Abstract
Tropomodulin, a tropomyosin-binding protein, caps the slow-growing (pointed) end of the actin filament regulating its dynamics. Tropomodulin, therefore, is important for determining cell morphology, cell movement, and muscle contraction. For the first time we show that one tropomodulin molecule simultaneously binds two tropomyosin molecules in a cooperative manner. On the basis of the tropomodulin solution structure and predicted secondary structure, we introduced a series of point mutations in regions important for tropomyosin-binding and actin-capping. Capping activity of these mutants was assayed by measuring actin polymerization using pyrene fluorescence. Using direct methods (circular dichroism and native gel-electrophoresis) for detecting tropomodulin/tropomyosin binding, we localized the second tropomyosin-binding site to residues 109-144. Despite previous reports that the second binding site is for erythrocyte tropomyosin only, we found that both short non-muscle and long muscle α-tropomyosins bind there as well, though with different affinities. We propose a model for actin capping where one tropomodulin molecule can bind to two tropomyosin molecules at the pointed end.
Keywords: tropomodulin, tropomyosin, actin filament, pointed end
Actin is a major component of the cell cytoskeleton and the sarcomeres of striated muscle, and plays a critical role in cell morphology, cell movement, and muscle contraction (1, 2). It is known that actin filaments form vastly different structures in different cell types and in different locations within the cell. Regulation of the dynamics at actin’s ends is of central importance in the assembly of these various structures. An actin filament has two distinct ends: a fast-growing barbed end and a slow-growing pointed end (3). The exclusive properties of these ends are exploited by over 160 distinct actin binding proteins, including those that cap, sever, or cross-link filaments (4). Of particular interest is tropomodulin (Tmod), a tropomyosin-binding protein, which regulates actin filament dynamics by capping the pointed end.
Tropomyosin (TM) is a two-chained coiled coil that binds head-to-tail (N to C terminus) along the length of both sides of actin filaments, spanning 6-7 actin monomers per TM molecule, depending on the isoform (5). There are a variety of TM isoforms encoded by different genes. The N-terminus of TM, which points towards the pointed end of the actin filament (6), is required for interaction with Tmod (7-9). Best known for its role in muscle contraction (10, 11), TM also regulates the stiffness of actin (12) and influences polymerization at the pointed end (13, 14). Tmod in combination with TM forms a cap at the pointed end, preventing addition or loss of G-actin (15-17).
Tropomodulin, originally discovered as a TM-binding protein in erythrocyte membranes (18), is the only known pointed end specific capping protein (15). Presently, there are four known Tmod isoforms (16, 19, 20). These isoforms are 60% identical and 70% similar in amino acid sequence.
Of the four isoforms, the most extensive work has been on Tmod1, a 359 residue protein. It is found in erythrocytes, the heart, and slow skeletal muscle. The molecule is elongated, with distinct N-terminal and C-terminal halves (21, 22). The structure of the N-terminal half shows the domain to be mostly extended, unstructured, and flexible (23, 24). The X-ray crystal structure of the C-terminal half (residues 160-344) reveals the domain to be compactly folded (25). Unique among all actin-binding proteins, the C-terminal half of Tmod1 contains five tandem leucine-rich repeat motifs.
In addition to structural differences, the N and C-terminal halves are functionally distinct. A C-terminal fragment (residues 160-359) binds and caps actin by itself (26) but does not bind TM (21). N-terminal fragments, Tmod11-130 or Tmod11-92, bind TM and contain a TM-dependent actin-capping activity, but do not bind or cap actin on their own (14, 26). In Tmod11-92, the N-terminal fragment of Tmod1, mutating Leu27 to glycine in the first TM-binding region caused a loss of TM binding; changing Thr59, Gly60, Pro61 to alanines resulted in a decrease in capping activity by decreasing flexibility; and mutating Leu71 to aspartate caused a loss of actin-capping activity (24, 27). The design of these mutations was based on information from the NMR solution structure of Tmod11-92 (24) and primary structure analysis (27) (Fig. 1). In the first 92 residues (Tmod11-92), the TM-binding domain for both muscle and non-muscle α-tropomyosins was localized to res. 1-38 (24) and the TM-regulated actin-capping site was localized to res. 48-92 (14).
Figure 1.

The amino acid sequence of Tmod1 aligned to the secondary structure obtained from NMR data (residues 1-38), secondary structure prediction (residues 39-159), crystal atomic structure (residues 160-344). Bold - helical regions where mutations were done. Underlined - residues that are mutated. Boxed residues represent fragments used for experiments.
The present study addresses three unanswered questions about Tmod by studying TM/Tmod interactions. The first question asks: How many TM-binding sites does a Tmod molecule have and what is the specificity? An earlier study suggested that residues 6-94 of Tmod1 contain the binding site for skeletal muscle TM, whereas residues 90 to 184 interact with erythrocyte TM (28). However, residues 95-359 exhibited a 160-fold increase in capping activity in the presence of skeletal muscle TM, inferring the presence of a second binding site in this region (26). The next question asks: How many molecules of TM does one Tmod molecule bind? Finally, how many molecules of Tmod are required to cap the pointed end of the actin filament?
In this paper, we have shown that full length Tmod1 has two binding sites for both short non-muscle and long muscle α-tropomyosins. We localized the second binding site to res. 109-144. We evaluated the influence of point mutations in both TM-binding sites and a TM-dependent actin-capping site on Tmod1 function. We found that one molecule of Tmod1 binds two molecules of TM and that this binding is cooperative.
Materials and methods
Plasmid Construction
Site-directed mutagenesis was performed using a QuikChange site-directed mutagenesis kit (Stratagene, La Jolla, CA). The plasmids were amplified by PCR using PfuTurbo DNA polymerase and two complementary sets of oligonucleotides. Mutated triplets are underlined.
To change Leu27 to glycine, Leu71 to aspartate, and Thr59/Gly60/Pro61 to alanines, the same oligonucleotides as in (24, 27) were used.
To change Ile131 to an aspartate, the following oligonucleotides were used, 5′ -GCGGAGCTCTGTGACGATGCTGCCATCCTTGGC -3′ and 5′- GCCAAGGATGGCAGCATCGTCACAGAGCTCCGC -3′.
