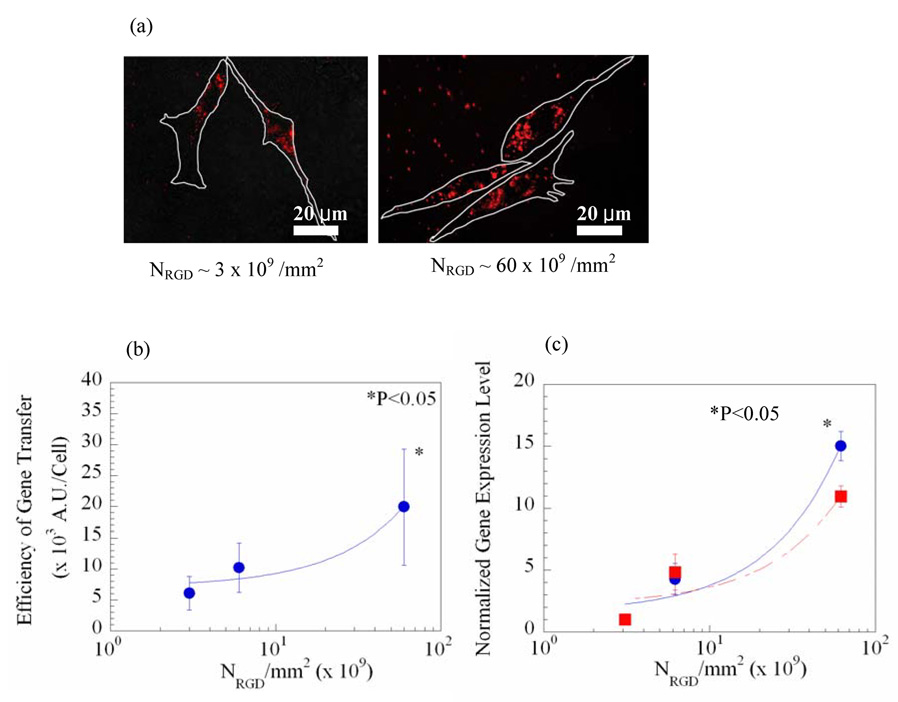
Next, the role of NRGD in gene therapy was examined by quantifying the efficiency of gene transfer and expression level in cells adhered to gel matrices presenting different NRGD. pDNA encoding for the luciferase protein was condensed with poly(ethyleneimine) (PEI) to yield nanosize pDNA condensates with enhanced cellular uptake, and delivered to cells.20 Interestingly, the efficiency of gene transfer quantified using pDNA labeled with rhodamine was significantly enhanced with NRGD, as illustrated with the linear increase in the fluorescence yield per cell (Fig. 2a–b). Subsequently, the gene expression level was quantified from the luciferase activity in the cells, and this linearly increased with NRGD (Fig. 2c). The gene expression level normalized to the average surface area of cells was also linearly related to NRGD.
Figure 2.
Gene delivery regulated with the total density of RGD peptides (NRGD) presented from the gel matrix to which cells adhered. Microscopic images show the increases in the cellular uptake of the rhodamine-labeled pDNA condensates with NRGD (a). White lines in photomicrographs represent the border of cells adhered to the gel matrix. The efficiency of gene transfer, quantified from the photomicrographs, linearly increased with NRGD (R2 = 0.94) (b). The gene expression level (● in c) and the gene expression level normalized to the average surface area of cells (■ in c) also linearly increased with NRGD (R2 = 0.94). More than 20 cells at each condition were used for the quantitative analysis of the efficiency of gene transfer. Differences in the values in (b)–(c) for cells adhered to gels with the highest NRGD versus the two other conditions were statistically significant (p < 0.05). The gene expression level in (c) was normalized to the gene expression level attained from cells cultured on gels with NRGD of 3 × 109/mm2. Data points and error bars represent the mean and standard deviation from four independent experiments.
