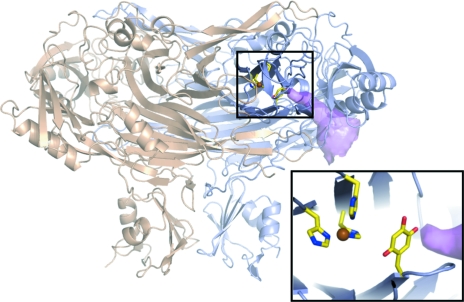
The structure of a xenon derivative of E. coli copper amine oxidase confirms the pathway of oxygen entry to the buried active site proposed for this class of enzymes.
Keywords: copper amine oxidase, xenon derivatives, oxygen entry
Abstract
The mechanism of molecular oxygen entry into the buried active site of the copper amine oxidase family has been investigated in several family members using biochemical, structural and in silico methods. These studies have revealed a structurally conserved β-sandwich which acts as a hydrophobic reservoir from which molecular oxygen can take several species-specific preferred pathways to the active site. Escherichia coli copper amine oxidase (ECAO) possesses an extra N-terminal domain that lies close to one entrance to the β-sandwich. In order to investigate whether the presence of this domain alters molecular oxygen entry in this enzyme, xenon was used as a molecular oxygen binding-site probe. The resulting 2.5 Å resolution X-ray crystal structure reveals xenon bound in similar positions to those observed in xenon-derivative crystal structures of other family members, suggesting that the N-terminal domain does not affect oxygen entry and that the E. coli enzyme takes up oxygen in a similar manner to the rest of the copper amine oxidase family.
1. Introduction
Copper amine oxidases (CuAOs) catalyse the oxidation of a wide range of amine substrates to generate a product aldehyde, ammonia and hydrogen peroxide (Mure et al., 2002 ▶). They are ubiquitous in nature and in humans are involved in diverse processes including cell adhesion and signalling (Salmi et al., 1998 ▶; Elmore et al., 2002 ▶; Yu et al., 2003 ▶). In bacteria, they play a role in general nitrogen catabolism (Parrott et al., 1987 ▶; Hacisalihoglu et al., 1997 ▶). They contain a protein-derived cofactor that is generated autocatalytically by post-translation modification of a conserved endogenous tyrosine to generate 2,4,5-trihydroxylphenylalanine quinone (TPQ), requiring only the presence of copper and molecular oxygen (Cai & Klinman, 1994 ▶; Kim et al., 2002 ▶). TPQ is situated in the core of the protein at the end of a well defined substrate-access channel next to a type II copper coordinated by three histidines (Fig. 1 ▶; Parsons et al., 1995 ▶; Duff et al., 2006 ▶; Wilce et al., 1997 ▶; Lunelli et al., 2005 ▶; Kumar et al., 1996 ▶; Airenne et al., 2005 ▶; Li et al., 1998 ▶). The Escherichia coli enzyme (ECAO) shows a preference for small aromatic amine substrates such as phenethylamine and tyramine (Roh et al., 1994 ▶). The catalytic reaction proceeds via two half-reactions; the aldehyde product is released at the end of the reductive half-reaction before reduction of molecular oxygen in the oxidative half-reaction (Fig. 2 ▶).
Figure 1.
The structure of ECAO (PDB code 1dyu) showing the substrate-entrance channel (magenta surface) and active site (Murray et al., 1999 ▶). The lower right panel is a more detailed view of the active site showing the TPQ, copper(II) ion, three coordinating histidine residues (sticks coloured by atom type) and the end of the substrate-entrance channel (magenta surface). The substrate-entrance channel was calculated using CAVER (Petrek et al., 2006 ▶) and the image was prepared using PyMOL (DeLano, 2002 ▶).
Figure 2.
Schematic catalytic cycle of CuAOs identifying the key intermediates in the reaction pathway. Shown are the type-2 copper centre, the protein-derived cofactor TPQ and the off-metal O2-binding site located in the vicinity of a conserved active-site methionine (Met699 in ECAO). I is the resting state of the enzyme, intermediate II is the product Schiff base, intermediate III is the reduced TPQ or aminoquinol form, intermediate IV shows the proposed off-metal-bound dioxygen species prior to oxygen activation and reduction and intermediate V is the iminoquinone species which hydrolyses to release ammonia, completing the catalytic cycle.