The plasmid for chicken Tmod1, pET(His)Tmod1, (21)was used as the template to obtain pET(His)Tmod1(L27G). pET(His)Tmod1(L27G) was then used as the template plasmid to construct pET(His)Tmod1(L27G/L71D). pET(His)Tmod1(L27G/L71D) was used as the template plasmid to construct pET(His)Tmod1(L27G/L71D/I131D).
To construct expression vectors of Tmod N-terminal fragments, res. 1-159, the plasmids pET(His)Tmod11-359 and pET(His)Tmod11-359 (L27G/L71D) were used as the templates.
To insert a stop codon instead of Gly160, the oligonucleotides used were 5′-GCACCATCGTGAACAAAGAATAACTTAACAGCGTGATTAAGCCC -3′ and 5′-GGG CTT AAT CAC GCT GTT AAG TTA TTC TTT GTT CAC GAT GGT GC -3′
The original plasmid was digested using DpnI, and the mixture was used to transform Escherichia coli (max efficiency DH5α). After plasmid purification, the presence of the mutations was confirmed by DNA sequencing. Synthesis of all oligonucleotides and sequence determination were done at the UMDNJ DNA Synthesis and Sequencing Facility (Robert Wood Johnson Medical School (RWJMS), Piscataway, NJ).
Protein Expression and Purification
Tmod mutants, Tmod1(L27G), Tmod1(L27G/L71D), Tmod1(T59A/G60A/P61A), and Tmod1(L27G/L71D/I131), were overexpressed in E.coli BL21(DE3)pLysE using the method described in (29) and purified according to (21). Tmod11-159 fragments were overexpressed in E.coli BL21(DE3)pLysS and additionally purified by reverse-phase high performance liquid chromatography according to (14). The yield of these peptides was low, their expression was poor and the purification was complicated due to the presence of a smaller peptide that was identified as a 1-146 residue fragment of Tmod1.
Molecular masses of Tmod1 mutants were checked with electrospray mass spectroscopy (Keck Biotechnology Resource Laboratory, Yale University, New Haven, CT). Tmod1109-144 was synthesized by the Tufts University Core Facility (Boston, MA).
Chicken pectoral skeletal muscle actin was purified from acetone powder as described (30). G-actin was purified on a Sephacryl S-300 column (31) and was stored in liquid nitrogen. Actin was labeled with pyrenyl-iodoacetamide and the labeling ratios were calculated according to (32, 33). The degree of the labeling was 80 - 99 %. Before experiments, G-actin (labeled or unlabeled) was defrosted in a 37°C water bath and then centrifuged at 100,000 rpm (TLA-100, Beckman) for 30 min at 4°C.
Protein purity was evaluated using SDS-PAGE (34). Concentrations of proteins were determined by using BCA protein assay kit (Pierce) or measuring their difference spectra in 6 M guanidine-HCl between pH 12.5 and 7.0 (35) using the extinction coefficients of 2357 per tyrosine and 830 per tryptophan (36).
TM peptides, AcTM1aZip and AcTM1bZip, are designed chimeric proteins that contain 14 residues of long rat α-tropomyosin encoded by exon 1a or 19 residues of short rat α-tropomyosin encoded by exon 1b correspondingly and the 18 C-terminal residues of the GCN4 leucine zipper domain (37); their structures were solved using NMR (38). These peptides and N-acetylated striated muscle α-tropomyosin, stTM, were a gift from Dr. Sarah Hitchcock-DeGregori (RWJMS, Piscataway, NJ). Recombinant human gelsolin was a generous gift from Dr. John H. Hartwig (Brigham and Women’s Hospital & Hematology Division, Boston, MA).
Binding Experiments
Binding was visualized using native polyacrylamide gel electrophoresis. 9 % polyacrylamide gels that were polymerized in the presence of 10% glycerol without SDS were used (24). To prepare complexes for loading onto gels, equimolar amounts of Tmod in the absence or presence of GlyTM1b1-19Zip were mixed in a 1:2 molar ratio. For titration, stock solutions of tropomodulin and GlyTM1b1-19Zip were mixed in different ratios in buffer containing 100 mM NaCl. Samples were analyzed using native polyacrylamide gels, stained with Coomassie R-250 and quantified using a Molecular Dynamics model 300A computing densitometer (Sunnyvale, CA).
Fluorescence measurements
The rates of actin polymerization were measured using the change in pyrene actin fluorescence (32) using a PTI fluorimeter (Lawrenceville, NJ) (excitation, 366 nm, and emission, 387 nm, with a 1 nm slit). To measure polymerization of actin at the pointed end, short filaments capped at the barbed ends with gelsolin were prepared by polymerization of 3 μM G-actin in the presence of 28 nM gelsolin according to (14). Polymerization was monitored by the increase in fluorescence when the filaments were diluted 5-fold with G-actin (10 % pyrenylactin) in F-buffer (100 mM KCl, 2 mM MgCl2, 1 mM EGTA, 0.5 mM DTT, 0.2 mM ATP, 0.2 mM CaCl2, 1 mM NaN3, 10 mM imidazole, pH 7.0) containing TM and Tmod1. The final concentrations of F- and G-actin after dilution were 0.6 μM and 1.1 μM respectively.
Gelsolin-capped filaments were prepared in sets of four and fluorescence measurements were carried out in parallel in a four cuvette holder with actin as a control in each set as described (14). Exponential curves were fit to the polymerization data using SigmaPlot, and initial rates (R) were calculated as the first derivatives at time zero.
Circular dichroism measurements
CD measurements were made using an Aviv model 215 spectropolarimeter (Lakewood, NJ) as previously described (38, 39). The protein concentrations were 10 μM in 100 mM NaCl, 10 mM sodium phosphate, pH 7. The dissociation constants of the Tmod1/TM peptide complexes, Kd, were determined using the relationship: Kd=exp(-ΔΔG/RT), where ΔΔG is the difference in free energy of folding of the complex minus that of the TM fragment alone, R is the gas constant and T is the temperature (40). The equations used for determining the dissociation constants and the thermodynamics of folding of the two-chain TM peptide and the three-chain TM/Tmod1 peptide complex from the changes in their circular dichroism as a function of temperature have been described in detail previously (9, 38).
Results
Influence of mutations in N-terminal TM-binding and actin-capping sites on function of full-length Tmod1
We introduced the same mutations previously used in Tmod fragment, Tmod11-92, into full-length Tmod1 to determine whether or not the mutations exhibit a similar effect on TM-binding and actin-capping activity.
