Abstract
Unraveling the effect of selection vs. drift on the evolution of quantitative traits is commonly achieved by one of two methods. Either one contrasts population differentiation estimates for genetic markers and quantitative traits (the Qst–Fst contrast) or multivariate methods are used to study the covariance between sets of traits. In particular, many studies have focused on the genetic variance–covariance matrix (the G matrix). However, both drift and selection can cause changes in G. To understand their joint effects, we recently combined the two methods into a single test (accompanying article by Martin et al.), which we apply here to a network of 16 natural populations of the freshwater snail Galba truncatula. Using this new neutrality test, extended to hierarchical population structures, we studied the multivariate equivalent of the Qst–Fst contrast for several life-history traits of G. truncatula. We found strong evidence of selection acting on multivariate phenotypes. Selection was homogeneous among populations within each habitat and heterogeneous between habitats. We found that the G matrices were relatively stable within each habitat, with proportionality between the among-populations (D) and the within-populations (G) covariance matrices. The effect of habitat heterogeneity is to break this proportionality because of selection for habitat-dependent optima. Individual-based simulations mimicking our empirical system confirmed that these patterns are expected under the selective regime inferred. We show that homogenizing selection can mimic some effect of drift on the G matrix (G and D almost proportional), but that incorporating information from molecular markers (multivariate Qst–Fst) allows disentangling the two effects.
IN spatially heterogeneous environments, diversifying selection and restricted gene flow between populations may lead to locally adapted populations (Lenormand 2002; Hedrick 2007). Since Levene (1953) and Dempster (1955), a large body of theoretical work showed how spatially heterogeneous environments, demography, and selection interact to determine local adaptation. Until now, empirical demonstration of local adaptation, i.e., a higher fitness of individuals in their native habitat, relative to alternative habitats, has relied on comparisons of phenotypes across populations, with or without additional genetic data from neutral markers. Phenotypic studies consist of either reciprocal transplants (cross-relocating individuals originating from different habitats) or common garden experiments, where a single environmental factor is tested for its selective effect among populations (reviewed in Kawecki and Ebert 2004). With the emergence of molecular quantitative genetics, new methods have arisen, allowing a better access to the genetic basis of local adaptation. This can be done either by pinpointing genomic regions that are responsible for local adaptation (QTL mapping, Lynch and Walsh 1998) or by comparing the genetic variance, within and among populations, for a set of neutral markers and for the traits under scrutiny (Qst–Fst comparisons, Spitze 1993). This last method provides a way to test for selection and local adaptation by comparing the distribution of neutral molecular variance among populations (Fst) with the same quantity for quantitative traits (Qst). Under neutrality and for an additive trait, Qst = Fst for any model of population structure (Whitlock 1999). The pattern Qst > Fst (i.e., the mean value of the trait being more divergent across populations than expected by neutral processes) is taken as evidence of local adaptation (Merilä and Crnokrak 2001). On the contrary, Qst < Fst (i.e., the mean of the trait is more similar across populations than expected under neutral processes) is taken as evidence for spatially homogeneous selection [although this last conclusion is less robust to nonadditivity than the former (Whitlock 1999; Goudet and Büchi 2006)].
Nevertheless, selection is unlikely to act only on single traits independently (Lande and Arnold 1983), and a multivariate approach is required to make more accurate evolutionary predictions and to study adaptation on several traits simultaneously. Lande (1979) introduced a multivariate equivalent of the breeder's equation to predict the response to selection on multiple traits: Δz = GP−1S, where Δz is the vector of change in population trait means, G is the matrix of genetic covariances, P is the matrix of phenotypic covariances, and S is the vector of selection differentials on all traits. In this context, the multivariate equivalent of heritability is given by the matrix GP−1. An important focus of evolutionary quantitative genetics in the last two decades has been the study of the evolution of G matrices through time and particularly the disentangling of the effects of drift and selection on G (for a review see Roff 2000, 2007; Steppan et al. 2002). These studies concluded that selection and drift have different impacts on the evolution of G (Phillips and Mcguigan 2006). Drift is expected to reduce all additive variances and covariances by the same factor, while selection should alter each trait differently depending on its impact on fitness. Therefore, proportionality between G matrices among distinct populations is often taken as evidence for the effect of drift alone (Roff 2000), while differences in matrix structure are typically taken to suggest selection (Cano et al. 2004). However, as recently demonstrated empirically (Phillips et al. 2001; Whitlock et al. 2002), under drift alone, proportionality is expected only on average (Lande 1979). Among a set of replicate populations, strong deviations from proportionality can be observed between any given population pairs. What is in fact expected under drift alone is proportionality between the within-populations (G) and among-populations (D) covariance matrices (Lande 1979). Yet Schluter (1996), directly using a comparison of the eigenvectors of G and D among three stickleback species, suggested that proportionality between these two matrices could also occur in the presence of selection (for a review of this type of study, see Merilä and Björklund 2004). Thus, as proportionality between D and G is expected under both a neutral and a selective regime, disentangling the effects of drift vs. selection is impossible from a test of matrix similarity alone. Merilä and Björklund (2004) therefore suggested combining the Qst–Fst approach with analyses of the structure of covariance matrices to distinguish the effect of these two forces on multivariate covariances. In the companion to this article (Martin et al. 2008), we develop a statistical framework to combine these two kinds of analyses into a single new neutrality test. The resulting test is a multivariate extension of the classic Qst–Fst comparison and allows testing for the relative importance of drift vs. selection on multiple traits simultaneously (and on G matrix evolution itself).
In this article, we apply this new method to a naturally subdivided population of the freshwater snail (Galba truncatula), to quantify and test for the impact of selection on G matrices among a set of life-history traits, both across populations and across habitats. G. truncatula is an ideal model for testing the relative impact of selection and drift (Chapuis et al. 2007) because it has been shown to be under the influence of both forces. Indeed, G. truncatula lives in a spatially heterogeneous environment consisting of permanent and temporary water habitats. The temporary water habitat is characterized by the possibility of completely drying out during the summer, while the water level in the permanent water habitat remains constant throughout the year (Trouvé et al. 2003, 2005; Chapuis et al. 2007). Directional selection for contrasted phenotypes between habitats has been demonstrated by univariate Fst–Qst comparisons (comparing means over several traits) in Chapuis et al. (2007). Over a large number of randomly distributed populations for both habitats, Qst was found to be higher than Fst between habitats, an indication that diversifying selection is acting at this level, while Qst < Fst was found at the among-populations within-habitat scale, suggesting homogeneous selection within habitats. On the other hand, G. truncatula populations were also shown to have extremely small effective population sizes in both temporary and permanent habitats (Trouvé et al. 2005) and therefore also undergo strong drift. Overall, these natural populations provide a suitable model to study the relative impacts of selection and drift on the G matrix since both forces have a priori a strong influence on the system. Chapuis et al. (2007) focused only on the average Qst over several traits compared to Fst and made no use of information from the covariances among traits. Here, we test for multivariate selection in natural populations of G. truncatula both between subpopulations within each habitat and between temporary and permanent habitats, using the new method developed by Martin et al. (2008). Because Chapuis et al. (2007) showed that early traits might not be under the same selective pressure as late traits, we separately tested early and late traits (as defined previously in Chapuis et al. 2007). We first show how the multivariate approach gives a more accurate picture of the impact of selection vs. drift on the system. Then, having ascertained an effect of selection, we show how it influences the orientation of G and D matrices.