Extensive structural and biochemical analyses of CuAOs from prokaryotic and eukaryotic sources have revealed that molecular oxygen binds on the opposite side of TPQ to the amine substrate (Fig. 2 ▶). There has been considerable interest in dissecting the pathway by which oxygen reaches and binds in the active site. Structural and mutagenesis studies have shown that oxygen does not enter via the substrate channel and that other possibly specific oxygen-access pathways must be present (Wilce et al., 1997 ▶; Li et al., 1998 ▶; Johnson et al., 2007 ▶; Wilmot et al., 1999 ▶; Goto & Klinman, 2002 ▶).
Several recent crystallographic studies of oxygen entry have been carried out using xenon as a probe to identify possible molecular oxygen-binding sites that may be part of the putative oxygen-entry pathway (Lunelli et al., 2005 ▶; Johnson et al., 2007 ▶; Duff et al., 2004 ▶). Xenon-complex crystal structures are now available for the CuAOs from Hansenula polymorpha (HPAO), Pisum sativum (PSAO), bovine serum (BSAO), Pichia pastoris (PPLO) and Arthobacter globiformis (AGAO) (Lunelli et al., 2005 ▶; Johnson et al., 2007 ▶; Duff et al., 2004 ▶). These structures have revealed a consistent pattern of xenon binding that has implicated a major pathway for oxygen entry through a conserved β-sandwich (Fig. 3 ▶). Very recently, in silico experiments that calculated likely low-energy pathways for oxygen entry in HPAO, PSAO, AGAO and PPLO were carried out that supported the major pathway identified by the structural studies (Johnson et al., 2007 ▶).
Figure 3.
ECAO subunit showing the substrate-entry channel as a magenta surface and the putative oxygen-entry pathway as a green surface (generated by placing dummy atoms along the channel). The xenon sites observed in PPLO, HPAO, AGAO and PSAO are shown as cyan spheres. TPQ and copper ligands are shown as sticks coloured by element and copper is shown as a bronze sphere. This image was generated using PyMOL (DeLano, 2002 ▶).
ECAO is distinct from Gram positive bacterial and yeast CuAOs in that it possesses an extra 9.3 kDa N-terminal domain, which gives ECAO a characteristic ‘mushroom stalk and cap’ shape (Parsons et al., 1995 ▶). Mammalian CuAOs also possess an N-terminal domain that is thought to act as a membrane anchor (Smith et al., 1998 ▶) but has not yet been visualized crystallographically (Airenne et al., 2005 ▶). Biochemical and structural characterization of ECAO indicate that the binding of oxygen in the active site is similar to that of the other family members (Smith & McPherson, 2008 ▶); however, it is possible that the presence of the additional N-terminal domain, which is located at one end of the structurally conserved β-sandwich, could alter or provide an alternate pathway for oxygen entry. In order to investigate this possibility, we have determined the crystal structure of ECAO in complex with xenon at 2.5 Å resolution.
2. Experimental
2.1. ECAO expression, purification and crystallization
ECAO expression and purification was carried out as previously described (Murray et al., 1999 ▶). ECAO was concentrated to 8 mg ml−1 and crystals were grown by vapour diffusion at 291 K. Large pink-coloured crystals grew over two weeks in 100 mM HEPES pH 7 and 1.2 M sodium citrate.
2.2. Xenon derivatization
ECAO crystals were harvested and transferred into a cryoprotectant solution comprising crystal-growth mother liquor containing 20% glycerol before being exposed to xenon for 8 min at 2.5 MPa in an Oxford Cryosystems Xcell. After derivatization, crystals were immediately flash-cooled in liquid nitrogen for X-ray data collection.