Three full-length mutants were expressed: Tmod1(T59A/G60A/P61A), Tmod1(L27G) and Tmod1(L27G/L71D). Their capping ability was assayed in the presence of a striated α-TM, stTM. Pointed end polymerization was nucleated by short actin filaments with gelsolin blocking the barbed ends. All these mutations, including the L27G/L71D mutation, which previously led to loss of capping ability in Tmod11-92, had no significant influence on the capping ability of full-length Tmod (Fig. 2).
Figure 2.
Pointed end elongation of gelsolin-capped actin filaments (5.6 nM) in the presence and absence of tropomodulin and stTM. 1.1 μM (10 % pyrene-labeled) G-actin mixed with 0.6 μM F-actin in F-buffer (100 mM KCl, 2 mM MgCl2, 1 mM EGTA, 0.5 mM DTT, 0.2 mM ATP, 0.2 mM CaCl2, 1 mM NaN3, 10 mM imidazole, pH 7.0). (●) control, no tropomyosin or tropomodulin; (○) with 0.75 μM stTM; (▼) with 0.75 μM stTM and 20 nM Tmod1; (▽) with 0.75 μM stTM and 20 nM Tmod1(T59A/G60A/P61A); (■) with 0.75 μM stTM and 20 nM Tmod1(L27G); (□) with 0.75 μM stTM and 20 nM Tmod1(L27G/L71D).
TM-binding properties of Tmod1 and mutants were studied using AcTM1bzip, a designed chimeric peptide that contains the N-terminus of short non-muscle α-tropomyosin encoded by exon 1b, and AcTM1azip, which contains the N-terminus of long skeletal muscle α-tropomyosin encoded by exon 1a (37, 38). AcTM1bzip and AcTM1azipboth form a coiled coil, melt as one cooperative unit, and have the major properties of full-length TM’s N-terminus. They bind tropomodulin (9, 14, 24, 26), bind C-terminal TM fragments, and form a ternary complex with troponin (41).
Binding of Tmods to AcTM1bZip was directly analyzed using native polyacrylamide gel electrophoresis (Fig. 3). An additional band corresponding to a complex between each Tmod and AcTM1bZip appeared when they were combined. Tmod1 mutants with the L27G mutation retain binding to the AcTM1bZip fragment as indicated by the presence of a Tmod/TM complex, implying binding of the AcTM1bZip peptide at a second TM-binding site. To our surprise, there was a noticeable difference in complex band position. The complex for Tmod1(L27G) or Tmod1(L27G/L71D) runs faster than that of wild-type Tmod1 and Tmod1(T59A/G60A/P61A).
Figure 3.

Complex formation between AcTM1bZip and Tmod1 mutants monitored by non-denaturing polyacrylamide gel-electrophoresis. AcTM1bZip is positively charged and does not enter the gel. Lane 1 and lane 3, Tmod1; lane 2 and lane 4, Tmod1 and AcTM1bZip; lane 5, Tmod1(L27G); lane 6, Tmod1(L27G) and AcTM1bZip; lane 7, Tmod1(L27G/ L71D); lane 8, Tmod1(L27G/ L71D) and AcTM1bZip; lane 9, Tmod1(L27G/ L71D/I131D); lane 10, Tmod1(L27G/ L71D/I131D) and AcTM1bZip; lane 11, Tmod1(T59A/G60A/P61A); lane 12, Tmod1(T59A/G60A/P61A) and AcTM1bZip. TM:Tmod molar ratio in lane 4 is 1:1, in all other lanes with AcTM1bZip it is 2:1. Arrows indicate TM/Tmod1 complexes, arrowhead - Tmod1.
We were unable to detect on native gels binding to another TM peptide, AcTM1aZip. Alternatively, another way to detect complex formation is by measuring the change in ellipticity during melting of a Tmod1 and TM peptide mixture. Unfortunately, the heat denaturation of Tmod1 is irreversible; Tmod1 begins aggregating at ∼64°C (23). Mixing Tmod1 with TM peptides does not prevent aggregation, but it does change the melting curves, indicating that there is binding of Tmod1 to the TM peptides (results not shown). However, it is impossible to analyze the unfolding curves because of the aggregation. Since aggregation is associated with the C-terminal LRR domain (res.160-359) (23), we decided to express fragments long enough to contain the second binding site, but which do not aggregate when melting. The C-terminal domain does not bind TM (21, 25) or contain TM-dependent actin-capping activity (26); therefore we expressed two fragments, Tmod11-159 and Tmod11-159(L27G/L71D), that do not aggregate during thermal denaturation.
TM-binding and actin-capping abilities of Tmod1 wild-type and mutated N-terminal fragments, res. 1-159
Binding of Tmod11-159 and Tmod11-159(L27G/L71D) with AcTM1bZip and AcTM1aZip was analyzed using circular dichroism(CD) spectroscopy (Fig. 4). The CD measurements at 222 nm indicated that there is no cooperativity in the melting of Tmod11-159, consistent with the absence of definite tertiary structure in this region (21-23). Unlike full-length Tmod1, thermal denaturation of complexes of Tmod11-159 fragments and TM peptides was reversible and highly cooperative. There was an increase in both the melting temperature, Tm, and helical content in the mixture relative to the sum. This indicates that there was binding of Tmod1 fragments to both α-TM peptides and that additional helices formed during the binding.