MATERIALS AND METHODS
Study sites and laboratory breeding:
G. truncatula is a freshwater snail reproducing mainly by selfing (>90%, Trouvé et al. 2003; Chapuis et al. 2007). It can be found in two types of habitats differing in water availability: either permanent ponds or temporary pools (see Chapuis et al. 2007 for details on the biology of the species and its ecology). Sixteen populations from Western Switzerland were sampled in 10 localities in June 2003. Ten populations were collected in permanent habitats and 6 in temporary ones. Within the same localities, we found populations from each habitat (see Chapuis et al. 2007). Thirty to 90 individuals were collected from each population and brought back to the laboratory (generation 0, G0). After field collection, G0 individuals were isolated into petri dishes (5 cm diameter) filled with water from Lake Geneva and fed with cereal flour used for snail breeding (TEXTIER). The photoperiod was set to 12 hr light:12 hr dark, and room temperature was maintained at 19° ± 1°. Every 10 days, water was changed and the petri dishes were moved at random to avoid any effect of their position in the rearing room. Egg capsules (G1) of G0 individuals were collected to constitute 10 families per population, on average (G1). Each G1 individual was kept alone in a petri dish and reared as detailed above for G0. Phenotypic measures were taken for 3 individuals per family on average (the different traits measured are detailed below).
Molecular variance estimation:
G0 individuals were genotyped at seven microsatellite loci (loci 9, 16, 20, 21, 29, 36, and 37) following the procedure described in Trouvé et al. (2000). To investigate the distribution of molecular variance between and within habitats we performed a hierarchical analysis. The hierarchical estimates of F-statistics were obtained from variance components of gene frequencies (Weir and Cockerham 1984; Weir 1996; Yang 1998) and were estimated with the package HIERFSTAT (Goudet 2005) implemented in the statistical environment R (R Development Core Team 2007). As explained in Wright (1969), this hierarchical decomposition allows us to estimate independently Fst (differentiation between populations and habitats) and Fis, which reflects the consequences of the species' mating system on genotypic proportions. The 95% confidence interval for each F-statistic and variance component was obtained by bootstrapping loci 1000 times using HIERFSTAT.
Quantitative trait measures:
A total of 12 quantitative traits were measured for each G1 snail, classified according to whether they were measured early in the life cycle (“early traits”) or later (after maturity, “late traits”). Shell length and width were measured at four dates: 3, 19, and 33 days after hatching (i.e., all early traits), and at 31 days after maturity (i.e., two late traits), to the nearest 0.01 mm, using an ocular micrometer on a binocular microscope. We also recorded age at maturity, which was determined when the first egg capsule was laid. We then estimated several measures of fecundity: the total number of eggs laid during the first 8 days after maturity, the average number of eggs per capsule over this same period, and the total number of egg capsules laid 30 days after maturity. These fecundity measures make up the subset of late traits, together with the age at maturity and size (length and width) 31 days after maturity.
Statistical analysis:
Transformation of the traits:
The G-matrices estimation and tests were performed after transformations of the trait values, to avoid spurious correlations among traits and to fulfill the requirements of the statistical analysis (Martin et al. 2008).
First, for early traits, sizes at different stages are artificially correlated, being the sum of the size at the previous stage plus recent growth. To eliminate this effect, we used differences between sizes at consecutive stages as our early trait measures. Second, the methods used to estimate (MANOVA) and compare G matrices (common principal analysis, CPC, a method determining the level of shared structure of matrices) (Flury 1988) both assume that the breeding values are normally distributed, which was not the case for the original data. Therefore, we used a power law transformation (zλ) to make the trait distributions as close as possible to Gaussian; this was performed automatically by maximum likelihood, using the Box–Cox method in R. Finally, as the measurements of the traits were on very different scales (e.g., a few millimeters for lengths and hundreds of days for age at maturity), traits were scaled to their mean, as suggested by Houle (1992) for any comparative analysis of variability between traits. The resulting distribution of transformed traits is fairly close to that of a Gaussian (see supplemental Figure 1, a and b).
Genetic (co)variance estimation:
A recent simulation study (Persson and Andersson 2004) showed that highly misleading genetic correlation estimates can be obtained from data sets that include missing records. To avoid this problem, we directly computed variances and covariances among family means for each trait or trait pair, instead of estimates of genetic covariances from individual data.
Considering the high selfing rate (>90%) and its constancy through time (Trouvé et al. 2003, 2005; Chapuis et al. 2007), the G0 were assumed completely inbred. Therefore, all G1 offspring from the same mother were considered genetically identical (inbred lines). Under this assumption, the differences between individuals within a family can be considered as only due to the environment (Bonnin et al. 1996, 1997), so that any genetic variance is given by the variance among family means ( ) and the environmental variance by the variance among individuals within families (
) and the environmental variance by the variance among individuals within families ( ), both estimated by MANOVA (see details below). Note that this is an approximation as sampling introduces a bias in such an estimate of genetic variance, within populations. Indeed, on average, the estimated covariance among family means is E(
), both estimated by MANOVA (see details below). Note that this is an approximation as sampling introduces a bias in such an estimate of genetic variance, within populations. Indeed, on average, the estimated covariance among family means is E(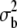 ) =
) = 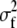 +
+ 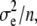 where
where  is the genetic variance between families,
is the genetic variance between families,  is the environmental variance, and n is sample size. However, because we use a MANOVA on the whole metapopulation to estimate G, this sample size is large (n is the degrees of freedom of the within-population level of the MANOVA: n = 244, see Table 1), so that the bias was neglected (E(
is the environmental variance, and n is sample size. However, because we use a MANOVA on the whole metapopulation to estimate G, this sample size is large (n is the degrees of freedom of the within-population level of the MANOVA: n = 244, see Table 1), so that the bias was neglected (E( ) ≈
) ≈  ≫
≫ 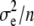 ). The sample sizes are smaller for the between-population covariances, but estimates from family means between populations are not biased by environmental variance within families. Other than that, the effect of sampling on covariance matrix estimates is of course accounted for in the test itself (see Martin et al. 2008).
). The sample sizes are smaller for the between-population covariances, but estimates from family means between populations are not biased by environmental variance within families. Other than that, the effect of sampling on covariance matrix estimates is of course accounted for in the test itself (see Martin et al. 2008).
TABLE 1.
Proportionality coefficients (ρ) between among- (D) and within- (G) population G matrices vs. their neutral expectation (Γ = F/(1 − F)) from genetic markers
| Value of Fst/(1 − Fst) | Value of ρst
|
|||
|---|---|---|---|---|
| Scale of comparison | d.f. | Early traits | Late traits | |
| One-level test | ||||
| Among populations uncorrected for habitat | DP: d.f. = 15 | ΓP = 3.14 | ρP = 0.19a | ρP = 0.24a |
| (2.73–3.66) | (0.15–0.28) | (0.18–0.35) | ||
| Two-level hierarchical test | ||||
| Between habitats | DH: d.f. = 1 | ΓH = −0.10 | ρH = 0.18 | ρH = 1.28a |
| (−0.22–0.03) | Randomization P = 0.48 | Randomization P = 0.004 | ||
| Among populations within habitats | DP/H: d.f. = 14 | ΓP/H = 3.19 | ρP/H = 0.20a | ρP/H = 0.18a |
| (2.80–3.71) | (0.15–0.29) | (0.14–0.26) | ||
Measures of phenotypic (ρ) and genetic (Γ) divergence are given for both early and late traits separately and for two alternative MANOVA models (see materials and methods for details). In the one-level test, G is compared either to D among populations without correcting for habitats (DP, ρP). In the two-level test, G is compared both to D among populations nested within habitats (DP/H, ρP/H) and to D between habitats and (DH, ρH). When available, the C.I. is given in parentheses. The degrees of freedom (d.f.) of the corresponding matrix estimates are given, and the corresponding within-population G matrix was estimated with d.f. = 244.