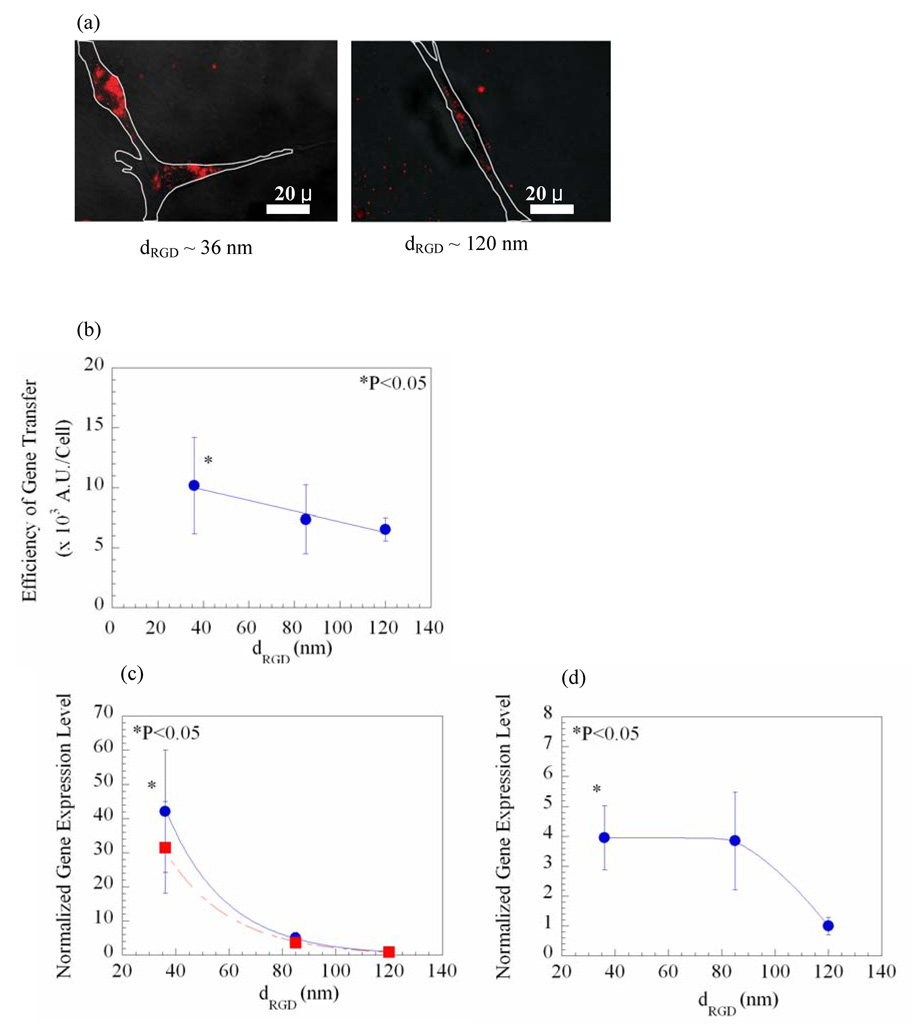
Manipulation of dRGD also modulated non-viral gene delivery, even as NRGD remained constant. Increasing dRGD linearly decreased the cellular uptake of pDNA condensates, as verified by quantifying the fluorescence yield per cell (Fig. 3a–b). Both overall gene expression levels and expression levels normalized to the average surface area of cells exponentially decreased with dRGD [gene expression level ∝ exp (−2dRGD)] (Fig. 3c). This inverse relationship between the gene expression level and dRGD was also found with cells maintained in 3D culture (cells encapsulated in the gel matrix) (Fig. 3d), although the magnitude of this effect was more evident with the larger dRGD.
Figure 3.
Gene delivery regulated with the distance between islands of RGD peptides (dRGD), as the total density of RGD peptides was kept constant at 6 × 109/mm2. Microscopic images show the decrease in the cellular uptake of the rhodamine-labeled pDNA condensates with dRGD (a). White lines in photomicrographs represent borders of cells adhered to the gel matrix. The efficiency of gene transfer quantified from photomicrographs was linearly decreased with dRGD (R2 = 0.96) (b). The gene expression level (● in c) and the gene expression level normalized to the average surface area of cells (■ in c) exponentially decreased with dRGD (R2 = 1). Increasing dRGD also down-regulated the gene expression level in cells cultured within the 3D gel matrix (d). Differences in the values in (b) for cells cultured on gels presenting the largest dRGD (~ 120 nm) versus those on gels presenting the smallest dRGD (~ 36 nm) were statistically significant (p < 0.05). The gene expression levels in (c) and (d) were normalized to the value attained from cells cultured on gels presenting dRGD of 120 nm. Data points and error bars represent the mean and standard deviation from four independent experiments.