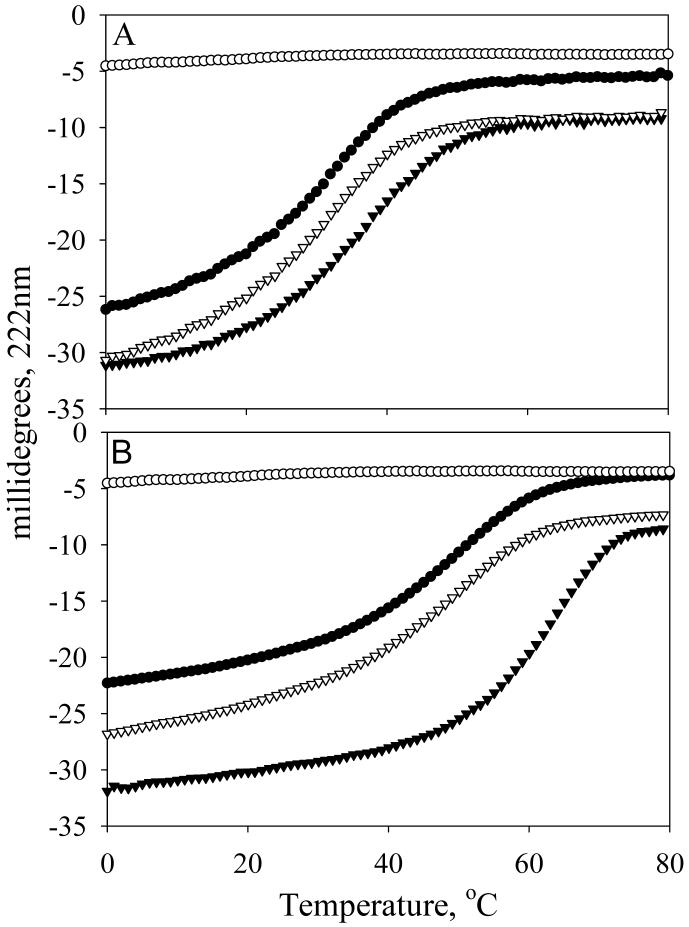
Figure 4.
Binding of Tmod11-159 mutants to a TM model peptides measured using circular dichroism spectroscopy. The temperature dependence of the ellipticity at 222 nm was measured for unmixed and mixed AcTM1bZip and AcTM1aZip. A: Tmod11-159WT and AcTM1aZip; and B: Tmod11-159WT and AcTM1bZip; C: Tmod11-159(L27G/L71D) and AcTM1aZip; D: Tmod11-159(L27G/L71D) and AcTM1bZip. (○) Tmod11-159 alone; (●) TM peptide; (▽) sum of the folding curves of the TM peptide and Tmod11-159 alone; (▼) the folding of the mixture of the TM peptide and Tmod11-159.
In the absence of TM, Tmod11-159 has no effect on actin polymerization at concentrations used for our experiments (5-175 nM). To compare the inhibitory activities of wild-type and mutated Tmod11-159, the dependence of the initial rate of polymerization on Tmod11-159 concentration was measured (Fig. 5). The TM-dependent capping ability of wild-type Tmod11-159 is comparable to full-length Tmod1 activity and higher than Tmod11-92 activity(14). The inhibitory activity of the Tmod11-159(L27G/L71D) reaches 50 % inhibition effect at a concentration of ∼30 nM while wild-type Tmod11-159 reaches 50 % inhibition effect at 3.5 nM. However, increasing the concentration further does not change the inhibition effect for this mutant.
Figure 5.

Dependence of inhibition of actin polymerization on Tmod11-159 concentration in the presence of 0.75 μM stTM. Initial rates (R) were calculated as the first derivatives at time zero after fitting. The inhibition of polymerization was calculated as Rexp / Rcontrol, where Rcontrol=1 (in the absence of TM and Tmod1 fragments). (●) Tmod11-159WT; (○) Tmod11-159(L27G/L71D). (■) The value shown for stTM in the absence of tropomodulin fragments (n=3).
Localization of the second TM-binding site
Our data indicates that there is a second binding site for muscle and non-muscle α-TMs within residues 93-159. Secondary structure analysis of the amino acid sequence of Tmod1 predicts the presence of several additional helices in this region (Fig. 1). Our hypothesis is that these helices might be important in forming the second TM-binding site. The longest predicted helix, residues 126-135, is mostly composed of hydrophobic amino acids; all of these residues are conserved except Cys139. Our previous data (24, 27) suggest that hydrophobic interactions are important for Tmod binding at the pointed end. The hydrophobic side of this helix is formed by Leu128, Ile131, Ala132, Ile134 and Leu135. We mutated a conserved Ile131 in the middle of the helix to Asp, destroying the hydrophobic surface. Tmod1(L27G/L71D), the mutant lacking the first TM-binding and actin capping sites, was used as a template. Tmod1(L27G/L71D/I131D) lost the ability to bind AcTM1bZip in a native gel (Fig. 3, lane 10). There was no change in melting temperature or helical content when this mutant was mixed with AcTM1aZip or AcTM1bZip. Therefore, Ile131 localizes the second TM-binding site. Studies of actin-capping ability for this mutant showed at least a 30-fold decrease in inhibitory activity (Fig. 6). Tmod1(L27G/L71D/I131) reaches a 50 % inhibition effect at a concentration of ∼60 nM, while wild-type Tmod1 reaches a 50 % inhibition effect at ∼2 nM. Increasing concentrations of Tmod1(L27G/L71D/I131D) up to 500 nM in the presence of TM increased inhibition of actin polymerization. However, it is difficult to distinguish what part of this inhibition is TM-dependent because at high concentrations Tmod1(L27G/L71D/I131D) itself inhibits polymerization even in the absence of TM (Fig. 6).
Figure 6.
A: Pointed end elongation of gelsolin-capped actin filaments (5.6 nM) in the presence and absence of Tmod1(L27G/L71D/I131D) and 0.75 μM stTM. 1.1 μM (10 % pyrene-labeled) G-actin mixed with 0.6 μM F-actin in F-buffer (100 mM KCl, 2 mM MgCl2, 1 mM EGTA, 0.5 mM DTT, 0.2 mM ATP, 0.2 mM CaCl2, 1 mM NaN3, 10 mM imidazole, pH 7.0). (●) control, no tropomyosin or tropomodulin; (○) with 0.75 μM stTM and 50 nM Tmod1(L27G/L71D/I131D); (▼) with 0.75 μM stTM and 200 nM Tmod1 (L27G/L71D/I131D); (▽) with 0.75 μM stTM and 500 nM Tmod1(L27G/L71D/I131D); (■) with 500 nM Tmod1(L27G/L71D/I131D). B: Dependence of inhibition of actin polymerization on Tmod1 concentration. Initial rates (R) were calculated as the first derivatives at time zero after fitting. The inhibition of polymerization was calculated as Rexp / Rcontrol, where Rcontrol=1 (in the absence of TM and Tmod1 fragments). Tmod1WT with (●) and without (◆) 0.75 μM stTM; Tmod1(L27G/L71D) in presence of 0.75 μM stTM (○); Tmod1(L27G/L71D/I131D) with (▼) and without (◇) 0.75 μM stTM. (■) The value shown for stTM in the absence of tropomodulin (n=3).