A significant difference, i.e., nonoverlapping C.I., or a significant randomization test for habitat effects.
Covariance matrices, summarizing all genetic covariances between traits, were estimated for the two distinct sets of traits described above (early and late traits) and at each level of population structure: within populations (G), among populations within habitats (DP/H), and between habitats (DH). DH, DP/H, and G were estimated by a MANOVA on family means, for all early and all late traits, and with populations nested within habitats as the explanatory factors. To estimate the effect of habitat on the phenotypic distribution, we also estimated the covariance among populations, without correcting for habitat (i.e., among populations, DP, and within populations, G), using a single-factor MANOVA.
Finally, as not all covariance matrices were positive definite (a prerequisite to apply the statistical tests developed), a matrix-bending method was used (Hill and Thompson 1987). This method consists of changing the null or negative eigenvalues to a very small positive value (10−6). Finally, we checked that the resulting covariances, after all the transformations, were very close to their values before transformations (supplemental Figure 2).
Principle of the neutrality test for G matrices:
We tested for departures from neutral divergence among each set of traits (early and late), using the method developed in our accompanying article (Martin et al. 2008). The method is fully detailed in that article so we here recall only its main principles. The method tests for departures from the expected relationship under neutrality: D = Fst/(1 − Fst)G, where D and G are the among-populations and within-populations covariance matrices. On the basis of estimates of these covariance matrices (by MANOVA), the method tests whether the departure from the neutral expectation is significant, given the sampling error. The method uses the maximum-likelihood framework of CPC analyses (Flury 1988): a maximum-likelihood estimate (MLE) of D and G under proportionality (D = ρstG) is obtained, with associated estimates of the proportionality coefficient ρst and of its confidence interval. The neutrality test then consists of jointly answering two questions (two tests): (i) Is the MLE of ρst significantly different from its neutral expectation Γst = Fst/(1 − Fst), estimated from the neutral markers?, and (ii) Is the proportionality itself the most likely relationship between D and G (as opposed to no relation at all), given their empirical estimates and the sampling error?
The first test was performed by comparing the confidence interval (C.I.) for the MLE of ρst with the C.I. for Γst, estimated from the molecular data by bootstrapping over microsatellite loci using the package HIERFSTAT (Goudet 2005). The second test is a test of proportionality between D and G, based on the CPC method (Flury 1988) and previously applied to the comparison of several within-population G matrices (Phillips and Arnold 1999). Here we applied this test to the comparison of D and G, using the corresponding mean square matrices, to comply with the distribution assumptions of the test (for details, see Martin et al. 2008). The test was performed using Eriksen's (1987) extension of the CPC method to compare sample covariance matrices, with a Bartlett correction of P-values for small samples. For simplicity, we refer to this test as a test of “proportionality between G and D” although the actual test is performed on the mean square matrix estimates. These two tests are independent but complementary in detecting different types of departure from the neutral pattern (discussed in Martin et al. 2008). The second test additionally gives some insight into the relative structure and orientation of G vs. D.
Note that the neutral expectation used here (D = Fst/(1 − Fst)G) corresponds to a haploid species or a diploid species with complete inbreeding [while D = 2Fst/(1 − Fst)G without inbreeding (Martin et al. 2008)]. This is to account for the very high inbreeding in G. truncatula, as demonstrated by a progeny array analysis and the very high Fis found by Trouvé et al. (2003) and Chapuis et al. (2007).
Extension of the test for hierarchical population structure:
Because individuals are hierarchically structured into populations nested within habitats, it is necessary to test for selection at these two levels. Consequently, the neutrality tests (D vs. G) were performed at the scale of the whole metapopulation (i.e., D among populations), but either ignoring habitat (a one-level test, see Table 1, as in Martin et al. 2008) or accounting for its effect (i.e., a two-level test with habitat as an extra factor, see Table 1). In the first case (denoted “Among populations uncorrected for habitat” in Table 1), DP and G estimates from a one-factor MANOVA were compared exactly as in Martin et al. (2008). Under neutrality, the observed proportionality coefficient between G and DP, denoted ρP, is ΓP = Fst/(1 − Fst). ΓP was estimated from the molecular data as  , where
, where 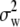 ,
,  , and
, and 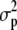 are the components of variance of marker allele frequencies within individuals, among individuals within populations, and among populations, respectively. The C.I. for ΓP was estimated by bootstrapping over loci 1000 times using HIERFSTAT, and C.I. overlap was used to test the neutral expectation ρP = ΓP.
are the components of variance of marker allele frequencies within individuals, among individuals within populations, and among populations, respectively. The C.I. for ΓP was estimated by bootstrapping over loci 1000 times using HIERFSTAT, and C.I. overlap was used to test the neutral expectation ρP = ΓP.
In the second case (two levels denoted “Among populations within habitat” and “Between habitats” in Table 1), a similar analysis was performed but G was compared to the covariance matrix among populations both within habitats (DP/H) and between habitats (DH), estimated from a nested MANOVA (see above). The expected neutral relationship between the three matrices is
 |
(1) |
Extending Equation 12 of Martin et al. (2008) to a hierarchical population structure, Eriksen's (1987) joint proportionality test between the three matrices was then used to test for neutrality. The corresponding expected coefficients of proportionality under neutrality were estimated as 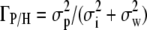 , among populations within habitats, and
, among populations within habitats, and  , between habitats, where
, between habitats, where 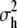 is the component of variance of marker allele frequencies between habitats, with the corresponding C.I. estimated by bootstrapping over loci using HIERFSTAT. The corresponding observed proportionality coefficients are denoted ρP/H and ρH for the population and habitat levels, respectively, and the neutral expectation in Equation 1 can be written ρP/H = ΓP/H and ρH = ΓH. C.I. overlap was used to test the first expectation ρP/H = ΓP/H, among populations within habitats, and a randomization test (described below) was used to test ρH = ΓH. Note that the two-level structure of the population is still completely represented in this case, because the test is no longer with two matrices (G vs. DP) but with three matrices (G vs. DP/H and G vs. DH), representing all levels of population structure, which are jointly estimated by the MANOVA.
is the component of variance of marker allele frequencies between habitats, with the corresponding C.I. estimated by bootstrapping over loci using HIERFSTAT. The corresponding observed proportionality coefficients are denoted ρP/H and ρH for the population and habitat levels, respectively, and the neutral expectation in Equation 1 can be written ρP/H = ΓP/H and ρH = ΓH. C.I. overlap was used to test the first expectation ρP/H = ΓP/H, among populations within habitats, and a randomization test (described below) was used to test ρH = ΓH. Note that the two-level structure of the population is still completely represented in this case, because the test is no longer with two matrices (G vs. DP) but with three matrices (G vs. DP/H and G vs. DH), representing all levels of population structure, which are jointly estimated by the MANOVA.