2.3. X-ray data collection and processing
A 2.5 Å diffraction data set was collected on beamline 14.1 at the SRS, Daresbury. Data collection was carried out at the highest beamline-accessible wavelength of 1.48 Å to maximize the anomalous signal from xenon (Xe f′′ = 6.9) and data were recorded using a Quantum4 ADSC CCD detector. The crystal was maintained at 100 K using a Cryostream (Oxford Cryosystems). 360° of data were collected with an oscillation angle of 1° and an exposure time of 30 s. The diffraction data were processed using MOSFLM and scaled using SCALA (Leslie, 1992 ▶; Evans, 1993 ▶; Table 1 ▶).
Table 1. Data collection and processing.
Values in parentheses are for the highest resolution shell.
| Wavelength (Å) | 1.48 |
| Temperature (K) | 100 |
| Unit-cell parameters (Å) | a = 166.9, b = 134.6, c = 80.3 |
| Space group | P212121 |
| Resolution (Å) | 50.0–2.5 (2.55–2.50) |
| Rmerge† (%) | 12.3 (44.8) |
| Completeness (%) | 98.6 (94.9) |
| Redundancy | 10.1 (7.4) |
| Mean I/σ(I) | 23.6 (4.4) |
R
merge = 
 , where Ii(hkl) is the observed intensity and 〈I(hkl)〉 is the average intensity for multiple measurements.
, where Ii(hkl) is the observed intensity and 〈I(hkl)〉 is the average intensity for multiple measurements.
2.4. Refinement and model building
All refinement and model building was carried out using the CCP4 software suite and Coot (Collaborative Computational Project, Number 4, 1994 ▶; Emsley & Cowtan, 2004 ▶). Initial maps were calculated by direct Fourier transform using the structure of native ECAO (PDB code 1dyu; Murray et al., 1999 ▶). An anomalous difference map was calculated and examined to locate possible xenon sites. 50 peaks were identified in the map with a σ level of 4 or above. Two of these (8.5 and 8.0 r.m.s.) were located at sites known to bind calcium in the native enzyme (Ca f′′ = 1.2 at 1.48 Å). Weaker peaks (3.6–4.6 r.m.s.) were located at the copper site and on S atoms, which is expected at this wavelength (Cu f′′ = 0.55 at 1.48 Å, S f′′ = 0.5 at 1.48 Å). The remaining peaks were assigned as xenon when they were higher than 4 r.m.s., did not overlap with copper, calcium or sulfur, corresponded to positive electron density in the F o − F c electron-density map and were located in mainly hydrophobic pockets. The occupancy of each xenon was modified such that after restrained refinement there was no further positive electron density in the F o − F c map and the B factors were similar to those of nearby atoms. The structure was further refined using iterative cycles of model building and restrained refinement to a final R factor of 16.1% and an R free of 21.9% (Table 2 ▶). Structure validation was carried out using SFCHECK and PROCHECK (Laskowski et al., 1993 ▶; Vaguine et al., 1999 ▶).
Table 2. Structure refinement and validation.
Values in parentheses are for the highest resolution shell.
| Rwork† | 16.1 (21.9) |
| Rfree‡ | 21.9 (30.1) |
| Total No. of reflections | 64552 |
| No. of reflections in the Rwork set | 61345 (3900) |
| No. of reflections in the Rfree set | 3207 (220) |
| No. of non-H atoms | 11947 |
| No. of protein atoms | 11348 |
| No. of ligand atoms | 17 |
| No. of solvent atoms | 582 |
| R.m.s.d. from ideality | |
| Bonds (Å) | 0.017 |
| Angles (°) | 1.7 |
| Average B factors (Å2) | |
| Main chain | 39.6 |
| Side chain | 41.5 |
| Ligands | 49.4 |
| Solvent | 44.7 |
| Ramachandran plot | |
| Allowed regions (%) | 99.8 |
| Disallowed regions (%) | 0.2 |
| PDB code | 2w0q |
R factor = 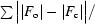
 , where |F
o| is the observed structure-factor amplitude and |F
o| is the calculated structure-factor amplitude.
, where |F
o| is the observed structure-factor amplitude and |F
o| is the calculated structure-factor amplitude.
R free is the R factor based on 5% of the data excluded from refinement.