As a contamination to the Tmod11-159 fragments, a peptide of smaller MW was co-purified. According to mass spectroscopy results this peptide corresponds to a 1-146 residue fragment of Tmod1. The Tmod11-146 fragments, both wild type and mutated, were still able to bind the TM peptide on a native gel showing the same difference in mobility of complexes as Tmod1 and Tmod11-159 fragments (data not shown). Therefore, residues 147-159 are not required for formation of the second TM-binding site. Based on sequence conservation and the probability to form secondary structure, we chose residues 109-144 for a synthetic peptide containing the second binding site. Binding of Tmod1109-144 to both TM peptides was measured using CD (Fig. 7). The dissociation constants of Tmod1109-144, estimated from the changes in ellipticity, were 2.5±1.1 nM for AcTM1bZip and 1.3±0.3 μM for AcTM1aZip. These data show that the second TM-binding site, earlier suggested to be for erythrocyte TM only (28), also binds other TM isoforms.
Figure 7.
Binding of Tmod1109-144 to TM model peptides measured using circular dichroism spectroscopy. The temperature dependence of the ellipticity at 222 nm was measured for unmixed and mixed AcTM1bZip and AcTM1aZip. A: Tmod1109-144 and AcTM1aZip; and B: Tmod1109-144 and AcTM1bZip. (○) Tmod1109-144 alone; (●) TM peptide; (▽) sum of the folding curves of the TM peptide and Tmod11-159 alone; (▼) the folding of the mixture of the TM peptide and Tmod1109-144.
One Tmod molecule binds two TM molecules
The question now is how many TM molecules does one Tmod molecule bind? There are several possibilities: one Tmod could bind two different TM molecules at the same time; both sites could bind the same TM molecule; the sites could be mutually exclusive so that TM binding to one site makes binding to the other site impossible. In the first case, one Tmod molecule would bind two TM molecules while in the other two cases a Tmod molecule would bind only one TM molecule.
On a native gel, the distance between the Tmod1 band and the TM/Tmod1 complex depends on the molecular weight (∼8.6 kDa) and positive charge (pI=10) of the TM peptide, which binds to Tmod. Complexes of the TM peptide with Tmod1 and its mutants migrate differently (Fig. 3). The distance between the Tmod1 band and complexes of the TM peptide with either wild-type Tmod1 or Tmod1((T59A/G60A/P61A) is twice the distance seen between the Tmod1 band and complexes of the TM peptide with Tmod1(L27G) or Tmod1(L27G/L71D), mutants with the first binding site destroyed. This drastic difference in mobility may be explained by one Tmod1 molecule binding two molecules of TM, while Tmod1 with the L27G mutation binds only one TM. However, mobility of proteins on native gels depends not only on molecular weight and charge but also on shape of a protein or protein complex. Since there is a probability that change in the protein shape may have caused the difference in mobility, we used titration to determine the stoichiometry.
Full-length Tmod1 was titrated with AcTM1bZip. The decrease of free Tmod in the mixtures was monitored by scanning and quantifying the Tmod band as well as the complex band in native polyacrylamide gels (Fig. 8A). In Fig 8B, the same samples are shown on a 15 % SDS gel. Standard deviation of the loaded Tmod1 is no more than 4 %.
Figure 8.
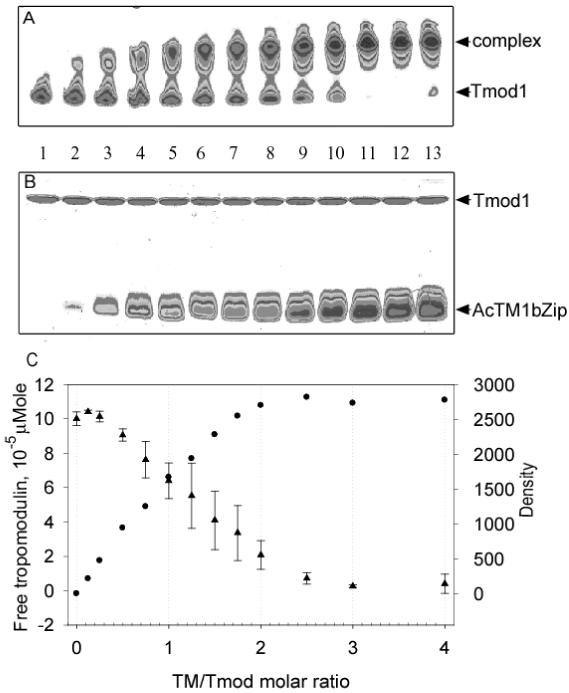
Titration of Tmod1 by AcTM1bZip. Stock solution of Tmod1 was diluted to a final concentration of 10 μM with different concentrations of TM peptide. The decrease of free tropomodulin and the increase of the complex were monitored by scanning and quantifying the Tmod1 band in native polyacrylamide gels. A: 9 % native gel and B: 15 % SDS gel scanned on densitometer; C: dependence of amount of free Tmod1 (●) and density of the complex band (▲) on amount of AcTM1bZip added (TM/Tmod molar ratio). Lanes on the gels correspond to points on the graph. Lanes 1-13 contain AcTM1bzip/Tmod1 in ratios 0, 0.125, 0.25, 0.5, 0.75, 1.0, 1.25, 1.5, 1.75, 2.0, 2.5, 3.0, 4.0. Error bars show standard deviation (n=4).
Quantification of complex bands was impeded because of the smear. According to the CD measurements, Tmod/TM binding is very tight. Thus, this smear is not a result of exchange between free and bound molecules, but is the result of partial dissociation of the complex due to heating during gel-electrophoresis. The density of the complex cannot be converted to moles and therefore cannot be averaged. The amount of unbound Tmod1 and the density of the complex are shown in Figure 8C as a function of the TM/Tmod ratio. At 1:1 ratio 63±10 % of Tmod is still free. Both curves show saturation between a ratio of 2:1 and 2.5:1. Titration of the Tmod1 fragment, containing only the first binding site, was done by Greenfield and Fowler and shows a 1:1 stoichiometry (9). Our data demonstrate that one mole of full-length Tmod binds not one but two moles of TM peptide.