In all these tests, the degrees of freedom associated with each matrix estimate were directly provided by the MANOVA degrees of freedom, but we used the corrected value for unbalanced designs given in Equation 9 of the accompanying article (Martin et al. 2008). These degrees of freedom are computed assuming no missing values, which should be approximately correct in our case, as there was only 2.8% (respectively 1.8%) of missing values in the early traits (respectively late traits) data set. Note that it was not possible to apply the test in each habitat separately, since the number of populations in the temporary habitat is less than the number of traits studied (five vs. six), which can lead to an incorrect P-value (Martin et al. 2008).
Randomization test for the habitat effect:
Although ρH between habitats can be estimated, its equality to the neutral expectation (ρH = ΓH) cannot be tested on the basis of confidence intervals, as there are only two habitats, and thus 1 d.f. left, a situation preventing the estimation of a confidence interval as explained in Martin et al. (2008). To circumvent this problem, we built a permutation test for ΓH as follows: we compared the observed value of the difference δobs = ρH − ΓH to values of the statistic δ obtained after randomizing populations between the two habitats (keeping the same number of populations per habitat). This randomization scheme allows generating the distribution of the statistic δ under the null hypothesis that the habitat type has no selective effect (ρH = ΓH, δ = 0). We tested this null hypothesis against the alternative ρH > ΓH (δ > 0), since we postulate that the two habitats should lead to different selective optima. If the null hypothesis is true, the observed δobs should be within the 95% lowest values of the null distribution of δ (for a test at the level α = 5%). On the other hand, if δobs is larger than the 95% lowest δ-values generated by randomization, the proportionality constant ρH is deemed significantly larger than expected under pure neutral processes. The probability (P-value) that the null hypothesis, δobs = 0, is true was thus estimated as the proportion of randomized δ-statistics larger than or equal to the observed δobs (Manly 1997).
Simulation checking:
We used individual-based simulations to check the validity of our interpretation of the results. We simulated a metapopulation subdivided into subpopulations, each pertaining to one of two alternative habitats (to simulate temporary or permanent habitats). The aim was to check whether selection for distinct habitats in this context could generate the observed pattern, i.e., a difference in the outcome of our tests with vs. without correction for habitat (see results). To do so, we used the exact same simulation method as in Martin et al. (2008) but with an additional habitat level (two habitats). The evolution of the metapopulation was simulated, and then we performed our tests on samples taken from the simulated metapopulation.
Following the simulation scheme of Martin et al., each population underwent mutation (pleiotropic on six traits), drift, and selection for a phenotypic optimum (on all six traits), for 50 generations. The additional habitat level was introduced by setting a different optimum for each habitat. The corresponding two optima differed only for two of the six traits simulated, as habitat type may affect the optimum value of some of the traits under study but not all of them. Of course, the optimum was the same for all populations of a given habitat (homogeneous selection within habitats). The metapopulation was a set of isolated subpopulations of small size N = 30 (10 subpopulations in one habitat and 6 in the other, as in our empirical design), to mimic the small effective sizes of our empirical G. truncatula populations. Each subpopulation was started from a single common polymorphic ancestral population generated by pure mutation, with initial average phenotype equally distant from each habitat optimum. The genetic setting was the same as in Martin et al. (2008): with mutational and selective covariances (randomly generated) scaled so that the average effect of mutations on fitness was E(s) = −0.1, with a genomewide mutation rate U = 0.1 and 30 loci. The individuals were haploid with a 10% recombination rate between loci. This was intended to mimic approximately the high inbreeding of G. truncatula, without having to simulate diploids.
The tests were then performed on the simulated populations, following our empirical sampling design. From each of the simulated subpopulations, 15 individuals were sampled (every five generations). From the phenotypic distribution (of the six traits) in these samples, covariance matrices G and D were estimated by MANOVA. The corresponding mean square matrix estimates were then used to test for proportionality between G and D (see materials and methods and Martin et al. 2008). As in our empirical study, two alternative MANOVA formulas were used, with D estimated either after correcting for a habitat effect (DP/H) or not (DP). The P-values of the test of proportionality of D vs. G (with or without correction for habitat) were recorded for 100 replicate simulations of the metapopulation (16 populations, two habitats). Results from these simulations are reported in Figure 4. Although not the main focus of our simulations, the proportionality coefficient (ρP/H or ρP with or without habitat correction, respectively) was also recorded in these simulations, to check the effect of habitat correction on the test of divergence (ρ vs. Γ). For this purpose, the corresponding ΓP = ΓP/H = Fst/(1 − Fst) was derived from the classic recursion for isolated populations (Martin et al. 2008), assuming no population structure at the habitat level, consistent with our empirical estimates (ΓH ≈ 0, see Table 1). The results of these simulations are given in supplemental Figure 3.
Figure 4.—
Effect of heterogeneous selection between habitats on the proportionality test between D and G. Using the same simulations as in Figure 3 with habitat heterogeneity (distinct optimum in each habitat for two traits of six), circles give the P-values of the proportionality test between the within-population covariance G and the among-population covariance DP (a, uncorrected for habitats) or DP/H (b, after habitat correction). The Bartlett corrected P-values are given as a function of Fst (indicated on the bottom horizontal axis). The dashed line gives the 5% α-level and the proportion of P-values above this α-level, for each Fst value, is indicated on the top horizontal axis. Correction for habitat type leads mostly to the acceptance of proportionality (G vs. DP/H) (b) whereas proportionality is often rejected in the absence of correction (G vs. DP) (a), in particular for high levels of population structure.
We also used these simulations to check the validity of the randomization test for heterogeneous selection between habitats (ρH vs. ΓH, see above). For each simulation run of the metapopulation with two contrasted habitats, we recorded the value of ρH after 50 generations, while ΓH was assumed null (see above). This gave us the distribution of ρH when there is selection for alternative optima in the two habitats. As a comparison, we also ran 100 replicate simulations with the same parameters but without habitat effect (selection for the same optimum in both habitats). This gave us the null distribution of ρH when there is no heterogeneity between habitats. Comparing the two distributions shows the effect of habitat heterogeneity on ρH. Finally, we applied our randomization approach to data from one randomly picked metapopulation (to mimic our empirical data set) and checked that the randomization distribution of ρH was similar to the null distribution obtained from simulations without habitat heterogeneity. This simulation check is reported in Figure 3.
Figure 3.—
Simulation check of the randomization test: detecting habitat heterogeneity. The distribution of δ = ρH − ΓH in 100 replicate simulations of a metapopulation (undergoing stabilizing selection, drift, and mutation) is given for parameters chosen to resemble our empirical data set and our study species (see the simulation check section in materials and methods). The selective optima were the same in all populations of a given habitat, but either differed between each habitat for two of the six traits (“heterogeneous habitats,” shaded histogram) or were the same (“null distribution,” solid line). As an example of the randomization test developed, circles give the randomization distribution of δ among 1000 randomization samples from one of the 100 simulated data sets after 50 generations (homogeneous selection among populations and heterogeneous selection between habitats). The null distribution is close to the randomization distribution as assumed in the test, and both strongly differ from the distribution with habitat heterogeneity.