3. Results and discussion
After refinement, 11 xenon-binding sites were found in the structure (Fig. 4 ▶ a). Although the occupancy of all sites was low (0.1–0.45), this is consistent with the xenon occupancies observed in other CuAO xenon complexes (Lunelli et al., 2005 ▶; Johnson et al., 2007 ▶; Duff et al., 2004 ▶). Xenons were numbered according to their occupancy (from high to low occupancy). The Xe atoms were bound in hydrophobic pockets located in either the amine substrate-entry channel or around the conserved β-sandwich oxygen-entry pathway (Table 3 ▶). Nearly identical sites were observed in both subunits of ECAO present in the asymmetric unit1.
Figure 4.
(a) ECAO monomer showing the substrate-entry channel as a magenta surface and the putative oxygen-entry pathway as a green surface (generated by placing dummy atoms along the channel). The Xe sites observed in ECAO are shown as magenta spheres. TPQ and copper ligands are shown as sticks coloured by element and copper is shown as a bronze sphere. (b) ECAO active site, coloured as before, showing xenons 1/2, 4/11, 7/5, 8/– and 9/–. This image was generated using PyMOL (DeLano, 2002 ▶).
Table 3. Xenon sites.
| Xenon site | Peak height | Occupancy/B factor (Å2) | Surrounding residues |
|---|---|---|---|
| 1 | 13 | 0.45/58 | Subunit A Leu189, Phe192, Phe387, Leu392 |
| 2 | 9.9 | 0.30/48 | Subunit B Leu189, Phe192, Tyr387, Leu392 |
| 3 | 9.9 | 0.30/39 | Subunit A His94, Ala140, Met322, Thr344 |
| 4 | 6.5 | 0.35/57 | Subunit A Phe192, Pro224, Leu225, Tyr381, Tyr387 |
| 5 | 5.8 | 0.15/48 | Subunit B Leu543, Ile570, Gln578, Leu638 |
| 6 | 5.7 | 0.3/57 | Subunit B His94, Ala140, Met322, Thr344 |
| 7 | 5.6 | 0.2/46 | Subunit A Leu543, Ile570, Gln578, Leu638 |
| 8 | 4.9 | 0.15/27 | Subunit A Ile396, Ile460 |
| 9 | 4.5 | 0.1/37 | Subunit A Leu543, Leu588, Ile605, Leu638, Val681 |
| 10 | 4.5 | 0.15/47 | Subunit B Pro458, Met563, Gln620 |
| 11 | 4.2 | 0.2/58 | Subunit B Phe192, Pro224, Leu225, Tyr381, Tyr387 |
Two Xe atoms are bound in the amine substrate-entry channel. Xenon 4/11 occupies a site between Tyr381 and Tyr387 that is equivalent to that previously observed in AGAO and HPAO. Tyr381 is proposed to act as a gate regulating substrate access to the TPQ and the aromatic rings of Tyr381 and Tyr387 allow stacking interactions with the aromatic moiety of aromatic amine substrates (Wilmot et al., 1997 ▶, 2004 ▶). Xenon 1/2 is bound further from the active site but is still close to the amine-substrate channel (Fig. 4 ▶ b).
Two xenons, xenon 7/5 and xenon 9/– (only in monomer A), are bound inside the ‘top’ entrance of the β-sandwich, close to the xenon-binding sites observed in PPLO and PSAO (Duff et al., 2004 ▶; Figs. 4 ▶ a and 4 ▶ b). We did not observe any xenon-binding sites inside the lower half of the β-sandwich near the ‘bottom’ entrance to the putative oxygen-entry pathway. However, two xenon-binding sites (3/6 and 8/–) flank the lower half of the β-sandwich, binding on its external face (Fig. 4 ▶ a). This is consistent with the other copper amine oxidase–xenon complexes, in which a xenon-binding site within the bottom half of the β-sandwich channel has only been reported for AGAO (Fig. 3 ▶). The lack of xenon-binding sites in this region in PPLO, PSAO and HPAO suggests that xenon binding is not as favourable in this region of the β-sandwich regardless of the presence or absence of an N-terminal domain. A final xenon, xenon –/10, is bound at the edge of ECAO in a hydrophobic pocket and is not close to any previously observed xenon position.