Discussion
We localized a second binding site for both short and long TM isoforms on Tmod1 within res. 109-144. This region contains a predicted helix, res. 126-135. The calculated dissociation constant of Tmod1109-144 for AcTM1bZip is 2.5±1.1 nM. This is two orders of magnitude tighter than the calculated dissociation constant of Tmod11-92 (0.22±0.10 μM) (14) and Tmod11-130 (0.23±0.15 μM) (9). The dissociation constant of Tmod1109-144 for AcTM1aZip is 1.3±0.3 μM, the same as the dissociation constant of Tmod11-130 (1.1±0.4 μM) (9). However, the difference in inhibitory activity of Tmod/TM on actin capping between short and long α-TMs is about 5-fold (14). It is possible that kinetic properties of the TM-binding sites, e.g. higher on-rate binding of long TM to this site or higher cooperativity of this binding, may minimize the difference in dissociation constants.
Figure 9A shows the location of the TM-binding and actin-capping sites identified in the present and previous studies. Our binding studies show that one Tmod1 molecule binds two TM molecules in a cooperative manner and lead to a new model for Tmod1 and TM binding at the pointed end of the actin filament (Fig. 9B).
Figure 9.

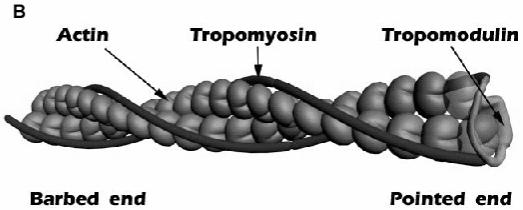
A: schematic model of the TM-binding and actin-capping sites. B: a cartoon representation of the pointed end. The N-terminal half of Tmod1 binds to both TMs and interacts with actin. Two actin molecules at the pointed end are shown as transparent.
The L27G mutation destroys α-helix formation in the helical region, res. 24-35, located in the first TM-binding site (24). This mutation causes a loss of TM-binding ability in Tmod11-92. However, Tmod11-159 and full-length Tmod1 with this mutation are still able to bind TM, because the second TM-binding site is intact. All these mutants inhibit actin elongation at the pointed end with effectiveness close to wild-type.
Mutation L71D inhibits formation of the hydrophobic surface in the amphipathic helix, res. 65-75, which we showed to be important for TM-dependent actin capping. Together, mutations L27G and L71D led to the loss of capping activity in Tmod11-92 (27). However, the effect of these mutations became less significant with the increasing size of the Tmod1 fragment. Tmod1-159(L27G/L71D) was still able to inhibit actin polymerization although ∼ten-fold less effectively than wild-type Tmod1-159. These mutations had no effect on the capping activity in full-length Tmod1. Therefore, destruction of the first actin-capping domain by the L71D mutation is inconsequential for full-length Tmod, which contains an intact C-terminal capping domain. Interestingly, Tmod1-159(L27G/L71D) never completely inhibits actin polymerization and is only ∼ 50 % effective compared to the wild-type fragment. Two of three sites crucial for TM-binding and actin-capping are destroyed in this fragment, so it cannot be well fixed at the pointed end. The fifty percent effectiveness may reflect the probability of the Tmod1-159(L27G/L71D) polypeptide chain to be found in a position where it prevents the next actin molecule from binding to the pointed end.
The third mutation, I131D, postulated to destroy the hydrophobic surface in a putative helix, res. 126-135, caused a loss of TM-binding in full-length Tmod1 and a 30-fold decrease of capping ability when L27G/L71D mutations were also present. This result confirms the location of the second TM-binding site within residues 109-144 and demonstrates that correct folding of the helix is crucial for the formation of this site. Our evidence that residues 109-144 contain a second TM-binding site is consistent with results from other studies. Original evidence of a second TM binding site was shown in the ability of 125I-labeled erythrocyte TM (a heterodimer of α and γ short TM) to bind to Tmod195-359 immobilized on nitrocellulose (28). Since we began our work, Vera et al. (29) mapped a binding site for TM5 (a short erythrocyte TM encoded by the γ-TM gene) to residues 105-127 and Kong et al (42) found that residues L134 and L135 are crucial for TM5 binding. However, all three studies used qualitative assays to measure interaction, methods that were unable to quantify the affinity or stoichiometry of the complex. Tmod binding to long α-TM is weaker than to short TMs and therefore difficult to detect with the methods used. Our assays appear to be more sensitive and allow detection of binding to both muscle and non-muscle α-TM isoforms (stTM and TM5a).
It was shown that a Tmod1 fragment consisting of residues 1-130 binds only one TM peptide (9). According to our data, the second binding site cannot be formed in this fragment because half of the putative helix, res. 126-135, is missing. Our data support the conclusion that full-length Tmod1 contains both TM-binding sites and binds two TM molecules. Several experimental results confirm this conclusion. In native gel-electrophoresis, the complex of wild-type Tmod1 and a TM peptide has lower mobility compared to Tmod1 with a mutated first TM-binding site. This difference in mobility is due to the binding of an additional TM peptide. The titration of full-length Tmod1 with AcTM1bZip showed that Tmod molecule binds two TM peptides.
At the lowest ratios on the native gel (Fig. 8A), the position of the complex band is intermediate between the position of the major density of the complex at higher ratios and the Tmod1 band alone. It corresponds to the position of a Tmod1 complex band with the L27G mutation (Fig. 3) and therefore indicates binding one TM peptide. When the ratio is 1/1, about half of wild-type Tmod1 is unbound, demonstrating that once Tmod1 binds one molecule of TM, its affinity for a second molecule is increased. Binding of AcTM1bZip to the second site is tighter than to the first site, however, we do not see an intermediate band corresponding to a complex with one TM fragment bound. Therefore, there is cooperativity in the binding of two moles of TM to one mole of Tmod1. The two-state transition in the melting curves for complexes of Tmod11-159 and TM peptides confirms this conclusion.
The presence of one of two TM-binding sites and one of two actin-capping sites is sufficient for successful functioning of Tmod1. The duplication of binding sites makes Tmod function more resistant to mutations. The mutation T59A/G60A/P61A in the flexible region of Tmod1, which caused a 10-fold decrease of capping ability in Tmod11-92, had no effect in full-length Tmod1. When the Tmod1 molecule is bound at both binding sites to two different TM molecules at the pointed end, a change in flexibility of the region between these binding sites is less important for capping than in a molecule with one binding site. Our data suggest that at the pointed end, one Tmod molecule binds both TM molecules. Therefore, one molecule of Tmod is necessary and sufficient for capping at the pointed end (Fig. 9). This completely changes the previous concept of pointed end organization, whereby one molecule of Tmod binds to one molecule of TM (4). This model should be subject to further investigations.