RESULTS
Phenotypic distribution of populations across habitats:
Because there are many traits measured in each population, graphically representing the distribution of multivariate phenotypes is difficult. To give a relevant visualization of this distribution, we present it in the two dimensions given by the first two principal components (PC) of the within-population covariance matrix (G), in Figure 1. The first two components of G account for 65 and 22%, respectively, of the within-population covariance of early traits and for 74 and 18%, respectively, of the within-population covariance of late traits. The orientation of G within each population (with a color code for habitat type) is given by solid ellipses, giving the 95% confidence region of the phenotypic distributions (under a bivariate Gaussian assumption for the data). For the sake of clarity, population means were dilated from the overall mean by a factor 30, so that phenotype distributions do not overlap. Note that this does not alter the orientation of G or D or the relative position of subpopulations from distinct habitats. Figure 1 illustrates the phenotypic distribution for late (Figure 1a) and early (Figure 1b) traits. Population mean phenotypes from the two habitats (gray vs. black ellipses) appear to form separate groups for late traits (Figure 1a), while they are widely overlapping for early traits (Figure 1b). This suggests that habitat type has led to the divergence of late traits but not of early traits, and the tests below confirm that this is an effect of selection for distinct optima.
Figure 1.—
Bivariate representation of phenotypic distributions along the main directions of the within-population covariance matrix. The (Gaussian transformed) phenotypic distribution of each population is represented in the coordinate system of the two principal components (PC1 and PC2) of the within-population covariance matrix (G, estimated from MANOVA), with the percentage of variance explained by each PC given in parentheses. Small ellipses give the 95% bivariate confidence intervals (C.I.) of each population's phenotypic distribution, with the gray corresponding to populations from permanent ponds and the black to populations from temporarily dried ponds. The means for each population (given by the small squares) were dilated by a factor of 30 for clarity of the plot. The thick solid line ellipse gives the bivariate C.I. for the observed distribution of population means (representing the among-population covariance matrix, DP). The thick dashed line ellipse gives the corresponding C.I. if DP was exactly proportional to G (the neutral expectation). (a) Late traits; (b) early traits.
To compare the orientation of G with that of the among-population covariance matrix (D), we represented the observed D by a solid ellipse in Figure 1 describing the overall distribution of the mean of each population. For comparison, a dashed ellipse in Figure 1 gives the expected orientation of D if it was proportional to G, i.e., parallel to the main PC and with length (resp. width) proportional to the eigenvalues of the first (resp. second) eigenvector (PC). In both cases, D appears to differ only slightly from the neutral expectation (D proportional to G, given by the dashed ellipse). We deal with the statistical test of these differences below, but it is clear that there is a fairly strong parallelism between G and D, even if they are not strictly proportional.
Neutral divergences in G:
First test—comparing multivariate phenotypic divergence to its neutral expectation:
Table 1 gives the value of neutral divergence (Γ) from molecular data and of the estimated proportionality coefficient ρ (divergence) between G and D for both early and late traits and for each level of structure (i.e., among populations uncorrected for habitats or among populations within habitats and between habitats). The divergence among populations is significantly lower than expected under neutrality, both with and without correcting for habitat (ρP < ΓP and ρP/H < ΓP/H, Table 1). This is true for both early and late traits, an indication that homogenizing selection is probably occurring at the population level on both trait sets. On the contrary, between habitats, for early and particularly late traits, the value of ρH is larger than its neutral expectation ΓH (ρH = 0.28 and 1.28 vs. ΓH = −0.02, Table 1). This effect cannot be tested by the C.I. overlap, but the randomization test shows that this difference is (i) not significant for early traits (P = 0.484, Table 1 and Figure 2b) but (ii) highly significant for late traits (P = 0.004, Table 1 and Figure 2a). Thus, there is strong evidence of selection for different optima between the two habitats, on late traits at least.
Figure 2.—
Randomization test of heterogeneous selection between habitats. Histograms of the distribution of δ = ρH − ΓH for randomized samples of the empirical data set (see materials and methods) are given for both (a) late and (b) early traits. These distributions give the null distribution of δ, if the habitat type had no effect on the phenotypic distribution (habitat type swapped between populations). The observed difference between ρH and ΓH, δobs, is given by the thick vertical lines. For late traits, δobs is larger than most δ-values (996 of 1000) under the null hypothesis of habitat homogeneity, which indicates the presence of heterogeneous selection between habitats. For early traits, no such discrepancy is observed: there is no significant departure from homogeneity between the two habitats for early traits.
Checking the validity of the randomization test of habitat heterogeneity:
In parallel, our simulations confirm the validity and efficiency of the randomization approach to detect heterogeneous selection between habitats. Figure 3 shows the distribution of δ = ρH − ΓH in replicate simulations of a metapopulation undergoing homogeneous selection among populations, with or without different optima in each habitat (see materials and methods). When there is habitat heterogeneity (shaded histogram), the distribution of δ is biased upward relative to that obtained when there is no habitat effect (null distribution of δ, solid line), which suggests that the test has the power to detect habitat effects. Furthermore, the randomization distribution of δ in a single simulated metapopulation with habitat heterogeneity (circles) is similar to the null distribution, which suggests that the randomization method efficiently estimates the distribution of the statistic under the null hypothesis.
Second test—proportionality between G and D:
The results for the test of proportionality between the within-population and among-population covariance matrices are given in Table 2. For early traits, proportionality between G and D, both with and without correction for habitat (DP and DP/H, respectively), cannot be rejected (P ≈ 0.2 and 0.29, respectively, Table 2). Conversely, for late traits, proportionality between G and D is rejected both with and without correction for habitat (P = 5.10−4 for DP and P = 0.04 for DP/H, respectively, Table 2). However, after Bonferroni correction for multiple tests, the test on DP/H becomes nonsignificant while the test DP remains strongly significant. These results show that although selection is occurring among populations in all traits (Table 1), proportionality between G and D is observed, except in the late traits for which this proportionality is restored only after correcting for habitat.
TABLE 2.
Proportionality test between the within-population (G) and among-population (D) covariance matrices for both early and late traits
|
P-value
|
||
|---|---|---|
| Scale of comparison | Early traits | Late traits |
| Among populations uncorrected for habitat (G proportional to DP) | 0.2 | 5.10−4a |
| Among populations within habitats (G proportional to DP/H) | 0.29 | 0.04 |
The P-value corresponds to a likelihood-ratio test (χ2) corrected by the Bartlett adjustment.
A significant P-value after a Bonferonni correction for multiple tests. Note that the tests were performed on mean square matrix estimates (see materials and methods).
At first glance, visual inspection of the phenotype distributions in Figure 1 did not lead us to expect a difference in the outcome of the proportionality test for late vs. early traits. Indeed, the difference in orientation between the observed D (solid ellipse) and that expected under proportionality (dashed ellipse) seemed quite similar for the two trait sets. However, for late traits (Figure 1a), the correlation between PC1 and PC2 is stronger (the ellipse is flatter) than for early traits (Figure 1b), which probably explains why the difference in orientation was detected in the former and not in the latter. With a more correlated distribution of population means (D), the test is more powerful and has more chances to reject proportionality. This increased correlation in D for late traits is due to the fact that mean phenotypes are in separate groups according to the habitat type of each population (gray vs. black in Figure 1a). This habitat effect orients the among-population covariance (solid line ellipse) along a direction different from that of G and disappears when correcting for habitat, which is confirmed by our statistical tests: for late traits proportionality is not rejected after correcting for habitat (Table 1).