The xenon-complex ECAO structure reported here shows that molecular oxygen is likely to access the ECAO active site by very similar pathways to those proposed by experimental and in silico studies in other CuAOs (Lunelli et al., 2005 ▶; Johnson et al., 2007 ▶; Duff et al., 2004 ▶). No xenon-binding sites are observed in the ECAO N-terminal domain itself nor does the presence of this domain appear to affect xenon binding in the lower half of the β-sandwich when compared with other CuAOs for which xenon complexes have been determined. These observations indicate that the N-terminal domain does not play a role in oxygen entry during ECAO catalysis, which is consistent with the lack of conservation of the N-terminal domain across the CuAOs.
Supplementary Material
PDB reference: E. coli copper amine oxidase, xenon derivative, 2w0q, r2w0qsf
Acknowledgments
PP is the recipient of a Wellcome Trust Studentship. This work was also supported by the Biotechnology and Biological Sciences Research Council (Grant No. BB/C00468X/1) and the RCUK (ARP). We thank Dr Thomas Edwards, Dr Chi Trinh and the beamline 14.1 local contacts for assistance during data collection.
Footnotes
As ECAO is a dimer in the asymmetric unit some xenon sites are observed in equivalent positions in both monomers, while others are observed in only one of the subunits. In the following discussion of individual xenon sites, those observed in both monomers are identified as ‘xenon A/B’, where A is the xenon number in monomer A and B that in monomer B. When a xenon is observed in only a single subunit, this is indicated as either ‘xenon A/–’ or ‘xenon –/B’.
References
- Airenne, T. T., Nymalm, Y., Kidron, H., Smith, D. J., Pihlavisto, M., Salmi, M., Jalkanen, S., Johnson, M. S. & Salminen, T. A. (2005). Protein Sci.14, 1964–1974. [DOI] [PMC free article] [PubMed]
- Cai, D. & Klinman, J. P. (1994). J. Biol. Chem.269, 32039–32042. [PubMed]
- Collaborative Computational Project, Number 4 (1994). Acta Cryst. D50, 760–763.
- DeLano, W. L. (2002). PyMOL Molecular Viewer. DeLano Scientific LLC, San Carlos, California, USA. http://www.pymol.org.
- Duff, A. P., Cohen, A. E., Ellis, P. J., Hilmer, K., Langley, D. B., Dooley, D. M., Freeman, H. C. & Guss, J. M. (2006). Acta Cryst. D62, 1073–1084. [DOI] [PubMed]
- Duff, A. P., Trambaiolo, D. M., Cohen, A. E., Ellis, P. J., Juda, G. A., Shepard, E. M., Langley, D. B., Dooley, D. M., Freeman, H. C. & Guss, J. M. (2004). J. Mol. Biol.344, 599–607. [DOI] [PubMed]
- Elmore, B. O., Bollinger, J. A. & Dooley, D. M. (2002). J. Biol. Inorg. Chem.7, 565–579. [DOI] [PubMed]
- Emsley, P. & Cowtan, K. (2004). Acta Cryst. D60, 2126–2132. [DOI] [PubMed]
- Evans, P. (1993). Proceedings of the CCP4 Study Weekend. Data Collection and Processing, edited by L. Sawyer, N. Isaacs & S. Bailey, pp. 114–122. Warrington: Daresbury Laboratory.
- Goto, Y. & Klinman, J. P. (2002). Biochemistry, 41, 13637–13643. [DOI] [PubMed]
- Hacisalihoglu, A., Jongejan, J. A. & Duine, J. A. (1997). Microbiology, 143, 505–512. [DOI] [PubMed]
- Johnson, B. J., Cohen, J., Welford, R. W., Pearson, A. R., Schulten, K., Klinman, J. P. & Wilmot, C. M. (2007). J. Biol. Chem.282, 17767–17776. [DOI] [PMC free article] [PubMed]
- Kim, M., Okajima, T., Kishishita, S., Yoshimura, M., Kawamori, A., Tanizawa, K. & Yamaguchi, H. (2002). Nature Struct. Biol.9, 591–596. [DOI] [PubMed]
- Kumar, V., Dooley, D. M., Freeman, H. C., Guss, J. M., Harvey, I., McGuirl, M. A., Wilce, M. C. & Zubak, V. M. (1996). Structure, 4, 943–955. [DOI] [PubMed]
- Laskowski, R. A., MacArthur, M. W., Moss, D. S. & Thornton, J. M. (1993). J. Appl. Cryst.26, 283–291.