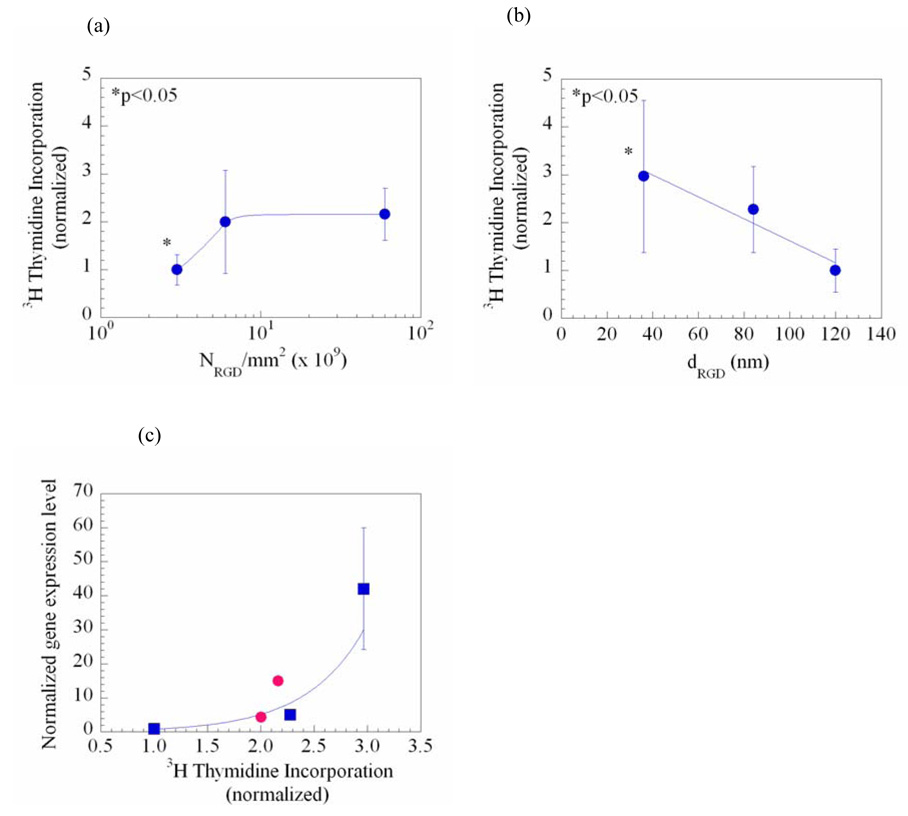
The relation between ECM control over gene delivery and cell growth was next assessed by quantifying 3H thymidine incorporation, as 3H thymidine is incorporated by cells synthesizing DNA. The magnitude of cell growth increased with NRGD (Fig. 4a), and, inversely, the magnitude of cell growth decreased with dRGD (Fig. 4b), as expected from past studies. Directly monitoring cell growth by quantifying cell number over time confirmed these effects of NRGD and dRGD on proliferation (data not shown). Both the efficiency of gene transfer (data not shown) and gene expression levels correlated in an exponential fashion to cell growth (Fig. 4c), although the dependency of gene expression on cell growth was distinct for the two material variables.
Figure 4.
The spatial organization of RGD peptides regulated the rate of cell proliferation, which was quantified with 3H thymidine incorporation. The amount of DNA synthesis was increased proportional to NRGD (a) or with the inverse of dRGD (b). Hence, the extent of cell proliferation was related to the normalized gene expression level following an exponential curve (c) (R2 = 0.94). In (a), differences in the values for cells cultured on the gel matrix presenting the smallest NRGD versus the two other conditions were statistically significant (p < 0.05). In (b), differences in the values for cells cultured on the gel matrix presenting the smallest dRGD versus the two other conditions were also statistically significant (p < 0.05). Data points and error bars represent the mean and standard deviation from four independent experiments. In (c), curves -●- and -■-represent conditions in which NRGD and dRGD, respectively, were varied.