Finally, note that the proportionality of G and D observed in our data set also confirms that the same selective regime is acting on all the traits in each set (i.e., within early or late traits). If different traits within a set were submitted to different selective regimes (i.e., a mixture of heterogeneous and homogeneous selection among populations), proportionality of G and D would be strongly rejected (as demonstrated in Figure 3c of the accompanying article, Martin et al. 2008). We now turn to the global interpretation of all these patterns.
Biological interpretation:
Our interpretation of these empirical patterns is as follows. For early traits, there is spatially homogeneous selection for a single optimum in all populations (the same in both habitats). Conversely, for late traits, distinct habitats correspond to distinct optima (heterogeneous selection between habitats), while keeping the same optimum for all populations within a given habitat (homogeneous selection among populations within habitats). This interpretation is first supported by examination of the multivariate phenotype distributions in Figure 1: for early traits, the population means are evenly distributed among permanent and temporary habitats, and there is no habitat effect. By contrast, population means tend to form two subgroups for late traits, denoting the effect of heterogeneous selection between habitats.
This interpretation also accounts for various results of our statistical tests. First, among-populations divergence (ρP and ρP/H) is always below its neutral expectation (homogeneous selection, Table 1). Second, divergence between habitats (ρH) is significantly higher than neutral for late but not early traits (heterogeneous selection between habitats for late traits, Table 1, Figure 2). Third, for early traits, G and D are proportional irrespective of habitat correction (both with DP and with DP/H, Table 2). Fourth, for late traits, G is proportional to D only after habitat correction (only with DP/H, Table 2).
The first two points are straightforward, but the third and fourth points deserve some further explanations. Previous simulations (Martin et al. 2008) have shown that homogeneous selection among populations tends to retain proportionality between G and D, thus mimicking the effect of pure drift (under neutrality, DP = ΓPG, so that the two matrices are proportional). This is why we observe proportionality between G and D for early traits, irrespective of habitat correction (with both DP and DP/H), as there is no habitat effect (our third point). By contrast, if each habitat selects for distinct optima, the distribution of population means forms two subgroups corresponding to each habitat (see Figure 1a), which breaks the proportionality between G and DP (over all populations, ignoring habitat). After correcting for habitat, this effect is removed and proportionality between G and D is restored (with DP/H), our fourth point.
To check the validity of the above interpretation, we used simulations of the scenario assumed above and checked whether these simulations could reproduce the patterns we obtained empirically, in particular regarding the effect of habitat correction on the proportionality test. This is detailed below.
Simulation check of the interpretation:
Figure 4 shows the effect of correcting for the habitat type before estimating the covariance matrices. In these simulations, the six traits are selected for a single optimum across all populations and habitats (homogeneous between habitats), while two of the six traits are under divergent selection between habitats (different optima for each habitat). In this context, proportionality is almost always rejected when correction for habitat is omitted, at least when Fst > 0.5 (Figure 4a, <9% of the P-values are above the α-level). By contrast, correcting for habitat (Figure 4b) leads to the frequent acceptation of the proportionality hypothesis: the P-values are often above the 5% rejection level in the corrected case (Figure 4b, >55% nonrejection, upper x-axis).
These simulations, although an idealized vision of the empirical situation, give support to our argument (fourth point mentioned previously) that when selection is homogeneous at the population level but heterogeneous at the habitat level, proportionality between G and D is observed only if a correction for habitat is performed when estimating D (i.e., with DP/H but not with DP).
Note that with the level of heterogeneous selection between habitats that we used in the simulations, habitat correction has almost no effect on the measures of phenotypic divergence (ρP ≈ ρP/H), although it does affect the proportionality test. Indeed, supplemental Figure 3 shows that ρP and ρP/H are almost equal, for the same simulations as in Figure 4. This is consistent with our observing few differences between ρP and ρP/H, for both early traits (ρP = 0.19 vs. ρP/H = 0.20, Table 1) and late traits (ρP = 0.24 vs. ρP/H = 0.18, Table 1).
DISCUSSION
We found evidence for the action of different forms of selection on the freshwater snail G. truncatula, both among populations within habitats and between the two habitats. To detect this selection, we applied a new neutrality test based on the comparison of the among-population and within-population covariance matrices of phenotypic distributions (Martin et al. 2008). We extended this neutrality test to a situation where populations are nested within two types of habitats. The evidence found for selection can be summarized as follows. First, among all populations, ignoring the habitat type, we found evidence of spatially homogeneous selection for the same optimum in both early and late life-history traits. This conclusion stems from (i) lower multivariate phenotypic divergence than expected under neutrality at the population level (ρP < ΓP and ρP/H < ΓP/H, Table 1) and (ii) strong parallelism between D and G (even proportionality in early traits, Table 2), which is consistent with the effect of homogeneous selection observed in simulations [see Figure 3a, test ii, of the accompanying article, Martin et al. 2008]. Second, for late traits only, we found strong evidence of selection for a distinct optimum in each habitat, as suggested by (i) a higher divergence than expected under neutrality at this level (ρH > ΓH, Table 1 and Figure 2) and (ii) the proportionality between D and G being observed only when a correction for habitat is used (Table 2 and Figure 4), which suggests that habitat heterogeneity introduced some departure from proportionality between D and G.
Overall, our data and analysis suggest that early traits are selected for the same optimum in all populations and habitats, while late traits are selected for the same optimum in all populations of the same habitat but this optimum differs between habitats. This interpretation of our data is supported by simulations of the above scenario (with parameters consistent with G. truncatula's biology) that gave the exact same patterns. The congruence between the experimental results and the simulations is definitely not a proof as these simulations are obviously a simplified model of the G. truncatula populations. However, they do suggest that our interpretation is plausible and fairly parsimonious.
One important methodological conclusion of this study is the importance of scaling to appropriately interpret G matrix comparison. In our data set, for example, age at maturity has a large variance compared to the others traits (data not shown). In the absence of scaling, G could appear artificially stable across populations because it would be mainly determined by this single trait with a large variance. Another methodological implication is the choice of relevant trait sets. We distinguished two types of traits: late and early traits, using the previous division made by Chapuis et al. (2007). Late traits are measures of fecundity, and age at maturity, which are a priori directly related to fitness, whereas early traits are growth traits, a priori less directly related to fitness (although they do appear to be under selection). Consistent with this, heritability and the genetic and environmental variances for these two categories of traits (supplemental table) show the pattern typically observed for fitness vs. nonfitness traits. Indeed, relative to early (nonfitness) traits, late traits have a low heritability but a high coefficient of environmental and genetic variation, as typically observed for fitness traits (reviewed in Merilä and Sheldon 1999). Thus, our results corroborate the common statement that fitness traits tend to have lower heritability than morphological traits (despite having larger genetic variances) because of their higher sensitivity to environmental variation (Houle 1992; Merilä and Sheldon 1999; Barton and Keightley 2002). Consistently, in our study, choosing either trait set has substantial impact on the outcome of the neutrality tests, at least for the effect of habitat where heterogeneity is detected only for late traits (Figure 2). This could be expected, if fitness traits are the first to respond to habitat heterogeneity. In any case, this illustrates that the nature of the traits chosen to form G matrices is important, so that comparisons between trait types may be useful as pointed out by Pigliucci (2006).