- Leslie, A. G. W. (1992). Jnt CCP4/ESF–EACBM Newsl. Protein Crystallogr.26
- Li, R., Klinman, J. P. & Mathews, F. S. (1998). Structure, 6, 293–307. [DOI] [PubMed]
- Lunelli, M., Di Paolo, M. L., Biadene, M., Calderone, V., Battistutta, R., Scarpa, M., Rigo, A. & Zanotti, G. (2005). J. Mol. Biol.346, 991–1004. [DOI] [PubMed]
- Mure, M., Mills, S. A. & Klinman, J. P. (2002). Biochemistry, 41, 9269–9278. [DOI] [PubMed]
- Murray, J. M., Saysell, C. G., Wilmot, C. M., Tambyrajah, W. S., Jaeger, J., Knowles, P. F., Phillips, S. E. & McPherson, M. J. (1999). Biochemistry, 38, 8217–8227. [DOI] [PubMed]
- Parrott, S., Jones, S. & Cooper, R. A. (1987). J. Gen. Microbiol.133, 347–351. [DOI] [PubMed]
- Parsons, M. R., Convery, M. A., Wilmot, C. M., Yadav, K. D., Blakeley, V., Corner, A. S., Phillips, S. E., McPherson, M. J. & Knowles, P. F. (1995). Structure, 3, 1171–1184. [DOI] [PubMed]
- Petrek, M., Otyepka, M., Banás, P., Kosinová, P., Koca, J. & Damborský, J. (2006). BMC Bioinformatics, 7, 316. [DOI] [PMC free article] [PubMed]
- Roh, J. H., Suzuki, H., Azakami, H., Yamashita, M., Murooka, Y. & Kumagai, H. (1994). Biosci. Biotechnol. Biochem.58, 1652–1656. [DOI] [PubMed]
- Salmi, M., Hellman, J. & Jalkanen, S. (1998). J. Immunol.160, 5629–5636. [PubMed]
- Smith, M. A. & McPherson, M. J. (2008). Unpublished work.
- Smith, D. J., Salmi, M., Bono, P., Hellman, J., Leu, T. & Jalkanen, S. (1998). J. Exp. Med.188, 17–27. [DOI] [PMC free article] [PubMed]
- Vaguine, A. A., Richelle, J. & Wodak, S. J. (1999). Acta Cryst. D55, 191–205. [DOI] [PubMed]
- Wilce, M. C., Dooley, D. M., Freeman, H. C., Guss, J. M., Matsunami, H., McIntire, W. S., Ruggiero, C. E., Tanizawa, K. & Yamaguchi, H. (1997). Biochemistry, 36, 16116–16133. [DOI] [PubMed]
- Wilmot, C. M., Hajdu, J., McPherson, M. J., Knowles, P. F. & Phillips, S. E. (1999). Science, 286, 1724–1728. [DOI] [PubMed]
- Wilmot, C. M., Murray, J. M., Alton, G., Parsons, M. R., Convery, M. A., Blakeley, V., Corner, A. S., Palcic, M. M., Knowles, P. F., McPherson, M. J. & Phillips, S. E. (1997). Biochemistry, 36, 1608–1620. [DOI] [PubMed]
- Wilmot, C. M., Saysell, C. G., Blessington, A., Conn, D. A., Kurtis, C. R., McPherson, M. J., Knowles, P. F. & Phillips, S. E. (2004). FEBS Lett.576, 301–305. [DOI] [PubMed]
- Yu, P. H., Wright, S., Fan, E. H., Lun, Z. R. & Gubisne-Harberle, D. (2003). Biochim. Biophys. Acta, 1647, 193–199. [DOI] [PubMed]
Associated Data
This section collects any data citations, data availability statements, or supplementary materials included in this article.
Supplementary Materials
PDB reference: E. coli copper amine oxidase, xenon derivative, 2w0q, r2w0qsf