Altogether, the results of this study demonstrate that the nanoscale organization of RGD peptides allows one to regulate non-viral gene delivery to both cells adhered to 2D substrates and those encapsulated in 3D matrices. Both NRGD and dRGD likely modulated the cell cycle by mediating cytoskeletal rearrangements and activation of intracellular signaling pathways.19 No significant influence of surface area of cells on the gene delivery and expression was observed over the range examined in these studies. Manipulation of cell mitosis may directly alter the cellular uptake of exogeneous genes and the resultant gene expression level, as cell division has been previously noted to facilitate the entry of pDNA into cells.9 It is likely that the greater dependency of the gene expression level on dRGD in 2D versus 3D cell culture is related to the decreased frequency of cell cycle progression in 3D. However, it is unclear whether the decreased proliferation is inherent to 3D culture generally, or if it relates to the ability of cells in 2D culture to spread more extensively than cells in this 3D culture, and more broadly probe their environment. One interesting finding from the current study is that the magnitude of the dependency of gene expression levels on the material variables was larger than the dependency of the efficiency of gene transfer on NRGD and dRGD. However, this result infers that material signals further influence events beyond pDNA uptake, and these additional control points may include the intracellular dissociation of pDNA condensates, their escape from lysosomes, and/or their trafficking to the cell nucleus.7 However, this issue must be further examined, as the dependencies of the efficiency of gene transfer and expression level on NRGD and dRGD may also relate to the different analytical methods used in their evaluation.
This study demonstrates for the first time that the number and nanoscale distribution of cell adhesion molecules in a material play critical roles in non-viral gene delivery. This is a previously undescribed material-based control point for gene transfer, and may significantly contribute to improving the quality of various gene-based therapies. We envisage that these results can be reproduced with diverse types of cells, including primary cells, and other biomaterials functionalized to promote cell adhesion.21 The results of this study may also be relevant to understand the varying degrees of gene transfer in cells within tissues under different conditions (e.g., normal versus pathologic ECM), as diseases and the state of development may alter the assembly and organization of the ECM22. In these situations, the concepts learned from this study may be useful in predicting the efficacy of gene transfer in a known ECM environment, and also in recreating, in vitro, more realistic tissue models to study these processes23. Most current model systems simply involve coating adhesion substrates with cell adhesion molecules, without regard to the critical effects of ligand presentation. Furthermore, the results of this study may be used to guide the design of gene activated matrices24, in which localized gene therapy is promoted in vivo by the migration of cells into a plasmid DNA-loaded material3. Proper ligand presentation in the material may be used to stimulate cell proliferation and subsequently promote the local and extended expression of target therapeutic molecules from both endogeneous and transplanted cells.
Supplementary Material
Description of the experimental procedures. This material is available free of charge via the Internet at http://pubs.acs.org
Acknowledgement
The authors thank the NIH/NIDCR (R 37 DE013033) and US Army Research Laboratories and Research Office (DAAD-19-03-1-0168) for financial support of this research.