Note that any study of quantitative traits is constrained to focus on a subset of all possible phenotypic traits of an organism. While this may be a limit for estimating multivariate selection (Lande and Arnold 1983), it is less problematic for testing multivariate neutrality as is done in this article. Indeed, under neutrality, covariances are randomly generated and are a priori little influenced by covariances with other unknown traits of the organism, so that they may be studied only on a sample of all traits. However, it remains true that, to provide meaningful interpretations, one must combine biologically coherent sets of traits, which is what we intended here by choosing early vs. late traits.
The fact that our study species shows a high level of inbreeding may affect our results because linkage disequilibrium (LD) may be significant between neutral markers and the loci determining the traits (QTL) under study. Linkage to selected loci has to be fairly high and selection fairly strong for it to impose a deviation of the distribution of Fst from its null expectation (Whitlock 2008). Yet, if such deviation was generated in our study species, this should favor the neutral hypothesis, as the estimated Fst at neutral markers would get closer to that at the QTL and thus closer to Qst (see, e.g., Le Corre and Kremer 2003; Porcher et al. 2006). Therefore, our conclusions (rejection of the neutral pattern, Table 1) are conservative in that they cannot have been generated by such linkage effects. The other potential issue arising from the high inbreeding in our study species is the effect of LD between neutral markers. Such LD is not overwhelming in our data set at least when measured at the scale of the whole metapopulation (14 of the 21 possible pairs of loci show no significant LD, as tested with F-Stat). However, it is almost necessary that there be some LD between some pairs of markers, in some populations. Such LD would imply that the bootstrap C.I. of ΓP, ΓH, and ΓP/H given in Table 1 might underestimate the actual C.I., as the bootstrap method implicitly assumes independence among marker loci. However, a set of partly linked markers should correspond to a smaller set of unlinked markers, so that the actual C.I. could be larger by no more than a factor of 2 or 3. Such an increase in the C.I. for Γ-estimates (from markers) would clearly not change our conclusions, as our ρ-estimates (from quantitative traits) deviate from the corresponding Γ-estimates by much larger orders of magnitude (see Table 1).
Our findings are consistent with those of Chapuis et al. (2007), based on Qst values obtained from the mean of several quantitative traits. However, the estimated C.I. in the latter study may have been incorrect since the different traits were not independent. By contrast, the method used here both accounts for covariance between traits and uses its information. Although qualitatively similar, the two methods yield slightly different estimates: in Chapuis et al. (2007) Qst within habitats was 0.22 on early traits (C.I.: 0.17–0.28) while ρst here is 0.20 (C.I.: 0.15–0.29). On late traits, Qst was 0.42 (C.I.: 0.31–0.54) while the estimated ρst here is 0.18 (C.I.: 0.14–0.26). As the range for Qst is [0, 1] while that for ρst is [0, +∞], it gives more power to the latter method. Furthermore, while a mixture of heterogeneous and homogeneous selection over the trait set could generate misleading results based on mean Qst, such a mixed regime would have been detected by the multivariate method (Figure 3c of Martin et al. 2008).
We now turn to how the observed patterns can shed light on the relative impacts of drift and selection on multivariate evolution. A strong effect of drift has been demonstrated from extremely small effective sizes in the G. truncatula populations studied here (Trouvé et al. 2005). Yet, evidence for selective effects is unequivocal for both early and late traits, suggesting that selection is strong enough to overcome these substantial drift effects. At the same time, we found proportionality between G and D among populations, in most cases (Table 1). Such proportionality has been diversely interpreted in the literature, taken as evidence of drift effects by some (e.g., Lande 1979; Rogers and Harpending 1983), while others interpreted it as evidence of directional selection (e.g., Schluter 1996); see the review by Merilä and Björklund (2004). Simulations indeed show that proportionality of G and D is rarely rejected when there is selection (either heterogeneous or homogeneous among populations, Figure 3 of Martin et al. 2008). From this proportionality only, it thus seems impossible to disentangle the effects of selection and drift. However, habitat heterogeneity for late traits is sufficient to break proportionality between G and D (Table 2, Figure 4), which suggests that at least heterogeneous selection can have a strong impact on proportionality. Overall, two general points can be deduced from the present study. First, we expect in general proportionality of G and D when Qst ≤ Fst (or ρst ≤ Γst), because both homogeneous selection and drift tend to produce this pattern. Second, when such proportionality is rejected, this might be because of divergent selection among habitats, which may have been neglected in the study. This method could therefore allow the detection of hidden habitat heterogeneity.
To conclude, this study provides strong evidence of multivariate selection in a heterogeneous environment using G matrices, with the possibility to detect both population and habitat selection levels. We observe proportionality between G and D despite evidence of a strong selective pressure, showing that both selection and drift tend to favor this pattern, as confirmed by simulations. Finally, this study illustrates how the joint use of molecular markers and multivariate tests on quantitative traits can allow accurate detection of the types of selection influencing populations at different hierarchical levels.
Acknowledgments
We thank Jean-Baptiste Ferdy and Patrice David for helpful comments on previous versions of this manuscript. We also acknowledge two anonymous reviewers for beneficial remarks. This is an Institut des Sciences de 1'Evolution de Montpellier publication (no. 2008-089). Funding for this study was provided by grants from the Swiss National Science Foundation (3100-68325.2) to J.G. G.M. was supported by a Swiss National Foundation postdoc grant (3100A0-108194/1) to J.G. and a Centre National de la Recherche Scientifique postdoc grant to Sylvain Gandon.
References
- Barton, N. H., and P. D. Keightley, 2002. Understanding quantitative genetic variation. Nature 3 11–21. [DOI] [PubMed] [Google Scholar]
- Bonnin, I., J. M. Prosperi and I. Olivieri, 1996. Genetic markers and quantitative variation in Medicago truncatula (Leguminosae): a comparative analysis of population structure. Genetics 143 1795–1805. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Bonnin, I., J. M. Prosperi and I. Olivieri, 1997. Comparison of quantitative genetic parameters between two natural populations of a selfing plant species, Medicago truncatula Gaertn. Theor. Appl. Genet. 94 641–651. [Google Scholar]
- Cano, J. M., A. Laurila, J. Palo and J. Merilä, 2004. Population differentiation in G matrix structure due to natural selection in Rana temporaria. Evolution 58 2013–2020. [DOI] [PubMed] [Google Scholar]
- Chapuis, E., S. Trouvé, B. Facon, L. Degen and J. Goudet, 2007. High quantitative and no molecular differentiation of a freshwater snail (Galba truncatula) between temporary and permanent habitats. Mol. Ecol. 16 3484–3496. [DOI] [PubMed] [Google Scholar]
- Dempster, E. R., 1955. Maintenance of heterogeneity. Cold Spring Harbor Symp. Quant. Biol. 20 25–32. [DOI] [PubMed] [Google Scholar]
- Eriksen, P. S., 1987. Proportionality of covariance matrices. Ann. Stat. 15 732–748. [Google Scholar]
- Flury, B., 1988. Common Principal Components and Related Multivariate Models. Wiley, New York.