Acknowledgements
We thank Dr. Sarah E. Hitchcock-DeGregori for her gift of tropomyosins and discussion, Dr. Norma J. Greenfield for discussing CD data and Abhishek Singh for discussion. We thank the UMDNJ Neuroscience Summer Undergraduate Research Program, the Aresty Research Center for Undergraduates, and the Henry Rutgers Scholars Program that allowed Andy Choy to work in the laboratory.
Supported by the American Heart Association grant 0535328N to Dr. Alla S. Kostyukova and in part by National Institutes of Health Grant GM63257 to Dr. Sarah E. Hitchcock-DeGregori.
Abbreviations
- Tmod
tropomodulin
- TM
tropomyosin
- CD
circular dichroism
References
- 1.Pollard TD, Borisy GG. Cellular motility driven by assembly and disassembly of actin filaments. Cell. 2003;112:453–65. doi: 10.1016/s0092-8674(03)00120-x. [DOI] [PubMed] [Google Scholar]
- 2.Rafelski SM, Theriot JA. Crawling toward a unified model of cell mobility: spatial and temporal regulation of actin dynamics. Annu Rev Biochem. 2004;73:209–39. doi: 10.1146/annurev.biochem.73.011303.073844. [DOI] [PubMed] [Google Scholar]
- 3.Carlier MF, Pantaloni D. Control of actin dynamics in cell motility. J Mol Biol. 1997;269:459–67. doi: 10.1006/jmbi.1997.1062. [DOI] [PubMed] [Google Scholar]
- 4.dos Remedios CG, Chhabra D, Kekic M, Dedova IV, Tsubakihara M, Berry DA, Nosworthy NJ. Actin binding proteins: regulation of cytoskeletal microfilaments. Physiol Rev. 2003;83:433–73. doi: 10.1152/physrev.00026.2002. [DOI] [PubMed] [Google Scholar]
- 5.Perry SV. Vertebrate tropomyosin: distribution, properties and function. J Muscle Res Cell Motil. 2001;22:5–49. doi: 10.1023/a:1010303732441. [DOI] [PubMed] [Google Scholar]
- 6.Ohtsuki I, Nagano K. Molecular arrangement of troponin-tropomyosin in the thin filament. Adv Biophys. 1982;15:93–130. doi: 10.1016/0065-227x(82)90006-5. [DOI] [PubMed] [Google Scholar]
- 7.Sung LA, Lin JJ. Erythrocyte tropomodulin binds to the N-terminus of hTM5, a tropomyosin isoform encoded by the gamma-tropomyosin gene. Biochem Biophys Res Commun. 1994;201:627–34. doi: 10.1006/bbrc.1994.1747. [DOI] [PubMed] [Google Scholar]
- 8.Vera C, Sood A, Gao KM, Yee LJ, Lin JJ, Sung LA. Tropomodulin-binding site mapped to residues 7-14 at the N-terminal heptad repeats of tropomyosin isoform 5. Arch Biochem Biophys. 2000;378:16–24. doi: 10.1006/abbi.2000.1802. [DOI] [PubMed] [Google Scholar]
- 9.Greenfield NJ, Fowler VM. Tropomyosin requires an intact N-terminal coiled coil to interact with tropomodulin. Biophys J. 2002;82:2580–91. doi: 10.1016/S0006-3495(02)75600-2. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 10.Geeves MA, Lehrer SS. Cooperativity in the Ca2+ regulation of muscle contraction. Results Probl Cell Differ. 2002;36:111–32. doi: 10.1007/978-3-540-46558-4_10. [DOI] [PubMed] [Google Scholar]
- 11.Lehrer SS. An overview of actin-based calcium regulation. Results Probl Cell Differ. 2002;36:107–9. doi: 10.1007/978-3-540-46558-4_9. [DOI] [PubMed] [Google Scholar]
- 12.Fujime S, Ishiwata S. Dynamic study of F-actin by quasielastic scattering of laser light. J Mol Biol. 1971;62:251–65. doi: 10.1016/0022-2836(71)90144-6. [DOI] [PubMed] [Google Scholar]
- 13.Broschat KO, Weber A, Burgess DR. Tropomyosin stabilizes the pointed end of actin filaments by slowing depolymerization. Biochemistry. 1989;28:8501–6. doi: 10.1021/bi00447a035. [DOI] [PubMed] [Google Scholar]
- 14.Kostyukova AS, Hitchcock-DeGregori SE. Effect of the structure of the N terminus of tropomyosin on tropomodulin function. J Biol Chem. 2004;279:5066–71. doi: 10.1074/jbc.M311186200. [DOI] [PubMed] [Google Scholar]
- 15.Weber A, Pennise CR, Babcock GG, Fowler VM. Tropomodulin caps the pointed ends of actin filaments. J Cell Biol. 1994;127:1627–35. doi: 10.1083/jcb.127.6.1627. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 16.Almenar-Queralt A, Lee A, Conley CA, de Pouplana LR, Fowler VM. Identification of a novel tropomodulin isoform, skeletal tropomodulin, that caps actin filament pointed ends in fast skeletal muscle. J Biol Chem. 1999;274:28466–75. doi: 10.1074/jbc.274.40.28466. [DOI] [PubMed] [Google Scholar]
- 17.Weber A, Pennise CR, Fowler VM. Tropomodulin increases the critical concentration of barbed end-capped actin filaments by converting ADP.P(i)-actin to ADP-actin at all pointed filament ends [In Process Citation] J Biol Chem. 1999;274:34637–45. doi: 10.1074/jbc.274.49.34637. [DOI] [PubMed] [Google Scholar]
- 18.Fowler VM. Identification and purification of a novel Mr 43,000 tropomyosin-binding protein from human erythrocyte membranes. J Biol Chem. 1987;262:12792–800. [PubMed] [Google Scholar]
- 19.Cox PR, Zoghbi HY. Sequencing, expression analysis, and mapping of three unique human tropomodulin genes and their mouse orthologs. Genomics. 2000;63:97–107. doi: 10.1006/geno.1999.6061. [DOI] [PubMed] [Google Scholar]
- 20.Conley CA, Fritz-Six KL, Almenar-Queralt A, Fowler VM. Leiomodins: larger members of the tropomodulin (tmod) gene family. Genomics. 2001;73:127–39. doi: 10.1006/geno.2000.6501. [DOI] [PubMed] [Google Scholar]