References
- 1.Glover DJ, Lipps HJ, Jans DA. Nat. Rev. Gene. 2005;6:299. doi: 10.1038/nrg1577. [DOI] [PubMed] [Google Scholar]
- 2.Niidome T, Huang L. Gene Ther. 2002;9:1647. doi: 10.1038/sj.gt.3301923. [DOI] [PubMed] [Google Scholar]
- 3.Huang YC, Kaigler D, Rice KG, Krebsbach PH, Mooney DJ. J. Bone Mineral Res. 2005;20:848. doi: 10.1359/JBMR.041226. [DOI] [PubMed] [Google Scholar]
- 4.Cavazzana-Calvo M, Lagresle C, Hacein-Bey-Abina S, Fischer SA. Annu. Rev. Med. 2005;56:585. doi: 10.1146/annurev.med.56.090203.104142. [DOI] [PubMed] [Google Scholar]
- 5.Mastrobattista E, van der Aa MAEM, Hennink WE, Crommelin DJA. Nat. Rev. Drug Discovery. 2006;5:115. doi: 10.1038/nrd1960. [DOI] [PubMed] [Google Scholar]
- 6.Park TG, Jeong JH, Kim SW. Adv. Drug Deliv. Rev. 2006;58:467. doi: 10.1016/j.addr.2006.03.007. [DOI] [PubMed] [Google Scholar]
- 7.Pack DW, Hoffman AS, Pun S, Stayton PS. Nat. Rev. Drug Discovery. 2005;4:581. doi: 10.1038/nrd1775. [DOI] [PubMed] [Google Scholar]
- 8.Segura T, Shea LD. Bioconju. Chem. 2002;13:621. doi: 10.1021/bc015575f. [DOI] [PubMed] [Google Scholar]
- 9.Fasbender A, Zabner J, Zeiher BG, Welsh MJ. Gene Ther. 1997;4:1173. doi: 10.1038/sj.gt.3300524. [DOI] [PubMed] [Google Scholar]
- 10.Discher DE, Jammey P, Wang YL. Science. 2005;310:1139. doi: 10.1126/science.1116995. [DOI] [PubMed] [Google Scholar]
- 11.Lee KY, Alsberg E, Hsiong S, Comisar W, Linderman J, Ziff R, Mooney DJ. Nano Lett. 2004;4:1501. doi: 10.1021/nl0493592. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 12.Keselowsky BG, Collard DM, Garcia A. Proc. Nat. Acade. Sci (USA) 2005;102:5953. doi: 10.1073/pnas.0407356102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 13.Koo LY, Irvine DJ, Mayes AM, Lauffenburger DA, Griffith LG. J. Cell Sci. 2002;115:1423. doi: 10.1242/jcs.115.7.1423. [DOI] [PubMed] [Google Scholar]
- 14.Kong HJ, Polte TR, Alsberg E, Mooney DJ. Proc. Nat. Acad. Sci. (USA) 2005;102:4300. doi: 10.1073/pnas.0405873102. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 15.Kong HJ, Liu J, Riddle K, Matsumoto T, Leach K, Mooney DJ. Nat. Mater. 2005;4:460. doi: 10.1038/nmat1392. [DOI] [PubMed] [Google Scholar]
- 16.Rowley J, Mooney DJ. J. Biomed. Mater. Res. 2002;60:217. doi: 10.1002/jbm.1287. [DOI] [PubMed] [Google Scholar]
- 17.Smetana K. Biomaterials. 1993;14:1046. doi: 10.1016/0142-9612(93)90203-e. [DOI] [PubMed] [Google Scholar]
- 18.Comisar WA, Hsiong S, Kong HJ, Mooney DJ, Linderman JJ. Biomaterials. 2006;27:2322. doi: 10.1016/j.biomaterials.2005.10.037. [DOI] [PubMed] [Google Scholar]
- 19.Cai S, Pestic-Dragovich L, O'Donnell ME, Wang N, Ingber D, Elson E, De Lanerolle P. Am. J. Physiol.-Cell Physiol. 1998;275:C1345. doi: 10.1152/ajpcell.1998.275.5.C1349. [DOI] [PubMed] [Google Scholar]
- 20.Boussif O, Lezoualch F, Zanta MA, Mergny MD, Scherman D, Demeneix B, Behr JP. Proc. Nat. Acade. Sci (USA) 1995;92:7297. doi: 10.1073/pnas.92.16.7297. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 21.Hersel U, Dahmen C, Kessler H. Biomaterials. 2003;23:4385. doi: 10.1016/s0142-9612(03)00343-0. [DOI] [PubMed] [Google Scholar]
- 22.Ingber DE. Ann. Med. 2003;35:564. doi: 10.1080/07853890310016333. [DOI] [PubMed] [Google Scholar]
- 23.Griffith LG, Swartz MA. Nat. Rev. Mol. Cell Biol. 2006;7:211. doi: 10.1038/nrm1858. [DOI] [PubMed] [Google Scholar]
- 24.Bonadio J. Adv. Drug Deliv. Rev. 2000;44:185. doi: 10.1016/s0169-409x(00)00094-6. [DOI] [PubMed] [Google Scholar]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
Description of the experimental procedures. This material is available free of charge via the Internet at http://pubs.acs.org