- Goudet, J., 2005. HIERFSTAT, a package for R to compute and test hierarchical F-statistics. Mol. Ecol. Notes 5 184–186. [Google Scholar]
- Goudet, J., and L. Büchi, 2006. The effects of dominance, regular inbreeding and sampling design on QST, an estimator of population differentiation for quantitative traits. Genetics 172 1337–1347. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Hedrick, P. W., 2007. Genetic polymorphism in heterogeneous environments: the age of genomics. Annu. Rev. Ecol. Evol. Syst. 37 67–93. [Google Scholar]
- Hill, W. G., and R. Thompson, 1987. Probabilities of non-positive definite between-groups or genetic covariance matrices. Biometrics 34 429–439. [Google Scholar]
- Houle, D., 1992. Comparing evolvability and variability of quantitative traits. Genetics 130 195–204. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Kawecki, T. J., and D. Ebert, 2004. Conceptual issues in local adaptation. Ecol. Lett. 7 1225–1241. [Google Scholar]
- Lande, R., 1979. Quantitative genetic analysis of multivariate evolution, applied to brain: body size allometry. Evolution 33 402–416. [DOI] [PubMed] [Google Scholar]
- Lande, R., and S. J. Arnold, 1983. The measurement of selection on correlated characters. Evolution 37 1210–1226. [DOI] [PubMed] [Google Scholar]
- Le Corre, V., and A. Kremer, 2003. Genetic variability at neutral markers, quantitative trait loci and trait in a subdivided population under selection. Genetics 164 1205–1219. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Lenormand, T., 2002. Gene flow and the limits to natural selection. Trends Ecol. Evol. 117 183–189. [Google Scholar]
- Levene, H., 1953. Genetic equilibrium when more than one niche is available. Am. Nat. 87 331–333. [Google Scholar]
- Lynch, M., and B. Walsh, 1998. Genetics and Analysis of Quantitative Traits. Sinauer Associates, Sunderland, MA.
- Manly, B. F. J., 1997. Randomization, Bootstrap and Monte Carlo Methods in Biology, Ed. 2. Chapman & Hall, London.
- Martin, G., E. Chapuis and J. Goudet, 2008. Multivariate Qst–Fst comparisons: a neutrality test for the evolution of the G matrix in structured populations. Genetics 180 2135–2149. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Merilä, J., and M. Björklund, 2004. Phenotypic integration as a constraint and adaptation, pp. 107–129 in The Evolutionary Biology of Complex Phenotypes, edited by M. Pigliucci and K. Preston. Oxford University Press, Oxford.
- Merilä, J., and P. Crnokrak, 2001. Comparison of genetic differentiation at marker loci and quantitative traits. J. Evol. Biol. 14 892–903. [Google Scholar]
- Merilä, J., and B. C. Sheldon, 1999. Genetic architecture of fitness and nonfitness traits: empirical patterns and development of ideas. Heredity 83 103–109. [DOI] [PubMed] [Google Scholar]
- Porcher, E., T. Giraud and C. Lavigne, 2006. Genetic differentiation of neutral markers and quantitative traits in predominantly selfing metapopulations: confronting theory and experiments with Arabidopsis thaliana. Genet. Res. 87 1–12. [DOI] [PubMed] [Google Scholar]
- Persson, T., and B. Andersson, 2004. Accuracy of single- and multiple-trait REML evaluation of data including non-random missing records. Silvae Genet. 53 135–139. [Google Scholar]
- Phillips, P. C., and S. J. Arnold, 1999. Hierarchical comparison of genetic variance-covariance matrices. I. Using the Flury hierarchy. Evolution 53 1506–1515. [DOI] [PubMed] [Google Scholar]
- Phillips, P. C., and K. L. Mcguigan, 2006. Evolution of Genetic Variance-Covariance Structure in Evolutionary Genetics: Concepts and Case Studies, edited by C. W. Fox and J. B. Wolf. Oxford University Press, Oxford.
- Phillips, P. C., M. C. Whitlock and K. Fowler, 2001. Inbreeding changes the shape of the genetic covariance matrix in Drosophila melanogaster. Genetics 158 1137–1145. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Pigliucci, M., 2006. Genetic variance-covariance matrices: a critique of the evolutionary quantitative genetics research program. Biol. Philos. 21 1–23. [Google Scholar]
- R Development Core Team, 2007. R: a language and environment for statistical computing, Vienna. http://www.R-project.org.
- Roff, D. A., 2000. The evolution of the G matrix: selection or drift. Heredity 84 135–142. [DOI] [PubMed] [Google Scholar]
- Roff, D. A., 2007. A centennial celebration for quantitative genetics. Evolution 61 1017–1032. [DOI] [PubMed] [Google Scholar]
- Rogers, A. R., and H. C. Harpending, 1983. Population structure and quantitative characters. Genetics 105 985–1002. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Schluter, D., 1996. Adaptive radiation along genetic lines of least resistance. Evolution 50 1766–1774. [DOI] [PubMed] [Google Scholar]
- Spitze, K., 1993. Population structure in Daphnia obtusa: quantitative genetic and allozymic variation. Genetics 135 367–374. [DOI] [PMC free article] [PubMed] [Google Scholar]
- Steppan, S. J., P. C. Phillips and D. Houle, 2002. Comparative quantitative genetics: evolution of the G matrix. Trends Ecol. Evol. 17 320–327. [Google Scholar]
- Trouvé, S., L. Degen, C. Meunier, C. Tirard, S. Hurtrez-Boussès et al., 2000. Microsatellites in the hermaphroditic snail, Lymnaea truncatula, intermediate host of the liver fluke, Fasciola hepatica. Mol. Ecol. 9 1661–1686. [DOI] [PubMed] [Google Scholar]
- Trouvé, S., L. Degen, F. Renaud and J. Goudet, 2003. Evolutionary implications of high selfing rate in the freshwater snail Lymnaea truncatula. Evolution 57 2303–2314. [PubMed] [Google Scholar]
- Trouvé, S., L. Degen and J. Goudet, 2005. Ecological components and evolution of selfing in the freshwater snail Galba truncatula. J. Evol. Biol. 18 358–370. [DOI] [PubMed] [Google Scholar]
- Weir, B. S., 1996. Genetic Data Analysis, Vol. II. Sinauer Associates, Sunderland, MA.
- Weir, B. S., and C. C. Cockerham, 1984. Estimating F-statistics for the analysis of population structure. Evolution 38 1358–1370. [DOI] [PubMed] [Google Scholar]
- Whitlock, M. C., 1999. Neutral additive genetic variance in a metapopulation. Genet. Res. 74 215–221. [DOI] [PubMed] [Google Scholar]
- Whitlock, M. C, 2008. Evolutionary inference from Qst. Mol. Ecol. 17 1885–1896. [DOI] [PubMed] [Google Scholar]
- Whitlock, M. C., P. C. Phillips and K. Fowler, 2002. Persistence of changes in the genetic covariance matrix after a bottleneck. Evolution 56 1968–1975. [DOI] [PubMed] [Google Scholar]
- Wright, S., 1969. Evolution and the Genetics of Populations, 2. The Theory of Gene Frequencies. University of Chicago Press, Chicago.
- Yang, R.-C., 1998. Estimating hierarchical F-statistics. Evolution 52 950–956. [DOI] [PubMed] [Google Scholar]