- 21.Kostyukova A, Yamauchi E, Maeda K, Krieger I, Maeda Y. Domain structure of tropomodulin: distinct properties of the N-terminal and C-terminal halves. Eur J Biochem. 2000;267:6470–5. doi: 10.1046/j.1432-1327.2000.01738.x. [DOI] [PubMed] [Google Scholar]
- 22.Fujisawa T, Kostyukova A, Maeda Y. The shapes and sizes of two domains of tropomodulin, the P-end-capping protein of actin-tropomyosin. FEBS Lett. 2001;498:67–71. doi: 10.1016/s0014-5793(01)02498-x. [DOI] [PubMed] [Google Scholar]
- 23.Kostyukova AS, Tiktopulo EI, Maeda Y. Folding properties of functional domains of tropomodulin. Biophys J. 2001;81:345–51. doi: 10.1016/S0006-3495(01)75704-9. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 24.Greenfield NJ, Kostyukova AS, Hitchcock-Degregori SE. Structure and tropomyosin binding properties of the N-terminal capping domain of tropomodulin 1. Biophys J. 2005;88:372–83. doi: 10.1529/biophysj.104.051128. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 25.Krieger I, Kostyukova A, Yamashita A, Nitanai Y, Maeda Y. Crystal structure of tropomodulin C-terminal half and structural basis of actin filament pointed-end capping. Biophysical J. 2002;83:2716–2725. doi: 10.1016/S0006-3495(02)75281-8. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 26.Fowler VM, Greenfield NJ, Moyer J. Tropomodulin contains two actin filament pointed end-capping domains. J Biol Chem. 2003;278:40000–9. doi: 10.1074/jbc.M306895200. [DOI] [PubMed] [Google Scholar]
- 27.Kostyukova A, Rapp B, Choy A, Greenfield NJ, Hitchcock-DeGregori SE. Structural requirements of tropomodulin for tropomyosin binding and actin filament capping. Biochemistry. 2005;44:4905–4910. doi: 10.1021/bi047468p. [DOI] [PubMed] [Google Scholar]
- 28.Babcock GG, Fowler VM. Isoform-specific interaction of tropomodulin with skeletal muscle and erythrocyte tropomyosins. J Biol Chem. 1994;269:27510–8. [PubMed] [Google Scholar]
- 29.Studier FW. Protein production by auto-induction in high density shaking cultures. Protein Expr Purif. 2005;41:207–34. doi: 10.1016/j.pep.2005.01.016. [DOI] [PubMed] [Google Scholar]
- 30.Spudich JA, Watt S. The regulation of rabbit skeletal muscle contraction. I. Biochemical studies of the interaction of the tropomyosin-troponin complex with actin and the proteolytic fragments of myosin. J Biol Chem. 1971;246:4866–71. [PubMed] [Google Scholar]
- 31.MacLean-Fletcher S, Pollard TD. Identification of a factor in conventional muscle actin preparations which inhibits actin filament self-association. Biochem Biophys Res Commun. 1980;96:18–27. doi: 10.1016/0006-291x(80)91175-4. [DOI] [PubMed] [Google Scholar]
- 32.Kouyama T, Mihashi K. Fluorimetry study of N-(1-pyrenyl)iodoacetamide-labelled F-actin. Local structural change of actin protomer both on polymerization and on binding of heavy meromyosin. Eur J Biochem. 1981;114:33–8. [PubMed] [Google Scholar]
- 33.Cooper JA, Walker SB, Pollard TD. Pyrene actin: documentation of the validity of a sensitive assay for actin polymerization. J Muscle Res Cell Motil. 1983;4:253–62. doi: 10.1007/BF00712034. [DOI] [PubMed] [Google Scholar]
- 34.Laemmli UK. Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature. 1970;227:680–5. doi: 10.1038/227680a0. [DOI] [PubMed] [Google Scholar]
- 35.Edelhoch H. Spectroscopic determination of tryptophan and tyrosine in proteins. Biochemistry. 1967;6:1948–54. doi: 10.1021/bi00859a010. [DOI] [PubMed] [Google Scholar]
- 36.Fasman GD. Practical handbook of biochemistry and molecular biology. CRC Press; Boca Raton, Florida: 1989. [Google Scholar]
- 37.Greenfield NJ, Huang YJ, Palm T, Swapna GV, Monleon D, Montelione GT, Hitchcock-DeGregori SE. Solution NMR structure and folding dynamics of the N terminus of a rat non-muscle alpha-tropomyosin in an engineered chimeric protein. J Mol Biol. 2001;312:833–47. doi: 10.1006/jmbi.2001.4982. [DOI] [PubMed] [Google Scholar]
- 38.Greenfield NJ, Montelione GT, Farid RS, Hitchcock-DeGregori SE. The structure of the N-terminus of striated muscle alpha-tropomyosin in a chimeric peptide: nuclear magnetic resonance structure and circular dichroism studies. Biochemistry. 1998;37:7834–43. doi: 10.1021/bi973167m. [DOI] [PubMed] [Google Scholar]
- 39.Greenfield NJ, Hitchcock-DeGregori SE. The stability of tropomyosin, a two-stranded coiled-coil protein, is primarily a function of the hydrophobicity of residues at the helix-helix interface. Biochemistry. 1995;34:16797–805. doi: 10.1021/bi00051a030. [DOI] [PubMed] [Google Scholar]
- 40.Greenfield NJ. Circular dichroism analysis for protein-protein interactions. Methods Mol Biol. 2004;261:55–78. doi: 10.1385/1-59259-762-9:055. [DOI] [PubMed] [Google Scholar]
- 41.Palm T, Graboski S, Hitchcock-DeGregori SE, Greenfield NJ. Disease-causing mutations in cardiac troponin T: identification of a critical tropomyosin-binding region. Biophys J. 2001;81:2827–37. doi: 10.1016/S0006-3495(01)75924-3. [DOI] [PMC free article] [PubMed] [Google Scholar]
- 42.Kong KY, Kedes L. Leucine-135 of tropomodulin-1 regulates its association with tropomyosin, its cellular localization and the integrity of sarcomeres. J Biol Chem. 2006;23:23. doi: 10.1074/jbc.M512064200. [DOI] [PubMed] [Google Scholar]